Abstract
Plastic pollution has become a serious issue of global concern, and biodegradation of plastic wastes is representing one attractive environment-friendly alternative to traditional disposal paths. It is known that insects are involved in the plastic polymer degradation process, with reported evidence of tenebrionid beetle larvae capable to degrade polystyrene (PS), one of the most used plastics worldwide. Recently, a ribosomal RNA based survey on the insect gut microbiota of the lesser mealworm Alphitobius diaperinus has revealed differentially abundant microbial taxa between PS-fed larvae and control group. Following these findings, an enrichment bacterial culture was set up in liquid carbon-free basal medium with PS film as sole carbon source using PS-fed larvae of A. diaperinus as inoculum. After two-months the culture was analysed both by molecular and culture-based methods. Isolated bacteria which had become prevalent under the selective enrichment conditions resulted ascribable to three taxonomic groups: Klebsiella, Pseudomonas, and Stenothrophomonas. The predominance of these groups in PS-fed larvae was confirmed by using bacterial 16S rRNA gene amplicon sequencing, and it was consistent with the results of previous reports. Isolated bacteria were able to attach to PS surfaces and SEM observations showed the presence of thin fibrillar structures connecting the bacterial cells to the abiotic surface.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Plastics are represented by a large family of polymeric materials deriving from hydrocarbons that, since 1950s, offer innovative solutions to human-society requirements. In 2019 global plastic production reached 368 million tons with Europe accounting for 16% of totality (Plastics Europe 2020). Among the most popular plastics in the world, polystyrene (PS) is still representing the fifth most used thermoplastic in Europe due to its excellent processability, low cost, resistance and insulating properties (Plastics Europe 2020); consequently, its end-of-life disposal is a source of particular concern. Nowadays, PS recycling rates are still low at global level (Takada and Bell 2021), and a consistent portion of PS waste ends up in the environment where its degradation is a complex and slow process which depends on chemical structure of the polymer and environmental conditions (Ho et al. 2018; Ali et al. 2021). The high durability of PS in the environment and its progressive fragmentation into submillimetre-sized particles can have a negative impact on different levels of biological complexity (Hollóczki and Gehrke 2019). Recently, a great interest has been directed towards some insect species involved in PS biodegradation with lepidopteran species, such as the greater wax moth Galleria mellonella (Jiang et al. 2021), and coleopteran species, such as the superworm Zophobas atratus (Peng et al. 2020), the dark mealworm Tenebrio obscurus (Peng et al. 2019), the yellow mealworm Tenebrio molitor (Yang et al. 2015a, 2015b) and the red flour beetle Tribolium castaneum (Wang et al. 2020), as candidate taxa to be tested for their ability to process plastics. Mealworms and superworms are provided with basic mandibulate mouthparts that enable them to chew and ingest the polymer. Moreover, suppression of their gut microbes with the antibiotic gentamicin indicated a gut-microbial dependence for PS biodegradation (Peng et al. 2019, 2020; Yang et al. 2021). A recent study carried out on another coleopteran species, Alphitobius diaperinus, concluded that also this beetle is capable of eating and degrading PS. In this species, comparative analyses of the gut microbiome have revealed several microbial taxa to be differentially abundant between PS-fed larvae and control groups, thus providing an initial list of microbial candidates involved in the degradation process (Cucini et al. 2020). The lesser mealworm A. diaperinus is a nocturnal cosmopolitan Tenebrionidae, commonly found in litter, manure and chicken coops. In chicken farms, the beetle feeds on manure, spilled feed, cracked eggs, and bird carcasses. A. diaperinus complete life-cycle lasts from 40 to 100 days. Adults, 5–8 mm long, usually lay their eggs (1.5 mm in length) in litter, droppings and grain hulls; larvae can reach 7–11 mm in length and, during the last larval instar, they leave the litter and make galleries into thermal insulation materials, frequently including PS, in order to construct pupal cells (Dunford and Kaufman 2006).
The aim of this study was to further expand the work started by Cucini et al. (2020) focusing on the gut-associated bacterial communities of PS-fed larvae of A. diaperinus. An enrichment procedure in liquid carbon-free basal medium with PS films as sole carbon source starting from PS-fed larvae of A. diaperinus was performed and analysed in order to (i) compare the composition of enriched communities by culture-dependent and culture-independent molecular methods and to (ii) obtain a collection of bacteria potentially exploitable for synthetic microbiota application.
