Abstract
Macroautophagy (hereafter called autophagy) is a highly conserved lysosomal pathway for catabolism of intracellular material in eukaryotic cells. Autophagy is also an essential homeostatic process through which intracellular components are recycled for reuse or energy production. The extremely regulated autophagy process begins with the formation of hallmarked double membrane bound organelles called autophagosomes which in turn fuse with lysosomes called autolysosomes and finally degrade the autophagic cargos. The multistages molecular machinery of autophagy is critically orchestrated by the action of a set of the autophagy proteins (Atg) and a supreme regulator, mTOR (mechanistic target of rapamycin). However, individual stages of autophagy are mechanistically complex and partially understood. In this review, the individual stages of autophagy are dissected, and the corresponding molecular regulation is discussed in view of current scientific knowledge of autophagy. This understanding of sequential events of autophagy machinery through this review may lead to great interest in the therapeutic potential for manipulating of autophagy in established diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
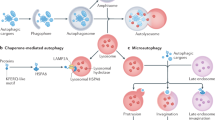
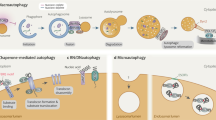
Autophagy is an evolutionarily conserved cellular homeostatic process that facilitates nutrient recycling via lysosomal degradation of potentially harmful cytoplasmic entities. As a housekeeping process, autophagy constitutively executed under basal condition that controls metabolic and cellular homeostasis [2]. Based on the mechanisms by which the target substrates are delivered to lysosomes for degradation, three types of autophagy have been clarified—macroautophagy, chaperone-mediated autophagy, and microautophagy. The best-characterized variant of autophagy is macroautophagy [17]; hereafter, it simply referring as autophagy.
General regulation of autophagy machinery
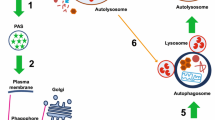
As an adaptive catabolic process, autophagy is activated in response to a wide variety of cellular stresses, including nutrient deprivation and hypoxia [17]. To date, more than forty-one Atg proteins have been identified for the dedication and execution of autophagy [92]. Most of them identified by genetic screens in yeast have mammalian homologs suggesting that are evolutionarily conserved in mammals. The core machinery of the classical and best-characterized autophagy involves the following steps: initiation, vesicle nucleation, elongation, and enclosure for the formation of an isolation membrane (IM, previously known as phagophore), cargo recruitment in IM, completion of the double-membrane vesicles called autophagosomes (APs) in the cytosol, transport of AP and lysosomes on the microtubules (MTs), docking and fusion of the AP with lysosome to form autolysosome (AL), autophagic cargo degradation for recycling, and finally autophagic lysosome reformation (ALR) cycle (Fig. 1). In general, canonical autophagy occurs via class III phosphoinositide 3-kinase (PI3K) complex (PI3KC3)/mechanistic target of rapamycin complex 1 (mTORC1) signaling axis [2, 80] which directly regulates the autophagic activity through the phosphorylation and inhibition of Ser/Thr protein kinase, UNC51-like kinase 1 (ULK1), a part of the first protein complex involved in autophagy initiation [1].
A simply form of mammalian autophagy process. Cellular stress such as nutrient deprivation, ER stress, and radiation or anticancer drug are well known autophagy inducers. Autophagy initiates with IM formation (also known as phagophore). The IM appears to have several sources such as organelles-endoplasmic reticulum (ER), Golgi apparatus, mitochondria, or PM or their contact sites such as ER exit sites (ERES). Through a series of chain reactions, numerous molecular machinery events of this IM are subsequently processed including nucleation, elongation, AP maturation, docking and fusion AP with lysosome, cargo degradation as well as ALR (autophagic lysosome reformation) cycle. Once the cargos are degraded, the degrading metabolites are exported to cytosol to be recycled for ATP production, biosynthesis new macromolecules as well as new organelles like lysosomes
Autophagy initiation: the activation of ULK1 complex
Autophagy is initiated by molecular network machineries that centrally involved a set of Atg proteins for modulating IM formation and AP maturation. Under cellular stress, ULK1 and PI3KC3 complexes are involved to initiate autophagy in mammalian cells [1, 2, 80]. The ULK1 complex consists of ULK1 itself and the noncatalytic subunits FIP200 (focal adhesion kinase (FAK) family kinase–interacting protein of 200 kD), Atg13 and Atg101 [84]. In starvation, ULK1 is activated by autophosphorylation at Thr180 in its kinase domain’s activation loop [49]. This autophosphorylation is promoted by coassembly with other subunits such as Atg13 of the complex and by the conditions that induce autophagy. This coassembly in turn enhances the local ULK1 concentration and promotes their mutual autophosphorylation. Since Atg13 bridges ULK1 to the scaffolding protein FIP200 [35, 51], the recruitment of ULK1 to FIP200 with the help of Atg13 may be a major mechanism for ULK1 transautophosphorylation.
The lipid kinase PI3KC3 phosphorylates the lipid head group of PI (phosphatidylinositol) to generate PI3P (phosphatidylinositol 3-phosphate) [3]. Formation of PI3P is an essential early event in autophagy initiation, occurring just downstream of ULK1 [17]. Both protein complexes are regulated by phosphorylation, mainly due to mTORC1, the master regulator of cell growth and metabolism, and AMPK (AMP-activated protein kinase) in nutrient-sensing pathways [1, 2, 80] as well as ubiquitination [29]. Under available growth factors and nutrients conditions in the extracellular space, the PI3K/mTORC1 pathway is highly active; mTORC1 phosphorylates Atg13 [2] and prevents ULK1 activation by phosphorylating it at Ser757 residue. This Atg13 phosphorylation by mTORC1 inhibits the assembly of Atg13 with ULK1, by introducing steric and electrostatic repulsion into the binding sites on ULK1 [23]. However, during nutrient deprivation or limited growth factors, mTORC1 becomes inactive and can no longer repress the ULK1 complex and triggers autophagy [2, 80]. In response to energy deficits (e.g., glucose starvation or amino-acid deprivation) AMPK can also modulate ULK1 activity via fine-tuning of the phosphorylation status of ULK1 (Fig. 2). The ULK1 kinase provides proautophagic transduction signals by phosphorylating many substrate proteins [43]. The numerous substrates of ULK1 include itself and other subunits of the ULK1 complex; other elements of the core autophagy machinery, including PI3KC3 subunits and the sole conserved transmembrane protein Atg9 [43].
Regulation of autophagy initiation by ULK1 complex. Under nutrient deprivation, autophagy is initiated by mTORC1 inhibition and AMPK activation, which in turn positively regulate ULK1 complex. Induction of ULK1 complex subsequently activates PI3KC3, which leads to PI3P synthesis in IMs. In nutrient rich conditions, mTORC1 represses autophagy activation by phosphorylation at Ser757 residue of ULK1 and at Ser258 of Atg13. In response to nutrient deprivation, mTORC1 is released from ULK1 complex, leading to dephosphorylation of ULK1 and Atg13 and induction of autophagy. Activated ULK1 phosphorylates both FIP200 and Atg13 that events are necessary for autophagy initiation. In contrast, AMPK-dependent phosphorylation of ULK1 activates autophagy induction under energy-depleted condition
The source(s) of isolation membrane
Although morphological features of APs are basically common to conventional and alternative autophagy, the origin(s) of IMs has long been an enigma and debated [35], because of the key questions where does the IM membrane material come from and in turn how does it form to AP. Although many microscopic techniques such as scanning electron microscopy (SEM) and live-cell imaging provide detailed morphological data, the images are unable to reveal the autophagy dynamics, particularly where and how rapidly does the sequential completion of APs with demonstrating kinetics [27, 57]. These may suggest the occurrence of AP formation from multiple IM assembly sites or by effective vesicular trafficking from existing compartments and membranes. In these cases, the AP membranes appear to expand by phospholipid membranes derived from various sources rather than by de novo lipid synthesis [6]..Several organelles and their contact sites such as ER (endoplasmic reticulum) exit sites (ERES), mitochondria, ER-mitochondria contact sites, ER-Golgi intermediate compartment (ERGIC), Golgi apparatus, and plasma membrane (PM) have been suggested to serve as membrane donors for the growing IM in mammalian cells, but the exact mechanism mediating this process remains ambiguous [91]. Dependent on the induction conditions and possible contribution in autophagy stages, different membrane sources are as shown in Table 1.
As the largest organelle in the cell, the ER is always in close contact with other endomembrane compartments and establishes membrane contact sites (MCS) for facilitating signaling events. It has been suggested that IM expansion and elongation are found to intimately association with ER membrane via physically connected to an ER subdomain called omegasome (an omega-shaped DFCP1 enriched) [5]. In mammalian cells, the IM begins at a tubular outgrowth [5] of the PI3P-positive domain of the ER omegasome [28]. Another evidence has revealed that ERES, the specialized ER regions are important mediators in the formation of APs [78]. Multipurpose membrane–trafficking factors such as coat complexes COPII and Atg9-containing cytoplasmic vesicles (Atg9 vesicles) seem to have central roles in mammal autophagy initiation [13]. After synthesis and translocation into the ER, Atg9 vesicle arrives to the Golgi possible by COPII-coated vesicles [1, 28]. From here, the Atg9 vesicle is transported via Golgi membranes into other vesicles and/or compartments that contribute to the formation of the IM. An evidence also suggests that AP is nucleated by the coalescence of Atg9 and COPII vesicles [6].
