Abstract
Snow crab (Chionoecetes opilio) by-products are a rich source of biomolecules, such as lipids, proteins, and chitin, which have not been extensively investigated. This study aims to identify antibacterial peptides to enhance the value of C. opilio by-products. After hydrolysis of different component parts using Protamex®, and concentration by solid-phase extraction, the resulting fractions were tested for antibacterial activity against Escherichia coli, Listeria innocua, and Vibrio parahaemolyticus. Hepatopancreas was the only tissue to display antibacterial activity detected using this protocol. Four fractions obtained with and without enzymatic hydrolysis of hepatopancreas followed by SPE C18 fractionation and elution with 50 and 80% acetonitrile demonstrated bacteriostatic activity against L. innocua HPB13, from concentrations of 0.30 to 43.05 mg/mL of peptides/proteins. Eleven peptides sharing at least 80% amino acid homology with four antimicrobial peptides were identified by mass spectrometry. Two peptides had homology to crustin-like and yellowfin tuna GAPDH antimicrobial peptides belonging to the marine organisms Penaeus monodon and Thunnus albacares, respectively. Other peptide sequence homologies were also identified: Odorranain-C7 from the frog Odorrana grahami and a predicted antibacterial peptide in the Asian ladybeetle Harmonia axyridis. These active peptides may represent a novel group of bioactive peptides deserving further investigation as food preservatives.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Canada’s long coastline from the Atlantic Ocean to the Pacific Ocean means access to a large variety of marine products for processing industries. According to the ministère de l’Agriculture des Pêcheries et de l’Alimentation Québec (MAPAQ), of all crustaceans, the snow crab, Chionoecetes opilio, represented the highest export value at about 55,360 t ($683 M) in 2014 [1]. C. opilio is principally found in three provinces: Newfoundland and Labrador, Nova Scotia, and Quebec [2]. Its exploitation generates important volumes of by-products, such as hemolymph (HE), shell (SH), hepatopancreas (HP), or gills (GI), corresponding to around 35% of the total crustacean biomass [3]. By-products are discarded, causing environmental concerns [4] which could be solved by finding new valorization channels. Otherwise, marine organisms are rich in bioactive compounds [5] that could represent a valuable source of food with high nutritional value [6]. These compounds could be used as functional food ingredients to increase positive effects on human health [5].
Previous studies have identified marine bioactive compounds, such as oligosaccharides [7], phospholipids [8], vitamins, carotenoids, minerals, fiber [9], and also steroids, terpenoids, proteins, and polypeptides [10]. Indeed, in marine invertebrates, antimicrobial peptides (AMPs) are the most abundant component involved in innate immunity, generating both humoral and cellular responses [11]. These AMPs can be native [12] or produced by enzymatic hydrolysis, which releases active peptides from their parent protein [13]. Antimicrobial peptides are divided into six classes (α-helices, small proteins and β-sheets, peptides with thio-ether rings, macrocyclic cystine knot peptides, lipopeptides, and peptides with one or two repeated amino acids) and have a molecular weight of less than 10 kDa [11]. Most AMPs are cationic and amphiphilic with both hydrophilic and hydrophobic surfaces [11]. Their antimicrobial effect is caused by forming pores in the microbial membrane or disturbing the membrane integrity of microorganisms [14].
Previous studies have reported the presence, in crustaceans, of AMPs in HE [15], respiratory organs, and gut because these tissues are regularly exposed to pathogens [11]. Crustacean AMPs exhibited antimicrobial activity mediated by positive charges, for example, crustins from Penaeus monodon hemocytes [16] and anti-lipopolysaccharide factor (ALF) isolated from Miyakea nepa hemocytes [17]. Other AMPs identified which were particularly active against bacteria include penaeidins from Litopenaeus vannamei HE, astacidin from the crayfish Pacifastacus leniusculus hemocyanin, and Ls-stylicin-1 from Litopenaeus stylirostris [18, 19] and fungi [17].
In crab, many AMPs have been identified and characterized, such as tachyplesin I, polyphemusins I and II, hyastatin, arasin-1, callectin, and crustin [15, 20,21,22]. Tachyplesin, a cationic peptide, and polyphemusins I and II have been isolated from hemocytes of the horseshoe crab, Tachypleus tridentatus. These peptides have been characterized by three tandem repeats of a tetrapeptide sequence, which contributes to their biological activity through antimicrobial effects against Gram-negative and Gram-positive bacteria and fungi, such as Candida albicans M9 [12, 15]. Thus, hyastatin, an 11.7-kDa glycine-rich peptide, and arasin 1, a proline-arginine-rich antimicrobial peptide, have been both isolated from Hyas araneus hemocytes and demonstrated antimicrobial activity against Gram-positive and Gram-negative bacteria [20, 21]. Callinectin, with 32 amino acids, is a proline-arginine-rich AMP with a primary structure similar to arasin and has been isolated and identified in the blue crab [23]. In addition, crustin, a cysteine-rich 11.5-kDa protein isolated from hemocytes of the shore crab, Carcinus maenas, exhibited antibacterial effects against Gram-positive marine strain bacteria, such as Micrococcus luteus and Aerococcus viridans [22].