2 Materials and methods
2.1 Insect material
Last instar larvae of A. diaperinus were purchased online from Agripet Garden (https://www.agripetgarden.it/; Padua, IT). Three-hundred grams (approximately 1700 individuals) were reared under controlled conditions (20 ± 2 °C; 12 L:12D photo-period; 50–70% RH) and fed with 20 g of PS as unique source of carbon. The PS foam (0.01 g/cm3), known as Extir® (CAS 9003-53-6), was purchased from Toscoespansi s.r.l. (http://www.toscoespansi.it/). According to the manufacturer information, no extra additives or catalysts were present. After 30 days, 50 larvae were collected, immersed 1 min in 75% ethanol and washed in Saline Water (SW) [0.85% NaCl] for the same time, then intestines were sterilely dissected under a Leica Wild M3C stereomicroscope.
2.2 Enrichment culture and bacteria isolation
Enrichment culture was established as essentially reported by Yang et al. (2014). Dissected intestines were harvested by centrifugation (7000×g, 20 min), and washed twice in SW. The pellet was resuspended in 40 ml of Liquid Carbon-Free Basal medium (LCFB) [0.7 g of KH2PO4, 0.7 g of K2HPO4, 1.0 g of NH4NO3, 0.7 g of MgSO4·7H2O, 0.005 g of NaCl, 0.002 g of FeSO4·7H2O, 0.002 g of ZnSO4·7H2O, and 0.001 g of MnSO4·H2O in 1000 ml of distilled water, pH 7.2] containing PS films 0.0125 g/ml. PS films were prepared dissolving 3.5 g of PS foam in 116 ml of trichloromethane (0.03 g/ml), then the solution was poured on glass plates and dried overnight under chemical fume hood (Yang et al. 2015b; Sekhar et al. 2016). Films were cut (10 mm × 10 mm; 9–10 mg/each), washed once with 40 ml of ethanol (>99%) and three times with sterile water then sterilized, both sides, under UV light. Serial dilutions of the suspension were performed in SW and spread-plated on Luria Bertani Agar (LBA) [10 g of bacto tryptone, 5 g of yeast extract, 5 g of NaCl and 16 g of agar to 1000 ml of distilled water, pH 7.2]. The plates were incubated under aerobic conditions at 27 °C for 24–48 h, and the total number of viable cells was determined. The enrichment culture was incubated on a shaker with gentle agitation (90 rpm) under aerobic conditions at room temperature (20–22 °C) for 60 days under natural light conditions. From the two-months old culture aliquots were prepared and stored at −80 °C for molecular analysis; at the same time, serial dilutions were performed in SW and spread-plated on LBA for bacterial enumeration and strain isolation. Plates were then incubated as reported above. Individual colonies representing isolates with different morphologies were selected according to differences in colour, size and appearance. They were picked from the different dilutions LBA plates and sub-cultured at least three times before identification. Bacterial isolates were then stored in LB broth with 30% glycerol (v/v) at −80 °C.
2.3 Phenotypic and phylogenetic characterization of bacterial isolates
Gram determination was carried out either by staining (Gram stain kit, Carlo Erba Reagents) or chemical method (Powers 1995). Catalase and oxidase activity were determined according to Smibert and Krieg (1981). The level of salt tolerance of the bacterial isolates was determined in LB broth supplemented with various concentrations of NaCl (5, 7.5, and 10%) inoculated with overnight grown bacteria and incubated at 27 °C for 48 h. LB broth without extra addition of NaCl was maintained as control. Optical density at 600 nm was measured using UV-Vis spectrophotometer (Novaspec, Amersham Biosciences, Amersham, UK). Sensitivity of the isolates to common antibiotics (all purchased from Sigma-Aldrich) was tested by spotting 4 μl drops of overnight cultures on LBA plates supplemented with carbenicillin (100 μg/ml), gentamicin (30 μg/ml), kanamycin (50 μg/ml), and antibiotic-free (control). Results of bacterial growth after incubation at 27 °C for 24–48 h were evaluated in comparison to positive (antibiotic-free) control. The ability of the isolates to produce lipolytic activity was tested on Tributyrin agar (Oxoid) [5 g of peptone, 3 g of yeast extract, 10 g of glyceryl tributyrate and 15 g of agar to 1000 ml of distilled water, pH 7.5]. Production of urease, lysine decarboxylase, ornithine decarboxylase, tryptophanase, hydrogen sulphide and acetoin, acid production from glucose, adonitol, lactose, arabinose, sorbitol and dulcitol, and assimilation of citrate were determined by the EnteroPluri-Test (Liofilchem®) according to the manufacturer’s instructions. All assays were repeated at least twice.