The Golgi apparatus is proposed to be a pivotal membrane source for AP growth due to the association of the Atg9 [99]. However, others argued that mammalian Atg9 is seen to associate with many other compartments, including recycling endosomes, early endosomes, and late endosomes [67]. It is possible that these organelles all participate in AP formation. In starvation, Atg9 vesicles are also mobilized to the IM by TRAPPIII complex [47, 99]. The amount of Atg9 expressed in cells appears to regulate the frequency of AP formation [38]. Phosphorylation of the N-terminal domain of Atg9 by an unknown kinase initiates autophagy and recruits Atg9 to the IM [20]. Thus, mammalian Atg9 is not a bulk component of APs but rather seems to recycle out of the IM prior to closure.
The generation of COPII transport-vesicles in ERES is also spatially, physically, and functionally linked to APs. Maday et al. [55] suggested that the generation of APs at DFCP1-positive subdomains of ER in the distal end of the axon is distinct from ER exit sites in primary neurons. Hence, ERES are the core elements in ER associated with IM formation and extension in mammalian cells [78]. However, emerging contradictory data indicates that only 30% of all APs are associated with the specialized regions of the ER [32]. In addition, the ERES protein TECPR2 (tectonin β-propeller–containing protein 2) which is mutated in HSP (hereditary spastic paraplegia) connects the COPII subunit SEC24D to microtubule (MT)-associated protein 1 (MAP1) light chain 3 (LC3) in human cells [39, 83].
Membrane traffic between the ER and the Golgi is bidirectional. COPII transport vesicles emerge from ERES, and most of the output of ERES is directed to the cis-Golgi via the ERGIC [78]. The transport of ERGIC structures from ERES to the Golgi apparatus may occur with the help of MTs for cargo sorting and recycling. As the donor membrane, the ERGIC acts as a sorting station for exchanging of dynamic membrane with COPI and COPII vesicles [78]. Thus, the generation of COPII vesicles from the ERGIC may participate to a special event for mobilization of autophagy-related membrane [24, 25]. For example, ERGIC supports the LC3 family protein lipidation [25, 83]. In addition, super-resolution imaging of mammalian autophagy initiation sites shows that Atg9 vesicles and the ULK1 complex subunit Atg13 coalesce with ERGIC components COPI and COPII [12].
In starvation-induced autophagy, mitochondria play a central role for translocation of Atg5 and LC3 to puncta localized in the outer membrane of mitochondria (OMM) [44] Moreover, ER-mitochondria contacts, also called mitochondria-associated ER membranes (MAMs) are claimed to be the origin of IM in mammalian cells [44]. MAMs spatially overlapping with omegasomes have been claimed to serve as a cradle of IM and AP vesicle formation in close to the ER [28, 32]. Upon starvation, phosphorylation of Beclin 1, one of the components in PI3KC3, promotes the local production of PI3P in omegasomes that recruit early autophagic effector proteins such as WIPI1/2 (WD-40 repeat domain containing protein 1 and 2 that interact with phosphoinositide) which are essential for the formation of LC3-positive APs [72]. The contradictory argument is that the sources of AP membrane are obscure in mitochondria due to presence of mammalian Atg9 in transGolgi network (TGN) and late endosomes, but not to mitochondria [82], as Atg9-containing vesicles are an important source of membranes for AP formation.
The large surface area of plasma membrane (PM) may act as a massive membrane store that allows cells to synthesize AP at much higher rates than under basic conditions [75]. Because autophagy contributes to the turnover of PM proteins such as connexins (Cx) [75] that act as an important mediator for transport of nutrient and growth factors from the extracellular environment as well as for sensing pH. The transmembrane proteins, Cx, assemble to form PM gap junctions [75]. In normal conditions, connexin 43 (Cx43) is associated with AP precursor proteins such as PI3K complex or Atg16L1 as Cx-Atg complex at the PM prior to the formation of AP [75]. Here, the Cx-Atg complex acts as a negative regulator of autophagic flux [4]. During starvation, Atg14L (autophagy-related14-like; also called BARKOR), a component of PI3KC3 and Atg9 reach to Cx43-enriched PM and activates class III PI3K kinase that triggers to neutralizes the Cx inhibitory effect through internalization of Cx-Atg complex in presence of EPS15 (epidermal growth factor receptor substrate 15), an endocytic adaptor protein and deliver the preAP components to the sites of AP formation [75]. ER-MP contact sites (ER-PMcs) are also essential for AP biogenesis. ER-PMcs can tether to extended synaptotagmins proteins which are important for PI3P synthesis at the cortical ER membrane and adjust the AP biogenesis [61]. In mammalian cells, the endosomal system is extremely dynamic and generates several structurally and functionally distinct compartments, namely recycling endosomes (REs), late endosomes, and lysosomes. REs emerge from vacuolar sorting endosomes and divert cargoes towards the cell surface, the Golgi, or lysosome-related organelles. Hence, REs may also implicate in AP formation through Atg9 trafficking from PM [67]. As a positive regulator of AP formation, synexin18 (SNX18) is also required for regulating Atg9 trafficking from REs [82].
Vesicle nucleation: the activation of class III PI3K complex
The class III PI3K complex, PI3KC3 consists of its catalytic subunit type 3 (best known as vacuolar protein sorting, VPS34 lipid kinase in yeast), Atg14L, p150 (a myristoylated Ser/Thr kinase), Beclin 1 (BECN1: coiled-coil, moesin-like BCL2-interacting protein), and DFCP1 [double FYVE (Fab1-YotB-Vac1p-EEA1) domain–containing protein 1] [3, 37]. In order to achieve complete enzymatic and biological activity, PI3KC3 has to coordinate with extraregulatory factors (Fig. 3). Here, PI3KC3 and Beclin 1, a multivalent adaptor protein form two distinct complexes, namely Beclin 1-PI3KC3 complex I and II. In Beclin 1-PI3KC3 complex I, Atg14L crosslinks between Beclin 1 and the PI3KC3-p150- NRBF2 (nuclear receptor binding factor 2)-complex for stimulation both initiation and maturation of autophagosome formation.
Regulation of autophagy by the complex Beclin 1 network. The PI3KC3 comprises a multiprotein complex PI3KC3-p150-Beclin 1 associated with Atg14L, UVRAG, AMBRA1, or Rubicon to perform distinct roles in AP formation. Rubicon binds with PI3KC3 through UVRAG and represses AP maturation. Under nutrient rich conditions, PI3KC3 mostly associates with UVRAG involved in endosomal transport. Upon starvation, ATG14L replaces UVRAG to induce autophagy. AMBRA1 translocates from mitochondria to ER and associates with Beclin 1-PI3KC3–p150–Atg14L, which is essential for autophagy activation. Microtubules (MTs) contribute to sequestration of Beclin 1 through their association with DLC1 in the two complexes such as DLC1-Beclin 1-AMBRA1, and the other DLC1-Beclin 1-BIM. Upon stimulation of autophagy, activated ULK1 phosphorylates AMBRA1, thus releasing it from the Beclin 1 complex and allowing the recruitment of Beclin 1 into PI3KC3. BIF-1 interacts with Beclin 1 through UVRAG and functions as a positive mediator of PI3KC3 and promoter of Atg9 trafficking. Phosphorylation of regulatory loop of BCL-2 and BIM by JNK1, releases Beclin 1. By binding BH3 domain of BCL-2, NAF-1 dissociates Beclin 1-BCL-2 complex within ER and positively regulates autophagy. Activated ROCK1 binds and phosphorylates at T119 of Beclin 1 and disrupts Beclin 1-BCL-2 interaction. DAPK phosphorylates at Thr119 in BH3 domain of Beclin 1 and promotes the dissociation of Beclin 1 from BCL-2. For dissociation of Beclin 1-BCL-2 complex, HMGB1 competes with BCL-2 for interaction with Beclin 1 and promotes ERK for phosphorylation of BCL-2. TRAF6 controls K63-linked ubiquitination of Beclin 1 to trigger autophagy. The full-length PINK1, Survivin and VMP1 interact with Beclin 1 and enhance autophagy. In addition, caspase-mediated cleavage of Beclin-1 inhibits autophagy. PI3KC3 phosphorylates PI (phosphatidylinositol) to synthesize PI3P in IMs, allowing for elongation of IMs and completion of APs. In addition, Rab5 also activates PI3KC3 for autophagy induction
The putative protein kinase p150 associates PI3KC3 with membranes via the interaction of its protein kinase domain to PI3KC3. Beclin 1 also interacts with a growing number of identified proteins in relatively unstable, transient or specific conditions (Table 2 and Fig. 3) for the localization and activity of PI3KC3-catalyzed PI3P production. The nucleation machinery of initial IM is critically dependent on local production of PI3P which is enriched in the inner surface of IM and marked by its binding protein, DFCP1 in the PI3KC3. This production of PI3P in the IM in turn engages the recruitment of scaffold proteins such as WIPI1/2 and becomes a membrane platform for accumulation of AP proteins and subsequent IM elongation and completion [18]. The functioning of PI3KC3 in autophagy requires its translocation to the IM, which is driven by its unique Atg14L subunit [33, 37]. Upon induction of autophagy, the ULK1 complex localizes to the IM site for regulation of the vesicle nucleation machinery (Fig. 3). The PI3KC3 catalytic subunit contains a major ULK1 phosphorylation site at Ser249 [51]. The ULK1 kinase transduces proautophagic signals by phosphorylating many substrate proteins including ULK1 itself and other subunits of ULK1 complex as well as the elements of core autophagy machinery subunits of PI3KC3 [43]. The PI3KC3-associated IDR (intrinsically disordered region) protein AMBRA1 (activating molecule in Beclin 1–regulated autophagy) acts as a ULK1 substrate [15]. Upon activation of autophagy, the kinase ULK1 phosphorylates AMBRA1 which is proposed to activate PI3KC3 by releasing Beclin 1 from its MTs [15, 54]. However, in Beclin 1-PI3KC3 complex II, Beclin 1 and the PI3KC3-p150 complex are bridged by UVRAG (UV radiation resistance–associated gene) for regulation of autophagy [58]. In addition, the small guanosine triphosphatase (GTPase) Rab5 also binds to and activates PI3KC3 for autophagy induction [75].
