Abstract
Drug resistance in cancer poses a serious challenge in finding an effective remedy for cancer patients, because of the multitude of contributing factors influencing this complex phenomenon. One way to counter this problem is using a more targeted and dose-limiting approach for drug delivery, rather than relying on conventional therapies that exhibit multiple pernicious side-effects. Stability and specificity have traditionally been the core issues of peptide-based delivery vectors. In this study, we employed a structural regression modelling approach in the design, synthesis and characterization of a series of peptides that belong to approximately same topological cluster, yet with different electrostatic signatures encoded as a result of their differential positioning of amino acids in a given sequence. The peptides tagged with the fluorophore 5(6)-carboxyfluorescein, showed higher uptake in cancer cells with some of them colocalizing in the lysosomes. The peptides tagged with the anti-cancer drug methotrexate have displayed enhanced cytotoxicity and inducing apoptosis in triple-negative breast cancer cells. They also showed comparable uptake in side-population cells of lung cancer with stem-cell like properties. The most-optimized peptide showed accumulation in the tumor resulting in significant reduction of tumor size, compared to the untreated mice in in-vivo studies. Our results point to the following directives; (i) peptides can be design engineered for targeted delivery (ii) stereochemical engineering of peptide main chain can resist proteolytic enzymes and (iii) cellular penetration of peptides into cancer cells can be modulated by varying their electrostatic signatures.
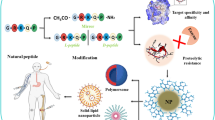
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ten million deaths and 19.3 million new cancer cases have been reported in the year 2020 with female breast cancer being the most diagnosed globally [1]. In higher-income countries, cancer has emerged as the leading cause of death exceeding the estimates of cardiovascular diseases indicating a new epidemiological transition. A major reason for this transition is the varied nature of cancer as a disease, with unknown risk factors [2]. Side-effects of conventional chemotherapy and radiotherapy seriously limit their future as dependable options, and more recent modes of treatment like immunotherapy and targeted therapy have also not been matured to the desired efficacy level [3]. This has given rise to the phenomenon of cancer drug resistance, where the patient becomes non-responsive towards therapeutic agents and therefore is a major reason for relapse.
Drug resistance in cancer can be intrinsic where the cancer cells are inherently resistant to a therapeutic agent before its administration or it can be extrinsic where the tumor cells develop resistance to the drug after treatment [4]. Numerous factors lead to drug resistance in cancer cells such as drug efflux, drug inactivation, drug target alteration, epigenetic mechanisms, inhibition of cell death, tumor microenvironment and tumor heterogeneity [5]. Current strategies to combat the problem of drug resistance in cancer include combinational therapy of two or more types of drugs, glucose deprivation, self-assembling prodrugs for reversing drug resistance, targeting the cellular and non-cellular components of the tumor microenvironment and targeting cytotoxicity of cancer stem cells [6,7,8,9,10].
It has been established that adoption of a structure is mediated by the interplay of short-range and long-range electrostatics in a peptide. The two interactions can be modulated in a de novo design attempt, by stereochemical engineering of the peptide backbone. This stereochemical sequence to structure relationship can be further extended by carefully choosing amino acids at designed positions to impart a desired function. This strategy has successfully been demonstrated in few earlier published reports [11,12,13,14,15,16]. This study is designed with two broad objectives. In the first, it proposes to systematically verify how effectively we can tune the structure and surface electrostatics in imparting specificity to a functional peptide while it encounters (i) cancerous and non-tumorigenic cell types and (ii) between different cancer cell types. Secondly, it investigates the possibility of using this design strategy to develop conformationally stable peptides for use as delivery vectors for the treatment of resistant cancers.
Cell-penetrating peptides (CPPs) are a novel class of peptides that possess the property of entering the cell, and their use as drug delivery vectors progressed after their discovery in 1988 [17]. They are short peptides with 5–30 amino acids and can be attached with a wide variety of cargo such as nucleic acids, proteins, drugs, quantum dots and metal chelators for therapeutic or imaging purposes [18,19,20]. CPPs can be cationic, amphipathic or hydrophobic in nature. They enter the cells by energy-independent direct penetration pathway or by an energy-dependent endocytosis pathway and the internalization route can vary depending on the physico-chemical nature of the CPP and the type of cargo attached [21, 22]. Use of syndiotactic CPPs containing alternating D and L amino acids have been reported for drug delivery in cancer cells having higher proteolytic stability and affinity to cancer cells [11, 12]. Tumor homing peptides (THPs) having RGD/NGR motifs have been studied for specifically targeting cancer cells, intended to reduce toxic side effects to non-cancerous cells [23]. Combining a cell-penetrating peptide with a tumor-homing domain conjugated with an anti-cancer drug molecule can increase the specificity to tumor cells many fold, thus enhancing the therapeutic action of drugs [14, 15].
In this study we have chosen the drug methotrexate to investigate the efficacy of the delivery vector. Farber et al. demonstrated that the drug methotrexate, a type of folate analogue, successfully induced remission in children suffering from acute lymphoblastic leukaemia (ALL) [24]. Since then, it has achieved widespread use in the treatment of many types of cancers such as leukemia, lymphoma, breast cancer, osteosarcoma, lung cancer and also for the treatment of autoimmune diseases such as rheumatoid arthritis [25]. Methotrexate enters the cell by active transport through the reduced-folate transporter 1 (RFT-1), gets converted to polyglutamate and inhibits the action of dihydrofolate reductase (DHFR) which ultimately leads to inhibition of purine and thymidylate synthesis [26]. Efficacy of using methotrexate as an anti-cancer drug is limited due to the development of methotrexate resistance by, for example triple-negative breast cancer cells. This resistance occurs due to many factors such as mutation in RFT-1, DHFR gene amplification and reduction in polyglutamation. Moreover, methotrexate has a very narrow therapeutic range and administering higher concentrations of the drug leads to serious side-effects [25]. Development of methotrexate resistance to breast cancer cells has been reported and new methods of targeting methotrexate resistance cells have been studied [27,28,29]. We attempt a strategy to deliver the drug methotrexate for therapeutic purposes in triple-negative breast cancer cells using cell-penetrating peptides with different electrostatic signatures and geometry. Many CPPs exist in solution as a random coil, but attain a secondary structural conformation on interaction with a biological membrane [30]. Secondary structure of the peptides has an impact on the uptake of CPPs in cells, wherein, it is observed that helical peptides have a higher tendency for uptake [16].
To achieve our designed objectives, we adopt a ‘structural regression model’ by systematically changing the diameter of helical peptide molecules, conserving their chemistry as much as possible. We have designed four peptides of different helical diameters, that may lead to four different electrostatic signatures which may further impart the much-needed specificity and enhanced efficacy that are critical in the selection of a delivery vector. Peptide YC101 is α-helical in nature. The non-proteinogenic amino acid 2-aminoisobutyric acid (AiB) is used for constraining the topology of the peptide to an α-helical configuration. YC102 has a syndiotactic sequence forming a Π(L, D) gramicidin helix. YC103 forms a polyproline helix and YC104 has an extended conformation without helical folds. These peptides have very distinct electrostatic signatures which is a vital factor for interacting with the biological membrane. The different helical diameters of the peptides play a role in the qualitative and quantitative aspects of cellular uptake in various cancerous and non-tumorigenic cell lines. The fixing of the peptide geometry is thus, extremely crucial for successful cell-penetration and drug delivery.