From bacterial cells grown overnight in LB, total DNA was extracted using the kit Wizard® SV Genomic DNA Purification System (Promega Corporation, Madison, WI, US). For each DNA sample the amplification of 16S rRNA gene was performed using universal primers: 16S_V3V4-341F (5’-CCTACGGGNGGCWGCAG-3′) and 16S_V3V4-785R (5’-GACTACHVGGGTATCTAATCC-3′) [Klindworth et al. 2013]. PCRs were executed in 25 μl reaction volume with 2.5 μl of DNA from each sample, 1.25 μl of both forward and reverse primers (10 mM), 2.5 μl of MgCl2 (25 mM), 2.5 μl of deoxynucleotides (10 mM), 5 μl of Green GoTaq Flexi Buffer (Promega, US), 0.125 μl of GoTaq Flexi DNA polymerase (Promega, US), and 9.875 μl of ddH2O. Amplifications were obtained in a GeneAmp® 2700 Thermal Cycler (Applied Biosystem, Foster City, CA, US), using the following conditions: (i) 95 °C for 3 min; (ii) 35 cycles of 95 °C for 1 min, 50 °C for 1 min, 72 °C for 90 sec; (iii) 72 °C for 5 min. The kit Wizard® SV Gel and PCR Clean-Up System (Promega Corporation) was employed for PCR products purification and their DNA concentration was assessed through Nanodrop ND1000UV vis device (NanoDrop Technologies, Wilmington, DE, US). Sanger sequencing was executed on a DNA Analyzer ABI 3730 at the BioFab Research core facility (Rome, IT). Electropherograms were manually checked with the Sequencher 4.4.2 software (Gene Codes, Ann Arbor, MI, US). The resulting sequences were then aligned against GenBank (NCBI) through the nucleotide Basic Local Alignment Search Tool - BLASTn (Altschul et al. 1990).
The sequence data were submitted to the DDBJ/EMBL/GenBank databases under accession numbers: OL470964, OL470965, and OL470969.
2.4 Sequencing of 16S rRNA amplicons
Total DNA extractions were carried out on three technical replicates from aliquots of the enrichment culture, using the Wizard® Genomic DNA Purification Kit according to manufacturer’s protocol. Final DNA concentrations were determined by NanoDrop ND1000V and Qubit® fluorometer (Life Technologies Corporation, Carlsbad, CA, US), while DNA quality was evaluated through 1% agarose gel electrophoresis. NGS metabarcoding was conducted with the same set of primers (V3V4: 341F-785R) used for Sanger sequencing at the BioFab Research core facility (Rome, IT). PCR reactions were performed in a 25 μl volume with 2.5 μl DNA, 5 μl 5x PCRBIO Reaction Buffer, 1 μl of both primers (10 μM), 0.25 μl PCRBIO HiFi Polymerase, 15.25 μl ddH2O with the following conditions: (i) 95 °C for 3 min; (ii) 25 cycles of 95 °C for 30 sec, 55 °C for 30 sec, 72 °C for 30 sec; (iii) 72 °C for 5 min. PCR products were purified with AMPure XP Beads (Beckman Coulter, High Wycombe, UK) according to the manufacturer’s specifications and each purification product was indexed using the Nextera XT Index Kit (Illumina) for library production. The latter reaction was carried out using the following conditions: (i) 95 °C for 3 min; (ii) 8 cycles of 95 °C for 30 sec, 55 °C for 30 sec, 72 °C for 30 sec; (iii) 72 °C for 5 min. Indexed samples were purified with Ampure XP Beads according to the PCR Clean-Up protocol suggested by Illumina; resulting products were quantified using a Bioanalyzer DNA 1000 (Agilent Technologies, Santa Clara, CA, US) and pooled in equal amounts for sequencing. A MiSeq Illumina sequencer was used to perform paired-end sequencing. Raw data, which amounted to a total of 207,858 reads (ranging from 56,899 to 83,230 per replicate) were filtered on a quality basis and primers trimmed using trimmomatic v.0.39 tool (Bolger et al. 2014). To investigate enriched bacterial community composition, sequences were processed using Quantitative Insights Into Microbial Ecology 2 (QIIME2 v.2021.4) (Boylen et al. 2019). In accordance with the software pipeline, through DADA2 plugin, already demultiplexed sequences were joined, filtered and dereplicated (Callahan et al. 2016). The vsearch plugin was employed both for chimera removal and clustering in the different Operational Taxonomic Units (OTUs) at 97% using the de novo strategy (Rognes et al. 2016). Furthermore, singletons and contaminants (i.e. mitochondrial and chloroplast sequences) were discarded. To train the classifier artifacts for taxonomic attribution of 16S sequences, the SILVA 132 database was employed (Quast et al. 2013), using feature-classifier plugin (Bokulich et al. 2018) [Protocol S1]. Resulting sequences were matched with those of Cucini et al. (2020) dataset though the NCBI BLAST+ tools (Camacho et al. 2009). Taxonomic distribution results were visualized QIIME2 view and also compared with those of the first 16S rRNA survey on PS-fed A. diaperinus larvae (Cucini et al. 2020). Raw sequencing reads were deposited in the Sequence Read Archive (SRA, NCBI) with the BioProject ID PRJNA781815.