Elongation: two ubiquitin-like conjugation systems
The elongation step of the IM is controlled by two evolutionary conserved ubiquitin-like conjugation systems. The first one involves Atg12-Atg5-Atg16 complex (called Atg12 conjugation system) and the other includes LC3-phosphatidylethanolamine (LC3-II) conjugation system. In Atg12 conjugation system, Atg12 is activated by Atg7 (an E1-like ubiquitin–activating enzyme), transferred to Atg10 (an E2-like ubiquitin–conjugating enzyme). Atg7 binds to the C-terminal glycine of Atg12 through its active cysteine site to form an intermediate complex via a thioester bond. Subsequently, Atg12 is induced by ATP hydrolysis and transferred to Atg10. Finally, the C-terminal glycine of Atg12 covalently binds to an internal lysine residue of Atg5 to form Atg12-Atg5 conjugate which is constitutive and irreversible process [43]. Then Atg12–Atg5 conjugation further interacts noncovalently with a coiled coil protein Atg16 to form Atg12-Atg5-Atg16 complex into a tetramer structure by self-oligomerization. This Atg12 complex coordinates with the formation of APs when LC3-phosphatidylethanolamine (PE) insertion is necessary in the membrane. The LC3-II conjugation system involves the cleavage and lipidation of MAP1-LC3 [43, 86]. In mammalian cells, LC3 is first processed by cleaving near the C-terminal glycine residue with cysteine protease, Atg4 to form the cytosolic soluble LC3-I. Atg7 activates LC3-I and delivers it to the E2-like Atg3. LC3-I is eventually cleaved and converted to a membrane-associated form LC3-II through covalent conjugating with the target lipid PE (known as “lipidation”) via an amide bond. Although LC3 primarily functions in the cytoplasm at the sites of autophagosome and autolysosome formation, several groups have reported that a distinct pool of LC3 also resides in the nucleus [63]. This nuclear LC3 serves as a reserve for the cytoplasmic pool and shifts to the cytosol when soluble cytoplasmic LC3 is transformed to the lipidated type and incorporated into autophagic membranes. During starvation, a nuclear pool of LC3 is deacetylated at K49 and K51 by the activated nuclear histone deacetylase SIRT1 [34] leading to its redistribution to the cytoplasm. This deacetylated LC3 interacts with TP53INP2 (tumor protein p53-inducible nuclear protein 2), also known as DOR (diabetes- and obesity-regulated gene), a nuclear cofactor of thyroid hormone receptors and returns to the cytoplasm with TP53INP2, where it is able to interact directly with ATG7 to form a LC3/TP53INP2/ATG7 complex in the cytoplasm and undergo PE conjugation to autophagosome formation [52, 95]. Amazingly, TP53INP2 regulates adiposity by promoting the sequestering of glycogen synthase kinase 3β (GSK3β) via an autophagy-dependent and endosomal sorting complexes required for transport (ESCRT) machinery pathway-dependent manner [76]. LC3-II, the cleaved product of LC3-I, is specifically inserted within the inner and outer elongating AP membrane and remains on completed APs until fusion with the lysosomes (Fig. 4). As a result, LC3 regulates the size of the AP due to its ability to determine membrane curvature. LC3-II on the cytoplasmic face of ALs can be delipidated by Atg4 and recycled [43].
Involvement of two ubiquitin-like conjugation systems in elongation of IM. In Atg12-Atg5-Atg16L1 conjugation system, Atg7 (an E1 enzyme) and Atg10 (an E2 enzyme) are responsible for conjugating Atg12 onto the acceptor lysine of Atg5. The conjugate of Atg12-Atg5 forms a complex with self-oligomerized Atg16L1 resulting in a formation of multimer Atg12-Atg5-Atg16L1 complex. In second conjugation system, LC3 is synthesized as a pro-LC3 and processed to cytosolic form LC3-I. LC3-I is activated by Atg7 and TP53INP2, transferred it to Atg3 (an E2 enzyme) and finally conjugated with PE, generating membrane bound form LC3-PE (LC3-II). Atg4 (cysteine protease) can convert LC3-II into LC3-I for its recycling
Cargo recruitment in autophagosomes
In response to starvation, nonselective bulk autophagy degrades the cytosolic materials such as damaged organelles or aggregated proteins by sequestrating the specific cargo with autophagy receptors in IM (Fig. 5). These receptors are able to bind specifically to cargos tagged with autophagy degradation signals through their LIR (LC3 interacting regions) leading to the engulfment of cargo by the AP membrane [100]. In general, LIR motifs can interact with autophagy regulatory proteins of the LC3/GABARAP family [22]. In mammalian cells, more than twenty autophagy receptors have been identified, and p62 [also known as sequestosome-1 (SQSTM1)] is one of the most common autophagy receptors [14]. p62 promotes the degradation of autophagic cargos in at least three ways: interacting with LC3 via its LIR motif, associating with early AP formation sites by recruiting Atg proteins through its PB1 domain, and binding to ubiquitinated proteins via its UBA (ubiquitin-binding domain) to form protein aggregates [11, 70]. Moreover, p62 may also provide ubiquitinated cargo recruitment to the AP via its interaction with both ubiquitin and LC3 for selective degradation [70]. Interestingly, some other proteins that have LIR and UBA domain such as FYCO1 (FYVE-coiled coil domain containing protein 1), a member of the RUFY (RUN) and FYVE domain–containing protein) family, HDAC6 (histone deacetylase 6), and ALFY (autophagy linked FYVE protein) interact with p62 [60, 65]. As a nuclear scaffold protein, the FYVE domain on ALFY may also recognize PI3P membranes and Atg12 complex and play a role in the assembly of IM recruitment. In addition, another p62 interacting partner, NBR1 (neighbor of BRCA1 gene 1 protein) is also needed ubiquitinated protein for constitutive autophagic degradation [36, 41].
