Abstract
Cytogenetic data showed a variation in diploid chromosome number in the genus Hyphessobrycon ranging from 2n = 46 to 52, and studies involving repetitive DNA sequences are scarce in representatives of this genus. The purpose of this paper was the chromosomal mapping of repetitive sequences (rDNA, histone genes, U snDNA and microsatellites) and investigation of the amplification of 5S rDNA clusters in the Hyphessobrycon eques genome. Two H. eques populations displayed 2n = 52 chromosomes, with the acrocentric pair No. 24 bearing Ag-NORs corresponding with CMA3+/DAPI−. FISH with a 18S rDNA probe identified the NORs on the short (p) arms of the acrocentric pairs Nos. 22 and 24. The 5S rDNA probe visualized signals on almost all chromosomes in genomes of individuals from both populations (40 signals); FISH with H3 histone probe identified two chromosome pairs, with the pericentromeric location of signals; FISH with a U2 snDNA probe identified one chromosome pair bearing signals, on the interstitial chromosomal region. The mononucleotide (A), dinucleotide (CA) and tetranucleotide (GATA) repeats were observed on the centromeric/pericentromeric and/or terminal positions of all chromosomes, while the trinucleotide (CAG) repeat showed signals on few chromosomes. Molecular analysis of 5S rDNA and non-transcribed spacers (NTS) showed microsatellites (GATA and A repeats) and a fragment of retrotransposon (SINE3/5S-Sauria) inside the sequences. This study expanded the available cytogenetic data for H. eques and demonstrated to the dispersion of the 5S rDNA sequences on almost all chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hyphessobrycon represents one of the largest genus in the family Characidae and it comprises 143 recognized species (Froese and Pauly 2019). Native to the Neotropical region, the species of the genus Hyphessobrycon are widely distributed from southern Mexico to Argentina (Rio de la Plata) (Lima et al. 2003). Data based on morphological and molecular characteristics of Characidae indicate that Hyphessobrycon does not form a monophyletic group (Mirande 2010; Javonillo et al. 2010; Oliveira et al. 2011) and presently, it is placed in the subfamily Stethaprioninae (Eschmeyer and Fong 2019). Approximately 30% of the Hyphessobrycon species are of commercial interest because they exhibit an attractive coloration pattern (Castro-Paz et al. 2014).
The genus Hyphessobrycon includes a large number of recognized species, but few of them have been cytogenetically studied. In spite of this fact, they have shown highly diverse karyotypes in terms of different both diploid number (2n) and karyotype structures. Among the studied representatives of this genus, the diploid number ranges from 2n = 46 in H. tropis (Sheel 1973) to 2n = 52 in H. eques (Martinez et al. 2012; Piscor and Parise-Maltempi 2015). Moreover, B chromosomes of the size of microchromosomes were revealed in genomes of H. eques individuals collected from the Paraná River basin (Piscor and Parise-Maltempi 2015).
The studies involving repetitive DNA sequences in the genus Hyphessobrycon are scarce, with data available only for 18S rDNA in H. anisitsi (Centofante et al. 2003; Mendes et al. 2011) and H. luetkenii (Mendes et al. 2011); and for 5S rDNA only in a single species, H. anisitsi (Centofante et al. 2003). Other repetitive DNA sequences such as histone genes, genes coding for small nuclear RNAs (U snDNA) and microsatellites have not been investigated in genomes of the species of this genus.
Considering that the patterns of repetitive DNA distribution are important for the study of chromosomal evolution, in fishes, this information has contributed substantially to the understanding of trends and dynamics of chromosomal evolution (Mestriner et al. 2000; Cioffi and Bertollo 2010; Symonová et al. 2013; Poltronieri et al. 2014; Sember et al. 2015; Yano et al. 2016; Fernandes et al. 2017; Glugoski et al. 2018; Soto et al. 2018). The purpose of this paper was the analyses of chromosomal locations of repetitive sequences (rDNA, histone genes, U snDNA and microsatellites) and investigation of the amplification of 5S rDNA clusters in the genome of H. eques.
Materials and methods
Samples and statement of ethics
Individuals from two populations of H. eques were collected for cytogenetic and molecular analyses: six males and four females from the Piracicaba River (Santa Maria da Serra—SP); seven males and four females from the Ribeirão Claro River (Rio Claro—SP).