2.5 Analysis of adhesion and growth of bacterial isolates in LCFB with PS
For each bacterial strain, cells were collected from overnight cultures in LB by centrifugation (10,000×g, 5 min), washed three times and suspended in sterile SW. A bacterial inoculum to give a final concentration of 5 × 105 Colony Forming Unit (CFU) per millilitre (ml) was added to sterile tubes containing 4 ml each of LCFB or Phosphate buffered saline (PBS) with 0.0125 g/ml PS films (10 mm × 10 mm). Bacterial suspensions were statically incubated under aerobic conditions at room temperature (20–22 °C) for 28 days.
For SEM observation, one PS film was collected from each culture and washed 1 h in 8 ml PBS on a shaker (90 rpm) to remove unattached cells. Films were fixed in 2.5% glutaraldehyde for 2 h and post-fixed in 1% osmium tetroxide for 1 h (Dallai et al. 2016). They were later washed in PBS and then dehydrated in a graded series of ethanol (50–100%), dried after terbutilic alcohol sublimation, and gold-coated with a Balzers MED 010 sputtering device. Samples were examined and photographed through a FEI Quanta 400 Scanning electron microscope operating at 20 kV.
For mechanic and enzymatic assay, the procedure reported by Banar et al. (2016) was modified as follows: three PS films from bacterial suspensions from both LCFB and PBS (control) were collected after 3 h and 28 days, respectively. Each film was individually transferred in a tube containing 8 ml of sterile PBS and left 1 h on a shaker (90 rpm). PBS was then removed and replaced with 1 ml of PBS containing 1 mg of trypsin (Sigma-Aldrich), vortexed 1 min, incubated 1 h at 37 °C, vortexed again for 1 min. Viable counts were determined through serial dilutions that were spread-plated on LBA. Plates were incubated at 27 °C for 24–48 h.
3 Results and discussion
3.1 Phenotypic and phylogenetic characterization of bacterial isolates
The liquid enrichment from PS-fed larvae of A. diaperinus in a carbon-free basal medium with PS film as sole carbon source was examined by the viable plate method at the beginning and after two months of incubation at the reported conditions. The estimated number of CFU/ml at the end of the trial showed that viable cells concentration increased from ~6.0 × 106 to ~1.5 × 108 CFU/ml (Table S1), most likely indicating that few bacteria became “enriched” by asserting their dominance through growth in such minimal medium over the considered period. In total three different morphologies were observed among colonies grown after 48 h in LBA from the two-months PS-enriched culture. Isolates representative of each morphotype were named 2 m/a, 2 m/b, and 2 m/c (Table S2). All three isolates resulted Gram-negative rod-shaped bacteria, and their phenotypic characterization, based on various biochemical and physiological parameters, is shown in Table 1. Within the set of whole determined phenotypic characteristics, some distinctive phenetic traits were considered by referring to a simplified dichotomic scheme for the identification of Gram-negative rods based on Bergey’s Manual of Determinative Bacteriology (Holt 1994; Fig. S1). Following this scheme, strain 2 m/a was included among lactose-fermenting Enterobacteriaceae; 2 m/b and 2 m/c, both unable to ferment glucose and lactose, but being different with respect to cytochrome oxidase production, were included within oxidase-positive (Pseudomonas), and oxidase-negative group (Acinetobacter, Burkholderia, Stenotrophomonas), respectively. Furthermore, the three isolates showed different level of resistance to a beta-lactam and aminoglycosides, a common trait among Klebsiella pneumoniae, Pseudomonas aeruginosa, and Stenotrophomonas maltophilia species which were included in the top ten list of multidrug-resistant bacteria (Rello et al. 2019). Crippen and Poole (2009) reported that the gastrointestinal tract of lesser mealworm larvae is capable of supporting the occurrence of horizontal transfer of antibiotic resistance genes between different species of bacteria under laboratory conditions, therefore A. diaperinus, as well as other edible and household insects, could constitute a reservoir of transferable resistance genes of human health concern (Gwenzi et al. 2021).
