Abstract
Corydoras is the richest genus of Corydoradinae, and many of its species have not been identified to date. We characterized Corydoras carlae and Corydoras sp. by performing cytogenetic, morphometric, and molecular analyses to facilitate correct identification and species delimitation and contribute an understanding of the evolutionary process of this group of fish. Individuals of C. carlae were collected in the Florido River, a tributary of the Iguassu River upstream of Iguassu Falls, and individuals of Corydoras sp. were collected in the Poço Preto River, a tributary of the Iguassu River downstream of Iguassu Falls. Corydoras sp. presented an extra rDNA 5S marker in an interstitial position on the short arm of one of the chromosomes of the submetacentric pair 15. Mitochondrial (COI) and nuclear (RAG1) sequences were efficient in discriminating C. carlae and Corydoras sp. Both species had exclusive haplotypes, which suggests the absence of gene flow between species. Furthermore, species delimitation analysis (GMYC and ABGD) suggested two MOTUs for Corydoras specimens from the Iguassu River. Differences in morphometric proportions were also observed. Considering the data gathered in this study, C. carlae and Corydoras sp. comprise distinct evolutionary lineages that are probably undergoing a recent speciation process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Corydoras is the richest genus of Corydoradinae and, consequently, of Callichthyidae and of Siluriformes, currently comprising 175 valid species (Tencatt et al., 2019; Lima & Britto, 2020; Fricke et al., 2021) distributed in the main rivers of South America. Although comprehensive studies of group systematics have published (e.g., Alexandrou et al., 2011; Britto, 2003; Eigenmann & Eigenmann, 1890; Ellis, 1913; Gosline, 1940; Nijssen, 1970; Nijssen & Isbrücker, 1967, 1980, 1983, 1986), knowledge about the taxonomy of many species and their phylogenetic relationships remains incipient (Tencatt & Ohara, 2016).
An unpublished taxonomic review of Corydoras paleatus (Jenyns, 1842) revealed the presence of four new species previously attributed to C. paleatus: Corydoras sp. A, Corydoras sp. B, Corydoras sp. C, and Corydoras sp. D (Tencatt, 2013). Subsequently, Tencatt et al. (2016), also in a review of C. paleatus, attributed to C. longipinnis, although not explicitly, the species previously identified as Corydoras sp. A, while Corydoras sp. C was described as C. froehlichi. Corydoras sp. B and Corydoras sp. D were not included by the authors in this review. For Corydoras sp. D, the scarcity of biological material (only five individuals) was the main limiting factor for its formal description, since the authors were able to clearly delimit this species morphologically (see Tencatt, 2013). In contrast, Corydoras sp. B had abundant material but could not be clearly diagnosed from its sympatric Corydoras carlae Nijssen & Isbrücker, 1983 (Tencatt personal observation). In fact, C. carlae and Corydoras sp. B present similar color and morphology patterns, having been diagnosed by Tencatt (2013) only by differences in the size of their dorsal and pectoral spines (dorsal spine 13.7–22.5% in standard length; pectoral spine 15.1–22.4% in standard length in Corydoras sp. B vs. 26.6–33.6 and 25.9–31.9 in C. carlae). Both species are apparently restricted to the Iguassu River basin (Ingenito et al., 2004; Rocha et al., 2016). Corydoras carlae was recorded upstream from Iguassu Falls, while Corydoras sp. B (hereinafter referred to as Corydoras sp.) was found only in the Poço Preto Stream, a tributary of the lower Iguassu River, downstream of Iguassu Falls and located in Iguassu National Park.
To contribute to knowledge regarding the diversity of species of Corydoras, different methods can be used for the correct delimitation of species. Most cytogenetic studies in Corydoras are restricted to conventional analysis and have demonstrated the existence of different diploid numbers, which can vary from 2n = 40 chromosomes in C. nattereri to 2n = 134 chromosomes in C. aeneus (Oliveira et al., 1990, 1992, 1993). These results suggest an intense polyploidy process in the diversification and evolutionary history of Corydoras (Oliveira et al., 1988, 1993; Turner et al., 1992). The distribution pattern of heterochromatin, as well as the location and quantity of chromosomes carrying Ag-NORs, is highly variable cytogenetic characteristics in the genus. However, little is known about the locations of the different types of rDNA. Thus, solving this gap in genetic knowledge is essential to better understand the relationships between species of Corydoras (Almeida et al., 2013; Artoni et al., 2006; Pazza et al., 2005).
Morphometry, similar to cytogenetics, is a tool that can also help elucidate systematic relationships within a group, providing precise interpretation and comparison of the variation patterns of quantitative characters (Blackith & Reyment, 1971; Cavalcanti & Lopes, 1990). This technique has been used to highlight differences in body shape in relation to fish size, which allows relationships between individuals to be detected and interpreted (Bemvenuti & Rodrigues, 2002; Shibatta & Hoffman, 2005; Almeida et al., 2012). In this context, fish populations isolated in headwater streams may present morphological divergences as a result of a change in gene frequency, leading to speciation through reproductive incompatibility (Castro, 1999).
Molecular techniques have strengthened the study of fish systematics in recent years, including Corydoras and other Callichthyidae genera (e.g., Shimabukuro-Dias et al., 2004). The development of molecular tools and methods for delimiting species makes it possible to more precisely estimate the existing biodiversity (Camargo & Sites, 2013; Pinacho-Pinacho et al., 2018). Thus, the use of DNA nucleotide sequences can help in the correct identification of species, especially in cases where real biodiversity cannot be detected by traditional taxonomy and systematic methods based on morphology (Bickford et al., 2007; Larson et al., 2016). In addition, the genetic characterization of C. carlae and Corydoras sp. can reveal important information about genus diversity. Analyses of different molecular markers of mitochondrial and nuclear DNA with different evolutionary rates are important for obtaining a better understanding of the evolutionary process in neotropical fishes (Fabrin et al., 2014).
Thus, in view of the morphological complexity of Corydoras fish species, the present study attempted to characterize Corydoras carlae and Corydoras sp. of the lower Iguassu River basin through cytogenetic, morphometric, and molecular analyses to help in the correct identification and delimitation of species in addition to contributing to an understanding of the evolutionary process of this group of fish.
Methods
Study area and sampling
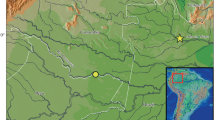
Seventeen Corydoras carlae specimens (Fig. 1a) were collected from the Florido River (26°00′04″S; 53°37′32″W; Fig. 2), a tributary of the Capanema River, which flows into the Iguassu River upstream of Iguassu Falls. In addition, 24 Corydoras sp. specimens (Fig. 1b) were collected at two sites (site 1: 25°36′45.5″S; 54°25′50.7″W and site 2: 25°37′19.7″S; 54°26′53.1″W; Fig. 2) in the Poço Preto Stream, a tributary that flows into the Iguassu River downstream from Iguassu Falls. Voucher specimens were deposited in the fish collection of the Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura (NUPELIA), Universidade Estadual de Maringá, Paraná, Brazil, as C. carlae (NUP 17885) and Corydoras sp. (NUP 14261 and NUP 17887).
This study was carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals, approved by the Committee on the Ethics of Animal Experiments of the Universidade Estadual do Oeste do Paraná (License Number: Protocol 13/09 – CEUA/Unioeste). Fish were collected with permission of IAP (Instituto Ambiental do Paraná, permit n° 43966/2015). Before the evisceration process, the individuals were euthanized by an overdose of clove oil (Griffiths, 2000).
Cytogenetic analysis
Cytogenetic analyses were performed on 10 individuals of C. carlae (four females and six males) and 15 individuals of Corydoras sp. (eight females and seven males). To obtain metaphasic chromosomes from cells extracted from the kidney, the methodology described by Bertollo et al. (1978) was used. The nucleolar organizer regions (NORs) were detected by means of silver nitrate staining (Ag-NORs), according to Howell and Black (1980), and analysis of C-positive heterochromatin (C-bands) followed the basic procedure of Sumner (1972), with some minor adaptations (Lui et al., 2012). Physical mapping of 5S rDNA and 18S rDNA was carried out by fluorescence in situ hybridization (FISH) according to Pinkel et al. (1986) and modifications suggested by Margarido and Moreira-Filho (2008) using DNA probes obtained from Megaleporinus obtusidens (cited as Leporinus elongates by Martins & Galetti, 1999) and Prochilodus argenteus (Hatanaka & Galetti, 2004), respectively. Probes were labeled by the nick translation method with digoxigenin-11-dUTP (5S rDNA) and biotin-16-dUTP (18S rDNA) (Roche®). Detection of signals was performed with antidigoxigenin-rhodamine (Roche®) as a probe for 5S rDNA and amplified avidin-FITC with biotinylated anti-avidin (Sigma-Aldrich) as a probe for 18S rDNA, with the chromosomes counterstained with 4′,6-diamidino-2-phenylindole (DAPI, 50 μg/mL).