All the institutional guidelines for the care and use of laboratory animals were followed. The animals were captured with permission from the Instituto Chico Mendes de Conservação da Biodiversidade—ICMBio (process number—43497-1) and used for laboratory experiments approved by the Animal Experimental Ethics Committee from the Universidade Estadual Paulista ‘Júlio de Mesquita Filho’—UNESP (protocol number—2335).
Conventional chromosome analyses
The chromosomes were prepared according to Foresti et al. (1981). The morphology of the chromosomes was determined according to the arm’s ratio (Levan et al. 1964). The fundamental number (FN) was calculated according to the chromosomal arm numbers (the m, sm and st chromosomes were considered biarmed—p and q arms—and a chromosomes were considered uniarmed—only q arm. Note that a correspond to acrocentric). The nucleolar organizer regions (NORs) were detected using the silver nitrate impregnation technique described by Howell and Black (1980). Heterochromatin was observed using the C-banding technique of Sumner (1972). CG- and AT-rich regions were identified by double-color Chromomycin A3/4’,6-diamidino-2-phenylindole (CMA3/DAPI) staining with denatured chromosomes, according to a technique commonly used in the Cytogenetic Laboratory of the Universidade Estadual Paulista ‘Júlio de Mesquita Filho’ (UNESP) in Rio Claro/SP (for more detail see, Piscor et al. 2015).
DNA extraction and production of probes
The genomic DNAs (gDNA) were extracted from fin samples of Astyanax and Hyphessobrycon individuals, as described by Sambrook and Russell (2001). The 18S rDNA, 5S rDNA, H3 histone and U2 snDNA probes were obtained by PCR using primers, described by White et al. (1990), Pendás et al. (1994), Cabral-de-Mello et al. (2010) and Bueno et al. (2013), respectively. Moreover, considering the high coincidence of many repetitive sequences, we have tested also FISH with 5S rDNA from other fish species (e.g. Astyanax altiparanae, A. fasciatus, Piabina argentea and Megaleporinus elongatus) in order to discard the possibility that our probe prepared from gDNA of H. eques was insufficient for the experiment due to diverse repetitive DNA content. The probes were labeled by PCR with digoxigenin-11-dUTP (Roche Applied Science, Penzberg, Germany) or biotin-16-dUTP (Roche Applied Science).
The (A)30, (CA)15, (CAG)10 and (GATA)8 microsatellites were amplified and labelled with biotin during synthesis as described by Milani and Cabral-de-Mello (2014). Microsatellites were donated by Prof. Dr. Diogo C. Cabral-de-Mello .
Technique of fluorescent in situ hybridization (FISH)
The FISH technique followed Pinkel et al. (1986), with modifications described by Margarido and Moreira-Filho (2008). Signals were detected using anti-digoxigenin–Rhodamine (Roche Applied Science) for digoxigenin-11-dUTP and avidin–FITC (Sigma Aldrich, St Louis, MO, USA) for biotin-16-dUTP (Roche Applied Science). The chromosomes were counterstained with DAPI. Metaphases were photographed using a BX 61 epifluorescence microscope, coupled with an Olympus DP 71 digital camera (Olympus America, Inc.) with Olympus DP Controller software 3.2.1.276.
Cloning, sequencing and analysis of 5S rDNA-NTS sequences
The PCR products of 5S rDNA amplification were cloned using competent bacteria according to chemical transformation with CaCl2. The DNA fragments were inserted into the plasmid vector with pGEM®-T kit (Promega, Madison, WI, USA) following the manufacturer's specifications. The clones were purified by treatment with ExoSAP-IT® (USB) and sent to sequencing service using the same primers of the PCR reaction as the international platform for sequencing, Macrogen Company (South Korea).
The sequences were edited and analyzed using the BioEdit (Hall 1999) program and the Clustal W algorithm (Thompson et al. 1994) for performing alignment of the sequences. For the identification of the sequences, the CENSOR tool (https://www.girinst.org) (Kohany et al. 2006) and nucleotide BLAST tool (NCBI - National Centre for Biotechnology Information) were used. Finally, the sequences were deposited in GenBank (access numbers: MN396769 to MN396772).