A partial sequence of 16S rRNA gene (464–475 bp) was determined for each of three bacterial isolates. A BLAST search with the 16S rDNA sequences against NCBI nucleotide databases, indicated that 2 m/a and 2 m/b shared 99.65% identity with Klebsiella aerogenes ATCC 13048 (GenBank: NR_118556.1; Zhang et al. 2014) and Pseudomonas aeruginosa DSM 50071 (GenBank: NR_117678.1; Swiderski 2012), respectively; while 2 m/c showed 97.59% identity with Stenotrophomonas maltophilia ATCC 13637 (GenBank: MT354120.1; Reid et al. 2021).
Members belonging to genus Klebsiella, Pseudomonas and Stenotrophomonas have been already found associated with plastic compounds and proposed as plastic eaters (Brandon et al. 2021), and, more specifically, in association to Z. atratus (Kim et al. 2020), and T. molitor (Urbanek et al. 2020; Brandon et al. 2021). The repeated finding of similar bacteria associated to PS-fed larvae strengthens then the hypothesis of their involvement in PS degradation process in A. diaperinus as well as in the other plastic-eater insects so far investigated.
3.2 Sequencing of 16S rRNA amplicons
From the Miseq Illumina sequencing of the 16S molecular marker (V3-V4 regions) a total amount of 207,858 reads, ranging from 56,899 to 83,230 per library, were obtained. After filtering, dereplication, clustering, singletons and chimeras’ removal, the three technical replicates had respectively 26,580, 24,920 and 36,302 sequences, for a total of 87,802, and 57 OTUs. Sequences shared an average length of 399 nucleotides. At family level, Xanthomonadaceae, Pseudomonadaceae and Enterobacteriaceae were largely dominant in the sample with regard to other taxa (Fig. 1), with the former owning the highest relative abundance. At genus level, two taxa predominated over the others, namely Stenotrophomonas and Pseudomonas (Fig. 1), with an average relative abundance of above 41% and 35.5%, respectively.
16S rRNA-based analysis of the enrichment culture from PS-fed Alphitobius diaperinus’ gut. Taxa bar plots exhibit OTUs diversity of bacteria at (a) Family level: left, present study; right, based on data from Cucini et al. (2020); (b) Genus level, showing their relative abundance in the three technical replicates (Meta1, Meta2, Meta3). Taxa are colour coded. In (a), OTUs with a relative abundance smaller than 1% were grouped and displayed as ‘Other’
Partial 16S rRNA gene sequences from strains 2 m/a, 2 m/b and 2 m/c were aligned on the 16S rRNA gene amplicons dataset, and all three found a meaningful correspondence (>99% identity). However, taxonomical association of the strain 2 m/a, namely Klebsiella sp., was assessed only up to OTU-family level – i.e. Enterobacteriaceae - probably due to a limitation of SILVA database used in the QIIME2 pipeline. This observation also explains the absence of Klebsiella genus in the taxonomic barplot (Fig. 1). From the initial 16S rRNA survey on A. diaperinus larval microbiome reported by Cucini et al. (2020), a total amount of 295 different OTUs were retrieved with only six as differentially abundant in PS group, namely: two Pseudomonas sp., Cronobacter sp., Kocuria sp., Pseudogracilibacillus sp. and Virgibacillus sp.. BLAST analysis of 16S rRNA gene sequences from strains 2 m/a, 2 m/b and 2 m/c against libraries from Cucini et al. (2020) retrieved significant matches (> 99% identity). Sequence of 16S rRNA gene from the three bacteria isolated in this study matched with OTUs that had a higher relative abundance in the previously reported PS libraries, meaning that they were also present with higher abundance in PS-fed A. diaperinus larvae (Fig. 1).
The alignment of 16S rRNA sequences obtained in this study against sequences differentially abundant in A. diaperinus’s PS-fed libraries from Cucini et al. (2020) provided confirmation of the presence of Lactococcus, Pseudomonas spp., and a group of Enterobacteriaceae (i.e. Klebsiella family). Other Enterobacteriaceae and Stenotrophomonas were also present in PS libraries, but their higher relative abundance with respect to control libraries was not significant by statistical analysis, (p-adj > 0.05), and then not indicative of differentially abundant bacterial taxa in the insect PS-fed group (Fig. 1).
3.3 Analysis of adhesion and growth of bacterial isolates in LCFB with PS
The ability of isolates Klebsiella sp. 2 m/a, Pseudomonas sp. 2 m/b, and Stenotrophomonas sp. 2 m/c to grow on PS films was evaluated by SEM observation and enzymatic/mechanical assay.