The slides were analyzed under an optical microscope, and chromosomal counts and more detailed observations of the metaphases were made with a 100 × objective. The best metaphases were captured with a DP 71 digital camera coupled to the BX 61 epifluorescence microscope using DP Controller software, version 3.2.1.276. After capturing the images, the chromosomes were classified as metacentric (m), submetacentric (sm), and subtelocentric (st) according to their arm ratio (Levan et al., 1964). For determination of the fundamental number (FN) or number of chromosome arms, the m, sm, and st chromosomes were considered to bear two arms, and the acrocentric chromosomes were considered to bear only one arm.
Morphometric analysis
Morphometric character measurements of 17 individuals of C. carlae and 24 individuals of Corydoras sp. were performed using a digital caliper. The measurements were obtained according to Reis (1998) with some additions, which are all listed hereafter: standard length, thorax length, abdomen length, body height at the origin of the dorsal fin, predorsal distance, prepelvic distance, preanal distance, preadipose distance, dorsal-fin spine length, pectoral-fin spine length, caudal peduncle height, adipose-fin spine length, distance between the end of the base of the dorsal fin and the origin of the spine of the adipose fin, dorsal-fin base length, anal-fin base length, maximum width of the cleiter, head length, maxillary barbell length, head height, interorbital distance, horizontal orbit diameter, snout length, and internareal distance. To navigate the effect of the size of the measured specimens, proportions were calculated using the measurement of each variable (mm) in relation to the standard length (for measurements referring to the post cephalic portion of the body) and in relation to the head length (measurements referring to the head).
To summarize the matrix of morphometric variables, we applied a principal component analysis (PCA) using PC-ORD 5.0 software (McCune & Mefford, 2007). To determine which principal components would be retained for interpretation, we used the broken-stick model as the criterion (Jackson, 1993). To test the null hypothesis that neither species showed morphological differences, PERMANOVA multivariate permutational variance analysis was used with the Bray–Curtis index obtained with 999 random permutations (Anderson, 2001). The level of significance adopted was p < 0.05.
Molecular analysis
The DNA of 22 Corydoras specimens was isolated using the Wizard® Genomic DNA Purification Kit (Promega), stored at − 20 °C, and quantified by electrophoresis on a 1% agarose gel compared to a 100 bp ladder molecular standard (Ludwig). Mitochondrial and nuclear DNA fragments were amplified via polymerase chain reaction (PCR). A partial region of the mitochondrial cytochrome c oxidase subunit I (COI) gene of approximately 700 base pairs (bp) was amplified with the primers Fish_F1 (5′ – TCA ACC AAC CAC AAA GAC ATT GGC AC – 3′) and Fish_R1 (5′ – TAG ACT TCT GGG TGG CCA AAG AAT CA – 3′) (Ward et al., 2005). For amplification of the nuclear gene recombination activating protein 1 (RAG1), the primer pair RAG1 F (5′ – AAG GAG AGG GGT ATA GAT GAT A – 3′) and RAG1 R (5′ – GCA AAA CGC TGA GAG TTG AA – 3′) (Alexandrou et al., 2011) was used, which resulted in a fragment of approximately 1,000 bp.
Mitochondrial and nuclear fragments were amplified in independent PCRs. The reaction mixture consisted of Tris-KCl buffer (20 mM Tris–HCl, pH 8.4, 50 mM KCl), 1.5 to 2 mM MgCl2, 0.6 µM of each primer, 0.4 mM of each dNTP, 3 U of Taq DNA polymerase (Invitrogen), 25 ng of DNA and filtered/deionized water (Milli-Q) for a final volume of 25 µL. The amplifications of the COI gene occurred in a thermocycler programmed for the following thermal profile: an initial cycle of 4 min at 94 °C, followed by 35 cycles of 1 min at 94 °C, 45 s at 58 °C, and 1 min at 72 °C, with an additional last step of 5 min at 72 °C. For amplification of the RAG1 fragment, an initial cycle of 4 min at 94 °C was used, followed by 40 cycles of 1 min at 94 °C, 45 s at 59 °C and 90 s at 72 °C, in addition to a last step of 7 min at 72 °C. The amplification efficiency was confirmed by electrophoresis on a 1% agarose gel and staining with SYBR Safe (Invitrogen).
Subsequently, the amplified samples were purified with a PureLink® PCR Purification Kit (Invitrogen). After purification, COI and RAG1 fragments were used in sequencing reactions, with the primers Fish_F1 and RAG1 F, respectively, using the BigDye® Terminator v3.1 Cycle Sequencing kit, in automatic sequencer ABI 3500 DNA Analyzer (Life Technologies), according to manufacturer’s instructions.
The nucleotide sequences obtained were edited using BioEdit software (Hall, 1999) and aligned with Clustal Omega software (Sievers et al., 2011). The genetic distance and the frequencies of nucleotide bases were estimated using MEGA X software (Kumar et al., 2018). COI and RAG1 sequences from other species of Corydoras available in BOLD Systems and GenBank were included in the analyses; Aspidoras sp. was used as an outgroup (Table S1).
The selection of the best-fit model of nucleotide evolution and the partitions were conducted using PartitionFinder 2.1 software (Lanfear et al., 2012). Maximum likelihood trees were reconstructed with raxmlGUI software (Silvestro & Michalak, 2012) using the partitions established by PartitionFinder (1st, 2nd, and 3rd bases, for COI; and 1st and 2nd codon, and 3rd codon RAG1) and the GTR + G model. A rapid bootstrap algorithm and autoMRE function for resamplings were implemented.
Bayesian ultrametric tree reconstructions were performed with BEAST 1.8.2 software with an input file generated in BEAUti 1.8.0 (Drummond et al., 2012); the birth–death process of speciation was used as a tree prior, and a strict molecular clock was used for both COI and RAG1. The COI region was partitioned according to codon bases (1st, 2nd, and 3rd bases) using the TN93 substitution model, whereas RAG1 was partitioned according to codon bases (1st and 2nd bases and 3rd bases) using the HKY model. Analysis ran for 20,000,000 (for COI) and 10,000,000 (for RAG1) generations with a sample frequency of 1,000. The final trees were calculated after 20% burn-in. The length of burn-in was determined by examining traces in Tracer 1.6 (Rambaut et al., 2014), considering > 200 as an appropriate effective sample size (ESS) value. Support for nodes was determined using posterior probabilities.
To estimate the time of divergence between C. carlae and Corydoras sp., an outgroup-rooted phylogenetic tree was built based on the sequences of the COI gene, and with the assumption of a calibrated molecular clock, which admits a constant mutation rate over time. The assumed calibrated molecular clock attributed an uncorrected mutation rate of 1.2% per million years (Mya) to the COI gene. This is an average mutation rate since geological and fossil data show a mutation rate in fish ranging from 1.1 to 1.3% per Mya (Bermingham et al., 1997; Near et al., 2003). For the construction of the tree, the same procedure described above was used for the ultrametric tree. Inferences of clade ages are presented as 95% highest posterior density (HPD).
Additionally, specifically for the COI gene, to identify molecular operational taxonomic units (MOTUs) (Hebert et al., 2003), methods for species delimitation were implemented to identify the specific boundaries in C. carlae and Corydoras sp. The Bayesian ultrametric tree was used for the general mixed Yule coalescent method (GMYC; Pons et al., 2006) using R Studio software (R Development Core Team, 2020) and the splits package (Fujisawa & Barraclough, 2013). The maximum likelihood gene tree was used for the Poisson tree process model (PTP; Zhang et al., 2013) delimitation test, which was performed online (http://species.h-its.org). The ABGD method was conducted on the online server http://wwwabi.snv.jussieu.fr/public/abgd using the default parameters and Kimura (K80) model of nucleotide substitution.