Results
The individuals of H. eques from the Piracicaba River 2n = 52 and karyotype composed of 10 m + 20sm + 8st + 14a chromosomes, with NF = 90 (Fig. 1a). Ag-NORs and CMA3 positive sites were located on the p arm of pair No. 24 (Fig. 1a, in box). The 18S rDNA clusters were located on the p arms of pairs Nos. 22 and 24 (Fig. 1b, in box). The heterochromatin regions were observed in the centromeric positions of almost all chromosomes (Fig. 1b).
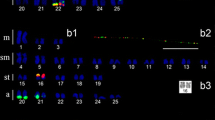
The 5S rDNA clusters were observed on almost all chromosomes (20 pairs) in pericentromeric positions in the individuals from both populations (Fig. 2). Four sites of H3 histone were observed on the pericentromeric regions in two pairs, where in one pair the histone H3 cluster was in a position adjacent to 5S rDNA (Fig. 3a). The U2 snDNA clusters were identified on the interstitial positions of the q arm of one chromosome pair, also adjacent to 5S rDNA (Fig. 3b). Microsatellites (A)30 and (GATA)8 were detected mainly in the centromeric regions of the chromosomes (Fig. 3c, f). The microsatellite (CAG)10 showed some signals on the short/long arms of chromosomes but no scatter pattern (Fig. 3d), with (CA)15 mainly on the terminal regions of chromosomes (Fig. 3e).
Repetitive DNA sequences in genome of Hyphessobrycon eques from the Piracicaba River (Santa Maria da Serra—SP). a Metaphase with 5S rDNA clusters in green and histone H3 in red. b Metaphase with 5S rDNA clusters in green and U2 snDNA in red. c Microsatellite (A)30 in green. d Microsatellite (CAG)10 in green. e Microsatellite (CA)15 in green. f Microsatellite (GATA)8 in green. Bar = 10 µm
The sequence clones of NTS regions (D1, D2, D3 and D4) showed around 300 bp (Fig. 4). Inside the sequences we identified fragments with 97 bp and approximately 97% similarity for the retrotransposon SINE3/5S-Sauria, SINE (Short Interspersed Nuclear Element), Non-LTR (Non-Long Terminal Repeat) and microsatellite repeats, e.g. GATA and A repeats (Fig. 4).
Discussion
Our results showed that individuals from populations of H. eques from the two analyzed locations had the same 2n = 52 and FN = 90. Although having the same 2n = 52, the individuals of H. eques from the Capivara River reported previously (Martinez et al. 2012) differed in their FN values. For example, the number of a chromosomes—14 elements (FN = 90) in the populations under study and 18 elements (FN = 86) from the Capivara River (Martinez et al. 2012). This may perhaps be due to different classification of the chromosome categories
Heterochromatin has been observed mainly on the centromeric/pericentromeric regions of almost all chromosomes of H. eques from the Ribeirão Claro River (Piscor and Parise-Maltempi 2015). In this paper, similar patterns of constitutive heterochromatin were identified on the chromosomes of H. eques from the Piracicaba River. Another species of the genus, Hyphessobrycon reticulatus (Carvalho et al. 2002), H. anisitsi and H. luetkenii (Mendes et al. 2011), also possessed karyotypes with constitutive heterochromatin in the pericentromeric regions of all or almost all chromosomes, indicating that this pattern may be a characteristic feature of their genomes.
Ag-NOR sites corresponded to CMA3+ signals on the p arms in the terminal position of the a chromosome pair No. 24 in individuals under this study. Similarly, the individuals from population from the Capivara River (Martinez et al. 2012) also had karyotypes one Ag-NOR site corresponding to CMA3+ in terminal position on the p arm of the pair No. 17, demonstrating that the NOR regions in H. eques were interspersed with GC-rich sequences, as well as in genomes of other fish species (Mayr et al. 1985; Amemiya and Gold 1986; Galetti et al. 1995; Fernandes and Martins-Santos 2004; Fernandes et al. 2015).
