Abstract
Acute myeloid leukemia (AML) is one of the most common hematological malignancy that has a high recurrence rate. FIBP was reported to be highly expressed in multiple tumor types. However, its expression and role in acute myeloid leukemia remains largely unknown. The aim of this study was to clarify the role and value of FIBP in the diagnosis and prognosis, and to analyze its correlation with immune infiltration in acute myeloid leukemia by The Cancer Genome Atlas (TCGA) dataset. FIBP was highly expressed in AML samples compared to normal samples. The differentially expressed genes were identified between high and low expression of FIBP. The high FIBP expression group had poorer overall survival. FIBP was closely correlated with CD4, IL-10 and IL-2. The enrichment analysis indicated DEGs were mainly related to leukocyte migration, leukocyte cell–cell adhesion, myeloid leukocyte differentiation, endothelial cell proliferation and T cell tolerance induction. FIBP expression has significant correlation with infiltrating levels of various immune cells. FIBP could be a potential targeted therapy and prognostic biomarker associated with immune infiltrates for AML.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Acute myeloid leukemia (AML) is the most common adult heterogeneous hematological malignancy that arises from clonal expansion of transformed hematopoietic stem and progenitor cells. It is associated with genomic alterations in cell proliferation and differentiation [1, 2]. It has a high incidence accounts for approximately 60% of all leukemia [3]. It seriously endangers human health and life. Chemotherapies are the main treatment for acute myeloid leukemia [3, 4]. However, there are still poor prognosis and short disease-free survival after chemotherapy. Therefore, it is urgent to find feasible molecular target for AML to complement existing therapeutic strategies.
FGF1 intracellular binding protein (FIBP) has been reported to be an intracellular protein and could bind to the acidic fibroblast growth factor (aFGF), which participated in cell proliferation by stimulating mitogenesis [5, 6]. FIBP might be involved in mitogenic activity and cell proliferation. The depletion of FIBP in breast cancer cells exhibited impaired proliferation and decreased cellular migration [7]. FIBP also increased tumorigenicity and induced chemotherapy resistance in colorectal cancer cells [8]. FIBP was highly expressed in tumors and a negative marker of antitumor T cells in solid tumors. FIBP KO enhanced T cell antitumor efficacy through downregulation of cholesterol metabolism [8, 9]. However, the role of FIBP in acute myeloid leukemia remains largely unknown.
Thus, we evaluated the prognostic value of FIBP expression in AML based on TCGA data. We investigated FIBP expression and its correlation with survival in AML patients to understand pathological process and aggressiveness in AML. We further investigated the hub genes and the important role of FIBP in the immune microenvironment through protein-protein interaction network and immune infiltration analysis. This study was expected to provide new targets for AML precise treatment and potential application in predicting AML prognosis.
2 Materials and methods
2.1 Data sources
The expression and clinical data of TCGA pan-cancer and GTEx data were downloaded from the UCSC Xena database [10] (https://xenabrowser.net/datapages/). AML clinical data were downloaded from TCGA database (https://portal.gdc.cancer.gov/). Patients with insufficient clinical information were not included. The RNA-Seq gene expression FPKM (Fragments Per Kilobase per Million) of 151 cases with AML and clinical data were retained and further analyzed. The HTSeq-FPKM data were transformed to TPM (transcription per million reads) for the following analysis. The healthy subjects and AML patient blasts used for ex vivo experiments were obtained from peripheral blood or bone marrow samples collected from Changzhi People’s Hospital, the Affiliated Hospital of Changzhi Medical College. The parents or guardians of each subject provided signed informed consent. The study protocol acquired approval from the ethics committee of Changzhi Medical College (No: RT2023001).
2.2 Analysis of differentially expressed genes
The patients with AML were divided into high or low expression groups according to the median expression value of FIBP in TCGA samples. Expression profiles (level 3 HTSeq-Counts) were compared between high and low FIBP expression groups to identify differentially expressed genes (DEGs) using R Package DESeq2. |logFC|>1and FDR < 0.05 were considered as DEGs [11].