SEM observations were carried out on PS films from 28 days-old liquid cultures in carbon-free mineral medium. Bacterial cells on PS films appeared healthy, rod-shaped with smooth intact surface. Moreover, the used magnification showed thin fibrillar structures connecting the bacterial cells to the abiotic surface (Fig. 2). For Stenotrophomonas sp., few bacterial aggregates were also observed (Fig. 2).
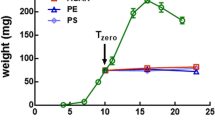
The number of attached viable cells on PS films after 3 h and 28 days incubation in mineral medium and phosphate-buffer was recorded for all strains. After 3 h no significative difference was observed between LCFB and PBS by considering the number of detached viable cells with enzymatic/mechanical treatment. After 28 days no viable cells of Klebsiella sp. 2 m/a and Stenotrophomonas sp. 2 m/c were recovered from PS films kept in PBS, on the contrary Pseudomonas sp. 2 m/b did not show any decline in viability accordingly with what was already stated for other members of the same genus which were found able to survive in PBS for at least 30 weeks (Liao and Shollenberger 2003). An increase of the number of viable cells detached from PS films maintained in LCFB was observed for all strains after 28 days when compared to the number of viable detached after 3 h, but the only significant difference (p value <0.05) was that exhibited by Stenotrophomonas sp. 2 m/c (Fig. 3a; Table S3).
Analysis of bacterial adhesion to PS film after enzymatic/mechanic assay. For each strain, histograms show viable counts after 3 h and 28 days (dd) incubation in LCFB medium and PBS, respectively. a: Cells detached from PS films; b: Planktonic cells. Error bars represent standard deviation. Effect sizes are presented in Table S3 and in Table S4
The viability of cells in planktonic form was also determined. The patterns of population changes in planktonic state were not significantly different in LCFB after 3 h and 28 days of incubation (Fig. 3b; Table S4). As already observed for the attached state, no viable planktonic cells were recovered from PBS for Klebsiella sp. 2 m/a and Stenotrophomonas sp. 2 m/c after 28 days; nonetheless, Klebsiella sp. 2 m/a, even if not increased significantly its concentration, survived well in mineral medium both in planktonic and attached state until the end of the observation period (Fig. 3b; Table S4). Results indicate that a pure culture made up of individuals of one species only, with the possible exception of Stenotrophomonas sp. 2 m/c (Fig. 3a; Table S3), does not show any significant change in LCFB with PS after an incubation period of four weeks. It must be highlighted that the approach followed in this study ignores the possibility that PS biodegradation may result from the cooperative action of different microbial species under natural conditions; in addition, the metabolic process of a single species may lead to the accumulation of intermediates or dead-end products with potentially toxic effects on bacterial growth.
4 Conclusion
This is the first study where culture-dependent and molecular analyses have been combined to the analysis of the bacterial population associated with plastic-eating A. diaperinus (Insecta: Coleoptera) after an enrichment phase in a medium with limited chemical composition and PS as sole carbon source. The results obtained indicate the predominance of three taxonomic groups, Klebsiella, Pseudomonas, and Stenothrophomonas, either by culture-dependent and -independent analysis of the enriched communities. Nonetheless, bacterial isolates of each taxon, when grown as monoculture in a synthetic medium with PS, revealed a weak metabolic activity. In nature, PS degradation process could be influenced by the interactions between members of a complex microbial community, the higher genotypic and phenotypic diversity present in microbial communities compared with a single strain population may mean that mixed communities are more efficient degraders of the pollutant. Therefore, the lack of cooperative microbial interactions at the applied experimental conditions could have been constraining. Notwithstanding these limitations, the study suggests that bacterial isolates could be representative members of microbial taxa involved in PS degradation as already reported (Brandon et al. 2021). Those findings could represent a valid starting point for the generation of synthetic bacterial communities to be used for microbiome transfer to gnotobiotic insects in order to better understand the web of interactions involving both heterospecific symbionts and the animal host during the process of PS degradation.
Data availability
All data generated or analysed during this study are included in this published article.