Cargo recruitment in IM and autophagy receptors. Although the sources of IM are contradicted, different membrane pools may contribute to the formation of IM. Atg9 vesicles dynamically traffic to and from IM. PI3P generated by PI3KC3 at IM can be sensed by WIPI1/2 and DFCP1 for regulation of Atg9 localization. Autophagy cargo receptors contain specific ligand-binding domain (e.g., UBA) for polyubiquitination and a short LIR sequence responsible for LC3 binding. Polyubiquitinated proteins are recognized by the receptor p62 that binds to LC3. ALFY sequestrates the specific cargo by bridging the cargo-receptor complex to core Atg proteins. By PI3P binding, ALFY associate with p62 and Atg5. In Atg-12-Atg5-Atg16L1 complex bound, ALFY stimulates lipidation of LC3 to LC3-II. Selective autophagy can distinguish and direct specific cargos to the lysosome. Autophagy receptors contain a short LIR sequence responsible for LC3 binding. Several cargo receptors have been described for mammalian mitophagy. For example, NIX acts as a mitophagy receptor; it has a LIR motif but lacks an UBA domain and is localized with in the OMM. Mitochondria depolarization promotes to PINK1 activation and phosphorylation of ubiquitin and Parkin. Receptors link mitochondria targeted for degradation to the IM. Domain architecture of autophagy receptors containing distinct ubiquitin-binding domains (light blue) and LC3-interacting motifs (LIR, yellow). The SLRs (sequestosome-1-like receptors) constitute of p62, NBR1, NDP52, TAX1BP, and OPTN (optineurin) as well as the known mitophagy receptors FUNDC1, BNIP3, and NIX (BNIP3L) in mammals. Here PB1, Phox and Bem1 domain; RIP, receptor-interacting protein; ZZ, ZZ-type zinc finger domain; NLS1 and NLS2, nuclear localization signals 1 and 2; NES, nuclear export signal; LIR, LC3 interacting region; KIR, Keap interacting region; UBA, ubiquitin-associated domain; TB, TRAF6-binding domain and TM, transmembrane domain
In case of selective autophagy, p62 is dependent on its PB1-domain–driven polymerization; thus, ubiquitin interactions with other proteins are also important for the delivery of p62-mediated cargos [46]. Following discovery of p62 as a sequestosome-1-like receptor (SLR) for selective autophagy, the NBR1 [41], the nuclear dot protein 52 kDa (NDP52), optineurin, and Tax1-binding protein 1 (TAX1BP1) [88] are also important selective autophagy receptors. These autophagy receptors use LIR-motif-dependent interactions to target their cargos for autophagic degradation (Fig. 5). Although selective autophagy receptors obscure a clear specialization, they often cooperate with each other in selecting a specific cargo, e.g., NBR1 interacts with p62 and plays an essential role in p62-dependent sequestration and degradation of aggregated proteins [41].
For higher organisms, mitophagy receptors and their mediated mechanisms are probably the best studied so far. When mitochondria are depolarized by uncouplers such as CCCP (carbonyl cyanide m-chlorophenyl hydrazone), PINK1 (PTEN-induced putative kinase 1) is activated and accumulated at the outer mitochondrial membrane (OMM) [48, 66]. The activated PINK1 recruits Parkin (ubiquitin E3 ligase) from the cytosol to damaged mitochondria leading to polyubiquitination [42], which are recognized and clustered through polymerization by the p62 [42, 64]. Following mitochondrial translocation, Parkin induces two types of polyubiquitin chain models. Firstly, Parkin-mediated K48 chains on specific OMM proteins stimulate proteasome-dependent degradation of Mitofusin1/2 and mitochondrial protein import receptor, TOM20 [16]. Secondly, Parkin also triggers K63 and K27 ubiquitin chains that initiate p62-mediated a signaling cascade to activate mitophagy [81]. The ion channel VDAC1 (voltage-dependent anion-selective channel 1) in the OMM has also been reported to be a target for Parkin-mediated K63 and K27 polyubiquitination and promotes mitophagy [12, 42]. Furthermore, three integral receptor proteins in the OMM such as two homologous BH3-only proteins, BNIP3 (BCl-2/adenovivus E1B 19-kDa interacting protein 3), and NIX (also known as BNIP3L) as well as FUNDC1 that all have a LIR motif in their cytosolic N-terminal domain can directly link the damaged mitochondria to AP membranes in Ub-independent mitophagy pathways [68] (Fig. 5).
Transport of autophagosomes and position of lysosomes
Autophagosomes (APs) are supposed to occur haphazardly throughout the cytoplasm and transport bidirectionally in their maturing process. During AP maturation, interconnected MT network often serves as tracks for intracellular movement. MTs powered by specific motor proteins such as kinesin or dynein families utilize the energy generated by ATP hydrolysis [40]. In general, kinesins move cargo to peripheral that is plus-end of MTs, whereas dyneins involve in AP movement towards the minus-end in perinuclear center. Simple schematic molecular assemblies of both MT-mediated movements of APs are illustrated in Fig. 6 a and b.
Transport of autophagosome (AP) on microtubule; positioning of lysosome and AP. Microtubules (MTs) organize the substantial organelle positioning throughout the autophagy steps. The extremities of MT have two sides namely minus-end in perinuclear region and plus-end in cell periphery. The movement of APs along MTs depends on a precise balance between dynein, a minus-end-directed motor protein and kinesin, a plus-end-directed motor (Fig. 6a). Under sufficient nutrient conditions, lysosomal localized mTORC1 is transported towards plus end cell periphery where it is kept active. Under starvation intracellular pH increases, and recruitment of ARL8B and kinesin KIF2A to lysosomal membrane are inhibited; this favors the centripetal movement of lysosomes and transport of APs in a perinuclear region close to the MT-organizing centre (MTOC). At the same time, mTORC1 is inhibited by ULK1, favoring the formation of new APs. Newly formed APs move along MTs in perinuclear directions as a result of opposing activities of dynein and kinesin/FYCO1. Thereafter, the APs cluster in a perinuclear region close to centrosome, where they fuse with lysosomes. Rab7 links APs to a MT motor through FYCO1 to facilitate kinesin driven movement towards the cell periphery. FYCO1 binds to LC3 and PI3P, a component of AP membrane as well as links to kinesin. Rab7 also binds to RILP and ORP1L in order to mediate dynein-driven movement towards the perinuclear region under normal cholesterol conditions
The Rab (Ras-related protein in brain) family small GTPases particularly Rab7 are the important coordinators of membrane trafficking in eukaryotic cells. Rab7 interlinks MT motors with the position of lysosomes and late APs through several downstream effector proteins such as RILP (Rab7-interacting lysosomal protein), ORP1L (oxysterol-binding protein (OSBP)-related protein 1 L), a cholesterol sensor, and FYCO1. Rab7 involves in both the minus-end and the plus-end direction transport on MTs based on the downstream interacting effector protein(s) and motor proteins such as RILP/dynein or FYCO1/kinesin [69] respectively. FYCO1 localizes on perinuclear cytosolic vesicles but upon starvation, it moves to the cell periphery in a MT-dependent manner [69]. In the plus-end directional movement along the MT, the C-terminal FYVE and Golgi dynamic (GOLD) domains of FYCO1 interacts with Rab7, LC3 and PI3P on AP membrane and its N-terminal coiled coil domain with the kinesin motor protein [69]. In the minus-end directional movement, Rab7 interacts with RILP, ORP1L, and dynein simultaneously in order to convey transport of APs to the perinuclear region [93] (Fig. 6a).
The proper positioning of lysosomes and APs at perinuclear area is required for efficient fusion between these two organelles. Starvation causes an increase in the intracellular pH which induces lysosome relocalization to perinuclear area and transports late APs to the same region of cell with the help of MTs [54]. Since lysosomes are predominantly found in the perinuclear regions, it is reasonable that the movement of mature APs along MT tracks towards the lysosomes is appeared to fuse [73]. Interestingly, mTORC1 signaling regulates the positioning of lysosomes [33, 54] and the intracellular pH (pHi) influences the docking of AP-lysosome in cells [59]. In response to amino acids, mTORC1 locates on peripheral lysosomes close to its upstream signaling elements, whereas in starvation associated higher pHi, the isolated mTORC1 and lysosomes are preferentially clustered in the perinuclear area. This elevated pHi condition also inhibits mTORC1 activity and favors the dissociation of ULK1 from mTORC1 complex and facilitates the docking of lysosomes with APs. More interestingly, pHi elevation also diminishes the recruitment of lysosomal attached KIF2B and KIF1B-β (kinesin family member 2B and 1B-β) as well as ARL8B [ADP ribosylation factor (ARF)-like GTPase 8B) to MTs [59]. Depletion of KIF2B and KIF1B-β disrupts centrifugal movement of lysosomes along MTs (Fig. 6b), and downregulation of ARL8B restricts lysosome transport to the cell periphery [59].