2.3 Functional enrichment analysis
The Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were analyzed for DEGs using the ggplot2 package for visualization and the cluster Profiler package for statistical analysis [12].
2.4 Diagnostic value analysis
The receiver operating characteristic (ROC) curve was used to assess the diagnostic value of FIBP in AML. The area value under the ROC curve is between 0.5 and 1. AUC in 0.5–0.7 has a low accuracy, AUC in 0.7–0.9 has a certain accuracy, and AUC above 0.9 has a high accuracy [13].
2.5 Immune infiltration analysis by ssGSEA
The single sample gene set enrichment analysis (ssGSEA) method was performed using R package GSVA to analyze the immune infiltration of AML for 24 types of immune cells in tumor samples [14]. The relative enrichment score of each immunocyte was quantified from gene expression profile for each tumor sample based on the signature genes of the 24 types immunocyte. The correlation between FIBP and these immune cells was analyzed by Spearman correlation.
2.6 Quantitative real-time PCR
The quantification of the expression of human genes was performed using real-time RT-PCR. The sequences of the primers used for detecting gene expression were as follows: FIBP, sense 5′-TGAGCTGGACATCTTCGTGG-3′, antisense 5′- GGTCACCGAGTAACCATCGAG-3′; GAPDH, sense 5′-TCGTCCCGTAGACAAA ATGG-3′, antisense 5′-TTGAGGTCA ATGAAGGGGTC-3′. For sample analysis, the threshold was set based on the exponential phase of products, and CT value for samples was determined. The resulting data were analyzed with the comparative CT method for relative gene expression quantification against GAPDH (house-keeping gene).
2.7 Western blot analysis
Western blot assay was done as described previously [15]. Antibodies were purchased from ABclonal Technology (Wuhan, China). Briefly, 50 µg of protein was loaded on 10% SDS-Page gel. Following blotting, the blots were incubated with appropriate primary antibodies at 4 °C overnight. Later, the blots were incubated with appropriate HRP conjugated secondary antibodies at room temperature for an hour. ECL reagent was used for imaging the blots.
3 Results
3.1 FIBP expression analysis in pan-cancer and LAML
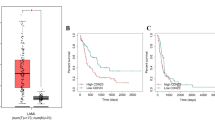
FIBP expression was explored in pan-cancer data from TCGA and GTEx. FIBP expression was significantly upregulated in 28 types of tumors than that in normal tissues, including BLCA, BRCA, CESC, CHOL, COAD, DLBC, ESCA, GBM, HNSC, KIRC, KIRP, LAML, LGG, LIHC, LUAD, LUSC, OV, PAAD, PCPG, PRAD, READ, SKCM, STAD, TGCT, THCA, THYM, UCEC, UCS (P < 0.05), while its expression was no significant difference between tumors and normal tissues including ACC, KICH, MESO, SARC and UVM (Fig. 1A). FIBP expression was further compared in 70 GTEx normal samples and 173 TCGA acute myeloid leukemia samples. FIBP was significantly upregulated in LAML samples (P < 0.05) (Fig. 1B). ROC analysis demonstrated that FIBP had a low diagnostic accuracy with AUC of 0.596 (Fig. 1C).
FIBP expression in pan-cancer and LAML. A FIBP expression between tumor tissues from TCGA and normal tissues from GTEx in pan-cancer. *P < 0.05, ***P < 0.001. B FIBP expression in GTEx normal samples and TCGA LAML samples. *P < 0.05. C ROC curve of FIBP. The area under the curve (AUC) values was considered as follows: AUC = 0.5 indicated noninformative; 0.5 < AUC ≤ 0.7 indicated low accurate; 0.7 < AUC ≤ 0.9 indicated moderately accurate; 0.9 < AUC < 1 indicated highly accurate; AUC = 1 indicated perfect
3.2 Analysis of differentially expressed genes
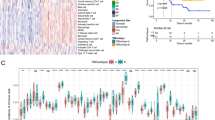
The differentially expressed genes (DEGs) were analyzed using TCGA cohort data and patients with LAML were divided into the high expression and the low expression group based on FIBP levels. A total of 720 differentially expressed genes were screened, including 411 upregulated genes and 309 downregulated genes (Fig. 2A). The gene expression heatmap was obtained for the top 20 differentially expressed genes in the high- and low FIBP-expression LAML patients (Fig. 2B, C).