Results
Cytogenetic data
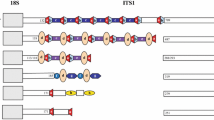
The diploid number, karyotype formula, and FN were the same for Corydoras carlae and Corydoras sp., with 2n = 46 chromosomes, composed of 22 metacentric chromosomes, 22 submetacentric chromosomes, and two subtelocentric chromosomes for both sexes and FN equal to 92 (Fig. 3a-d). Silver nitrate impregnation identified Ag-NOR in a terminal position in the long arm of metacentric pair six in the two species (Box Fig. 3a, d). FISH with 18S rDNA coincided with the marking of silver nitrate in C. carlae and Corydoras sp., featuring a simple NOR system (Fig. 3c, f). In addition, 5S rDNA cistrons colocalized with 18S rDNA were observed for C. carlae and Corydoras sp. (Fig. 3c, f and Fig. 4). However, Corydoras sp. presented an extra marker of 5S rDNA located in an pericentromeric position on the short arm of one of the chromosomes of submetacentric pair 15 (Fig. 3f).
Heterochromatins were observed in the centromeric/pericentromeric region of most of chromosomes of the complement, in addition to being associated with NORs in both species (Fig. 3b, e).
Morphometric data
The proportions analysis calculated from the morphometric measurements of C. carlae and Corydoras sp. demonstrated differences in body height/standard length, average of 29.1 mm in C. carlae and 34.2 mm in Corydoras sp., interorbital distance/head length, average of 52.6 mm in C. carlae and 39.3 mm in Corydoras sp. and the horizontal diameter of the orbit/head length, average of 30.0 mm in C. carlae and 23.4 mm in Corydoras sp. (Table 1).
The first two axes of the principal component analysis showed eigenvalues greater than the eigenvalues of the broken-stick and were retained for interpretation. These two axes showed an accumulated explained variance of 35% (Table 2). The results of PERMANOVA indicated that C. carlae and Corydoras sp. showed significant differences in morphology (Pseudo-F = 1.98; p < 0.01). Thus, Corydoras sp. have a body height in relation to the standard length greater than that found for C. carlae. On the other hand, the interorbital distance and the horizontal diameter of the orbit in relation to the head length are greater in C. carlae than in Corydoras sp. (Fig. 5).
Principal component analysis scores (mean and maximum and minimum values) for the morphometric data of Corydoras carlae samples in the Florido River, and Corydoras sp. sampled in the Poço Preto Stream, lower Iguassu River basin. Proportions that showed a significant difference between species: (a) body height/standard length; (b) interorbital distance/head length; (c) horizontal diameter of the orbit/head length
Molecular data
A total of 76 sequences of the Corydoras mitochondrial COI gene were used in this study, including 22 sequences of 634 bp (base pair) of specimens collected in the Iguassu River basin (C. carlae and Corydoras sp.) and 54 COI sequences of Corydoras species obtained from GenBank (Table S1). For the nuclear RAG1 gene, 51 nucleotide sequences were obtained, 20 sequences of 839 bp referring to the Iguassu River specimens, and 31 Corydoras RAG1 sequences obtained from BOLD and GenBank (Table S1). All sequences generated in this study were deposited in GenBank (GenBank accession numbers = MT846090—MT846111 for COI sequences; MT855475—MT855494 for RAG1 sequences). The nucleotide composition of the COI fragment for specimens of Corydoras sp. and C. carlae was 27.7% (T), 28.8% (C), 26.2% (A), and 17.3% (G), while for RAG1, it was 25.1% (T), 20.8% (C), 27.9% (A), and 26.2% (G). Polymorphic and species-specific nucleotide sites are described in Table S2. A single mitochondrial haplotype COI was identified among the specimens of C. carlae, and there were two haplotypes among Corydoras sp. For the RAG1 fragment, six haplotypes were observed in C. carlae and three in Corydoras sp. No shared haplotypes were observed between C. carlae and Corydoras sp. In some regions of the nuclear sequence of RAG1, mainly in specimens of C. carlae, sites were observed in heterozygous states, which were identified as double strong peaks of the same height or very close heights seen in the chromatograms.
Mitochondrial (COI) and nuclear (RAG1) nucleotide sequences were efficient in discriminating Corydoras species, revealing high values of posterior probabilities or bootstraps supporting the clades in both dendrograms (Figs. 6 and 7). Corydoras carlae and Corydoras sp. from the Iguassu River basin were grouped into two distinct clades in both analyses. The clustering of the specimens was performed according to their morphological identification. Some species of Corydoras, such as C. nattereri, C. paleatus, and C. aeneus, according to data regarding the COI sequences, and C. diphyes and C. ehrhardti, for RAG1, formed more than one clade, and some were even nonmonophyletic. One of the clades formed by specimens of C. diphyes was allocated within the larger clade constituted by the species of Corydoras from the Iguassu River, suggesting nonmonophyletic conditions for C. carlae and Corydoras sp.
Calibrated Bayesian tree based on the cytochrome c oxidase I gene sequence of Corydoras species. Values in parentheses indicate sample number for each species. Values near branches indicate Bayesian (posterior probability; above) and maximum likelihood (bootstrap; below) support values for each node. The dashed lines indicate the delimitations of molecular operational taxonomic units (MOTUs) according to the GMYC, PTP, and ABGD approaches
Bayesian phylogenetic tree based on the recombination activating protein 1 (RAG1) gene of Corydoras species. Values in parentheses indicate the sample number for each species. Values near branches indicate Bayesian (posterior probability; above) and maximum likelihood (bootstrap; below) support values for each node
According to the results of divergence time estimation, the approximate origin of the species of Corydoras evaluated here was 22.1 Mya (95% HPD 26.1-18.3 Mya) (Fig. 6). The clade formed by Corydoras sp., C. carlae, C. nattereri, C. paleatus, C. ehrhardti, and C. sterbai was estimated to have originated 8.7 Mya (95% HPD 10.8-6.7), in the early Miocene. The event that originated the clade of Corydoras sp. and C. carlae occurred more recently; in the Pleistocene, 1.1 Mya (95% HPD 1.7-0.5 Mya). The average values of genetic distance ranged from 3.8% (between C. nattereri and C. ehrhardti) to 18.4% (between C. sterbai and C. flaveolus) for COI sequences and from 0.7% (between C. ehrhardti and C. natttereri) to 4.3% (between C. difluviatilis and C. sterbai) for RAG1. The average genetic distance between C. carlae and Corydoras sp. was 1.1% for COI and 0.5% for RAG1 (Table 3).
The ultrametric Bayesian tree was subjected to the GMYC delimitation method, and 19 MOTUs were obtained. Using the PTP method, based on the maximum likelihood tree, 17 MOTUs were delimited, while according to the ABGD method, 16 MOTUs were defined (Fig. 6). The difference in the number of MOTUs delimited by the three methods was related to Corydoras sp., C. carlae, C. paleatus, C. ehrhardti, and C. aeneus. Regarding the species in the Iguassu River basin, two MOTUs were delimited by the GMYC and ABGD methods, with each MOTU referring to a species (Corydoras sp. and C. carlae), while a single MOTU was defined by the PTP method (Fig. 6).
Discussion
Cytogenetics
Our cytogenetic analyses revealed that the two Corydoras species studied shared the same diploid number (2n = 46) and karyotype formula, including both species within group 4 (2n = 40–52 chromosomes, with many metacentric and submetacentric chromosomes) according to a classification based on molecular data and variation in the diploid number (Oliveira et al., 1992). Considering our results, Corydoras sp. (2n = 46, 22 m + 22sm + 2st) is the third karyotyped species of this group occurring in the Iguassu River basin: C. carlae (2n = 46, 22 m + 22sm + 2st) collected in the Lower Iguassu River (Rocha et al., 2016), C. paleatus (2n = 44, 20 m + 24sm) collected in the Upper Iguassu River (Oliveira et al., 1993), and C. aff. paleatus (2n = 44, 18 m + 26sm) collected in the Upper Iguassu River (Barbosa et al., 2017) were analyzed early. Thus, the similar karyotypic macrostructure between C. carlae and Corydoras sp. probably indicates that these species are undergoing a recent speciation process.
The studied specimens of C. carlae and Corydoras sp. had only one Ag-NOR situated on the long arm of the sixth-largest metacentric pair. Ag-NORs were also evident in a single metacentric pair in a terminal position of the long arm in C. ehrhardti and C. paleatus (Artoni et al., 2006). According to Oliveira and Gosztonyi (2000), the presence of simple Ag-NORs in the terminal location is a possible basal condition for Siluriformes. Thus, C. carlae and Corydoras sp., presenting simple Ag-NORs in the terminal location, seem to maintain this basal condition.