Our FISH results, represented the first physical mapping of repetitive sequences (rDNA, histone genes, U snDNA and microsatellites) in genome of H. eques. H3 histone clusters were observed on two chromosome pairs, one pair holds syntenic sequences of H3 histone and 5S rDNA. Synteny of H3 histone and 5S rDNA clusters was also described in genome of Astyanax (Piscor and Parise-Maltempi 2016). The H. eques also had a syntenic location of U2 snDNA and 5S rDNA in one pair of chromosomes. On the other hand, the chromosomal locations of the 5S rDNA and U2 snDNA clusters in genome of Astyanax were not consistently linked (Piscor et al. 2016). This spatial separation of 5S rDNA and U2 snDNA clusters appears to be the most frequent pattern in fish chromosomes (Merlo et al. 2012; Yano et al. 2017; Sember et al. 2018; Piscor et al. 2018). However, a synteny of these repetitive sequences in genome of H. eques could be related to the high number of 5S rDNA clusters .
The 18S rDNA clusters were observed in four chromosomes of karyotype of H. eques from the Piracicaba River, while in that of H. anisitsi, 18S rDNA signals were observed in ten chromosomes (Centofante et al. 2003). The transposable elements can move via transposition and/or ectopic recombination taking of rDNA sequences to other sites (Raskina et al. 2004, 2008). Similarly, we demonstrated an extensive dispersion of 5S rDNA sequences in individuals from two H. eques populations, which might also be influenced by the activity of transposable elements.
Physical mapping of 5S rDNA in the genome of H. eques showed these clusters distributed on 40 chromosomes in individuals from both analyzed populations, while in formerly investigated H. anisitsi, 5S rDNA were observed in four chromosomes (Centofante et al. 2003). Other studies showed the 5S rDNA clusters on several chromosome pairs of another fish species (see, for example, Cioffi et al. 2010; Nakajima et al. 2012; Sember et al. 2015; Silva et al. 2016). In Gymnotus mamiraua (Gymnotiformes, Gymnotidae) with 2n = 54, more than half of them had syntenic localisation with Tc1/Mariner transposon and 5S rDNA signals, possibly indicating a pseudogene (Silva et al. 2016). In other examples, the spreading of 5S rDNA clusters have led subsequently to development of specific centromeric satellite DNA in genome of Hoplias malabaricus (Erythrinidae) (Martins et al. 2006), while highly amplified 5S rDNA loci in two sister species of Esox (Esociformes, Esocidae) seem to retained functionality, as their sequence is not degenerated into pseudogenes; and the expression of additional copies seems to be regulated by DNA methylation (Symonová et al. 2017).
The NTS regions are subject to intense modification and rapid evolution, resulting in deletions, insertions and substitutions, as well as the inclusion of pseudogenes and microsatellites (Eickbush and Eickbush 2007; Pinhal et al. 2011; Rebordinos et al. 2013; Silva et al. 2016). In this paper, microsatellite repeats were also shown in NTS regions in two populations de H. eques (Ribeirão Claro River and Piracicaba River), as well as fragments of transposable element (Non-LTR, SINE3/5S-Sauria), probably responsible for spreading 5S rDNA sequences on almost all chromosomes. Merlo et al. (2013) showed that different NTS types may contain pseudogenes, LTR-Gypsy, non-LTR (LINE—Long Interspersed Nuclear Element) and microsatellites in Diplodus sargus (Sparidae ). Thus, the latter authors suggest that the concerted evolution model does not explain the 5S rDNA variability of D. sargus; however, the birth-and-death evolution model could explain it.
Mechanisms such as concerted and birth-and-death models have been proposed for explaining the evolution of multigene families (Nei and Rooney 2005). Similarly, Pinhal et al. (2011) pointed out that 5S rDNA molecular evolution in fish genomes is driven by a mixed mechanism that integrates birth-and-death and concerted evolution models. Both these mechanisms could explain the evolution of 5S rRNA genes in genome of H. eques.
Our data of the physical mapping of repetitive sequences in the genome of H. eques genome showed one pair that holds syntenic sequences of H3 histone and 5S rDNA and the other pair holds syntenic sequences of U2 snDNA and 5S rDNA. The 18S rDNA clusters were observed on four chromosomes, while we demonstrated an extensive dispersion of 5S rDNA sequences, with microsatellite sequences and transposable elements identified in the NTS regions. Thus, the dispersion of 5S rDNA clusters on almost all chromosomes was an indication that this form of organization may be favorable for the ongoing elevated genome dynamic in H. eques .