The differential gene expression map in the TCGA-LAML database. A The volcano plot of DEGs. Each point represents one gene; blue color indicated downregulation and red color indicated upregulation. B The heatmap of the top 20 differentially expressed genes in the high FIBP-expression LAML patients. The blue represent downregulated genes and the red represent upregulated gene. ***P < 0.001. C The heatmap of the top 20 differentially expressed genes in the low FIBP-expression LAML patients. Blue represents low expression, and red represents high expression. The blue represent downregulated genes and the red represent upregulated gene. ***P < 0.001
3.3 GO and KEGG enrichment analysis and PPI network
The GO and KEGG enrichment analysis of DEGs were conducted and the primary BP contained leukocyte migration, extracellular matrix organization, signal release, leukocyte cell-cell adhesion, regulation of blood circulation, tissue remodeling, leukocyte chemotaxis, myeloid leukocyte differentiation, endothelial cell proliferation, granulocyte migration, positive regulation of endothelial cell proliferation, lymphocyte apoptotic process and T cell tolerance induction. The CC was mainly enriched in transporter complex, transmembrane transporter complex, membrane region and membrane microdomain. The MF was primarily involved in G protein-coupled receptor binding, cytokine activity, cytokine receptor binding, growth factor binding, cytokine receptor activity and extracellular matrix binding. The KEGG pathway enrichment was mainly related to cytokine-cytokine receptor interaction, cell adhesion molecules, complement and coagulation cascades and renin-angiotensin system (Fig. 3A, B). Furthermore, the top 10 hub genes of 720 DEGs were identified including HGF, SELE, IL-2, LEP, CD4, HMOX1, MMP2, FN1, CXCL10 and IL-10 (Fig. 3C). The top 5 hub genes were IL-2, IL-10, CXCL10, CD4 and FN1 among them (Fig. 3D). The top 3 hub genes were CD4, IL-10 and IL-2 (Fig. 3E). The relationship between FIBP and the top 10 genes was also explored and the result showed that FIBP had a significant positive correlation with CD4, CXCL10 and HMOX1, whereas FIBP was a significantly negatively correlated with HGF, LEP and SELE. However, no significant correlation was found between FIBP and FN1, IL2, IL10 and MMP2 (Supplementary Fig. S1A). Compared with the normal group, the expression level of HGF, CD4, HMOX1, MMP2 and IL-10 was significantly increased in AML group, whereas the expression level of LEP and FN1 was significantly decreased. There was no significant difference in IL-2 and CXCL10 expression between AML and normal group (Supplementary Fig. S1B).
3.4 Association between FIBP expression and clinicopathological characteristics
Correlation analysis revealed that FIBP expression was significantly associated with WBC count (p < 0.05), PB blasts (p < 0.01), FAB classifications (p < 0.01) and Cytogenetic risk (p < 0.001). No correlation was found between FIBP expression and other clinicopathologic characteristics (Table 1). Univariate logistic regression analysis revealed that FIBP upregulation in LAML was significantly associated with WBC count (p < 0.05), PB blasts (p < 0.01), Cytogenetic risk (p < 0.001), and NPM1 mutation (p < 0.05) (Table 2). The higher FIBP expression was significantly correlated with age (p < 0.05), cytogenetic risk (Favorable vs. Intermediate, Favorable vs. Poor, p < 0.01), FAB classifications (M0 vs. M5, p < 0.01; M2 vs. M5, p < 0.05; M3 vs. M5, p < 0.001), OS event (p < 0.001) and PB blasts (p < 0.05) (Fig. 4).