FISH analyses revealed that the two Corydoras species studied shared the same location and number of 18S sites, with terminal markings in two metacentric chromosomes and syntenic marks with 5S ribosomal sites. The occurrence of one chromosome pair bearing 18S rDNA in C. carlae and Corydoras sp. is similar to what has been found in C. ehrhardti (Artoni et al., 2006; Barbosa et al., 2017) and C. aff. paleatus (Barbosa et al., 2017). However, only the 18S ribosomal sites of C. ehrhardti were syntenic with 5S, albeit in independent clusters, while in C. carlae and Corydoras sp., the 5S rDNA was interspersed along with the clusters of 18S rDNA (colocalization). Synteny is a rare trait in fish and is recorded here for the first time in Corydoras sp., highlighting the originality of these results. Syntenic 18S and 5S markings have already been described for Callichthyidae, Callichthys callichthys (Konerat et al., 2014), C. carlae (Rocha et al., 2016), and C. ehrhardti (Barbosa et al., 2017). Thus, our results expand the synteny information for the 18S and 5S ribosomal sites of the family. The colocalization of 18S and 5S has also been described for other groups of fish, such as Mugil incilis (Hett et al., 2011), Psalidodon fasciatus, P. scabripinnis (Almeida-Toledo et al., 2002), and Salea senegalensis (Cross et al., 2006).
In contrast, 5S rDNA is a chromosomal marker that is specific to the species analyzed in this study. The localization of the 5S rDNA sites is divergent among C. carlae and Corydoras sp. In Corydoras sp., the in situ analysis of the 5S rDNA sequences revealed signals on three chromosomes, while only one chromosome pair bearing 5S rDNA was present in C. carlae. The syntenic marking of the 18S and 5S rDNA in pair 6 was shared between the two species studied. However, in Corydoras sp. additional 5S rDNA cistrons located in pericentromeric position on the short arm of one homolog of pair 15 were detected. From an evolutionary point of view, these data are intriguing because the insertion of transposable elements in sequences of the 5S rDNA of the metacentric pair could have led to the dispersion of these sequences to the submetacentric chromosome (par 15) of Corydoras sp. According to Raskina et al. (2004), one of the mechanisms responsible for the process of moving rDNA sequences to new sites would be due to the action of transposable elements. The action of transposable elements was suggested to justify the difference in the number and location of 5S rDNA cistrons in three species of Bryconamericus (Piscor et al., 2013) and appears to be responsible for the dispersion of 5S rDNA in almost all chromosomes of Hyphessobrycon eques (Piscor et al., 2020).
Corydoras species share a heterochromatin distribution pattern that is very similar, preferably centromeric and pericentomeric, and in most cases is associated with NORs. In C. carlae and Corydoras sp., this pattern was also observed, with centromeric and pericentromeric heterochromatic blocks displayed on many chromosomes. Thus, the heterochromatin distribution pattern was not an effective marker in the delimitation of the Corydoras species analyzed here. Corydoras britskii from the Miranda River also showed a large amount of pericentromeric heterochromatin, but with terminal heterochromatic blocks (Takagui et al., 2014), which were not observed in this study.
Morphometry
Our results revealed differences between species in morphometric proportions, especially for body height/standard length, interorbital distance/head length, and the horizontal diameter of the orbit/head length. On the other hand, Tencatt (2013) morphologically compared Corydoras sp. with C. carlae collected in the Tormenta, Adelaide, and Guarani rivers and found that these species can be differentiated by differences in the lengths of their dorsal and pectoral clusters (dorsal cluster 13.7–22.5% in the SL; pectoral cluster 15.1–22.4% in the SL in Corydoras sp. vs. 26.6–33.6 and 25.9–31.90 in C. carlae).
The differences found make it possible to speculate that these are different species, possibly because both have different geographic distributions. Corydoras carlae were collected in the Florido River upstream of Iguassu Falls, while Corydoras sp. were captured only in the Poço Preto Stream, downstream of Iguassu Falls. In this way, the two species analyzed are separated by a natural geographic barrier (Iguassu Falls) formed approximately 22 million years ago, which is considered to be one of the main causes of isolation and allopatric speciation of fish species for the Iguassu River basin (Agostinho & Gomes, 1997; Baumgartner et al., 2012; Garavello et al., 1997; Mezzaroba et al., 2021). However, populations apparently corresponding to C. carlae were recorded in the Río Urugua-í basin, a tributary of the lower Paraná River in Misiones, Argentina, and thus, considering that they are in fact fragmented populations (probably relictual) of the same species, C. carlae would no longer be restricted to tributaries of the Iguassu River upstream of Iguassu Falls.
In a study carried out with C. paleatus from different basins, including the Iguassu River (Shibatta & Hoffmann, 2005), it was proposed that the differences found between the populations occurred due to the uplift of Iguassu Falls, which separated the species from the Paraná River and the Iguassu River. On the other hand, Florentino and Súarez (2014) attributed and correlated the differences between populations of C. aeneus to the characteristics of the environment that, over evolutionary time, selected the individuals with the greatest adaptation. According to the evidence presented by Tencatt et al. (2016), these populations, although often separated by well-defined geographical barriers, apparently correspond to a single and variable species, C. longipinuunis.
Molecular analysis
Molecular analyses revealed a separation between C. carlae and Corydoras sp. (Figs. 5 and 6), presented exclusive haplotypes (or groups of haplotypes), allowed correct discrimination of species and provided evidence for the nonsharing of haplotypes, which also suggested the absence of gene flow. In addition, two of the three species delimitation methods (GMYC and ABGD) assigned two different MOTUs to Corydoras species from the Iguassu River. Although these data show a clear separation of the two species, the average value of genetic distance was not sufficient for their discrimination, according to the threshold stipulated by the DNA barcoding methodology.
DNA barcoding is one of the most commonly used tools today in the identification of species based on DNA sequences (Hebert et al., 2003). The methodology is based on a standardized region of the mitochondrial cytochrome c oxidase I (COI) gene for the identification of animal species based on differences in their COI sequences (Hebert et al., 2003). A threshold value of 2% divergence is normally used in the delimitation of species (Carvalho et al., 2011; Pereira et al., 2011; Ward, 2009; Ward et al., 2009). Although the method is highly efficient in identifying a large number of animal species, including fish (e.g., Carvalho et al., 2011; Hubert et al., 2008; Pereira et al., 2011, 2013), criticisms have been made regarding the use of a single gene to delimit species and, mainly, regarding the established cutoff value. Most likely, the 2% COI divergence threshold is not suitable for all groups of fish, especially for some pairs of species that may naturally have low interspecific values, as is the case with fish from the Neotropical region (Pereira et al., 2013).
Although C. carlae and Corydoras sp. have an average value of interspecific distance K2P (1.13%; Table 3) below the barcode threshold of 2%, the species formed cohesive groups of haplotypes and presented diagnostic nucleotides (i.e., species-specific; Table S2) (Wong et al., 2009), which allowed the correct identification of species based not only on COI sequences but also on RAG1. In addition, even though the average interspecific K2P genetic distance values were low (1.13% for COI; 0.49% for RAG1), the intraspecific average values were comparatively many times smaller (0–0.02% for COI; 0–0.01% for RAG1). Furthermore, the nonmonophyletic conditions observed for C. carlae and Corydoras sp. (data based on RAG1 sequences; Fig. 6) reinforce the hypothesis that they are different species.
Similarly, Pereira et al. (2013) found several pairs of neotropical fish species with low genetic distance values for the COI gene (< 2%). However, correct delimitation of the species was possible due to the formation of cohesive groups of haplotypes, as well as the occurrence of diagnostic nucleotides. Maia (2014), in a study aimed at molecular identification of fish specifically from Corydoradinae, obtained the correct discrimination of 85% of 94 species analyzed using DNA barcoding methodology. However, seven pairs of species had a genetic distance value of less than 2% (Maia, 2014), reinforcing the hypothesis that some groups of fish have low genetic distance values. In this study, all pairs of Corydoras species analyzed had genetic distance values above the 2% threshold for the barcode sequences (Table 3). The only exception was the distance between C. carlae and Corydoras sp.
A possible explanation for the low values of genetic distance found may be related to recent radiation. Based on previous studies, most of the diversification of Neotropical ichthyofauna occurred recently, 10-3 Mya (Hubert et al., 2007; Lovejoy & Araújo, 2000; Montoya-Burgos, 2003; Pereira et al., 2013). According to the estimated divergence time obtained, C. carlae and Corydoras sp. started to differentiate approximately 1.1 Mya. The recently estimated origin for C. carlae and Corydoras sp. is consistent with the low genetic distance values identified between species. These species are probably undergoing a recent speciation process, which has prevented further accumulation of polymorphisms in the DNA. According to Queiroz (2007), the speciation process is not uniform, and depending on the character evaluated, it is possible not to reach a precise conclusion regarding the existence of one or more species. This moment of uncertainty during speciation is called the gray zone (Queiroz, 2007), and C. carlae and Corydoras sp. may be going through this period. According to the results obtained so far, C. carlae and Corydoras sp. appear to correspond to independent evolutionary lineages.