Experimental procedures
Peptide synthesis
The designed peptides were chemically synthesized using Solid Phase Peptide Synthesis (SPPS) method using rink amide resin and Fmoc-chemistry. The resin is weighed according to the scale of synthesis and peptides are synthesized sequentially from C-terminus to N-terminus. Fmoc group is cleaved using 20% piperidine solution in DMF for 30 min. pH of the solution is reduced by washing the resin 8–10 times using DMF. Fmoc-protected amino acids along with activators HoBT, HBTU and DIPEA are added in 3 times excess. Attachment of amino acid is performed twice for a period of 1 h and second time for 30 min. After attachment and washing with DMF, the resin is again subjected to treatment with 20% piperidine and the next amino acids are added sequentially in a similar manner. Once all the amino acids are attached, we proceed for the attachment of the fluorophore 5(6)-carboxyfluorescein and the drug methotrexate. Once the synthesis is complete, the peptides are deprotected and cleaved from the resin using a mixture of 1,2-ethanedithiol, m-cresol, thioanisole, and trifluoroacetic acid. The peptides are filtered using glass wool and precipitated and washed several times with ice-cold diethyl ether.
Purification and characterization of synthesized peptides
Synthesized peptides were purified by reverse-phase High Performance Liquid Chromatography (RP-HPLC) using a C-18 semi-preparative column (250 × 10 mm, particle size = 10 μm, pore size = 120 Å). Gradient using 10% acetonitrile in ultrapure water with 0.1% trifluoroacetic acid as solvent A and 100% acetonitrile with 0.1% trifluoroacetic acid as solvent B was used. HPLC peaks were collected and analysed using mass spectrometry. MALDI-TOF (Bruker, Autoflex Speed) machine in reflectron mode is used to determine the molecular mass of the peptides using α-cyano-4-hydroxycinnamic acid (HCCA) matrix.
Characterization by circular dichroism (CD) spectroscopy
Purified peptides at 50 µM concentration were prepared by dissolving in ultrapure water. CD spectra of the peptides was recorded using Jasco J-1500 spectropolarimeter by scanning between 190 nm and 280 nm using a quartz cuvette of 1 mm pathlength. Each spectrum has an average of 8 scans and the data was converted to mean residual ellipticity as described [31].
Electrostatic profiling of the peptides
Delphi software was used to calculate the Poisson-Boltzmann electrostatic potentials (PB-EP) using the Finite Difference Poisson-Boltzmann equation [32]. The output file was saved into phi format. The PB-EP was mapped on the molecular surface of peptides, analysed and rendered using Pymol. The potentials are in the units of kT per unit charge (e) where k is the Boltzmann constant and T is the absolute temperature. Each peptide has a unique electrostatic fingerprint that correlates with its functional properties.
Molecular dynamics simulation studies
GROMACS program suite was used to perform Molecular Dynamics Simulations [33]. GROMOS96 forcefield with 53a6 parameter set together with Berger lipids forcefield were used for simulations [33, 34]. The 1-palmitoyl-2-oleoylglycero-3-phosphoglycerol (POPG) lipid bilayer structure was obtained from the MemBuilder server [35]. The POPG forcefield parameter from former research work by Kukol et al. was employed [36]. The lipid bilayer was positioned in the XY plane, while its thickness was determined in the Z plane. Four peptide molecules were positioned on top of the lipid bilayer in the Z plane. The system was then packed with water molecules, followed by a 200 ns production run [11, 12, 37].
Briefly, a 128-membered POPG lipid bilayer was generated. Four peptide molecules were positioned close to one of the outer edge of the bilayer. The system was subsequently solvated with water and its energy was minimized. Water was used as a solvent (simple point-charge model, SPC). Pressure coupling was achieved using the Parrinello-Rahman barostat, and the isothermal compressibility was set at 4.5 × 10− 5 bar − 1. Energy minimization (steepest descent) until a 1000 KJ 1000 KJ mol− 1 nm− 1 tolerance was the next step. Subsequently, it was equilibrated for one nanosecond (ns) in NPT (conserved Number of particles, Pressure, and Temperature) conditions.
A 200 ns production run followed equilibration with four peptides on a 128-membered POPG bilayer. Water (SPC) was used as the solvent. The Root Mean Square Deviation and radius of gyration were determined for the conformations of the peptides using the g_rms and g_gyrate tools of the GROMACS package. Conformational analysis and free energy calculations were carried out based on the output of g_cluster program of the GROMACS package.
Cell culture
Human cancer cell lines MDA-MB-231 (breast cancer) and A549 (lung cancer) were procured from National Cancer Institute (NCI), USA. HeLa (cervical cancer) was obtained from Central cell line repository of RGCB. These cells were cultured in Dulbecco’s Modified Eagle Media (DMEM) with the addition of 10% fetal bovine serum (FBS) and 1% antibiotic mixture (10 mg/mL streptomycin, 10,000 units/mL penicillin and 25 µg/mL Amphotericin B) [31]. Non-tumorigenic breast epithelial cell line MCF-10 A was cultured in Mammary Epithelial Basal Medium supplemented with 10% fetal bovine serum, bovine pituitary extract (BPE), hydrocortisone, hEGF, insulin and gentamicin/amphotericin-B. Lifetime imaging facility of RGCB is acknowledged for FRET-based Caspase sensor cells MDA-MB-231 SCAT3 NLS. All the cells were grown in a humidified incubator at 37 ºC with 5% CO2.
Comparative uptake of peptides by flow cytometry
70,000 cells of MDA-MB-231, A549, HeLa and MCF-10 A were seeded in 24-well plates and placed in an incubator at 37 ºC overnight. On the following day, the cells were treated with 5 µM of peptides tagged with 5(6)-carboxyfluorescein (peptide-CF) for 3 h. After 3 h, the cells were rinsed with PBS and treated with 0.04% of Trypan Blue dye for 5 min. Next the wells were washed thrice with PBS and cells were subjected to trypsinization. The cell pellet was then suspended in 200 µL of ice- cold PBS and subjected to flow cytometry analysis [38].
Cellular uptake of peptides by confocal microscopy
10,000 cells of MDA-MB-231, A549, HeLa and MCF-10 A were seeded in 96-well plates and placed in the incubator for 24 h. The following day, 5 µM of peptide-CF conjugates in serum-free and phenol red-free DMEM and MEBM media was added to the cells for 3 h in a 37 ºC incubator. Once the incubation period is complete the cells were rinsed twice with PBS and stained with 5 µg/ml Hoechst 33342 stain for 5 min. The cells in the 96-well plate were visualized in a confocal laser scanning microscope (Zeiss LSM 980 Airyscan 2) to confirm cellular uptake using 20X objective with 0.8 numerical aperture.
Colocalization of peptides by confocal microscopy
10,000 cells of MDA-MB-231, A549, HeLa and MCF-10 A were seeded in 96-well plates and kept in the incubator for 24 h. The following day, cells were incubated with 5 µM of peptide-CF conjugates in serum-free and phenol red-free DMEM and MEBM media for 3 h in a 37 ºC incubator. After 3 h, two PBS washes were given and incubated with either 125nM of MitoTracker Deep Red dye or 40nM of Lysotracker Deep Red dye for 1 h. After another PBS wash, the cells were stained with 5 µg/ml Hoechst 33,342 stain for 5 min. They were visualized in a confocal laser scanning microscope (Zeiss LSM 980 Airyscan 2) to study the organelle (mitochondria or lysosomes) colocalization using 40X objective with 0.95 numerical aperture. Values for Pearson’s correlation coefficient for colocalization was determined using the colocalization module in the Zeiss Zen Blue software.