Fusion of autophagosome with lysosome
After acquiring docking machinery, APs fuse with endocytic vesicles and lysosomes. The double-membranous AP docks and fuses with lysosome to form single-membranous autolysosome (AL) [79]. It is also evident that prior to fusion with lysosome, APs merge with late endosomes to form amphisomes that deliver cargo in autophagic vesicle [59]. The machinery aspect of this fusion process is coordinated by three sets of protein families: Rab GTPases, HOPS (homotypic vacuole fusion and protein sorting) tethering complexes, and autophagosomal SNAREs (soluble N-ethylmaleimide-sensitive factor-attachment protein receptors), e.g., syntaxin17 (STX17). Late endosome- and lysosome-localized Rab7 is essential for membrane trafficking from late endosome to lysosome as well as AP-lysosome fusion [59]. Several Rab7 downstream effectors such as RILP, ORP1L, FYCO1, and PLEKHM1 (pleckstrin homology domain–containing family M member 1) have been characterized [69]. PLEKHM1 involves in AP-lysosome fusion through complex formation with Rab7, HOPS, and LC3 on late endosomes and lysosomes [62]. ORP1L binding to Rab7 in the presence of RILP negatively regulates fusion [93]. RUFY4 (RUN and PI3P-interacting FYVE domain–containing protein 4) promotes both autophagic flux and the tethering of APs with lysosomes [87]. Rubicon and UVRAG involve in endocytic transport and AP-lysosomal fusion through Rab7 [99] although they have opposite effects. OATL1 (ornithine aminotransferase-like 1, also called TBC1D25) is recruited onto APs through direct interaction with LC3, and OATL1 overexpression has been shown to inhibit AP-lysosome fusion. The generation of phosphatidylinositol 4-phosphate (PI4P) by PI3P phosphatases on APs is crucial for AP-lysosome fusion in mammals [89]. PI4KIIα (phosphatidylinositol 4 kinase type 2-alpha) normally localizes to both perinuclear region and TGN. Upon starvation, PI4KIIα exits from TGN, disperses into cytoplasm and some PI4KIIα localize to APs in a palmitoylation dependent manner for AP-lysosomal fusion [89]. In addition, INPP5E (inositol polyphosphate-5 phosphatase E) [31] also promotes the fusion step in autophagy (Fig. 7). The lysosomal membrane–localized INPP5E mediates the conversion of phosphatidylinositol (3,5)-bisphosphate [PI(3,5)P2] to PI3P which is required for fusion. Lysosomal INPP5E activity has shown to be ultimately required for cortactin (CTTN) phosphorylation which leads to polymerize actin followed by AP-lysosome fusion [31]. Since the cytoskeleton plays a role in AL formation, several evidences suggest that actin involves in AP-lysosome fusion. For example, HDAC6 recruits the CTTN-dependent actin remodeling machinery, which in turn assembles the actin-network that stimulates the fusion step. PIKFYVE (1-phosphatidylinositol 3-phosphate 5- kinase) resided on lysosomes reverses the action of INPP5E and generates PI(3,5)P2 from PI3P.
Docking and fusion of autophagosome with lysosome. Cortactin (CTTN) binds with lysosomal localized PI(3,5)P2. During fusion step, cellular INPP5E are partially localized on lysosomes where it converts PI(3,5)P2 to PI3P. As a result, INPP5E decreases the amount of PI(3,5)P2 level and releases CTTN from trapped lysosomal membrane. This released CTTN in turn is phosphorylated by other kinases and the activated CTTN acts as a promoter for stabilization of actin polymerized filaments on lysosome, an event required for fusion step. On the other hand, PIKFYVE resided on lysosomes reverses the action of INPP5E and generates PI(3,5)P2 from PI3P. TECPR1 on lysosome interacts with Atg12-Atg5 complex and binds to PI3P on the outer membrane of AP, thus functioning as a tethering factor for AP-lysosome docking. LC3/GABARAP recruits palmitoylated PI4KIIα from TGN to APs, and this PI4KIIα produces PI4P, an essential phospholipid for APs-lysosome fusion. In machinery aspect of AP-lysosome fusion process, three sets of protein families involve: Rab GTPases, HOPS tethering complexes and autophagosomal SNAREs such as STX17, SNAP-29, and VAMP8. The recruitment of EPG5 on late endosomes/lysosomes together with Rab7 and VAMP8 causes a tethering action by binding to LC3-II and STX17/SNAP29. The adaptor protein PLEKHM1 interacts with Rab7, HOPS–SNARE complexes and LC3 and/or GABARAP proteins to facilitate AP-lysosome fusion. Under normal conditions, SNAP-29 is O-GlcNAcylated which is generated by O-linked β-N-acetylglucosamine (O-GlcNAc) transferase (OGT). O-GlcNAcylated SNAP-29 has a decrease binding affinity for its partner SNAREs. This modification is suppressed by starvation and a reduced level of O-GlcNAcylated SNAP-29 acts as a signal for the assembly of SNAP-29-containing transSNARE complexes, thus stimulating autophagy
The AP-anchored SNARE proteins such as syntaxin17 (STX17), SNAP-29 (synaptosome associated protein 29), and VAMP8 (vesicle associated membrane protein 8) can interact with each other to form a highly energetically favorable complex. Moreover, several tethering factors such as HOPS complex, PI3P-binding protein, Atg14L, TECPR1 (tectonin β-propeller-repeat–containing protein 1), and EPG5 (Ectopic P-granules autophagy protein 5) help SNARE proteins for physically driving the AP-lysosomal fusion (Fig. 7). As a tethering factor, Atg14L acts by directly binding to a binary STX17-SNAP29 complex on APs and facilitates its interaction with VAMP8 for promoting STX17-SNAP29-VAMP8–mediated fusion. Absence of TECPR1 in cells results in impaired autophagic flux and accumulated APs which apparently unable fuse efficiently with lysosomes. TECPR1 also interacts with the subunits of lipidated LC3 (LC3-II) complex in an Atg12-Atg5-conjugation dependent manner. EPG5 identified as a Rab7-VAMP7/8 interactor on endosomes/lysosomes determines the specificity of fusion between APs with endosomes/lysosomes [71, 90]. Interestingly, EPG5 also binds to LC3 through its LIR motif and assembles the STX17-SNAP29 complex on APs [45, 59]. Upon starvation, the recruitment of STX17 from the cytosol to matured APs mediates fusion by binding partners HOPS complex, SNAP29, and lysosomal R-SNARE VAMP8 [37]. Furthermore, modification of O-GlcNAc (O-linked N-acetylglucosamine) in SNAP29 negatively regulates SNARE-dependent fusion [30]. Consequently, knockdown of O-GlcNAc transferase or mutating SNAP29O-GlcNAc sites promotes SNAP29-containing SNARE complex formation and enhances the fusion step.
Autophagic lysosome reformation
After fusion with matured lysosome, the intraAP constituents get degraded and released the degraded end products such as amino acids and fatty acids into the cytosol, thereby, causing a local rise in nutrient availability for energy production. These phenomena lead to reactivation of mTORC1 signaling pathway and regeneration of mature lysosomes from ALs in a dynamic remodeling process known as autophagic lysosome regeneration (ALR) [9, 10]. During ALR, tubular structures are extruded from AP membrane and several small vesicles are generated from these tubules called protolysosomes. Initially, the protolysosomes are in neutral pH and absent in lysosomal luminal proteins. Subsequently, the protolysosomes convert into nascent lysosomes by acquiring acidity and lysosomal luminal proteins through ALR, and lysosomal homeostasis is maintained [9, 10]. Although mTORC1 is inhibited and autophagy is induced in short-term starvation, prolonged starvation, and/or nutrient availability by degrading autophagic cargo, reactivation of mTORC1 signaling acts as a negative feedback mechanism to avoid excess autophagy. As part of mTORC1 mechanism, the protolysosomal expansions dynamically emerge from ALs and eventually detaches from the AL as renewed lysosome pool by converting into functional mature lysosomes. Based on proteomic analysis of purified ALs, three key molecular regulatory pathways have been identified to regulate ALR and lysosome homeostasis such as (i) clathrin and phosphatidylinositol (4,5)-bisphosphate, [PI(4,5)P2] mediating budding of tubules, (ii) motor protein KIF5B (kinesin family member 5B) driving tubule structure elongation, and (iii) large GTPase Dynamin 2 (Dyn2) mediating scission of protolysosomes (Fig. 8).