References
Ali SS, Elsamahy T, Al-Tohamy R, Zhu D, Mahmoud YAG, Koutra E, Metwally AM, Kornaros M, Sun J (2021) Plastic wastes biodegradation: mechanisms, challenges and future prospects. Sci Total Environ 780:146590. https://doi.org/10.1016/j.scitotenv.2021.146590
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Banar M, Emaneini M, Satarzadeh M, Abdellahi N, Beigverdi R, Leeuwen WB, Jabalameli F (2016) Evaluation of mannosidase and trypsin enzymes effects on biofilm production of Pseudomonas aeruginosa isolated from burn wound infections. PLoS One 11. https://doi.org/10.1371/journal.pone.0164622
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Boylen E, Knight R, Huttley GA, Caporaso JG (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6:90. https://doi.org/10.1186/s40168-018-0470-z
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Boylen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Brandon AM, Garcia AM, Khlystov NA, Wu W, Criddle CS (2021) Enhanced bioavailability and microbial biodegradation of polystyrene in an enrichment derived from the gut microbiome of Tenebrio molitor (mealworm larvae). Environ Sci Technol 55:2027–2036. https://doi.org/10.1021/acs.est.0c04952
Callahan B, McMurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Crippen TL, Poole TL (2009) Conjugative transfer of plasmid-located antibiotic resistance genes within the gastrointestinal tract of lesser mealworm larvae, Alphitobius diaperinus (Coleoptera: Tenebrionidae). Foodborne Pathog Dis 6:907–915. https://doi.org/10.1089/fpd.2008.0260
Cucini C, Leo C, Vitale M, Frati F, Carapelli A, Nardi F (2020) Bacterial and fungal diversity in the gut of polystyrene-fed Alphitobius diaperinus (Insecta: Coleoptera). Animal Gene 17-18:200109. https://doi.org/10.1016/j.angen.2020.20010
Dallai R, Mercati D, Mashimo Y, Machida R, Beutel RG (2016) The fine structure of the rectal pads of Zorotypus caudelli Karny (Zoraptera, Insecta). Arthropod Struct Dev 45:380–388. https://doi.org/10.1016/j.asd.2016.06.004
Dunford J, Kaufman P (2006) Lesser mealworm, litter beetle, Alphitobius diaperinus (Panzer) (Insecta: Coleoptera: Tenebrionidae). EDIS. https://doi.org/10.32473/edis-in662-2006
Gwenzi W, Chaukura N, Muisa-Zikali N, Teta C, Musvuugwa T, Rzymski P, Abia A (2021) Insects, rodents, and pets as reservoirs, vectors, and sentinels of antimicrobial resistance. Antibiotics 10:68. https://doi.org/10.3390/antibiotics10010068
Ho B, Roberts T, Lucas S (2018) An overview on biodegradation of polystyrene and modified polystyrene: the microbial approach. Crit Rev Biotechnol 38:308–320. https://doi.org/10.1080/07388551.2017.1355293
Hollóczki O, Gehrke S (2019) Nanoplastics can change the secondary structure of proteins. Sci Rep 9:16013. https://doi.org/10.1038/s41598-019-52495-w
Holt JG (1994) Bergey’s Manual of Determinative Bacteriology, 9th edn. Lippincott Williams and Wilkins, Baltimore
Jiang S, Su T, Zhao J, Wang Z (2021) Isolation, identification, and characterization of polystyrene-degrading bacteria from the gut of Galleria mellonella (Lepidoptera: Pyralidae) larvae. Front Bioeng Biotechnol 9:736062. https://doi.org/10.3389/fbioe.2021.736062
Kim HR, Lee HM, Yu HC, Jeon E, Lee S, Li J, Kim DH (2020) Biodegradation of polystyrene by Pseudomonas sp. isolated from the gut of superworms (larvae of Zophobas atratus). Environ Sci Technol 54:6987–6996. https://doi.org/10.1021/acs.est.0c01495
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:1. https://doi.org/10.1093/nar/gks808
Liao C-H, Shollenberger L (2003) Survivability and long-term preservation of bacteria in water and in phosphate-buffered saline. Lett Appl Microbiol 37:45–50. https://doi.org/10.1046/j.1472-765x.2003.01345.x
Peng B, Li Y, Fan R, Chen Z, Chen J, Brandon AM, Criddle C, Zhang Y, Wu W (2020) Biodegradation of low-density polyethylene and polystyrene in superworms, larvae of Zophobas atratus (Coleoptera: Tenebrionidae): broad and limited extent depolymerization. Environ Pollut 266:115206. https://doi.org/10.1016/j.envpol.2020.115206
Peng B, Su Y, Chen Z, Chen J, Zhou X, Benbow M, Criddle C, Wu W, Zhang Y (2019) Biodegradation of polystyrene by dark (Tenebrio obscurus) and yellow (Tenebrio molitor) mealworms (Coleoptera:Tenebrionidae). Environ Sci Technol 59:5256–5265. https://doi.org/10.1021/acs.est.8b06963