Cytotoxicity assay
Cytotoxicity in MDA-MB-231 and MCF-10 A cells was determined using the tetramethyl rhodamine methyl ester (TMRM) dye. 10,000 cells were seeded in a 96-well plate and placed in an incubator overnight. The next day, cells were washed once with PBS and stained using 5 µg/ml Hoechst 33342 for 5 min. Next, the cells were treated with media containing 100nM TMRM dye and incubated at 37 ºC for 10 min. Cells were rewashed with PBS once more. Cells were then treated with peptides conjugated with the drug methotrexate (peptide-MTX) at different concentrations (5, 10, 15 and 20 µM), prepared in media containing 2% FBS and 20nM of TMRM. This was subjected to incubation for 48 h in an incubator. Fluorescent images using confocal microscope (Nikon A1R HD) were taken in red channel for mitochondrial staining and in blue channel for nuclear staining using 20X objective with 0.75 numerical aperture.
Apoptosis assay
10,000 MDA-MB-231 cells possessing stable expression of FRET-based caspase sensor (DEVD) were used to determine the apoptosis inducing potential of the peptide-drug conjugates. After seeding cells, peptide-MTX treatment at various concentrations (5, 10 and 15 µM) was given. After incubation for 48 h, imaging of CFP/YFP FRET ratio was performed using fluorescence microscope (Nikon). The images were analysed to generate quantitative data using the color code of CFP/YFP FRET ratio ranging from scale 0 to 2.
Serum biocompatibility assay
70,000 cells of MDA-MB-231, A549, HeLa and MCF-10 A were cultured in 24-well plates and placed in an incubator at 37 ºC overnight. The following day, 5 µM of peptide-CF conjugates were mixed with equal volume of Fetal Bovine Serum (FBS) and incubated at 37 ºC for 1 h. After one PBS wash, cells were subjected to treatment with serum-incubated and serum-free peptide-CF for 2 h. After 2 h, the cells are washed once and treated with 0.04% of Trypan Blue dye for 5 min. Multiple PBS washes were given and cells were subjected to trypsinization. The cell pellet was then suspended in 200 µL of ice- cold PBS and subjected to flow cytometry analysis.
Serum stability assay
25 µl of peptides at a concentration of 50µM was taken and mixed with an equal volume of sterile human serum (Sigma). This was incubated at 37 ºC for 1 h. After the incubation, 25 µl of 0.6% trichloroacetic acid was added for precipitation of serum proteins. The mixture is placed at 4 ºC for 30 min and then centrifuged at 13,000 rpm for 5 min. The supernatant was collected and 50 µl was injected in reverse-phase High Performance Liquid Chromatography (RP-HPLC) for analysis. Similar procedure was repeated for serum-untreated peptides and only serum without peptide, as control.
Hemolysis assay
2 ml of blood was drawn from a healthy individual in a tube containing sodium heparin. The whole blood was centrifuged at 2000 rpm for 5 min to separate the plasma from the RBCs. 2 ml of Phosphate buffer saline (PBS, pH 7.4) was added to the layer of RBCs and this was subjected to centrifugation at 2000 rpm for 5 min. This step of washing with PBS was repeated thrice until a clear supernatant was observed. Next, a 20% haematocrit was prepared from the RBCs by mixing 2 ml of PBS with 1 ml of RBCs. In a microcentrifuge tube, 50 µl of 20% haematocrit was mixed with an equal volume of 40µM peptide-MTX conjugate and subjected to incubation for 2 h at 37 ºC in an incubator with shaking conditions. Complete hemolysis of RBCs and no hemolysis was obtained using 0.5% Triton X-100 and PBS respectively. The tubes were then centrifuged at 800 g for 5 min and 75 µl of the supernatant was added in triplicate to the wells in a 96-well plate. Absorbance of the samples was read at 540 nm wavelength, and hemolysis (percent) was calculated as following [31]:
Cancer stem cell isolation by side-population assay
Side-population cells can be isolated using flow cytometry by exploiting their ability to efflux out the DNA-binding dye Hoechst 33342 by ABC transporters. The side-population cells are enriched in cancer stem cells that induce tumors, cause malignancies and resistance to therapy. This technique is very useful to isolate and culture cancer stem cells and study the effect of therapeutic molecules in-vitro [39]. A549 cells were seeded in 100-mm tissue culture plates to reach a confluency of 50–70%. Cells were detached by trypsinization and washed with PBS. 1 × 106 cells were subjected to Hoechst staining (5 µg/ml) for a duration of 90 min in a water bath and shaking the cells intermittently every 15 min. One set of cells were also treated with 100µM of the drug Verapamil along with Hoechst staining, and incubated in a similar way. After 1.5 h, the cells were centrifuged and sorted by flow cytometry. The Hoechst dye is excited with a UV laser at 355 nm and its fluorescence emission is measured with both 505 long-pass 670/40 filter (Hoechst Red) and 450/50 filter (Hoechst Blue). The side-population cells and the main population cells were collected through cell sorting and seeded in 96-well plates.
Comparison of cellular uptake of peptides in side-population and main-population cells
5000 side-population and main-population cells of A549 were seeded in a 96-well black-coloured clear bottom tissue culture plate. The following day, cells were treated with 5µM peptide-CF conjugates for three hours. Wells were washed with PBS twice and stained with 5 µg/ml Hoechst 33342 nucleic acid stain for 10 min at 37 ºC. Imaging was performed using confocal laser scanning microscopy (Zeiss LSM 980 Airy Scan 2). The uptake of peptides in side-population and main-population cells was compared by measuring the intensity in the green channel.
Tumor regression studies
All animal experiments were performed as per the ethical guidelines under the approval of Institutional animal ethical committee of RGCB (IAEC/813/TRSK/2020). For in-vivo experiments, NSG (NOD SCID gamma mouse) female mice were maintained at the Animal Research Facility of RGCB in a pathogen free condition and used at the age of 6–8 weeks. 100 µl of 106 MDA-MB-231 cells showing stable expression of Luc2-TdTomato (matrigel 50:50) were injected to the right fourth mammary fat pad in a subcutaneous manner. Tumor formation was confirmed by palpation and mice were segregated into three groups based on the treatment-control, MTX treated and YC102-MTX treated (N = 3). The peptide drug conjugate YC102-MTX, MTX or normal saline were administered as 100 µl injection once every 14 days after 50 days of tumor cell inoculation, when tumor reaches the size ready for imaging. YC102-MTX and MTX were administrated at a concentration of 2 mg/kg body weight of the mice. After the end of 4 weeks, the mice were placed under anesthesia using 2% isoflurane and D-luciferin potassium salts was injected by the intra-peritoneal route. Bioluminescence imaging was performed using IVIS Spectrum in vivo imaging system (Perkin Elmer). Every fifth day the tumor volume was measured using digital caliper and calculated using the equation V = 0.5 X L X W2., where V is the tumor volume, L is the tumor length and W is the tumor width. The animals were also euthanized and organs such as lung, spleen, heart, kidney and tumor tissues were collected for histopathology studies.