Autophagic lysosome reformation cycle. At autolysosome (AL), PI4KB converts PI to PI4P, which suppresses uncontrolled lysosomal tubulation from ALs and facilitates lysosome budding organized by clathrin and the GTPase Dyn2. The localization of spastizin-spatacsin complex on lysosome/AL is governed by interaction of spastizin FYVE domain and PI3P, and this complex is an important component for initiation of lysosome budding. The conversion of PI(4,5)P2 from PI4P by PIP5K1B is an essential event for tubule elongation. It is also known that Dyn2 is phosphorylated by starvation as well as Src kinase, a nonreceptor protein tyrosine kinase and this activated Dyn2 acts as scissor for separation of proto-lysosome from ALR [50]. In prolonged starvation, energy sources are rapidly generated by degradation of autophagy cargo in ALR pathway which in turn activate mTORC1 and reform lysosomes efficiently. The phosphorylated mTORC1 acts as a regulator of UVRAG phosphorylation and PI3K complex activation in order to increase lysosomal PI3P formation that appears to be critical for tubule scission, a process regulated by Dyn2. In starvation, mTORC1 is inhibited and dissociated from its lysosomal surface. As a result, TFEB is no longer mTORC1-associated phosphorylation, diffuse it from lysosomal surface to nuclear translocation and induces lysosome-autophagy gene expressions
Phosphatidylinositol (PI) such as PI(4,5)P2 and PI4P play essential roles in controlling lysosomal tubule initiation. Clathrin and its associated adapter protein complexes AP2 and AP4 mediate the formation of PI(4,5)P2-enriched microdomains on ALs. Here, one clathrin molecule recruits two AP2 proteins and one AP2 molecule recruits two PI(4,5)P2 molecules. Thus, AP2 links clathrin to PI(4,5)P2 in order to form PI(4,5)P2-enriched microdomains in the clathrin lattices on AL membrane. Intriguingly, PI(4,5)P2 appears on the main body of ALs and on reformation tubules by two different PI4P kinases such as PI4P 5-kinase type 1 beta (PIP5K1β) and PIP5K1α. Here, PIP5K1β initiates AL tubulation by generating membrane PI(4,5)P2-enriched microdomains from PI4P at the AL surface, and induces membrane budding by recruiting clathrin and its associated proteins AP2 and AP4. Another related kinase, PIP5K1α localized in the leading edges of tubules produce PI(4,5)P2 in protolysosomes aided for tubule scission. Besides PI(4,5)P2, PI4P, and PI3P also involve in different stages of ALR. PI4K variant PI4KIIIβ produced PI4P on lysosomes also regulate lysosomal membrane tubule production. In ALR, the generated PI(4,5)P2 from PI4P by PIP5Ks acts in both restricted clathrin recruitment at lysosomal membrane and vesicular scission at the tips of tubules. MT-based motor protein, KIF5B, plays essential roles in transport of lysosome and tubulation of AL [19]. Lysosomal PI3P produced by PI3KC3 complex plays an important role in the scission of protolysosomes in ALR [58]. HSP autosomal recessive gene products, SPG15 (spastic gait 15 protein, also called spastizin) and SPG11 protein (also called spatacsin) also act as PI3P-binding proteins that may pivotally regulate lysosomal tabulation in ALR [7]. Moreover, spastizin and spatacsin are also essential components for the initiation of lysosomal tubulation [9]. However, it remains unclear the detailed mechanisms of how these phospholipids and their kinases regulate ALR. Dyn2 involves in scission of protolysosome through binding to PI(4,5)P2 and forming microtubule bundles. Genetic depletion or treatment of cells with dynasore, a chemical inhibitor of dynamins, Dyn2 prevent lysosomal tubular scission resulting in decreased lysosomal pools. Mechanistically, a set of elegant studies demonstrate that mTOR regulates generation of lysosomal PI3P produced by PI3KC3 complex, and this PI3P in turn regulates the scission of ALR tubules [9, 10, 58]. When mTOR activity is inhibited, the omegasomal PI3P pool increases, and autophagy is initiated [33]. However, when mTOR is reactivated at the termination of autophagy, the lysosomal pool of PI3P increases due to activation of PI3KC3 complex and results in scission of ALR tubules [74]. Thus, PI3KC3 with different complexes in different localizations acts as a regulatory node for coordinating initiation and termination of autophagy (Fig. 8). However, lysosome homeostasis and autophagic function are also regulated by transcription factor EB (TFEB), a protein belonging to the MiTF/TFE family of transcription factors. In addition to inhibiting autophagy, mTORC1 regulates lysosomal biogenesis through the TFEB-dependent pathway [56]. In normal feeding conditions, TFEB is associated with Rag GTPases at the lysosomal surface and undergoes phosphorylation (at residues Ser211 and Ser142) by mTORC1. mTORC1 also promotes the secretion of phosphorylated TFEB to the cytosol and the interaction with chaperones,14-3-3 proteins [56], thus stabilizing TFEB in the cytosol and preventing its nuclear translocation [56]. Conversely in starvation, mTORC1 and Rag GTPase are inhibited, mTORC1 dissociates from its lysosomal surface, and TFEB no longer undergoes mTORC1-associated phosphorylation. As a result, phosphorylated TFEB-14-3-3 complex is no longer available, TFEB diffuses from the surface of lysosomes to undergo nuclear translocation and TFEB induces lysosomal-autophagy gene expressions [56].
Conclusions and future perspectives
In this review, it is summarized the findings regarding several sequential steps of autophagy process. Understanding about the variation of IM sources are in cell-, tissue specific, or in different contexts to environmental or pathological conditions. Thus, different conclusions reached by the different laboratories might be in part due to different experimental approaches and techniques used in the various laboratories. The tissue and cell specificity as well as time-dependent manner of the dissection of autophagy must be addressed in future studies. Answering these questions will provide the opportunity to develop new therapeutic strategies in order to manipulate autophagy for combating established diseases including cancer and neurodegeneration. Moreover, the precise involvement of individual steps of autophagy in human individual diseases remains partially understood. Thus, precise dissecting of autophagy process in specific disease contexts and determining whether manipulation of these steps in autophagy are more beneficial future goals.
References
Abada A, Elazar Z (2014) Getting ready for building: signaling and autophagosome biogenesis. EMBO Rep 15:839–852
Al-Bari MAA, Xu P (2020) Molecular regulation of autophagy machinery by mTOR-dependent and -independent pathways. Ann N Y Acad Sci 1467:3–20. https://doi.org/10.1111/nyas.14305
Backer JM (2016) The intricate regulation and complex functions of the class III phosphoinositide 3kinase Vps34. Biochem J 473:2251–2271
Bejarano E, Yuste A, Patel B, Stout Jr RF, Spray DC, Cuervo AM (2014) Connexins modulate autophagosome biogenesis. Nat Cell Biol 16:401–414
Biazik J, Yla-Anttila P, Vihinen H et al (2015) Ultrastructural relationship of the phagophore with surrounding organelles. Autophagy 11:439–451
Carlsson SR, Simonsen A (2015) Membrane dynamics in autophagosome biogenesis. J Cell Sci 128:193–205
Chang J, Lee S, Blackstone C (2014) Spastic paraplegia proteins spastizin and spatacsin mediate autophagic lysosome reformation. J Clin Invest 124(12):5249–5262
Chen K, Shi W (2016) Autophagy regulates resistance of non-small cell lung cancer cells to paclitaxel. Tumour Biol 37:10539–10544
Chen Y, Yu L (2017) Recent progress in autophagic lysosome reformation. Traffic 18(6):358–361
Chen Y, Yu L (2018) Development of research into autophagic lysosome reformation. Mol Cells 41(1):45–49
Conway O, Kirkin V (2017) Love laughs at locksmiths: Ubiquitylation of p62 unlocks its autophagy receptor potential. Cell Res 27(5):595–597
Cook KL, Soto-Pantoja DR, Abu-Asab M, Clarke PAG, Roberts DD, Clarke R (2014) Mitochondria directly donate their membrane to form autophagosomes during a novel mechanism of parkin- associated mitophagy. Cell Biosci 4:16. https://doi.org/10.1186/2045-3701-4-16
Davis S, Wang J, Zhu M (2016) Sec24 phosphorylation regulates autophagosome abundance during nutrient deprivation. eLife 5. https://doi.org/10.7554/eLife.21167