The fact that C. carlae and Corydoras sp. do not occur together must have contributed to establishing the morphological, cytogenetic, and genetic differences found between them, since the lack of gene flow between species supposedly resulted in a process of speciation. The differences found between the two species of Corydoras analyzed are probably due to the geographic isolation caused by the uplift of Iguassu Falls. In addition to this notable geographical barrier, others occur along the Iguassu River, such as Salto Saicanga, Salto Grande, Salto Santiago, Salto Osório, and Salto Caxias (Maack, 2012), almost all flooded by the formation of reservoirs (Baumgartner et al., 2012). However, the differentiation of the species must have occurred after the uplift of Iguassu Falls, since low values of interspecific genetic distance and recent divergence times were detected (1.1 Mya). Geographic isolation is also suggested to explain cytogenetic differences, which is reflected in the diploid number and banding patterns of fish species such as Characidium in the Paraná River basin (Pucci et al., 2014), Psalidodon scabripinnis (= Astyanax scabripinnis) (Moreira-Filho & Bertollo, 1991), and Astyanax lacustris (= Astyanax altiparanae) (Hashimoto et al., 2008).
In this context, the evolution of fish species confined to different hydrographic systems is the result of a close relationship between the histories of the basins and the evolutionary histories of their species (Kavalco & Moreira-Filho, 2003). An interesting feature is that small fish species tend to be more susceptible to speciation, since populations located in small streams can diverge genetically from the others more quickly than the typical species of large rivers (Weitzman et al., 1998). However, from a genetic perspective, speciation caused by reproductive isolation is a property of a few individual loci or genomic regions and not of the genome as a whole (Lexer & Widmer, 2008; Qvarnstron & Bailey, 2009).
Although C. carlae and Corydoras sp. have the same diploid number, karyotype formula, number of Ag-NORs, and the same pattern distribution of constitutive heterochromatin, differences in the number of chromosomes carrying 5S rDNA cistrons were observed between species. In addition, the synteny and colocalization of 5S rDNA with 18S rDNA represent unprecedented results for Corydoras. Nevertheless, analysis of the morphometric proportions also confirmed significant differences between the species. The combined analyses of cytogenetic, morphometric, and molecular results were important for characterization of the two species and made it possible to differentiate C. carlae from Corydoras sp. via an allopatric speciation process, indicating they are distinct cytogenetically, molecularly, and morphologically. Therefore, there is evidence that C. carlae and Corydoras sp. of the Iguassu River basin comprise distinct evolutionary lineages that are probably undergoing a recent process of speciation.
Availability of data and material
Data generated or analyzed during this study consist of 24 fish specimens. Voucher specimens were deposited in the fish collection of the Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura (NUPELIA), Universidade Estadual de Maringá, Paraná, Brazil, as Corydoras carlae (NUP 17885) and Corydoras sp. (NUP 14261 and NUP 17887). In addition, all nucleotide sequences generated in this study were deposited in GenBank (GenBank accession numbers = MT846090—MT846111 for COI sequences; MT855475—MT855494 for RAG1 sequences); all GenBank accession numbers used in our analysis are also listed individually in the Table S1.
Code availability
Not applicable.
References
Agostinho, A. A., & Gomes, L. C. (1997). Manejo e monitoramento de recursos pesqueiros: perspectivas para o reservatório de Segredo. In A. A. Agostinho, & L. C. Gomes (Eds.), Reservatório de Segredo: bases ecológicas para o manejo. Eduem. [In Portuguese]. 319–364.
Alexandrou, M. A., Oliveira, C., Maillard, M., Mcgill, R. A. R., Newton, J., Creer, S., & Taylor, M. I. (2011). Competition and phylogeny determine community structure in Müllerian co-mimics. Nature, 469, 84–89. https://doi.org/10.1038/nature09660
Almeida, J. S., Affonso, P. R. A. M., & Dias, A. L. (2012). Remarkable karyotypic homogeneity in a widespread tropical fish species: Hoplosternum littorale (Siluriformes, Callichthyidae). Journal of Fish Biology, 81(4), 1415–1421. https://doi.org/10.1111/j.1095-8649.2012.03387.x
Almeida, J. S., Affonso, P. R. A. M., Diniz, D., Carneiro, P. L. S., & Dias, A. L. (2013). Chromosomal variation in the tropical armoured catfish Callichthys callichthys (Siluriformes, Callichthyidae): Implications for conservation and taxonomy in a species complex from a Brazilian hotspot. Zebrafish, 10(4), 451–458. https://doi.org/10.1089/zeb.2013.0885
Almeida-Toledo, L. F., Ozouf-Costaz, C., Foresti, F., Bonillo, C., Porto-Foresti, F., & Daniel-Silva, M. F. Z. (2002). Conservation of the 5S-bearing chromosome pair and co-localization with major rDNA clusters in five species of Astyanax (Pisces, Characidae). Cytogenetic and Genome Research, 97(3–4), 229–233. https://doi.org/10.1159/000066609
Anderson, M. J. (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecology, 26, 32–46. https://doi.org/10.1111/j.1442-9993.2001.01070
Artoni, R. F., Terêncio, M. L., Vicari, M. R., Matiello, M. C. A., Cestari, M. M., & Bertollo, L. A. C. (2006). Cytogenetics of two sympatric Corydoras species (Pisces, Siluriformes, Callichtyidae) of Southern Brazil. Brazilian Journal of Biology, 66(1B), 191–198. https://doi.org/10.1590/S1519-69842006000200002
Barbosa, P., Pucci, M. B., Nogaroto, V., Almeida, M. C., Artoni, R. F., & Vicari, M. R. (2017). Karyotype analysis of three species of Corydoras (Siluriformes: Callichthyidae) from southern Brazil: Rearranged karyotypes and cytotaxonomy. Neotropical Ichthyology, 15(1), e160056. https://doi.org/10.1590/1982-0224-20160056
Baumgartner, G., Pavanelli, C. S., Baumgartner, D., Bifi, A. G., Debona, T., & Frana, V. A. (2012). Peixes do baixo rio Iguaçu. Eduem. [In Portuguese].
Bemvenuti, M. A., & Rodrigues, F. L. (2002). Análise comparativa entre técnicas morfométricas aplicadas a Odontesthes bonariensis (Valenciennes) e Odontesthes humensis De Buen (Osteichthyes, Atherinopsidae). Revista Brasileira De Zoologia, 19(3), 789–796. https://doi.org/10.1590/S0101-81752002000300017.[InPortuguese]
Bermingham, E., McCafferty, S. S., & Martin, A. P. (1997). Fish biogeography and molecular clocks: Perspectives from the Panamanian Isthmus. In T. D. Kocher & C. A. Stepien (Eds.), Molecular systematics of fishes (pp. 113–128). Academic Press.
Bertollo, L. A. C., Takahashi, C. S., & Moreira-Filho, O. (1978). Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Brazilian Journal of Genetics, 1, 103–120.
Bickford, D., Lohman, D. J., Sodhi, N. S., Ng, P. K. L., Meier, R., Winker, K., Ingram, K. K., & Das, I. (2007). Cryptic species as a window on diversity and conservation. Trends in Ecology and Evolution, 22(3), 148–155. https://doi.org/10.1016/j.tree.2006.11.004
Blackith, R. E., & Reyment, R. A. (1971). Multivariate morphometrics. Academic Press.
Britto, M. R. (2003). Phylogeny of the subfamily Corydoradinae Hoedeman, 1952 (Siluriformes: Callichthyidae), with a definition of its genera. Proceedings of the Academy of Natural Sciences of Philadelphia, 153(1), 119–154. https://doi.org/10.1635/0097-3157(2003)153[0119:POTSCH]2.0.CO;2
Camargo, A., & Sites, J. W. (2013). Species delimitation: A decade after the Renaissance. In I. Pavlinov (Ed.), The species problem-ongoing issues. IntechOpen. https://doi.org/10.5772/52664. from: https://www.intechopen.com/books/the-species-problem-ongoing-issues/species-delimitation-a-decade-after-the-renaissance.
Carvalho, D. C., Oliveira, D. A. A., Pompeu, P. S., Leal, C. G., Oliveira, C., & Hanner, R. (2011). Deep barcode divergence in Brazilian freshwater fishes: The case of the São Francisco River Basin. Mitochondrial DNA, 22(S1), 80–86. https://doi.org/10.3109/19401736.2011.588214
Castro, R. M. C. (1999). Evolução da ictiofauna de riachos sulamericanos: padrões gerais e possíveis processos causais. In E. P. Caramaschi, R. Mazzoni, & P. R. Peres-Neto (Eds.), Ecologia de Peixes de Riachos. PPGEUFRJ Série Oecologia Brasiliensis. [In Portuguese]. 139–155.