Biodistribution assay
To check for the biodistribution in mice, peptide YC102 was tagged with the fluorophore cyanine 7 (Cy7) at the N-terminus. Female NSG mice were segregated in 2 groups, healthy mice and tumor-bearing mice (N = 3). YC102-Cy7 conjugate was administered as 100µL injection through the tail vein at a concentration of 2 mg/kg body weight. Mice were placed under isoflurane anesthesia and imaged at the time intervals of 3, 24, 48, 72, 96 and 120 h using IVIS Spectrum in vivo imaging system (Perkin Elmer). Time dependent tumour accumulation of YC102-Cy7 conjugate was calculated as Total Radiant Efficiency (p/s) / (µW/cm²) using in vivo imaging software, Living Image® (64-bit) after background subtraction.
Histopathology studies
Organs such as liver, lung, spleen and tumor xenograft were collected, sectioned and subjected to H&E staining after performing dewaxing, dehydration and washing. H&E images were captured using 40X objective (oil immersion, NA 1.3) in Nikon ECLIPSE Ni-E microscope.
Statistical analysis
All experiments were performed with a minimum sample size of three with data represented as mean ± SEM. Statistical analysis was done using OriginPro 2018 software (OriginLab Corporation, USA). Comparison of data between groups was done using using one-way analysis of variance (ANOVA) and Tukey’s test with significance value set at p-value < 0.05 (*).
Results and discussion
Design, electrostatic profiling and chemical synthesis of the peptide-based delivery vectors
A set of four peptide-based delivery vectors having varying helical diameters with diversified geometrical shapes were designed. Peptide YC101 has an α-helical configuration, YC102 is a Π(L, D) gramicidin helix, YC103 forms a poly-proline helix and YC104 has an extended conformation. YC102 has the largest helical diameter, followed by α-helical peptide YC101. YC103 has the smallest diameter among all the helical peptides with 3 residues per turn. The designed peptides were synthesized at laboratory scale by employing conventional solid-phase peptide synthesis (Fmoc chemistry). Peptides were conjugated with 5(6)-carboxyfluorescein or methotrexate on the N-terminus for cellular uptake and cytotoxicity studies respectively. Schematics for solid-phase peptide synthesis is shown in Fig S1. The synthesized peptides were purified using reverse-phase HPLC with 95% purity and further characterized by mass spectrometry (MALDI-TOF) (Fig S2). The electrostatic fingerprints of the peptides were generated by electrostatic potential mapping using Delphi software (Fig. 1). The designed peptides are amphipathic, having distinct electrostatic fingerprints. Variations in the electrostatic potential distributions is an important aspect that modulates the interaction of the peptides with the membrane.
Structure and electrostatics of the designed peptides. Electrostatic fingerprints of the designed peptides are generated using DELPHI software. Poisson-Boltzmann electrostatic potential (PB-EP) maps on the peptide surface, as well as the cylindrical dimensions (diameter and height) of the four peptides (YC101: α-helix, YC102: syndiotactic gramicidin helix, YC103: poly-proline helix, and YC104: extended conformation) are shown. The color bar represents the color-coded PB-EP values on a scale of -10 (red) to + 10 (blue) in kT/e units. The difference among peptides in the spatial PB-EP and dimensions are a result of differences in stereochemistry and amino acid sequences. Amino acid sequence of the designed peptides are depicted with L-amino acids written in uppercase letters and D-amino acids written in lowercase letters and underlined
Molecular dynamics simulation and circular dichroism spectroscopy of the designed peptides
The interaction of the designed peptides with lipid bilayer in the presence of water was studied with Molecular Dynamics simulation experiments performed using POPG bilayer in water. All MD simulations were carried out in GROMACS program suite using the GROMOS96 force field with 53a6 parameter set and Berger lipids parameters. Structure of YC102 was designed with structural parameters similar to a gramicidin helix alternating L and D chiral amino acids at successive positions. Stereochemical distributions for the designed peptide molecules were validated with ProChiral program in bPE Toolkit [40].
To study the peptide-lipid bilayer interaction, the four peptide molecules were positioned close to one end of the POPG bilayer in water. The system was equilibrated, followed by a 200 ns production run in NPT conditions. After the 200 ns production run, Root Mean Square Deviations, Radius of Gyration distributions, cluster formation and free energy was calculated (Fig. S3). The peptides successfully penetrated the lipid bilayer’s outer layer during the 200 ns simulation (Supp. Video 1–4). Additionally, the relative positions of the peptides concerning the lipid head groups, specifically phospho-glycerol, at both the beginning and end of the 200 ns production run provides further evidence of this penetration (Fig. 2A, Fig. S3). The peptide molecules remained within the boundaries of lipid bilayer throughout the simulation.
Following the 200 ns production run, an assessment was conducted on each designed peptide to determine whether they maintained their intended structures throughout the simulation. This evaluation was based on Root Mean Square Deviations and Radius of Gyration distributions (Fig. S3). While the RMSD of the four designed peptides was quite comparable to each other (within 0.5 nm distance of deviations), YC102 and YC103 peptides have deviated relatively to a greater extent, especially after 100 ns. The radius of gyration estimates during the course of the 200 ns simulation suggests that gramicidin helical peptide YC102 and poly-proline helical peptide YC103 have deviated slightly from the designed fold. The Gibbs free energy landscape of YC101, YC102 and YC104 lipid bilayer simulations presents a relatively distinct Global Minimal Energy Conformation (GMEC), while the energy landscape of the YC103 peptide does not show any clear conformational directive (Fig. S3).
The topological changes are evident from the secondary structural changes verified using circular dichroism spectroscopy (CD). The secondary conformations of the peptides were confirmed by their signature peaks in CD spectroscopy experiments (Fig. S4).
Cellular uptake of the designed peptides. (A) Position of the peptide YC102 when interacting with POPG bilayer in water at the start, 0 ns, (dotted) and end (solid) of a 200 ns run in MD simulation. (B) Quantitative uptake of the peptides tagged with 5,6-carboxyfluorescein was compared in cancer cell-lines namely MDA-MB-231, A549, HeLa and non-tumorigenic breast epithelial cell-line MCF-10 A. All the cells were treated with 5µM of peptide-CF conjugates for 3 h and analysed using flow cytometry. Error bars represent mean ± SEM of three independent experiments and the statistical significance was set at p < 0.05 (*) and was assessed in comparison with the cellular uptake of the positive control TAT peptide. (C) Cellular uptake of the peptides in cells, visualized and imaged through confocal laser scanning microscopy. The cells were incubated with 5µM of peptide-CF conjugates for 3 h, washed with PBS and imaged using confocal microscope. Peptide-CF staining and nucleus stained with Hoechst 33342 are denoted by the green and blue channels respectively. Scale bar corresponds to 20 μm
Estimation of the cellular uptake of peptide-fluorophore conjugates in cancer and non-tumorigenic cells
Intracellular uptake of the peptides conjugated to 5(6)-carboxyfluorescein was assessed in four cell lines namely MDA-MB-231 (triple negative breast cancer), A549 (lung cancer), HeLa (cervical cancer) and MCF-10 A (non-tumorigenic mammary epithelial cells). The cells were treated with 5 µM of peptide-CF conjugates for 3 h. Multiple PBS washes and trypsinization of cells preceded flow cytometry. As shown in Fig. 2B, the peptides show differential uptake in different cell lines. The peptide YC102 showed enhanced uptake in MDA-MB-231 cells. The other three peptides YC101, YC103 and YC104 showed maximal uptake in cervical cancer cells. A well-established cell-penetrating peptide, TAT, is used as a positive control in our experiments. All the peptides showed better uptake than the control TAT peptide as shown in Fig. 2B.