Associations between the FIBP expression and clinicopathological characteristics. A age (≤ 60 and > 60), B cytogenetic risk (Favorable, Intermediate and Poor), C FAB classifications (M0, M1, M2, M3, M4 and M5), D OS event (Alive and Dead), E PB blasts (≤ 70% and > 70%). *P < 0.05, **P < 0.01, ***P < 0.001
3.5 Prognostic value of FIBP in LAML
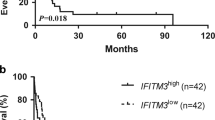
To confirm the correlation between FIBP expression and LAML prognosis, survival rates were compared between the high and low FIBP level groups. The Kaplan–Meier survival analysis indicated that the LAML patients with high FIBP expression had poorer overall survival (HR = 3.77(2.39–5.95), p < 0.001) (Fig. 5A). Multivariate analyses showed that FIBP remained independently associated with overall survival (HR = 3.571(2.191–5.821), p < 0.001), along with age (p < 0.001) in Table 3. The age and FIBP expression were included in the nomogram based on Cox proportional hazards regression model (Fig. 5B). The calibration plots were constructed to evaluate the agreement between predicted and actual OS for the prognosis model, and the results showed that the predicted results of the nomogram were reliable (Fig. 5C).
Analysis of prognostic value of FIBP in LAML. A Overall survival curve of LAML patients with high and low FIBP expression levels. HR: hazard ratio. B Nomogram for predicting the probability of 1-, 3-, and 5-year OS for LAML patients. C Calibration plot of the nomogram for predicting the probability of OS at 1, 3, and 5 years
3.6 Relationship between FIBP expression and tumor-infiltrating immune cells
To confirm whether FIBP expression was associated with tumor-infiltrating immune cells in LAML, Spearman correlation was performed to show the association between the expression of FIBP and the GSVA enrichment scores of immune cell infiltration calculated from RNA-seq in LAML tumor microenvironment. FIBP was positively correlated with aDC, eosinophils, iDC, neutrophils, NK CD56dim cells, NK CD56birght cells, NK cells, TFH and Treg (Fig. 6A–I), whereas it was negatively correlated with Tcm, T cells and T helper cells (Fig. 6J–L).
Relationship between FIBP expression and tumor-infiltrating immune cells. A aDC, B Eosinophils, C iDC, D Neutrophils, E NK CD56dim cells, F NK CD56bright cells, G NK cells, H TFH, I Treg, J Tcm, K T cells and L T helper cells. r: spearman’s correlation coefficient, r < 0 was considered as a negative correlation, and r > 0 was considered a positive correlation. P < 0.05 means statistically significant
3.7 Expression validation for FIBP gene in human acute myeloid leukemia
To further investigate FIBP expression in AML patients, qPCR and Western blot were performed and showed FIBP high expression in AML patients compared with the healthy control (Fig. 7A, B and Supplementary Fig.S2).
4 Discussion
FIBP was an intracellular protein binding selectively to acidic fibroblast growth factor (aFGF), which regulated cell proliferation for multiple cell types by stimulating mitogenesis or inducing morphological changes [6, 16]. Studies have shown that FIBP increased tumorigenicity and was highly expressed in colon carcinoma [17]. FIBP knockdown increased sensitization of chemoresistant cells and attenuated cancer stemness [9, 18]. Moreover, it was showed that FIBP was also highly expressed in skin carcinogenesis and was involved in tumor cell cycle processes by regulating the key downstream target cyclin D1 [19]. To date, the role of FIBP in acute myeloid leukemia has not been investigated.
In this study, bioinformatics analysis based on TCGA data demonstrated that the expression of FIBP was significantly higher in AML samples than normal samples, indicating that FIBP played a role in tumorigenesis and progression. In addition, ROC analysis showed that FIBP might be a potential diagnostic biomarker. The relationship between FIBP expression and clinicopathological factors was further explored, and high FIBP protein expression was significantly associated with age (p < 0.05), cytogenetic risk (p < 0.01), FAB classifications (p < 0.001), OS event (p < 0.001) and PB blasts (p < 0.05). Kaplan–Meier survival analysis indicated that the high expression of FIBP was correlated with poorer overall survival times. Multivariate Cox regression analysis showed that FIBP was an independent prognostic factor affecting survival of AML patients (P < 0.001).