Plastics Europe (2020) Plastics - The facts 2020. https://www.plasticseurope.org/it/resources/publications/4312-plastics-facts-2020. Accessed 3 September 2021
Powers EM (1995) Efficacy of the Ryu nonstaining KOH technique for rapidly determining Gram reactions of food-borne and waterborne bacteria and yeasts. Appl Environ Microbiol 61:3756–3758. https://doi.org/10.1128/aem.61.10.3756-3758.1995
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Rello J, Kalwaje Eshwara V, Lagunes L, Alves J, Wunderink RG, Conway-Morris A, Rojas JN, Alp E, Zhang Z (2019) A global priority list of the TOp TEn resistant Microorganisms (TOTEM) study at intensive care: a prioritization exercise based on multi-criteria decision analysis. Eur J Clin Microbiol Infect Dis 38:319–323. https://doi.org/10.1007/s10096-018-3428-y
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:2584. https://doi.org/10.7717/peerj.2584
Sekhar VC, Nampoothiri KM, Mohan AJ, Nair NR, Bhaskar T, Pandey A (2016) Microbial degradation of high impact polystyrene (HIPS), an e-plastic with decabromodiphenyl oxide and antimony trioxide. J Hazard Mater 318:347–354. https://doi.org/10.1016/j.jhazmat.2016.07.008
Smibert RM, Krieg NR (1981) General characterization. In: Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Krieg NR, Philips GB (eds) Manual of methods for general bacteriology. American Society for Microbiology, Washington, pp 411–442
Takada H, Bell L (2021) Plastic waste management hazards. IPEN. https://ipen.org/sites/default/files/documents/ipen-plastic-waste-management-hazards-en.pdf. Accessed 3 September 2021
Urbanek AK, Rybak J, Wróbel M, Leluk K, Mirończuk AM (2020) A comprehensive assessment of microbiome diversity in Tenebrio molitor fed with polystyrene waste. Environ Pollut 262:114281. https://doi.org/10.1016/j.envpol.2020.114281
Wang Z, Xin X, Shi X, Zhang Y (2020) A polystyrene-degrading Acinetobacter bacterium isolated from the larvae of Tribolium castaneum. Sci Total Environ 726:138564. https://doi.org/10.1016/j.scitotenv.2020.138564
Yang J, Yang Y, Wu W, Zhao J, Jiang L (2014) Evidence of polyethylene biodegradation by bacterial strains from the guts of plastic-eating waxworms. Environ Sci Technol 48:13776–13784. https://doi.org/10.1021/es504038a
Yang L, Gao J, Liu Y, Zhuang G, Peng X, Wu W, Zhuang X (2021) Biodegradation of expanded polystyrene and low-density polyethylene foams in larvae of Tenebrio molitor Linnaeus (Coleoptera: Tenebrionidae): broad versus limited extent depolymerization and microbe-dependence versus independence. Chemosphere 262:127818. https://doi.org/10.1016/j.chemosphere.2020.127818
Yang Y, Yang J, Wu W, Zhao J, Song Y, Gao L, Yang R, Jiang L (2015a) Biodegradation and mineralization of polystyrene by plastic-eating mealworms: part 1. Chemical and physical characterization and isotopic tests. Environ Sci Technol 49:12080–12086. https://doi.org/10.1021/acs.est.5b02661
Yang Y, Yang J, Wu W, Zhao J, Song Y, Gao L, Yang R, Jiang L (2015b) Biodegradation and mineralization of polystyrene by plastic-eating mealworms: part 2. Role of gut microorganisms. Environ Sci Technol 49:12087–12093. https://doi.org/10.1021/acs.est.5b02663
Acknowledgements
This work was supported by funding from the University of Siena (project number 2264-2020-BL-PSR2020_001) to LM, (project number 2264-2021-BL-PSR2021_001) to A.C. and L.M.
Funding
Open access funding provided by University of Siena within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
C.C. and L.M. planned the research; L.M. supervised the work; C.C., L.M., and R.F. conducted the experiments, D.M. performed SEM microscopy analysis; C.C. L.M., and R.F. analysed the results; A.C. and F.N. provided advice in manuscript compilation; L.M. prepared the original draft. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 145 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cucini, C., Funari, R., Mercati, D. et al. Polystyrene shaping effect on the enriched bacterial community from the plastic-eating Alphitobius diaperinus (Insecta: Coleoptera). Symbiosis 86, 305–313 (2022). https://doi.org/10.1007/s13199-022-00847-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-022-00847-y