Intracellular uptake of peptides was visualized by confocal laser scanning microscopy (Fig S5-S8). Peptides were observed to differentially internalize in the cells based on the type of cells as well as the overall geometry and surface electrostatics of the peptide. Peptide YC102 shows higher uptake in triple-negative breast cancer cells MDA-MB-231 in comparison to non-tumorigenic mammary epithelial cells MCF-10 A. It also displays higher uptake in HeLa and A549 cells (Fig. 2C).
Determination of peptide localization in sub-cellular organelles
Colocalization of peptide-CF conjugates were studied in two organelles namely mitochondria (Fig. S9-S12) and lysosomes (Fig. S13-S16) using commercially available MitoTracker Deep Red and Lysotracker Deep Red dyes. The degree of colocalization can be quantified by Pearson’s correlation coefficient whose values range from − 1 to + 1. The value of + 1 indicates perfect linear correlation between the fluorophores, the value − 1 denotes perfect negative correlation and the value 0 denotes uncorrelation between the fluorophore probes [41, 42]. As mentioned in Table S1, the designed peptide-CF conjugates show correlation values 0 or less than 0 with the mitochondria staining dye. This means that the peptides do not co-localize in the mitochondria but are distributed elsewhere in the cytoplasm or in other organelles. As seen in Fig. 3B and Fig. S13-S16, it can be observed that some of the peptides are localized in the lysosomes. This can be due to their uptake by the endosomal pathway. As mentioned in Table S2, peptide YC102 shows a correlation coefficient value of 0.85 and 0.87 with the lysosome staining dye in MDA-MB-231 and HeLa respectively. Such a high positive linear correlation indicates its colocalization in the lysosomes. The peptides YC101 and YC103 also show colocalization but to a lesser extent in the lysosomes of MDA-MB-231, as can be judged from their coefficient values ranging from 0.48 to 0.6. Further verification of localization is beyond the scope of this paper and hence not attempted.
Localization studies of the peptide YC102-CF in mitochondria and lysosome. (A) Cancer cells and non-tumorigenic cells after treatment with 5µM of peptide-CF conjugate for 3 h, were washed and stained with 125nM MitoTracker deep red dye and Hoechst 33342. (B) Cells after treatment with 5µM of peptide-CF conjugate for 3 h, were washed and stained with 40nM LysoTracker deep red dye and Hoechst 33342. Blue colour denotes nucleus, green denotes YC102-CF and red denotes mitochondria in panel A and lysosomes in panel B
Estimation of cytotoxic and apoptotic potential of peptide-drug conjugates in triple-negative breast cancer cells
The principal aim of a delivery vehicle is to successfully deliver the drug molecule to the cancer cells. Tetramethylrhodamine methyl ester (TMRM) based assay was performed to estimate cytotoxicity potential of the designed peptide-methotrexate (MTX) conjugates against triple negative breast cancer MDA-MB-231 cells and mammary epithelial MCF-10 A cells. Tetramethylrhodamine, methyl ester (TMRM) is a cell-permeant dye that accumulates in active mitochondria with intact membrane potentials. Healthy mitochondrial membranes maintain a difference in electrical potential between the interior and exterior of the organelle, referred to as membrane potential. Healthy cells with intact mitochondria show a bright red signal but in cells undergoing death there is a loss in the mitochondrial membrane potential, and the TMRM signal diminishes (Fig.S17, Fig. S18). Figure 4A, shows loss of cell viability of MDA-MB-231 cells by the peptide-MTX conjugates, with minimal loss to MCF-10 A cells.
Cytotoxicity studies for peptide-MTX conjugates. (A) Quantitative cell viability studies of the peptide-drug conjugates on MDA-MB-231 and MCF-10 A cells. MDA-MB-231 and MCF-10 A cells were treated with MTX and peptide-MTX conjugates at 5µM, 10µM, 15µM and 20µM concentrations for 48 h and stained with TMRM and Hoechst 33342 dye. The cells with diminished TMRM red signal and enhanced blue signal of nuclear condensation were undergoing cell death. (B) MDA-MB-231 SCAT3 NLS cells were treated with peptide-MTX conjugates and the cells undergoing apoptosis displayed a change in fluorescence CFP-YFP ratio corresponding to the activation of Caspase-3. The graph represents percentage of MDA-MB-231 cells undergoing apoptotic cell death on treatment with peptide-MTX conjugates
Cellular apoptosis of cancer cells is a therapeutic target for many drugs and molecules. Apoptosis, also known as programmed cell death is under tight regulation by the cells and is characterized by certain morphological hallmarks. It causes minimal inflammation as compared to necrosis and therefore it is targeted by cancer therapeutics. Apoptosis is initiated by activation of cysteine proteases called caspases. Two initiation pathways of apoptosis, namely extrinsic and intrinsic, finally converge into a common pathway by activation of caspase-3 [43]. To study the apoptotic potential of our peptide-drug conjugates, we have used MDA-MB-231 cells expressing stable CFP-YFP FRET-based probe. The two fluorophores, CFP and YFP, are joined together by a peptide-based caspase-3 cleavage site. Upon treatment with the peptide-drug conjugates, the cells undergo apoptosis which can be observed by a change in the CFP-YFP fluorescence ratio (Fig. S19). As shown in Fig. 4B, treatment of the peptide YC102-MTX at a concentration of 15µM causes more than 30% of cells to undergo irreversible apoptosis.
Estimation of hemotoxicity, serum stability and serum biocompatibility of the peptides
Hemolysis assay was performed to estimate the toxicity of the drug delivery vehicle. RBCs were isolated and treated with 20µM peptide-MTX conjugate. Heme release for all the peptide drug conjugates was less than 3% indicating negligible hemotoxicity (Fig.S20).
Peptides are susceptible to serum protease degradation and is one of the important limiting factors for the use of peptide-based delivery vehicles. To address this, we incorporated D-amino acids and non-proteinogenic amino acid 2-Aminoisobutyric acid (AiB) into the peptide sequence based on their structure. As shown in Fig. S21, cellular uptake of serum treated and untreated peptides is comparable thus indicating stability of the peptides in serum. We have extensively studied this in our earlier published reports, and this is the most important feature that qualifies heterochiral peptide sequences as promising drug delivery vehicles.
Stability of the peptides in serum was checked by incubating the peptides with human serum, followed by characterization with reverse phase-HPLC. As shown in Fig. S22, similar HPLC profile resulted for untreated and serum-treated peptides, indicating stability of peptides in serum and resistance to protease degradation. This indicates that the designed peptides can be utilized for in-vivo studies and as a future therapeutic agent.