To explore the biological functions of FIBP, DEGs were analyzed based on AML patients with high or low FIBP expression from TCGA data. A total of 720 differentially expressed genes were identified and the functional enrichment analysis of these DEGs was performed in AML samples. The results demonstrated that these DEGs were mainly enriched in BP terms associated with leukocyte migration, extracellular matrix organization, signal release, leukocyte cell–cell adhesion, regulation of blood circulation, tissue remodeling, leukocyte chemotaxis, myeloid leukocyte differentiation, endothelial cell proliferation, granulocyte migration, positive regulation of endothelial cell proliferation, lymphocyte apoptotic process and T cell tolerance induction. MF was primarily involved in G protein-coupled receptor binding, cytokine activity, cytokine receptor binding, growth factor binding, cytokine receptor activity and extracellular matrix binding. It has been reported that Interactions between AML blasts and their adjacent endothelial cells in the bone marrow microenvironment were important for chemotherapy sensitivity [20]. AML cells have been confirmed to secrete angioregulatory mediators for stimulating endothelial cell proliferation and inducing angiogenesis [21, 22]. Moreover, the chemotherapy-resistant leukemic cells were surrounded by stromal cells, which promote AML cells survival by enabling them to evade immune destruction [23]. Therefore, FIBP may be essential for promoting AML proliferation and angiogenesis by these biological processes and pathways.
AML is highly dependent on the immune microenvironment for survival and growth [24, 25]. Therefore, the difference in immune cell infiltration between patients with high and low FIBP expression was compared in this study. FIBP was negatively correlated with Tcm (R = − 0.290, p < 0.001), T cells (R = − 0.180, p = 0.027) and T helper cells (R = − 0.232, p = 0.004), while it was positively correlated with aDC (R = 0.239, p = 0.003), Eosinophils (R = 0.214, p = 0.008), iDC (R = 0.214, p = 0.008), Neutrophils (R = 0.163, p = 0.045), NK CD56dim cells (R = 0.290, p < 0.001), NK CD56 bright cells (R = 0.390, p < 0.001), NK cells (R = 0.176, p = 0.031), TFH (R = 0.240, p = 0.003) and Treg (R = 0.310, p < 0.001). Multiple clinical studies have demonstrated various disruptions of T cell immunity in AML. T cell numbers and functions are altered to favor the progression of acute myeloid leukemia. A higher frequency of Tregs could impair the cell-mediated anti-leukemia immune response and was considered as a pivotal regulator of immune escape [26,27,28,29]. FIBP high expression may inhibit T cells and T helper cells numbers and increase the frequency of Treg cells to promote AML development. FIBP knockout consistently promoted T cell-mediated cancer killing and significantly reduced tumor size [8]. On the other hand, it has been reported that AML was also capable of inhibiting NK cell maturation and effector function and the loss of peripheral CD56 bright NK cells were found in AML patients [30, 31]. Importantly, NK cells are a type of innate lymphoid cell (ILC) and AML microenvironment creates the possibility of disrupting this balance of ILCs to drive the development of other ILC subsets at the expense of cytotoxic NK cells [32]. Thus, FIBP high expression was positively correlated with NK cells, but FIBP expression possibly increased NK cells with developmental defects.
In conclusion, these findings in this study determined FIBP may be a potential poor prognostic biomarker, which could aid clinicians in clinical application, assessment and therapeutics for AML. Future researches are required to include experiments in vivo and in vitro and enroll more patients to further verify these conclusions.
Data availability
All relevant data are within the paper and TCGA database: https://portal.gdc.cancer.gov/.
References
Piotr O, Anna K, Jakub W, Julia T, Monika L, Zawitkowska JZ. Molecular-targeted therapy of Pediatric Acute myeloid leukemia. Molecules. 2022;27(12):3911.
Huang XB, Zhou D, Ye XJ, Jin J. A novel ferroptosis-related gene signature can predict prognosis and influence immune microenvironment in acute myeloid leukemia. Bosn J Basic Med Sci. 2022;22(4):608–28.