References
Amemiya CT, Gold JR (1986) Chromomycin A3 stains nucleolus organizer regions of fish chromosomes. Copeia 1:226–231. https://doi.org/10.2307/1444915
Bueno D, Palacios-Gimenez OM, Cabral-de-Mello DC (2013) Chromosomal mapping of repetitive DNAs in Abracris flavolineata reveal possible ancestry for the B chromosome and surprisingly H3 histone spreading. PLoS ONE 8:e66532. https://doi.org/10.1371/journal.pone.0066532
Cabral-de-Mello DC, Moura RC, Martins C (2010) Chromosomal mapping of repetitive DNAs in the beetle Dichotomius geminatus provides the first evidence for an association of 5S rRNA and histone H3 genes in insects, and repetitive DNA similarity between the B chromosome and A complement. Heredity 104(4):393–400. https://doi.org/10.1038/hdy.2009.126
Carvalho ML, Oliveira C, Foresti F (2002) Cytogenetics analysis of five species of the subfamily Tetragonopterinae (Teleostei, Characiformes, Characidae). Caryologia 55:181–188. https://doi.org/10.1080/00087114.2002.10589275
Castro-Paz FP, Batista JdS, Porto JIR (2014) DNA barcodes of Rosy Tetras and allied species (Characiformes: Characidae: Hyphessobrycon) from the Brazilian Amazon basin. PLoS ONE 9(5):e98603. https://doi.org/10.1371/journal.pone.0098603
Centofante L, Bertollo LAC, Miyazawa CS, Moreira-Filho O (2003) Chromosomal differentiation among allopatric populations of Hyphessobrycon anisitsi (Pisces, Tetragonopterinae). Cytologia 68:283–288. https://doi.org/10.1508/cytologia.68.283
Cioffi MB, Bertollo LAC (2010) Initial steps in XY chromosome differentiation in Hoplias malabaricus and the origin of an X(1)X(2)Y sex chromosome system in this fish group. Heredity 105:554–561. https://doi.org/10.1038/hdy.2010.18
Cioffi MB, Martins C, Bertollo LAC (2010) Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol Biol 10:217. https://doi.org/10.1186/1471-2148-10-271
Eickbush TH, Eickbush DG (2007) Finely orchestrated movements: evolution of the ribosomal RNA genes. Genetics 175:477–485. https://doi.org/10.1534/genetics.107.071399
Eschmeyer WN, Fong JD (2019) Species of fishes by family/subfamily. http://research.calacademy.org/research/ichthyology/catalog/SpeciesByFamily.asp. Accessed 18 Mar 2019
Fernandes CA, Martins-Santos IC (2004) Cytogenetic studies in two populations of the Astyanax altiparanae (Pisces, Characiformes). Hereditas 141:328–332. https://doi.org/10.1111/j.1601-5223.2004.01832.x
Fernandes CA, Alves DS, Guterres ZR, Martins-Santos IC (2015) Cytogenetic analysis of two locariid species (Teleostei, Siluriformes) from Iguatemi River (Parana River drainage) in Brazil. Comp Cytogenet 9(1):67–78. https://doi.org/10.3897/CompCytogen.v9i1.8804
Fernandes CA, Paiz LM, Baumgärtner L, Margarido VP, Vieira MMR (2017) Comparative cytogenetics of the black ghost knifefish (Gymnotiformes: Apteronotidae): evidence of chromosomal fusion and pericentric inversions in karyotypes of two Apteronotus species. Zebrafish 14:471–476. https://doi.org/10.1089/zeb.2017.1432
Foresti F, Almeida-Toledo LF, Toledo-Filho SA (1981) Polymorphic nature of nucleolus organizer regions in fishes. Cytogenet Cell Genet 31:137–144. https://doi.org/10.1159/000131639
Froese R, Pauly D (2019) FishBase. World Wide Web electronic publication. https://www.fishbase.in/search.php. Accessed 18 Mar 2019
Galetti PM, Mestriner CA, Monaco P, Rasch EM (1995) Post-zygotic modifications and intra- and inter-individual nucleolar organizer region variations in fish: report of a case involving Leporinus friderici. Chromosome Res 3:285–290. https://doi.org/10.1007/BF00713066