Estimation of cellular uptake of peptides in side-population cells having stem cell-like properties
Drug resistance in cancer can occur due to a wide array of factors like drug efflux, drug inactivation, epigenetic phenomena, DNA repair mechanisms, inhibition of cell death and involvement of components from the tumor microenvironment. Cancer stem cells which are a part of the heterogenous tumor population show chemotherapy resistance and play a significant role in drug resistance. Side-population assay is a technique to identify and isolate cancer cells having stem cell-like properties. This assay is based on the Hoechst dye efflux properties of the side-population cells through the ATP-binding cassette (ABC) transporters. To confirm the specificity of the dye efflux, the ABC transporter inhibition assay using the drug verapamil, a known inhibitor of ABC transporter is performed simultaneously.
Side-population cells are less prevalent in basal subtype of breast cancer cells as compared to the luminal subtypes [44]. Therefore, MDA-MB-231 cells, which are of the basal subtype showed undetectable side population cells on treatment with Hoechst 33,342 [45]. A549 lung cancer cells showed significant side-population, which were isolated by flow cytometry and cultured in 96-well plates. The cellular uptake potential of peptides tagged with carboxyfluorescein was estimated in the side-population and main population cells using confocal laser scanning microscopy.
As observed in the flow cytometry profile of Fig. 5A, the percentage of side-population cells that are sorted by Hoechst staining is approximately 4% of the entire cellular population of A549. A tight smaller gate was used for sorting based on Verapamil treated sample to ensure extreme drug efflux population. This percentage reduces to almost a negligible 0.05% after Verapamil treatment. Thus, this method provides an effective way to isolate and study side-population cells enriched in cancer stem cells.
Cellular uptake of peptides in side-population cells (A) Representative flow cytometric profiles obtained after staining A549 lung carcinoma cells with 5 µg/mL Hoechst 33342 in the absence or presence of verapamil. The side-population (SP) and main-population (MP) fractions are gated. (B) Comparative uptake of the peptides tagged with 5,6-carboxyfluorescein in SP and MP cells of A549 cell line. The cells were treated with 5µM of peptide-CF conjugates for 3 h and the uptake was quantified by measuring the intensity in the green channel. (C) Uptake of the peptide-CF conjugates in the SP and MP cells, visualized and imaged through confocal laser scanning microscopy. The cells were incubated with 5µM of peptide-CF conjugates for 3 h, washed with PBS and imaged with a 20x objective using a confocal microscope. Nucleus stained with Hoechst 33342 and peptide-CF conjugates are denoted by the blue and green channels respectively. Scale bar corresponds to 20 μm
After isolating the side population and main population cells, the cellular penetration and uptake of the peptides was determined. The side-population and main-population cells isolated after Hoechst staining assay were plated in equal number in 96-well plates and made to attach and grow overnight. The following day, the cells were treated with peptide-CF conjugates for 3 h and the intracellular cellular uptake was studied using confocal laser scanning microscopy. The extent of uptake was quantified by measuring the intensity of the green channel for the carboxyfluorescein-tagged peptides. Panel C in Fig. 5, displays the qualitative images for cellular uptake, where the peptides are denoted by green colour and the nucleus stained with Hoechst 33342 is denoted by blue colour. Based on the quantitative measurement of the intensity of the green channel shown in panel B, it is suggested that the cellular uptake of the peptides is similar in side-population and main-population cells of A549. Also, the cellular uptake of the designed peptides is 30–40 folds higher than the uptake of TAT peptide. This indicates that the peptides can be used to therapeutically target cancer stem-cell like population of the tumors.
Estimation of tumor regression in triple-negative breast cancer tumor bearing mice
For pre-clinical breast cancer studies, mice models are routinely used for their human genetic similarities, quick results and cost-effectiveness [46]. The conjugate YC102-MTX was used to test the efficacy in mouse xenograft models. Figure 6A shows bioluminescence images taken from tumor bearing control mice and tumor bearing mice subjected to MTX or YC102-MTX conjugate treatment. Tumor regression was observed in MTX and YC102-MTX treated mice in comparison with the untreated control. Maximum tumor regression was observed in YC102-MTX treated mice. Considerable reduction in the size of tumor excised from mice treated with YC102-MTX was observed (Fig. 6B) in comparison to the tumor excised from control and MTX treated mice. Tumor regression by measurement of tumor volume is shown in Fig. S23. Splenic enlargement is a characteristic observed in metastatic breast cancer [47]. As can be seen in the Fig. 6B, the spleen from untreated tumor-bearing control mice is found to be considerably enlarged. Spleen from mice treated with only MTX is also observed to be enlarged but the size of the spleen excised from mice treated with YC102-MTX was significantly smaller. This also confirms tumor regression in mice treated with YC102-MTX conjugate.
Estimation of peptide-fluorophore biodistribution in-vivo mice models
To study the biodistribution in healthy and tumor-bearing mice, YC102 peptide was conjugated with the fluorophore cyanine-7 (excitation = 750 nm, emission = 773 nm). YC102-Cy7 was administered at a concentration of 2 mg/kg body weight of the mice. The mice were imaged at the time intervals of 3, 24, 48, 72, 96 and 120 h (Fig. 6C). In healthy mice, fluorescence was observed from the renal and hepatic system within 3 h which suggests excretion and metabolism respectively. In tumor-bearing mice, YC102-Cy7 started accumulating at the tumor site after 3 h of treatment, which continued till the end of the experiment (120 h). Time dependent tumoral accumulation of YC102-Cy7 is shown in Fig. S24. This indicates that the peptide YC102 has affinity for the cancerous tumor whereas in the absence of tumor it gets metabolized and excreted.
Tumor regression and Biodistribution in mouse xenograft model. (A) Representative bioluminescence image from NOD/SCID female mice showing tumor regression with MTX and YC102-MTX treatment compared with untreated control (N = 3). In vivo, bioluminescence images were obtained using an IVIS Lumina system to record the tumor size at the end of 4 weeks of MTX and YC102-MTX treatment. Bioluminescence images showed reduction in tumor size in YC102-MTX treatment compared to MTX and untreated control. (B) Photographic images of excised tumors from the untreated control, MTX, and YC102-MTX treatment groups. YC102-MTX treatment showed a maximum reduction in tumor size in comparison with other groups. The spleens were also observed to be enlarged in the untreated control and MTX-treated group, but significantly small in the YC102-MTX treatment group. (C) Biodistribution of YC102-Cy7 conjugate. Healthy and tumor-bearing mice were injected with YC102-Cy7 conjugate (2 mg/kg body weight, N = 3). Fluorescent images in healthy mice showed peptide accumulation in the renal and hepatic system within 3 h of injection. On the other hand, in tumor-bearing mice, the peptide started accumulating at the site of tumor after 3 h of treatment till the end (120 h)
Histopathology of tissues of internal organs and tumor xenograft
Tumor histology from the control untreated mice showed proliferating tumor mass with increased proliferating front infiltration into the adjacent skin with necrotic areas (Fig. 7). Proliferation induced necrosis was also evident in tumor tissue from MTX treated mice; whereas, the tumor from the mice treated with YC102-MTX did not show necrosis. Liver histology from the control and MTX treated mice displayed micrometastasis which was absent in mice treated with YC102-MTX. Overall, the study demonstrated that tumour retention by YC102-MTX conjugate suppressed tumour growth and secondary metastasis in comparison to MTX alone.