Su YZ, Wang CB, Zhou Y, Sun NT. Effects of changes in serum endostatin and fibroblast growth factor 19 on the chemotherapeutic sensitivity in acute myeloid leukemia patients. Genet Mol Res. 2015;14(2):5181–7.
Martner A, Thorén FB, Aurelius J, Hellstrand K. Immunotherapeutic strategies for relapse control in acute myeloid leukemia. Blood Rev. 2013;27(5):209–16.
Kolpakova E, Wiedlocha A, Stenmark H, Klingenberg O, Falnes PO, Olsnes S. Cloning of an intracellular protein that binds selectively to mitogenic acidic fibroblast growth factor. Biochem J. 1998;336:213–22.
Tassi E, AI-Attar A, Aigner A, Swift MR, McDonnell K, Karavanov A, et al. Enhancement of fibroblast growth factor (FGF) activity by an FGF-binding protein. J Biol Chem. 2001;276(43):40247–53.
Xu SB, Li X, Gong ZH, Wang WQ, Li YJ, Nair BC, et al. Proteomic analysis of the human cyclin-dependent kinase family reveals a novel CDK5 complex involved in cell growth and migration. Mol Cell Proteomics. 2014;13(11):2986–3000.
Zhang Y, Vu T, Palmer DC, Kishton RJ, Gong LQ, Huang J, et al. A T cell resilience model associated with response to immunotherapy in multiple tumor types. Nat Med. 2002;28(7):1421–31.
Huang YF, Niu WN, Hu R, Wang LJ, Huang ZY, Ni SH, et al. FIBP knockdown attenuates growth and enhances chemotherapy in colorectal cancer via regulating GSK3β-related pathways. Oncogenesis. 2018;7(9):77.
Vivian J, Rao AA, Nothaft FA, Ketchum C, Armstrong J, Novak A, et al. Toil enables reproducible, open source, big biomedical data analyses. Nat Biotechnol. 2017;35(4):314–6.
Ritchie ME, Phipson B, Wu D, Hu YF, Law CW, Shi W, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47.
Yu G, Wang LG, Han YY, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7.
Ding B, Lou W, Liu JX, Li RH, Chen J, Fan WM. Silico analysis excavates potential biomarkers by constructing miRNA-mRNA networks between non-cirrhotic HCC and cirrhotic HCC. Cancer Cell Int. 2019;19:186.
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNAseq data. BMC Bioinformatics. 2013;14:7.
Kondo H, Ratcliffe Colin DH, Hooper S, Ellis J, MacRae JI, Hennequart M, et al. Single-cell resolved imaging reveals intra-tumor heterogeneity in glycolysis, transitions between metabolic states, and their regulatory mechanisms. Cell Rep. 2021;34(7):108750.
Schulze D, Plohmann P, Hobel S, Aigner A. Anti-tumor effects of fibroblast growth factor-binding protein (FGF-BP) knockdown in colon carcinoma. Mol Cancer. 2011;10:144.
Tassi E, Henke RT, Bowden ET, Swift MR, Kodack DP, Kuo AH, et al. Expression of a fibroblast growth factor–binding protein during the development of adenocarcinoma of the pancreas and colon. Cancer Res. 2006;66(2):1191–8.
Jeter CR, Liu B, Liu X, Chen X, Liu C, Calhoun-Davis T, et al. NANOG promotes cancer stem cell characteristics and prostate cancer resistance to androgen deprivation. Oncogene. 2011;30(36):3833–45.
Bhoumik A, Fichtman B, Derossi C, Breitwieser W, Kluger HM, Davis S, et al. Suppressor role of activating transcription factor 2 (ATF2) in skin cancer. Proc Natl Acad Sci USA. 2008;105(5):1674–9.
Zhang JR, Ye JJ, Ma DX, Liu N, Wu H, Yu S, et al. Cross-talk between leukemic and endothelial cells promotes angiogenesis by VEGF activation of the Notch/Dll4 pathway. Carcinogenesis. 2013;34(3):667–77.
Reikvam H, Hatfield KJ, Oyan AM, Kalland KH, Kittang AO, Bruserud O. Primary human acute myelogenous leukemia cells release matrix metalloproteases and their inhibitors: release profile and pharmacological modulation. Eur J Haematol. 2010;84(3):239–51.