Glugoski L, Giuliano-Caetano L, Moreira-Filho O, Vicari MR, Nogaroto V (2018) Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of robertsonian fusion in armored catfish. Gene 650:49–54. https://doi.org/10.1016/j.gene.2018.01.099
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Howell WM, Black DA (1980) Controlled silver-staining of nucleolus organizer regions with the protective colloidal developer: a 1-step method. Experientia 36(8):1014–1015. https://doi.org/10.1007/BF01953855
Javonillo R, Malabarba LR, Weitzman SH, Burns JR (2010) Relationships among major lineages of characid fishes (Teleostei: Ostariophysi: Characiformes), based on molecular sequence data. Mol Phylogenet Evol 54(2):498–511. https://doi.org/10.1016/j.ympev.2009.08.026
Kohany O, Gentles AJ, Hankus L, Jurka J (2006) Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and Censor. BMC Bioinform 7:474. https://doi.org/10.1186/1471-2105-7-474
Levan A, Fredga K, Sandberg AA (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lima FCT, Malabarba LR, Buckup PA et al (2003) General Incertae sedis in characidae. In: Reis RE, Kullander SO, Ferraris CJ (eds) Checklist of the freshwater fishes of South and Central America, 1st edn. Edipucrs, Porto Alegre, pp 106–168
Margarido VP, Moreira-Filho O (2008) Karyotypic differentiation through chromosome fusion and number reduction in Imparfinis hollandi (Ostariophysi, Heptapteridae). Genet Mol Biol 31:235–238. https://doi.org/10.1590/S1415-47572008000200012
Martinez ERM, Alves AL, Silveira SM, Foresti F, Oliveira C (2012) Cytogenetic analysis in the incertae sedis species Astyanax altiparanae Garutti and Britzki, 2000 and Hyphessobrycon eques Steindachner, 1882 (Characiformes, Characidae) from the upper Paraná river basin. Comp Cytogenet 6(1):41–51. https://doi.org/10.3897/compcytogen.v6i1.1873
Martins C, Ferreira IA, Oliveira C, Foresti F, Galetti PM (2006) A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica 127:133–141
Mayr B, Rab P, Kalat M (1985) Localisation of NORs and counterstain-enhanced fluorescence studies in Perca fluviatilis (Pisces, Percidae). Genetica 67:51–56. https://doi.org/10.1007/BF02424460
Mendes MM, Rosa R, Giuliano-Caetano L, Dias AL (2011) Karyotype diversity of four species of the incertae sedis group (Characidae) from different hydrographic basins: analysis of Ag-NORs, CMA3 and 18S rDNA. Genet Mol Res 10(4):3596–3608. https://doi.org/10.4238/2011.November.22.5
Merlo MA, Cross I, Palazón JL, Úbeda-Manzanaro M, Sarasquete C, Rebordinos L (2012) Evidence for 5S rDNA horizontal transfer in the toadfish Halobatrachus didactylus (Schneider, 1801) based on the analysis of three multigene families. BMC Evol Biol 12(1):201. https://doi.org/10.1186/1471-2148-12-201
Merlo MA, Cross I, Manchado M, Cárdenas S, Rebordinos L (2013) The 5S rDNA high dynamism in Diplodus sargus is a transposon-mediated mechanism. Comparison with other multigene families and sparidae species. J Mol Evol 76:83–97. https://doi.org/10.1007/s00239-013-9541-8
Mestriner CA, Galetti PM, Valentini SR, Ruiz IRG, Abel LDS, Moreira-Filho O, Camacho JP (2000) Structural and functional evidence that a B chromosome in the characid fish Astyanax scabripinnis is an isochromosome. Heredity 85:1–9. https://doi.org/10.1046/j.1365-2540.2000.00702.x
Milani D, Cabral-de-Mello DC (2014) Microsatellite organization in the grasshopper Abracris flavolineata (Orthoptera: Acrididae) revealed by FISH mapping: remarkable spreading in the A and B chromosomes. PLoS ONE 9:e97956. https://doi.org/10.1371/journal.pone.0097956
Mirande JM (2010) Phylogeny of the family characidae (Teleostei: Characiformes): from characters to taxonomy. Neotrop Ichthyol 8(3):385–568. https://doi.org/10.1590/S1679-62252010000300001