Histopathology of tumor and internal organs. Histopathological H&E staining was performed on tumor xenograft, liver, lung and spleen. Tumor from the untreated control and MTX treated mice indicated signs of necrosis which was absent in YC102-MTX treated mice. The liver from the untreated mice also showed micro metastasis. Images were captured using 40X objective (oil immersion, NA 1.3) in Nikon ECLIPSE Ni-E microscope
Conclusion
In this paper, we proposed a structural regression modelling approach to verify the efficacy of such a design algorithm in peptide functional design and to further explore the possibility of employing it in the generation of delivery vectors for the treatment of resistant cancer. Such ‘dimensionality reduction’ approaches have already been used in other computational methods, especially in data science and machine learning [48].
Conventional chemotherapy for cancer treatment causes toxic side-effects in patients necessitating the need for employment of drug delivery techniques, to deliver therapeutic molecules specifically targeted to cancer cells. These techniques include a wide-variety of delivery molecules such as lipids, proteins, polymers, nanoparticles, organic and inorganic materials. In this study we use cell-penetrating peptides having distinct electrostatic signatures and varying helicity to deliver the anti-cancer drug methotrexate. Among the four designed peptides, the syndiotactic peptide YC102 with a gramicidin Π(L, D) helix showed enhanced cellular uptake in MDA-MB-231 triple negative breast cancer cells. YC102 conjugated to methotrexate displayed enhanced cytotoxicity and apoptosis in triple negative breast cancer cells with negligible toxicity in non-tumorigenic breast epithelial cells. Moreover, the designed peptides also showed cellular uptake in side-population cells having stem-cell like properties which indicates their application in targeting cells with drug efflux property.
The efficacy of peptide-drug conjugate YC102-MTX was further tested in mice models which showed significant tumor regression in the treated mice, while compared with the control. Tumor size was also found to be significantly reduced in the case of treatment with YC102-MTX, compared to same concentration of MTX. Biodistribution studies of YC102 tagged with the fluorophore cyanine-7 showed continued accumulation in tumor. In contrast, the peptide was found to accumulate in the liver and kidneys of the control mice indicating its excretion and metabolism. Enlargement of spleen a characteristic feature of metastasis in breast cancer, was observed in the case of untreated and methotrexate treated tumor bearing mice. But the spleen was found to be approximately normal-sized in the case of mice treated with YC102-MTX.
The results obtained from our study indicate that the cell-penetrating peptide YC102 can be a promising potential candidate for drug delivery for anti-cancer therapeutics in triple negative breast cancer which is an aggressive subtype of breast cancer with negligible possible therapeutic opportunity. This may be regarded as one of the key translational advantage of such a multidisciplinary investigation.
Data availability
The datasets generated and analyzed from this study can be made available upon request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49.
Dagenais GR, Leong DP, Rangarajan S, Lanas F, Lopez-Jaramillo P, Gupta R, Diaz R, Avezum A, Oliveira GBF, Wielgosz A, Parambath SR, Mony P, Alhabib KF, Temizhan A, Ismail N, Chifamba J, Yeates K, Khatib R, Rahman O, Zatonska K, Kazmi K, Wei L, Zhu J, Rosengren A, Vijayakumar K, Kaur M, Mohan V, Yusufali A, Kelishadi R, Teo KK, Joseph P, Yusuf S. Variations in common diseases, hospital admissions, and deaths in middle-aged adults in 21 countries from five continents (PURE): a prospective cohort study. Lancet. 2020;395(10226):785–94.
Mun EJ, Babiker HM, Weinberg U, Kirson ED, Von Hoff DD. Tumor-treating fields: a fourth modality in cancer treatment. Clin Cancer Res. 2018;24(2):266–75.
Prieto-Vila M, Takahashi RU, Usuba W, Kohama I, Ochiya T. Drug resistance driven by cancer stem cells and their niche. Int J Mol Sci. 2017;18(12):2574.
Housman G, Byler S, Heerboth S, Lapinska K, Longacre M, Snyder N, Sarkar S. Drug resistance in cancer: an overview. Cancers (Basel). 2014;6(3):1769–92.
Suraweera A, O’Byrne KJ, Richard DJ. Combination therapy with histone deacetylase inhibitors (HDACi) for the treatment of cancer: achieving the full therapeutic potential of HDACi. Front Oncol. 2018;8:92.
Abdel-Wahab AF, Mahmoud W, Al-Harizy RM. Targeting glucose metabolism to suppress cancer progression: prospective of anti-glycolytic cancer therapy. Pharmacol Res. 2019;150:104511.
Yang C, Tu K, Gao H, Zhang L, Sun Y, Yang T, Kong L, Ouyang D, Zhang Z. The novel platinum(IV) prodrug with self-assembly property and structure-transformable character against triple-negative breast cancer. Biomaterials. 2020;232:119751.
Mo R, Gu Z. Tumor microenvironment and intracellular signal-activated nanomaterials for anticancer drug delivery. Mater Today. 2016;19(5):274–83.
Sun HR, Wang S, Yan SC, Zhang Y, Nelson PJ, Jia HL, Qin LX, Dong QZ. Therapeutic strategies targeting cancer stem cells and their microenvironment. Front Oncol. 2019;9:1104.
Jerath G, Goyal R, Trivedi V, Santhoshkumar TR, Ramakrishnan V. Syndiotactic peptides for targeted delivery. Acta Biomater. 2019;87:130–9.
Jerath G, Darvin P, Christian Y, Trivedi V, Kumar TRS, Ramakrishnan V. Delivery of small molecules by syndiotactic peptides for breast cancer therapy. Mol Pharm. 2022;19(8):2877–87.
Goyal R, Jerath G, Akhil R, Chandrasekharan A, Puppala ER, Ponneganti S, Sarma A, Naidu VGM, Santhoshkumar TR, Ramakrishnan V. Geometry encoded functional programming of tumor homing peptides for targeted drug delivery. J Control Release. 2021;333:16–27.
Goyal R, Jerath G, Chandrasekharan A, Christian Y, Kumar TRS, Ramakrishnan V. Molecular hybridization combining tumor homing and penetrating peptide domains for cellular targeting. Drug Deliv Transl Res. 2022;12(5):1285–92.
Goyal R, Jerath G, Chandrasekharan A, Kumar TRS, Ramakrishnan V. Peptide-based delivery vectors with pre-defined geometrical locks. RSC Med Chem. 2020;11(11):1303–13.
Jerath G, Goyal R, Trivedi V, Santhoshkumar TR, Ramakrishnan V. Conformationally constrained peptides for drug delivery. J Pept Sci. 2020;26(4–5):e3244.
Frankel AD, Pabo CO. Cellular uptake of the tat protein from human immunodeficiency virus. Cell. 1988;55(6):1189–93.
Chen B, Yoo K, Xu W, Pan R, Han XX, Chen P. Characterization and evaluation of a peptide-based siRNA delivery system in vitro. Drug Deliv Transl Res. 2017;7(4):507–15.
Schneider AFL, Wallabregue ALD, Franz L, Hackenberger CPR. Targeted subcellular protein delivery using cleavable cyclic cell-penetrating peptides. Bioconjug Chem. 2019;30(2):400–4.
Ruan G, Agrawal A, Marcus AI, Nie S. Imaging and tracking of tat peptide-conjugated quantum dots in living cells: new insights into nanoparticle uptake, intracellular transport, and vesicle shedding. J Am Chem Soc. 2007;129(47):14759–66.
Guidotti G, Brambilla L, Rossi D. Cell-penetrating peptides: from basic research to clinics. Trends Pharmacol Sci. 2017;38(4):406–24.