Hatfield K, Øyan AM, Ersvaer E, Kalland KH, Lassalle P, Gjertsen BT, et al. Primary human acute myeloid leukaemia cells increase the proliferation of microvascular endothelial cells through the release of soluble mediators. Br J Haematol. 2009;144(1):53–68.
Milojkovic D, Devereux S, Westwood NB, Mufti GJ, Thomas NSB, Buggins AGS. Antiapoptotic microenvironment of acute myeloid leukemia. J Immunol. 2004;173(11):6745–52.
Guerrouahen BS, Al-Hijji I, Tabrizi AR. Osteoblastic and vascular endothelial niches, their control on normal hematopoietic stem cells, and their consequences on the development of leukemia. Stem Cells Int. 2011; 2011: 375857.
Vijay V, Miller R, Vue GS, Pezeshkian MB, Maywood M, Ast AM, et al. Interleukin-8 blockade prevents activated endothelial cell mediated proliferation and chemoresistance of acute myeloid leukemia. Leuk Res. 2019;84:106180.
Hao F, Sholy C, Wang C, Cao M, Kang XL. The role of T cell immunotherapy in Acute myeloid leukemia. Cells. 2021;10:12.
Ustun C, Miller JS, Munn DH, Weisdorf DJ, Blazar BR. Regulatory T cells in acute myelogenous leukemia: is it time for immunomodulation? Blood. 2011;118(19):5084–95.
Pyzer AR, Stroopinsky D, Rajabi H, Washington A, Tagde A, Coll M, et al. MUC1-mediated induction of myeloid-derived suppressor cells in patients with acute myeloid leukemia. Blood. 2017;129(13):1791–801.
Wan YL, Zhang CX, Xu YX, Wang M, Rao Q, Xing HY, et al. Hyperfunction of CD4 CD25 regulatory T cells in de novo acute myeloid leukemia. BMC Cancer. 2020;20(1):472.
Lordo MR, Wu KG, Altynova E, Shilo N, Kronen P, Nalin AP, et al. Acute myeloid leukemia alters Group 1 innate lymphoid cell differentiation from a common precursor. J Immunol. 2021;207(6):1672–82.
Mundy-Bosse BL, Scoville SD, Chen L, McConnell K, Mao HC, Ahmed EH, et al. MicroRNA-29b mediates altered innate immune development in acute leukemia. J Clin Invest. 2016;126(12):4404–16.
Scoville SD, Nalin AP, Chen LX, Chen L, Zhang MH, McConnell K, et al. Human AML activates the aryl hydrocarbon receptor pathway to impair NK cell development and function. Blood. 2018;132(17):1792–804.
Funding
This work was supported by Scientific and Technologial Innovation Programs of Higher Education Institutions in Shanxi (STIP) [Grant number: 2021L341, 2020L0385] and Fund Program for the Scientific Activities of Selected Returned Overseas Professionals in Shanxi Province [Grant number: 20220035].
Author information
Authors and Affiliations
Contributions
Conceptualization: Gang Chi, Muya Ma. Data curation: Muya Ma, Lingling Xu. Formal analysis: Wenhua Cui, Yan Huang. Funding acquisition: Gang Chi. Supervision: Gang Chi, Wenhua Cui, Yan Huang. Validation: Gang Chi. Visualization: Muya Ma, Lingling Xu. Writing—original draft: Muya Ma, Lingling Xu, Gang Chi. Writing—review & editing: Muya Ma, Gang Chi.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All procedures involving human participants were performed in accordance with the Declaration of Helsinki (as revised in 2013). This study was approved by the ethics committee of Changzhi Medical College (No: RT2023001).
Consent for publication
All authors agree to publish this paper.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, M., Xu, L., Cui, W. et al. FIBP is a prognostic biomarker and correlated with clinicalpathological characteristics and immune infiltrates in acute myeloid leukemia. Discov Onc 14, 97 (2023). https://doi.org/10.1007/s12672-023-00723-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-023-00723-1