Cavalcanti, M. J., & Lopes, P. R. D. (1990). Morfometria comparada de Ctenosciaena gracilicirrhus, Paralonchurus brasiliensis e Micropogonias furnieri (Teleostei: Sciaenidae) pela análise multivariada de redes de treliças. Revista Brasileira De Zoologia, 7(4), 627–635. https://doi.org/10.1590/S0101-81751990000400016.[InPortuguese]
Cross, I., Merlo, A., Manchado, M., Infante, C., Cañavate, J. P., & Rebordinos, L. (2006). Cytogenetic characterization of the sole Solea senegalensis (Teleostei: Pleuronectiformes: Soleidae): Ag- NOR, (GATA)n, (TTAGGG)n and ribosomal genes by one-color and two-color FISH. Genetica, 128(1–3), 253–259. https://doi.org/10.1007/s10709-005-5928-9
Drummond, A. J., Suchard, M. A., Xie, D., & Rambaut, A. (2012). Bayesian phylogenetics with BEAUti and the BEAST 1.7. Molecular Biology and Evolution, 29(8), 1969–1973. https://doi.org/10.1093/molbev/mss075.
Eigenmann, C. H., & Eigenmann, R. S. (1890). A revision of the South American Nematognathi or cat-fishes. California Academy of Sciences.
Ellis, M. D. (1913). The plated nematognaths. Annals of the Carnegie Museum, 8, 384–413.
Fabrin, T. M. C., Simone, I., Prioli, S. M. A. P., Prioli, A. J., & Gasques, L. S. (2014). A utilização de marcadores na filogenia dos ciclídeos (Teleostei: Perciformes): Uma análise cienciométrica. Enciclopédia Biosfera, 10(18), 3118–3128. [In Portuguese].
Florentino, A. C., & Súarez, Y. R. (2014). Diferenciação Morfológica entre Populações de Corydoras aeneus Gill (1858) (Siluriformes, Callichthyidae) em riachos das bacias hidrográficas dos Rios Paraná e Paraguai. Biota Amazônia, 4(3), 95–99. https://doi.org/10.18561/2179-5746/biotaamazonia.v4n3p95-99. [In Portuguese].
Fricke, R., Eschmeyer, W. N. & van der Laan R. (Eds.) (2021). Eschmeyer’s Catalog of Fishes: Genera, Species, References. (http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp). Electronic version accessed 07 May 2020.
Fujisawa, T., & Barraclough, T. G. (2013). Delimiting species using singlelocus data and the generalized mixed yule coalescent approach: A revised method and evaluation on simulated data sets. Systematic Biology, 62(5), 707–724. https://doi.org/10.1093/sysbio/syt033
Garavello, J. C., Pavanelli, C. S., & Suzuki, H. I. (1997). Caracterização da ictiofauna do rio Iguaçu. In A. A. Agostinho & L. C. Gomes (Eds.), Reservatório de Segredo: Bases ecológicas para o manejo (pp. 61–84). Eduem.
Gosline, W. A. (1940). Rediscovery and redescription of Pariolius armillatus, a genus and species of pimelodid catfishes described by E. D. Cope from the Peruvian Amazon in 1872. Copeia, 2, 78–80. https://doi.org/10.2307/1439046
Griffiths, S. P. (2000). The use of clove oil as an anaesthetic and method for sampling intertidal rockpool fishes. Journal of Fish Biology, 57(6), 1453–1464. https://doi.org/10.1111/j.1095-8649.2000.tb02224.x
Hall, T. A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41(1), 95–98.
Hashimoto, T., Arion, D., Unger, T., Maldonado-Aviles, J. G., Morris, H. M., Volk, D. W., & Lewis, D. A. (2008). Alterations in GABA-related transcriptome in the dorsolateral prefrontal cortex of subjects with schizophrenia. Molecular Psychiatry, 13(2), 147–161. https://doi.org/10.1038/sj.mp.4002011
Hatanaka, T., & Galetti, P. M. (2004). Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus, Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica, 122, 239–244. https://doi.org/10.1007/s10709-004-2039-y
Hebert, P. D. N., Ratnasingham, S., & Waard, J. R. (2003). Barcoding animal life: Cytochrome c oxidase subunit 1 divergences among closely related species. Proceedings of the Royal Society London B Biological Sciences, 270(Suppl_1), S96–S99. https://doi.org/10.1098/rsbl.2003.0025.
Hett, A. S., Nirchio, M., Oliveira, C., Siccha, Z. R., Rossi, A. R., & Sola, L. (2011). Karyotype characterization of Mugil incilis Hancock, 1830 (Mugiliformes: Mugilidae), including a description of an unusual co-localization of major and minor ribosomal genes in the family. Neotropical Ichthyology, 9(1), 107–112. https://doi.org/10.1590/S1679-62252011005000005
Howell, W. M., & Black, D. A. (1980). Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: A 1-step method. Experientia, 36, 1014–1015. https://doi.org/10.1007/BF01953855
Hubert, N., Duponchelle, F., Nuñez, J., Garcia-Davila, C., Paugy, D., & Renno, J. F. (2007). Phylogeography of the piranha genera Serrasalmus and Pygocentrus: Implications for the diversification of the Neotropical ichthyofauna. Molecular Ecology, 16(10), 2115–2136. https://doi.org/10.1111/j.1365-294X.2007.03267.x
Hubert, N., Hanner, R., Holm, E., Mandrak, N. E., Taylor, E., Burridge, M., Watkinson, D., Dumont, P., Curry, A., Bentzen, P., Zhang, J., April, J., & Bernatchez, L. (2008). Identifying Canadian freshwater fishes through DNA barcodes. PLoS ONE, 3(6), e2490. https://doi.org/10.1371/journal.pone.0002490
Ingenito, L. F. S., Duboc, L. F., & Abilhoa, V. (2004). Contribuição ao conhecimento da ictiofauna da bacia do alto Rio Iguaçu, Paraná, Brasil. Arquivos De Ciências Veterinárias e Zoologia Da UNIPAR, 7(1), 23–36. [In Portuguese].
Jackson, D. A. (1993). Stopping rules in principal components analyses: A comparison of heuristical and statistical approaches. Ecology, 74(8), 2204–2214. https://doi.org/10.2307/1939574
Jenyns, L. (1842). Part IV. Fish. In C. R. Darwin (Ed.), The zoology of the voyage of H. M. S. Beagle, under the command of Captain Fitzroy, R.N. during the years 1832 to 1836. Smith, Elder, and Co.
Kavalco, K. F., & Moreira-Filho, O. (2003). Cytogenetical analyses in four species of the genus Astyanax (Pisces, Characidae) from Paraíba do Sul river basin. Caryologia, 56(4), 453–461. https://doi.org/10.1080/00087114.2003.10589358
Konerat, J. T., Bueno, V., Martins-Santos, I. C., & Margarido, V. P. (2014). Karyotypic diversity and chromosome evolution in the armored catfishes Callichthyinae (Siluriformes, Callichthyidae). Caryologia, 67(2), 140–148. https://doi.org/10.1080/00087114.2014.931635
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549. https://doi.org/10.1093/molbev/msy096
Lanfear, R., Calcott, B., Ho, S. Y. W., & Guindon, S. (2012). PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molecular Biology and Evolution, 29(6), 1695–1701. https://doi.org/10.1093/molbev/mss020
Larson, E. R., Castelin, M., Williams, B. W., Olden, J. D., & Abbott, C. L. (2016). Phylogenetic species delimitation for crayfishes of the genus Pacifastacus. PeerJ, 4, e1915. https://doi.org/10.7717/peerj.1915
Levan, A., Fredga, K., & Sandberg, A. A. (1964). Nomenclature for centromeric position on chromosomes. Hereditas, 52(2), 201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lexer, C., & Widmer, A. (2008). The genic view of plant speciation: Recent progress and emerging questions. Philosophical Transactions of the Royal Society London B Biological Sciences, 363(1506), 3023–3036. https://doi.org/10.1098/rstb.2008.0078
Lima, F. C. T., & Britto, M. R. (2020). A new Corydoras (Ostariophysi: Siluriformes: Callichthyidae) with an unusual sexual dimorphism from the rio Juruena basin, Brazil. Zootaxa, 4742(3), 518–530. https://doi.org/10.11646/zootaxa.4742.3.6.