Copolovici DM, Langel K, Eriste E, Langel Ü. Cell-penetrating peptides: design, synthesis, and applications. ACS Nano. 2014;8(3):1972–94.
Davoodi Z, Shafiee F, Internalizing RGD. A great motif for targeted peptide and protein delivery: a review article. Drug Deliv Transl Res. 2022;12(10):2261–74.
Farber S, Diamond LK. Temporary remissions in acute leukemia in children produced by folic acid antagonist, 4-aminopteroyl-glutamic acid. N Engl J Med. 1948;238(23):787–93.
Koźmiński P, Halik PK, Chesori R, Gniazdowska E. Overview of dual-acting drug methotrexate in different neurological diseases, autoimmune pathologies and cancers. Int J Mol Sci. 2020;21(10):3483.
Chabner BA, Roberts TG Jr, Timeline: chemotherapy and the war on cancer. Nat Rev Cancer. 2005;5(1):65–72.
Volk EL, Farley KM, Wu Y, Li F, Robey RW, Schneider E. Overexpression of wild-type breast cancer resistance protein mediates methotrexate resistance. Cancer Res. 2002;62(17):5035–40.
Lindgren M, Rosenthal-Aizman K, Saar K, Eiríksdóttir E, Jiang Y, Sassian M, Ostlund P, Hällbrink M, Langel U. Overcoming methotrexate resistance in breast cancer tumour cells by the use of a new cell-penetrating peptide. Biochem Pharmacol. 2006;71(4):416–25.
Böhme D, Krieghoff J, Beck-Sickinger AG. Double methotrexate-modified Neuropeptide Y Analogues Express increased toxicity and overcome drug resistance in breast cancer cells. J Med Chem. 2016;59(7):3409–17.
Medina SH, Miller SE, Keim AI, Gorka AP, Schnermann MJ, Schneider JP. An intrinsically disordered peptide facilitates non-endosomal cell entry. Angew Chem Int Ed Engl. 2016;55(10):3369–72.
Vibin Ramakrishnan. Biophysical characterization of functional peptides. Humana New York, NY, 2023.
Li L, Li C, Sarkar S, Zhang J, Witham S, Zhang Z, Wang L, Smith N, Petukh M, Alexov E. DelPhi: a comprehensive suite for DelPhi software and associated resources. BMC Biophys. 2012;5:9.
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 2015;1–2:19–25.
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF. A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem. 2004;25(13):1656–76.
Ghahremanpour MM, Arab SS, Aghazadeh SB, Zhang J, van der Spoel D. MemBuilder: a web-based graphical interface to build heterogeneously mixed membrane bilayers for the GROMACS biomolecular simulation program. Bioinformatics. 2014;30(3):439–41.
Kukol A. Lipid models for united-atom molecular dynamics simulations of proteins. J Chem Theory Comput. 2009;5(3):615–26.
Redkar AS, Ramakrishnan V. Molecular dynamics simulations. In: Ramakrishnan V, editor. Biophysical characterization of functional peptides. Humana, New York, NY: Springer Protocols Handbooks; 2023.
Christian Y, Ramakrishnan V. Comparative uptake of peptides by flow cytometry. In: Ramakrishnan V, editor. Biophysical characterization of functional peptides. New York, NY: Springer Protocols Handbooks. Humana; 2023.
Shimoda M, Ota M, Okada Y. Isolation of cancer stem cells by side population method. Methods Mol Biol. 2018;1692:49–59.
Jerath G, Hazam PK, Ramakrishnan V. bPE toolkit: toolkit for computational protein engineering. Syst Synth Biol. 2014;8(4):337–41.
Dunn KW, Kamocka MM, McDonald JH. A practical guide to evaluating colocalization in biological microscopy. Am J Physiol Cell Physiol. 2011;300(4):C723–42.
Adler J, Parmryd I. Quantifying colocalization by correlation: the Pearson correlation coefficient is superior to the Mander’s overlap coefficient. Cytometry A. 2010;77(8):733–42.
Wong RS. Apoptosis in cancer: from pathogenesis to treatment. J Exp Clin Cancer Res. 2011;30(1):87.
Nakanishi T, Chumsri S, Khakpour N, Brodie AH, Leyland-Jones B, Hamburger AW, Ross DD, Burger AM. Side-population cells in luminal-type breast cancer have tumour-initiating cell properties, and are regulated by HER2 expression and signalling. Br J Cancer. 2010;102(5):815–26.
Patrawala L, Calhoun T, Schneider-Broussard R, Zhou J, Claypool K, Tang DG. Side population is enriched in tumorigenic, stem-like cancer cells, whereas ABCG2 + and ABCG2- cancer cells are similarly tumorigenic. Cancer Res. 2005;65(14):6207–19.
Boix-Montesinos P, Soriano-Teruel PM, Armiñán A, Orzáez M, Vicent MJ. The past, present, and future of breast cancer models for nanomedicine development. Adv Drug Deliv Rev. 2021;173:306–30.
Dunn MA, Goldwein MI. Hypersplenism in advanced breast cancer: report of a patient treated with splenectomy. Cancer. 1975;35(5):1449–52.
Van Der Maaten L, Postma E, van den Herik J. Dimensionality reduction: a comparative review. J Mach Learn Res. 2009;10(66–71):13.
Acknowledgements
The authors thank the Central Instrument facility, IIT Guwahati for mass spectrometry facility and Bio-imaging facility at Rajiv Gandhi Centre for Biotechnology for confocal and flow cytometry facilities.
Funding
This work was supported by funding from Central Council for Research in Ayurvedic Sciences, Ministry of AYUSH, Government of India [F.No. 3–27/2021-CCRASAdmn./Coll. (O.O. No. 347/2021-22)], Department of Biotechnology, Ministry of Science & Technology, Government of India [BT/PR25526/NER/95/1238/2017] and Indian Institute of technology, Guwahati.
Author information
Authors and Affiliations
Contributions
Yvonne Christian- experimental work, data analysis, manuscript writing; Amay Sanjay Redkar- experimental work (in silico); Naveen Kumar- experimental work (in silico); Shine Varghese Jancy- experimental work (in vivo); Aneesh Chandrasekharan- confocal imaging; Thankayyan Retnabai Santhoshkumar- supervision and inputs in in vitro and in vivo experiments, resources, manuscript review; Vibin Ramakrishnan- project idea and conceptualization, experimental supervision, manuscript review and editing.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All animal experiments were performed under the supervision of institutional animal care and experimental committee of Rajiv Gandhi Centre for Biotechnology (RGCB), Thiruvananthapuram, India. Informed consent was obtained from all the human participants of this work.
Consent for publication
Consent for publication was obtained from all the participants.
Conflict of interest
No conflicts to declare. However, a patent has been filed on the discussed content of this paper at IPO Kolkata (Application number: 202431019764, date: 17/03/2024).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1: Additional experimental schematics, graphs and results along with supplementary video files to this article are attached with this submission and can be found under Supplementary Information
Supplementary Material 2
Supplementary Material 3
Supplementary Material 4
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Christian, Y., Redkar, A.S., Kumar, N. et al. Structural regression modelling of peptide based drug delivery vectors for targeted anti-cancer therapy. Drug Deliv. and Transl. Res. (2024). https://doi.org/10.1007/s13346-024-01674-y
Accepted:
Published:
DOI: https://doi.org/10.1007/s13346-024-01674-y