Deng Z, Purtell K, Lachance V, Wold MS, Chen S, Yue Z (2017) Autophagy receptors and neurodegenerative diseases. Trends Cell Biol 27(7):491–504
Di Bartolomeo S, Corazzari M, Nazio F et al (2010) The dynamic interaction of AMBRA1 with the dynein motor complex regulates mammalian autophagy. J Cell Biol 191(1):155–168
Di Maio R, Barrett PJ, Hoffman EK et al (2016) Alpha-Synuclein binds to TOM20 and inhibits mitochondrial protein import in Parkinson’s disease. Sci Transl Med 8(342):342ra378
Dikic I, Elazar Z (2018) Mechanism and medical implications of mammalian autophagy. Nat Rev Mol Cell Biol 19(6):349–364
Dooley HC, Razi M, Polson HE et al (2014) WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Mol Cells 55:238–252
Du W, Su QP, Chen Y et al (2016) Kinesin 1 drives autolysosome tubulation. Dev Cell 37(4):326–336
Feng YC, Backues SK, Baba M, Heo JM, Harper JW, Klionsky DJ (2016) Phosphorylation of Atg9 regulates movement to the phagophore assembly site and the rate of autophagosome formation. Autophagy 12:648–658
Fernandez AF, Sebti S, Wei Y et al (2018) Disruption of the beclin 1-BCL2 autophagy regulatory complex promotes longevity in mice. Nature 558(7708):136–140
Fracchiolla D, Sawa-Makarska J, Martens S (2017) Beyond Atg8 binding: the role of AIM/LIR motifs in autophagy. Autophagy 13(5):978–979
Fujioka Y, Suzuki SW, Yamamoto H, Kondo-Kakuta C, Kimura Y, Hirano H, Akada R, Inagaki F, Ohsumi Y, Noda NN (2014) Structural basis of starvation-induced assembly of the autophagy initiation complex. Nat Struct Mol Biol 21:513–521
Ge L, Wilz L, Schekman R (2015) Biogenesis of autophagosomal precursors for LC3 lipidation from the ER-Golgi intermediate compartment. Autophagy 11:2372–2374
Ge L, Zhang M, Kenny SJ, Liu D, Maeda M, Saito K, Mathur A, Xu K, Schekman R (2017) Remodeling of ER-exit sites initiates a membrane supply pathway for autophagosome biogenesis. EMBO Rep 18:1586–1603
Ge L, Zhang M, Schekman R (2014) Phosphatidylinositol 3-kinase and COPII generate LC3 lipidation vesicles from the ER-Golgi intermediate compartment. Elife 3:04135
Geng J, Klionsky DJ (2017) Direct quantification of autophagic flux by a single molecule-based probe. Autophagy 13:639–641
Graef M, Friedman JR, Graham C, Babu M, Nunnari J (2013) ER exit sites are physical and functional core autophagosome biogenesis components. Mol Biol Cell 24:2918–2931
Grumati P, Dikic I (2018) Ubiquitin signaling and autophagy. J Biol Chem 293:5404–5413
Guo B, Liang Q, Li L, Hu Z, Wu F, Zhang P, Ma Y, Zhao B, Kovács AL, Zhang Z, Feng D, Chen S, Zhang H (2014) O-GlcNAc-modification of SNAP-29 regulates autophagosome maturation. Nat Cell Biol 16(12):1215–1226
Hasegawa J, Iwamoto R, Otomo T, Nezu A, Hamasaki M, Yoshimori T (2016) Autophagosome-lysosome fusion in neurons requires INPP5E, a protein associated with Joubert syndrome. EMBO J 35(17):1853–1867
Hayashi-Nishino M, Fujita N, Noda T, Yamaguchi A, Yoshimori T, Yamamoto A (2009) A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat Cell Biol 11:1433–1437
Hong Z, Pedersen NM, Wang L, Torgersen ML, Stenmark H, Raiborg C (2017) PtdIns3P controls mTORC1 signaling through lysosomal positioning. J Cell Biol 216(12):4217–4233
Huang R, Xu Y, Wan W, Shou X, Qian J, You Z, Liu B, Chang C, Zhou T, Lippincott-Schwartz J, Liu W (2015) Deacetylation of nuclear LC3 drives autophagy initiation under starvation. Mol Cell 57:456–466. https://doi.org/10.1016/j.molcel.2014.12.013
Hurley JH, Young LN (2017) Mechanisms of autophagy initiation. Annu Rev Biochem 86:225–244
Isakson P, Holland P, Simonsen A (2013) The role of ALFY in selective autophagy. Cell Death Differ 20(1):12–20
Itakura E, Kishi-Itakura C, Mizushima N (2012) The hairpin-type tail-anchored SNARE syntaxin 17 targets to autophagosomes for fusion with endosomes/lysosomes. Cell 151(6):1256–1269
Jin M, He D, Backues SK, Freeberg MA, Liu X, Kim JK, Klionsky DJ (2014) Transcriptional regulation by Pho23 modulates the frequency of autophagosome formation. Curr Biol 24:1314–1322
Karanasios E, Walker SA, Okkenhaug H, Manifava M, Hummel E, Zimmermann H, Ahmed Q, Domart MC, Collinson L, Ktistakis NT (2016) Autophagy initiation by ULK complex assembly on ER tubulovesicular regions marked by ATG9 vesicles. Nat Commun 7:12420
Kast DJ, Dominguez R (2017) The cytoskeleton-autophagy connection. Curr Biol 27(8):R318–R326
Kenific CM, Debnath J (2016) NBR1-dependent selective autophagy is required for efficient cell-matrix adhesion site disassembly. Autophagy 12(10):1958–1959
Koyano F, Okatsu K, Kosako H, Tamura Y, Go E, Kimura M, Kimura Y, Tsuchiya H, Yoshihara H, Hirokawa T, Endo T, Fon EA, Trempe JF, Saeki Y, Tanaka K, Matsuda N (2014) Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510(7503):162–166
Kraft LJ, Dowler J, Manral P, Kenworthy AK (2016) Size, organization, and dynamics of soluble SQSTM1 and LC3-SQSTM1 complexes in living cells. Autophagy 12:1660–1674
Krols M, Bultynck G, Janssens S (2016) ER-mitochondria contact sites: a new regulator of cellular calcium flux comes into play. J Cell Biol 214:367–370
Kumar S, Jain A, Farzam F (2018) Mechanism of Stx17 recruitment to autophagosomes via IRGM and mammalian Atg8 proteins. J Cell Biol 217(3):997–1013
Lamark T, Svenning S, Johansen T (2017) Regulation of selective autophagy: the p62/SQSTM1 paradigm. Essays Biochem 61(6):609–624
Lamb CA, Nühlen S, Judith D, Frith D, Snijders AP, Behrends C, Tooze SA (2016) TBC1D14 regulates autophagy via the TRAPP complex and ATG9 traffic. EMBO J 35:281–301
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, Sideris DP, Fogel AI, Youle RJ (2015) The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524(7565):309–314
Lazarus MB, Novotny CJ, Shokat KM (2015) Structure of the human autophagy initiating kinase ULK1 in complex with potent inhibitors. ACS Chem Biol 10:257–261
Li Y, Ding W-X (2017) Impaired Rab7 and Dynamin2 block fat turnover by autophagy in alcoholic fatty livers. Hepatol Commun 1:473–476
Lin MG, Hurley JH (2016) Structure and function of the ULK1 complex in autophagy. Curr Opin Cell Biol 39:61–68
Liu X, Klionsky DJ (2015) TP53INP2/DOR protein chaperones deacetylated nuclear LC3 to the cytoplasm to promote macroautophagy. Autophagy 11(9):1441–1442
Luo S, Garcia-Arencibia M, Zhao R (2012) Bim inhibits autophagy by recruiting Beclin 1 to microtubules. Mol Cell 47(3):359–370
Mackeh R, Perdiz D, Lorin S, Codogno P, Pous C (2013) Autophagy and microtubules - new story, old players. J Cell Sci 126:1071–1080
Maday S, Holzbaur ELF (2014) Autophagosome biogenesis inprimary neuronsfollows anordered and spatially regulated pathway. Dev Cell 30:71–85
Martina JA, Chen Y, Gucek M, Puertollano R (2012) MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8(6):903–914
Morishita H, Kaizuka T, Hama Y, Mizushima N (2017) A new probe to measure autophagic flux in vitro and in vivo. Autophagy 13:757–758
Munson MJ, Allen GF, Toth R et al (2015) mTOR activates the VPS34-UVRAG complex to regulate autolysosomal tubulation and cell survival. EMBO J 34(17):2272–2290
Nakamura S, Yoshimori T (2017) New insights into autophagosome-lysosome fusion. J Cell Sci 130(7):1209–1216
Nakashima H, Nguyen T, Goins WF, Chiocca EA (2015) Interferon-stimulated gene 15 (ISG15) and ISG15-linked proteins can associate with members of the selective autophagic process, histone deacetylase 6 (HDAC6) and SQSTM1/p62. J Biol Chem 290(3):1485–1495
Nascimbeni AC, Giordano F, Dupont N, Grasso D, Vaccaro MI, Codogno P, Morel E (2017) ER-plasma membrane contact sites contribute to autophagosome biogenesis by regulation of local PI3P synthesis. EMBO J 36:2018–2033
Nguyen TN, Padman BS, Usher J, Oorschot V, Ramm G, Lazarou M (2016) Atg8 family LC3/GABARAP proteins are crucial for autophagosome-lysosome fusion but not autophagosome formation during PINK1/Parkin mitophagy and starvation. J Cell Biol 215(6):857–874
Nowak J, Archange C, Tardivel-Lacombe J, Pontarotti P, Pébusque MJ, Vaccaro MI, Velasco G, Dagorn JC, Iovanna JL (2009) The TP53INP2 protein is required for autophagy in mammalian cells. Mol Biol Cell 20:870–881
Okatsu K, Kimura M, Oka T, Tanaka K, Matsuda N (2015) Unconventional PINK1 localization to the outer membrane of depolarized mitochondria drives Parkin recruitment. J Cell Sci 128(5):964–978
Olsvik HL, Lamark T, Takagi K, Larsen KB, Evjen G, Øvervatn A, Mizushima T, Johansen T (2015) FYCO1 contains a C-terminally extended, LC3A/B-preferring LC3-interacting region (LIR) motif required for efficient maturation of autophagosomes during basal autophagy. J Biol Chem 290(49):29361–29374