Lovejoy, N. R., & Araújo, M. L. (2000). Molecular systematics, biogeography and population structure of Neotropical freshwater needlefishes of the genus Potamorrhaphis. Molecular Ecology, 9(3), 259–268. https://doi.org/10.1046/j.1365-294x.2000.00845.x
Lui, R. L., Blanco, D. R., Moreira-Filho, O., & Margarido, V. P. (2012). Propidium iodide for making heterochromatin more evident in the C-banding technique. Biotechnic & Histochemistry, 87(7), 433–438. https://doi.org/10.3109/10520295.2012.696700
Maack, R. (2012). Geografia Física do Estado do Paraná (4th ed.). Editora UEPG. [In Portuguese].
Maia, G. M. G. (2014). Identificação Molecular de Espécies da Subfamília Corydoradinae (Siluriformes: Callichthyidae). Dissertação (mestrado) – Universidade Estadual Paulista, Unesp. Instituto de Biociências de Botucatu. [In Portuguese].
Margarido, V. P., & Moreira-Filho, O. (2008). Karyotypic differentiation through chromosome fusion and number reduction in Imparfinis hollandi (Ostariophysi, Heptapteridae). Genetics and Molecular Biology, 31(1), 235–238. https://doi.org/10.1590/S1415-47572008000200012
Martins, C., & Galetti, P. M. (1999). Chromosomal localization of 5S rDNA genes in Leporinus fish (Anostomidae, Characiformes). Chromosome Research, 7(5), 363–367. https://doi.org/10.1023/A:1009216030316
McCune, B., & Mefford, M. J. (2007). Multivariate analysis on the PC-ORD system. MjM Software.
Mezzaroba, L., Debona, T., Frota, A., Graça, W. J., & Gubiani, É. A. (2021). From the headwaters to the Iguassu Falls: Inventory of the ichthyofauna in the Iguassu River basin shows increasing percentages of nonnative species. Biota Neotropica, 21(2), e20201083. https://doi.org/10.1590/1676-0611-BN-2020-1083
Montoya-Burgos, J. I. (2003). Historical biogeography of the catfish genus Hypostomus (Siluriformes: Loricariidae), with implications on the diversification of Neotropical ichthyofauna. Molecular Ecology, 12(7), 1855–1867. https://doi.org/10.1046/j.1365-294X.2003.01857.x
Moreira-Filho, O., & Bertollo, L. A. C. (1991). Astyanax scabripinnis (Pisces, Characidae): A “species complex.” Brazilian Journal of Genetics, 14, 331–357.
Near, T. J., Kassler, T. W., Koppelman, J. B., Dillman, C. B., & Philipp, D. P. (2003). Speciation in North American Black Basses, Micropterus (Actinopterygii: Centrarchidae). Evolution, 57(7), 1610–1621. https://doi.org/10.1111/j.0014-3820.2003.tb00368.x
Nijssen, H., & Isbrücker, I. J. H. (1980). A review of the genus Corydoras Lacépède, 1803. Bijdragen Tot De Dierkunde, 50(1), 190–220.
Nijssen, H., & Isbrücker, I. J. H. (1967). Notes on the Guiane species of Corydoras Lacépède, 1803, with descriptions of seven new species and designation of a neotype for Corydoras punctatus (Bloch, 1794)-(Pisces, Cypriniformes, Callichthyidae). Zoologische Mededelingen, 42(5), 21–50.
Nijssen, H., & Isbrücker, I. J. H. (1983). Review of the genus Corydoras from Colombia, with descriptions of two new species (Pisces, Siluriformes, Callichthyidae). Beaufortia, 33(5), 53–71.
Nijssen, H., & Isbrücker, I. J. H. (1986). Review of the genus Corydoras from Peru and Ecuador (Pisces, Siluriformes, Callichthyidae). Studies on Neotropical Fauna and Environment, 21(1–2), 1–68. https://doi.org/10.1080/01650528609360697
Nijssen, H. (1970). Revision of the Surinam catfishes of the genus Corydoras Lacépède, 1803 (Pisces, Siluriformes, Callichthyidae). Beaufortia, 18(230), 1–75.
Oliveira, C., & Gosztonyi, A. E. (2000). A cytogenetic study of Diplotnystes mesembrinus (Teleostei, Siluriformes, Diplomystidae) with a discussion of chromosome evolution in siluriformes. Caryologia, 53(1), 31–37. https://doi.org/10.1080/00087114.2000.10589178
Oliveira, C., Toledo, L. F. A., Foresti, F., Britski, H. Á., & Toledo-Filho, S. A. (1988). Chromosome formulae of Neotropical freshwater fishes. Revista Brasileira De Genética, 11(3), 577–624.
Oliveira, C., Toledo, L. F. A., Mori, L., & Toledo-Filho, S. A. (1993). Cytogenetic and DNA content studies on armoured catfishes of the genus Corydoras (Pisces, Siluriformes, Callichyidae) from the southeast coast of Brazil. Revista Brasileira De Genética, 16(3), 617–629.
Oliveira, C., Toledo, L. F. A., Mori, L., & Toledo-Filho, S. A. (1992). Extensive chromosomal rearrangements and nuclear DNA content changes in the evolution of the armoured catfishes genus Corydoras (Pisces, Siluriformes, Callichthyidae). Journal of Fish Biology, 40(3), 419–431. https://doi.org/10.1111/j.1095-8649.1992.tb02587.x
Oliveira, C., Toledo, L. F. A., & Toledo-Filho, S. A. (1990). Comparative cytogenetic analysis of three cytotypes of Corydoras nattereri (Pisces, Siluriformes, Callichthyidae). Cytologia, 55(1), 21–26.
Pazza, R., Kavalco, K. F., Toledo, L. F. A., & Bertollo, L. A. C. (2005). Hoplosternum littorale (Teleostei, Callichthyidae) from a Coastal River basin in Brazil - Cytogenetic analysis and gene mapping of 5S and 18S rDNA. Caryologia, 58(4), 339–344. https://doi.org/10.1080/00087114.2005.10589472
Pereira, L. H. G., Hanner, R., Foresti, F., & Oliveira, C. (2013). Can DNA barcoding accurately discriminate megadiverse Neotropical freshwater fish fauna? BMC Genetics, 14, 20. https://doi.org/10.1186/1471-2156-14-20
Pereira, L. H. G., Maia, G. M. G., Hanner, R., Foresti, F., & Oliveira, C. (2011). DNA barcodes discriminate freshwater fishes from the Paraíba do Sul River Basin, São Paulo. Brazil. Mitochondrial DNA, 22(Suppl 1), 71–79. https://doi.org/10.3109/19401736.2010.532213
Pinacho-Pinacho, C. D., García-Varela, M., Sereno-Uribe, A. L., & León, G. P. P. (2018). A hyper-diverse genus of acanthocephalans revealed by tree-based and non-tree based species delimitation methods: Ten cryptic species of Neoechinorhynchus in Middle American freshwater fishes. Molecular Phylogenetics and Evolution, 127, 30–45. https://doi.org/10.1016/j.ympev.2018.05.023
Pinkel, D., Straume, T., & Gray, J. W. (1986). Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proceedings of the National Academy of Sciences of the United States of America, 83(9), 2934–2938. https://doi.org/10.1073/pnas.83.9.2934
Piscor, D., Paiz, L. M., Baumgärtner, L., Cerqueira, F. J., Fernandes, C. A., Lui, R. L., Parise-Maltempi, P. P., & Margarido, V. P. (2020). Chromosomal mapping of repetitive sequences in Hyphessobrycon eques (Characiformes, Characidae): A special case of the spreading of 5S rDNA clusters in a genome. Genetica, 148(1), 25–32. https://doi.org/10.1007/s10709-020-00086-3
Piscor, D., Ribacinko-Piscor, D. B., Fernandes, C. A., & Parise-Maltempi, P. P. (2013). Cytogenetic analysis in three Bryconamericus species (Characiformes, Characidae): First description of the 5S rDNA-bearing chromosome pairs in the genus. Molecular Cytogenetics, 6, 13. https://doi.org/10.1186/1755-8166-6-13
Pons, J., Barraclough, T. G., Gomez-Zurita, J., Cardoso, A., Duran, D. P., Hazell, S., Kamoun, S., Sumlin, W. D., & Vogler, A. P. (2006). Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Systematic Biology, 55(4), 595–609. https://doi.org/10.1080/10635150600852011
Pucci, M. B., Barbosa, P., Nogaroto, V., Almeida, M. C., Artoni, R. F., Pansonato-Alves, J. C., Foresti, F., Moreira-Filho, O., & Vicari, M. R. (2014). Population differentiation and speciation in the genus Characidium (Characiformes: Crenuchidae): Effects of reproductive and chromosomal barriers. Biological Journal of the Linnean Society, 111(3), 541–553. https://doi.org/10.1111/bij.12218