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152. https://doi.org/10.1146/annurev.genet.39.073003.112240
Nakajima RT, Cabral-de-Mello DC, Valente GT, Venere PC, Martins C (2012) Evolutionary dynamics of rRNA gene clusters in cichlid fish. BMC Evol Biol 12:198. https://doi.org/10.1186/1471-2148-12-198
Oliveira C, Avelino GS, Abe KT, Mariguela TC, Benine RC, Ortí G, Vari RP, e Castro RMC (2011) Phylogenetic relationships within the speciose family Characidae (Teleostei: Ostariophysi: Characiformes) based on multilocus analysis and extensive ingroup sampling. BMC Evol Biol 11(1):275. https://doi.org/10.1186/1471-2148-11-275
Pendás AM, Morán P, Freije JP, García-Vázquez E (1994) Chromosomal location and nucleotide sequence of two tandem repeats of the Atlantic salmon 5S rDNA. Cytogenet Cell Genet 67:31–36. https://doi.org/10.1159/000133792
Pinhal D, Yoshimura TS, Araki CS, Martins C (2011) The 5S rDNA family evolves through concerted and birth-and-death evolution in fish genomes: an example from freshwater stingrays. BMC Evol Biol 11:151. https://doi.org/10.1186/1471-2148-11-151
Pinkel D, Straume T, Gray JW (1986) Citogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA 83(9):2934–2938. https://doi.org/10.1073/pnas.83.9.2934
Piscor D, Parise-Maltempi PP (2015) First description of B chromosomes in the Hyphessobrycon (Characiformes, Characidae) genus: a hypothesis for the extra element of Hyphessobrycon eques Steindachner, 1882. Comp Cytogenet 9(3):325–333. https://doi.org/10.3897/CompCytogen.v9i3.5224
Piscor D, Parise-Maltempi PP (2016) Chromosomal mapping of H3 histone and 5S rRNA genes in eight species of Astyanax (Pisces, Characiformes) with different diploid numbers: syntenic conservation of repetitive genes. Genome 59(3):67–172. https://doi.org/10.1139/gen-2015-0112
Piscor D, Alves AL, Parise-Maltempi PP (2015) Chromosomal microstructure diversity in three Astyanax (Characiformes, Characidae) species: comparative analysis of the chromosomal locations of the 18S and 5S rDNAs. Zebrafish 12(1):81–90. https://doi.org/10.1089/zeb.2014.1036
Piscor D, Centofante L, Parise-Maltempi PP (2016) Highly similar morphologies between chromosomes bearing U2 snRNA gene clusters in the group Astyanax Baird and Girard, 1854 (Characiformes, Characidae): an evolutionary approach in species with 2n = 36, 46, 48, and 50. Zebrafish 13(6):565–570. https://doi.org/10.1089/zeb.2016.1292
Piscor D, Fernandes CA, Parise-Maltempi PP (2018) Conserved number of U2 snDNA sites in Piabina argentea, Piabarchus stramineus and two Bryconamericus species (Characidae, Stevardiinae). Neotrop Ichthyol 16(1):e170066. https://doi.org/10.1590/1982-0224-20170066
Poltronieri J, Marquioni V, Bertollo LAC, Kejnovsky E, Molina WF, Liehr T, Cioffi MB (2014) Comparative chromosomal mapping of microsatellites in Leporinus species (Characiformes, Anostomidae): unequal accumulation on the W chromosomes. Cytogenet Genome Res 142:40–45. https://doi.org/10.1159/000355908
Raskina O, Belyayev A, Nevo E (2004) Quantum speciation in Aegilops: molecular cytogenetic evidence from rDNA cluster variability in natural populations. Proc Natl Acad Sci USA 101(41):14818–14823. https://doi.org/10.1073/pnas.0405817101
Raskina O, Barber JC, Nevo E, Belyayev A (2008) Repetitive DNA and chromosomal rearrangements: speciation-related events in plant genomes. Cytogenet Genome Res 120:351–357. https://doi.org/10.1159/000121084
Rebordinos L, Cross I, Merlo A (2013) High evolutionary dynamism in 5S rDNA of fish: state of the art. Cytogenet Genome Res 141:103–113. https://doi.org/10.1159/000354871