Olszewska DA, Lynch T (2016) PINK1, parkin, and autophagy receptors: a new model of mitophagy. Mov Disord 31(11):1628–1629
Orsi A, Razi M, Dooley HC, Robinson D, Weston AE, Collinson LM, Tooze SA (2012) Dynamic and transient interactions of Atg9 with autophagosomes, but not membrane integration, are required for autophagy. Mol Biol Cell 23:1860–1873
O'Sullivan TE, Johnson LR, Kang HH et al (2015) BNIP3- and BNIP3L-mediated mitophagy promotes the generation of natural killer cell memory. Immunity 43(2):331–342
Pankiv S, Alemu EA, Brech A, Bruun JA, Lamark T, Øvervatn A, Bjørkøy G, Johansen T (2010) FYCO1 is a Rab7 effector that binds to LC3 and PI3P to mediate microtubule plus end-directed vesicle transport. J Cell Biol 188(2):253–269
Peng H, Yang J, Li G, You Q, Han W, Li T, Gao D, Xie X, Lee BH, du J, Hou J, Zhang T, Rao H, Huang Y, Li Q, Zeng R, Hui L, Wang H, Xia Q, Zhang X, He Y, Komatsu M, Dikic I, Finley D, Hu R (2017) Ubiquitylation of p62/sequestosome1 activates its autophagy receptor function and controls selective autophagy upon ubiquitin stress. Cell Res 27(5):657–674
Piano Mortari E, Folgiero V, Marcellini V, Romania P, Bellacchio E, D'Alicandro V, Bocci C, Carrozzo R, Martinelli D, Petrini S, Axiotis E, Farroni C, Locatelli F, Schara U, Pilz DT, Jungbluth H, Dionisi-Vici C, Carsetti R (2018) The Vici syndrome protein EPG5 regulates intracellular nucleic acid trafficking linking autophagy to innate and adaptive immunity. Autophagy 14(1):22–37
Proikas-Cezanne T, Takacs Z, Donnes P et al (2016) WIPI proteins: essential PtdIns3P effectors at the nascent autophagosome. J Cell Sci 128:207–217
Pu J, Guardia CM, Keren-Kaplan T, Bonifacino JS (2016) Mechanisms and functions of lysosome positioning. J Cell Sci 129(23):4329–4339
Rabanal-Ruiz Y, Otten EG, Korolchuk VI (2017) mTORC1 as the main gateway to autophagy. Essays Biochem 61(6):565–584
Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC (2010) Plasma membrane contributes to the formationof pre-autophagosomal structures. Nat Cell Biol 12:747–757
Romero M, Sabaté-Pérez A, Francis VA, Castrillón-Rodriguez I, Díaz-Ramos Á, Sánchez-Feutrie M, Durán X, Palacín M, Moreno-Navarrete JM, Gustafson B, Hammarstedt A, Fernández-Real JM, Vendrell J, Smith U, Zorzano A (2018) TP53INP2 regulates adiposity by activating β-catenin through autophagy-dependent sequestration of GSK3β. Nat Cell Biol 20:443–454. https://doi.org/10.1038/s41556-018-0072-9
Kang R, Livesey KM, Zeh HJ et al (2010) HMGB1: a novel Beclin 1-binding protein active in autophagy. Autophagy 6(8):1209–1211. https://doi.org/10.4161/auto.6.8.13651
Sanchez-Wandelmer J, Ktistakis NT, Reggiori F (2015) ERES: sites for autophagosome biogenesis and maturation? J Cell Sci 128:185–192
Sasaki T, Lian S, Khan A, Llop JR, Samuelson AV, Chen W, Klionsky DJ, Kishi S (2017) Autolysosome biogenesis and developmental senescence are regulated by both Spns1 and v-ATPase. Autophagy 13(2):386–403
Saxton RA, Sabatini DM (2017) mTOR signaling in growth, metabolism, and disease. Cell 169(2):361–371
Song P, Li S, Wu H, Gao R, Rao G, Wang D, Chen Z, Ma B, Wang H, Sui N, Deng H, Zhang Z, Tang T, Tan Z, Han Z, Lu T, Zhu Y, Chen Q (2016) Parkin promotes proteasomal degradation of p62: implication of selective vulnerability of neuronal cells in the pathogenesis of Parkinson’s disease. Protein Cell 7(2):114–129
Søreng K, Munson MJ, Lamb CA et al (2018) SNX18 regulates ATG9A trafficking from recycling endosomes by recruiting Dynamin-2. EMBO Rep 19(4). https://doi.org/10.15252/embr.201744837
Stadel D, Millarte V, Tillmann KD et al (2015) TECPR2 cooperates with LC3C to regulate COPII-dependent ER export. Mol Cell 60:89–104
Suzuki H, Kaizuka T, Mizushima N, Noda NN (2015) Structure of the Atg101-Atg13 complex reveals essential roles of Atg101 in autophagy initiation. Nat Struct Mol Biol 22(7):572–580
Tait SW, Ichim G, Green DR (2014) Die another way--non-apoptotic mechanisms of cell death. J Cell Sci 127(Pt 10):2135–2144
Tanida I, Ueno T, Kominami E (2004) LC3 conjugation system in mammalian autophagy. Int J Biochem Cell Biol 36:2503–2518
Terawaki S, Camosseto V, Pierre P, Gatti E (2016) RUFY4: Immunity piggybacking on autophagy? Autophagy 12(3):598–600
Tumbarello DA, Manna PT, Allen M, Bycroft M, Arden SD, Kendrick-Jones J, Buss F (2015) The autophagy receptor TAX1BP1 and the molecular motor myosin VI are required for clearance of Salmonella Typhimurium by autophagy. PLoS Pathog 11(10):e1005174
Wang H, Sun HQ, Zhu X, Zhang L, Albanesi J, Levine B, Yin H (2015) GABARAPs regulate PI4P-dependent autophagosome:lysosome fusion. Proc Natl Acad Sci U S A 112(22):7015–7020
Wang Z, Miao G, Xue X, Guo X, Yuan C, Wang Z, Zhang G, Chen Y, Feng D, Hu J, Zhang H (2016) The Vici syndrome protein EPG5 is a Rab7 effector that determines the fusion specificity of autophagosomes with late endosomes/lysosomes. Mol Cell 63(5):781–795
Wei Y, Liu M, Li X, Liu J, Li H (2018) Origin of the autophagosome membrane in mammals. Biomed Res Int 2018:1012789–1012789. https://doi.org/10.1155/2018/1012789
Wen X, Klionsky DJ (2016) An overview of macroautophagy in yeast. J Mol Biol 428(9 Pt a):1681–1699
Wijdeven RH, Janssen H, Nahidiazar L, Janssen L, Jalink K, Berlin I, Neefjes J (2016) Cholesterol and ORP1L-mediated ER contact sites control autophagosome transport and fusion with the endocytic pathway. Nat Commun 7:11808
Li X, Wu X-Q, Deng R, Li DD, Tang J, Chen WD, Chen JH, Ji J, Jiao L, Jiang S, Yang F, Feng GK, Senthilkumar R, Yue F, Zhang HL, Wu RY, Yu Y, Xu XL, Mai J, Li ZL, Peng XD, Huang Y, Huang X, Ma NF, Tao Q, Zeng YX, Zhu XF (2017) CaMKII-mediated Beclin 1 phosphorylation regulates autophagy that promotes degradation of Id and neuroblastoma cell differentiation. Nat Commun 8:1159. https://doi.org/10.1038/s41467-017-01272-2
You Z, Xu Y, Wan W, Zhou L, Li J, Zhou T, Shi Y, Liu W (2019) TP53INP2 contributes to autophagosome formation by promoting LC3-ATG7 interaction. Autophagy 15(8):1309–1321. https://doi.org/10.1080/15548627.2019.1580510
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L, Brunner T, Simon HU (2006) Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol 8:1124–1132
Yu Y, Zhao J (2019) Modulated autophagy by microRNAs in osteoarthritis chondrocytes. Biomed Res Int 2019:1484152
Zalckvar E, Berissi H, Mizrachy L, Idelchuk Y, Koren I, Eisenstein M, Sabanay H, Pinkas-Kramarski R, Kimchi A (2009) DAP-kinase-mediated phosphorylation on the BH3 domain of Beclin 1 promotes dissociation of beclin 1 from Bcl-XL and induction of autophagy. EMBO Rep 10:285–292
Zhong Y, Wang QJ, Li X, Yan Y, Backer JM, Chait BT, Heintz N, Yue Z (2009) Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat Cell Biol 11:468–476
Zientara-Rytter K, Subramani S (2018) AIM/LIR-based fluorescent sensors-new tools to monitor mAtg8 functions. Autophagy 14(6):1074–1078. https://doi.org/10.1080/15548627.2018.1454238
Acknowledgements
The author gives special thanks to the Institute of Biophysics, Chinese Academy of Sciences (CAS), Beijing, China, for technical support of this manuscript. The author strongly apologizes to the investigators for not citing here all original publications due to space limitations.
Author information
Authors and Affiliations
Contributions
AAB contributed in the writing, editing, and revised of whole manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author declares that he has no competing interests.
Ethics approval and consent to participate
The MS is a review paper only and does not involve human participants, human data, or human tissue. So, ethics approval and consent to participate are “Not applicable”.
Consent for publication
“Not applicable”.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key points
• Autophagy is a conserved lysosomal pathway in eukaryotic cells.
• Multistages of autophagy are orchestrated by a set of autophagy proteins.
• Regulation of individual stages of autophagy is complex to understand.
• This review may lead to the treatment options for established diseases.
Rights and permissions
About this article
Cite this article
Al-Bari, M.A.A. A current view of molecular dissection in autophagy machinery. J Physiol Biochem 76, 357–372 (2020). https://doi.org/10.1007/s13105-020-00746-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13105-020-00746-0