Queiroz, K. (2007). Species concepts and species delimitation. Systematic Biology, 56(6), 879–886. https://doi.org/10.1080/10635150701701083
Qvarnström, A., & Bailey, R. I. (2009). Speciation through evolution of sex-linked genes. Heredity, 102(1), 4–15. https://doi.org/10.1038/hdy.2008.93
Rambaut, A., Suchard, M. A., Xie, D., & Drummond, A. J. (2014). Tracer v1.6. http://tree.bio.ed.ac.uk/software/tracer/
Raskina, O., Belyayev, A., & Nevo, E. (2004). Quantum speciation in Aegilops: Molecular cytogenetic evidence from rDNA cluster variability in natural populations. Proceedings of the National Academy of Sciences of the United States of America, 101(41), 14818–14823. https://doi.org/10.1073/pnas.0405817101
R Core Team. (2020). R: A language and environment for statistical computing. R Foundation for Statistical Computing. www.r-project.org/
Reis, R. E. (1998). Anatomy and phylogenetic analysis of 343 the Neotropical Callichthyidae catfish (Ostariophysi, Siluriformes). Zoological Journal of the Linnean Society, 124(2), 105–168. https://doi.org/10.1111/j.1096-3642.1998.tb00571.x
Rocha, R. H., Baumgätner, L., Paiz, L. M., Margarido, V. P., Fernandes, C. A., & Gubiani, É. A. (2016). An uncommon co-localization of rDNA 5S with major rDNA clusters in Callichthyidae (Siluriformes): A report case in Corydoras carlae Nijssen & Isbrücker, 1983. Comparative Cytogenetics, 10(4), 603–613. https://doi.org/10.3897/CompCytogen.v10i4.9507
Shibatta, O., & Hoffmann, A. C. (2005). Variação geográfica em Corydoras paleatus (Jenyns) (Siluriformes, Callichtyidae) do sul do Brasil. Revista Brasileira De Zoologia, 22(2), 366–371. https://doi.org/10.1590/S0101-81752005000200010.[InPortuguese]
Shimabukuro-Dias, C. K., Oliveira, C., Reis, R. E., & Foresti, F. (2004). Molecular phylogeny of the armored catfish Family Callichthyidae (Ostariophysi, Siluriformes). Molecular Phylogenetics and Evolution, 32(1), 152–163. https://doi.org/10.1016/j.ympev.2003.11.010
Sievers, F., Wilm, A., Dineen, D., Gibson, T. J., Karplus, K., Li, W., Lopez, R., McWilliam, H., Remmert, M., Söding, J., Thompson, J. D., & Higgins, D. G. (2011). Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular Systems Biology, 7(1), 539–544. https://doi.org/10.1038/msb.2011.75
Silvestro, D., & Michalak, I. (2012). RaxmlGUI: A graphical frontend for RAxML. Organisms Diversity & Evolution, 12, 335–337. https://doi.org/10.1007/s13127-011-0056-0
Sumner, A. T. (1972). A simple technique for demonstrating centromeric heterochromatin. Experimental Cell Research, 75(1), 304–306. https://doi.org/10.1016/0014-4827(72)90558-7
Takagui, F. H., Venturelli, N. B., Sampaio, T. R., Dias, A. L., & Giuliano-Caetano, L. (2014). Cytogenetic study in Corydoras britskii (Siluriformes: Callichthyidae), using different chromosomal banding and fluorescence hybridization in situ with rDNA probes. Ichthyological Research, 61, 201–206. https://doi.org/10.1007/s10228-014-0392-0
Tencatt, L. F. C., Graça, W. J., & Pavanelli, C. S. (2013). First record of Megalechis picta (Muller and Troschel, 1849) (Siluriformes: Callichthyidae) in the upper Rio Paraná basin, Brazil. Check List, 9(5), 1081–1083. https://doi.org/10.15560/9.5.1081.
Tencatt, L. F. C., Lima, F. C. T., & Britto, M. R. (2019). Deconstructing an octogenarian misconception reveals the true Corydoras arcuatus Elwin 1938 (Siluriformes: Callichthyidae) and a new Corydoras species from the Amazon basin. Journal of Fish Biology, 95(2), 453–471. https://doi.org/10.1111/jfb.13980
Tencatt, L. F. C., & Ohara, W. M. (2016). Two new species of Corydoras Lacépède, 1803 (Siluriformes: Callichthyidae) from the rio Madeira basin. Brazil. Neotropical Ichthyology, 14(1), e150063. https://doi.org/10.1590/1982-0224-20150063
Turner, C. H. (1992). On Wolff’s law of trabecular architecture. Journal of Biomechanics, 25(1), 1–9. https://doi.org/10.1016/0021-9290(92)90240-2
Ward, R. D., Hanner, R., & Hebert, P. D. N. (2009). The campaign to DNA barcode all fishes. FISH-BOL. Journal of Fish Biology, 74(2), 329–356. https://doi.org/10.1111/j.1095-8649.2008.02080.x
Ward, R. D., Zamlak, S. Z., Innes, B. H., Last, P. R., & Hebert, P. D. N. (2005). DNA barcoding Australia’s fish species. Philosophical Transactions of the Royal Society London B Biological Sciences, 360(1462), 1847–1857. https://doi.org/10.1098/rstb.2005.1716
Ward, R. D. (2009). DNA barcode divergence among species and genera of birds and fishes. Molecular Ecology Resources, 9(4), 1077–1085. https://doi.org/10.1111/j.1755-0998.2009.02541.x
Weitzman, S. H., Menezes, N. A., & Weitzman, M. J. (1988). Phylogenetic biogeography of the Glandulocaudini (Teleostei: Characiformes, Characidae) with comments of the distributions of other freshwater fishes in Eastern and Southeastern Brazil. In P. E. Vanzolini, & W. R. Heyer (Eds.), Proceedings of a workshop on Neotropical Distribution Patterns. Academia Brasileira de Ciências. 379–427.
Wong, E. H. K., Shivji, M. S., & Hanner, R. H. (2009). Identifying sharks with DNA barcodes: Assessing the utility of a nucleotide diagnostic approach. Molecular Ecology Resources, 9(s1), 243–256. https://doi.org/10.1111/j.1755-0998.2009.02653.x
Zhang, J., Kapli, P., Pavlidis, P., & Stamatakis, A. (2013). A general species delimitation method with applications to phylogenetic placements. Bioinformatics, 29(22), 2869–2876. https://doi.org/10.1093/bioinformatics/btt499
Acknowledgements
The authors thank the Ineo/Gerpel, specially Poliana, Guido, Angélica, and Werike by assistance in sample collection. The authors are also grateful to C. D. Pisol and J. M. Vargas for the assistance in obtaining the sequences COI and RAG1. The authors also thank I. J. Oliveira for assistance with analysis of time divergence estimation.
Funding
This study was funded partially by the Coordination of Superior Level Staff Improvement – Finance Code 001. We would also like to express our gratitude to Brazilian agency Fundação Araucária for scholarship to first author. We also thank the National Council for Scientific and Technological Development (CNPq) for the continuous research productivity grants to ÉAG (PQ Process Number: 308578/2017–1) and WJG (PQ Process Number: 305200/2018–6). The CNPq also provided grants to LFCT (process #160674/2019–0).
Author information
Authors and Affiliations
Contributions
ÉAG, LFCT, WJG and RHR conceived the study for this manuscript with input from CAF and VPM. RHR and ÉAG collected data and resources. Most raw data analysis was performed by RHR, TSB, VPM, LFCT and WJG. RHR wrote the first draft, and all authors contributed to writing.
Corresponding author
Ethics declarations
Ethics approval
This study was carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals, approved by the Committee on the Ethics of Animal Experiments of the Universidade Estadual do Oeste do Paraná (License Number: Protocol 13/09 – CEUA/Unioeste).
Consent for publication
All authors approved the final version of the manuscript for publication.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
da Rocha, R.H., Fernandes, C.A., Bignotto, T.S. et al. Integrated analysis reveals a new species of Corydoras Lacépède, 1803 (Siluriformes: Callichthyidae) in the lower Iguassu River, Brazil. Org Divers Evol 22, 457–474 (2022). https://doi.org/10.1007/s13127-021-00534-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-021-00534-8