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sember A, Bohlen J, Šlechtová V, Altmanová M, Symonová R, Ráb P (2015) Karyotype differentiation in 19 species of river loach fishes (Nemacheilidae, Teleostei): extensive variability associated with rDNA and heterochromatin distribution and its phylogenetic and ecological interpretation. BMC Evol Biol 15:251. https://doi.org/10.1186/s12862-015-0532-9
Sember A, Bohlen J, Šlechtová V, Altmanová M, Pelikánová Š, Ráb P (2018) Dynamics of tandemly repeated DNA sequences during evolution of diploid and tetraploid botiid loaches (Teleostei: Cobitoidea: Botiidae). PLoS ONE 13(3):e0195054. https://doi.org/10.1371/journal.pone.0195054
Sheel JJ (1973) Fish chromosome and their evolution. Internal Report of Danmarks Akuarium, Charlottenlund
Silva M, Barbosa P, Artoni RF, Feldberg E (2016) Evolutionary dynamics of 5S rDNA and recurrent association of transposable elements in electric fish of the family Gymnotidae (Gymnotiformes): the case of Gymnotus mamiraua. Cytogenet Genome Res 149:297–303. https://doi.org/10.1159/000449431
Symonová R, Majtánová Z, Sember A, Staaks GB, Bohlen J, Freyhof J, Rábová M, Ráb P (2013) Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol 13:42. https://doi.org/10.1186/1471-2148-13-42
Symonová R, Ocalewicz K, Kirtiklis L, Delmastro GB, Pelikánová Š, Garcia S, Kovařík A (2017) Higher-order organisation of extremely amplified, potentially functional and massively methylated 5S rDNA in European pikes (Esox sp.). BMC Genomics 18:391. https://doi.org/10.1186/s12864-017-3774-7
Soto M, Castro JP, Walker LI, Malabarba LR, Santos MH, Almeida MC, Moreira-Filho O, Artoni RF (2018) Evolution of trans-Andean endemic fishes of the genus Cheirodon (Teleostei: Characidae) are associated with chromosomal rearrangements. Rev Chil Hist Nat. https://doi.org/10.1186/s40693-018-0078-5
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75(1):304–306. https://doi.org/10.1016/0014-4827(72)90558-7
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680. https://doi.org/10.1007/978-1-4020-6754-9_3188
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal genes for phylogenetics. In: Innis MA, Gelfand DH, Shinsky JJ, White TJ (eds) PCR Protocols, a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yano CF, Bertollo LAC, Ezaz T, Trifonov V, Sember A, Liehr T, Cioffi MB (2016) Highly conserved Z and molecularly diverged W chromosomes in the fish genus Triportheus (Characiformes, Triportheidae). Heredity 118:276–283. https://doi.org/10.1038/hdy.2016.83
Yano CF, Bertollo LAC, Rebordinos L, Merlo MA, Liehr T, Portela-Bens S, Cioffi MB (2017) Evolutionary dynamics of rDNAs and U2 small nuclear DNAs in Triportheus (Characiformes, Triportheidae): high variability and particular syntenic organization. Zebrafish 14(2):146–154. https://doi.org/10.1089/zeb.2016.1351
Piscor D, Fernandes CA, Parise-Maltempi PP, (2018) Conserved number of U2 snDNA sites in Piabina argentea, Piabarchus stramineus and two Bryconamericus species (Characidae, Stevardiinae). Neotrop Ichthyol 16(1):e170066. https://doi.org/10.1590/1982-0224-20170066
Acknowledgements
This study was supported by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). The authors are grateful to Dr. Diogo Cavalcanti Cabral-de-Mello for the microsatellite DNA probes.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Piscor, D., Paiz, L.M., Baumgärtner, L. et al. Chromosomal mapping of repetitive sequences in Hyphessobrycon eques (Characiformes, Characidae): a special case of the spreading of 5S rDNA clusters in a genome. Genetica 148, 25–32 (2020). https://doi.org/10.1007/s10709-020-00086-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-020-00086-3