Other AMPs have been found in C. opilio by-products. In fact, a recent study has shown that these by-products represent a source of antibacterial peptide fractions with copper residues and possible glycosides part [24]. These hydrophobic peptides, generated after enzymatic hydrolysis of C. opilio by-products, were negatively charged with a low molecular weight around 800 Da and had antibacterial effects against Gram-positive and Gram-negative bacteria [24]. However, the antimicrobial peptides and the tissues producing them have not yet been elucidated.
This work highlights, for the first time, the protein precursors of antibacterial peptides originating from C. opilio by-products, specifically from HP. Fractions obtained from hydrolyzed component parts were characterized to determine the degree of hydrolysis (DH), total protein content, peptide concentration in an extracted aqueous fraction (soluble peptides), and antibacterial activity using indicative strains. Active fractions were analyzed by liquid chromatography coupled with mass spectrometry (LC-MS/MS) to identify active peptides.
Materials and Methods
Snow Crab Samples
Nine crabs fished from the Gulf of St. Lawrence (Eastern coast Canada) were purchased from a fish shop in Rimouski (QC, Canada) in May 2015. The crabs were dissected on ice, and the component parts, HE, HP, SH, GI, claws (CL), and legs (LE), were separated, collected immediately, and treated as follows. The HE was filtered through cheesecloth, poured into a tube, and frozen in liquid nitrogen before being stored at − 80 °C. The other component parts were lightly blotted to remove water, immersed in liquid nitrogen for 30 s, then homogenized by cryomilling using a 12-mm stainless steel grinding ball in a Mixer Mill MM 400 system (Retsch, Germany) [25]. Each powdered sample was transferred into a cryotube and stored at − 80 °C.
Protein and Moisture Content
Each component part (100 mg) obtained from snow crab dissection was analyzed in duplicate by the Dumas method [26] using a LECO FP-2000 carbon and nitrogen analyzer (Truspec N, Leco®, St. Joseph, Michigan, USA) to characterize the total nitrogen (Ntot). A nitrogen conversion factor of 6.25 was used to calculate the total protein content (% P), as shown in the following equation [6]:
The percentage of chitin (%Chitin) in SH was obtained using the Spinelli method [27] with extraction and demineralization treatment using 2% NaOH (v/v) and 5% HCl (v/v), respectively [28]. The percentage of nitrogen in SH was measured by subtracting the percentage of nitrogen in chitin (NChitin) multiplied by the %Chitin from Ntot. Analyses were performed in duplicate, and the total protein content was expressed as a percentage of nitrogen multiplied by the protein conversion factor 6.25 [6].
The moisture of each component part (100 mg) was determined by weighing before and after incubation at 60 °C overnight. The moisture content analysis was performed in duplicate and calculated according to the following equation [29]:
Hydrolysis of Samples
Enzymatic Hydrolysis
Snow crab component parts were hydrolyzed separately. Briefly, commercial enzyme Protamex® (Novozymes, Bagsvaerd, Denmark; 1 mg/mL) was added to each sample at a ratio of 1:1 (v/w). Depending on the nature and the viscosity of each component part, it was diluted in different volumes of distilled water (1:2 for HE, 1:10 for SH, and 1:6 for the other component parts, all (w/v)). Samples with Protamex® were incubated at 40 °C and constantly agitated at 325 rpm (Innova 44R incubator shaker, New Brunswick Scientific, Edison, NJ, USA). After one hour, the hydrolysis was stopped by transferring samples to a water bath at 85 °C for 10 min. Soluble peptides/proteins were extracted overnight at 4 °C using 5 mM HCl and centrifuged at 21,000 ×g for 10 min to collect the aqueous fraction [30]. The supernatant was kept at 4 °C for 2 h until use in antibacterial assays. For each component part, controls, corresponding to the non-hydrolyzed component parts (without Protamex®), were subjected to the same treatments.
Degree of Hydrolysis
The DH was calculated in triplicate according to Church (1983) using o-phthaldialdehyde to measure the amino acids generated after sample hydrolysis [31]. The calculation was performed as described in the following equation [30]:
where htot corresponds to the total number of peptide bonds of protein equivalent and h is the number of hydrolyzed bonds. htot = 8 and h = [((ODsample − ODblank) / (ODstandard − ODblank)) × 0.9516 × ((V × 100) / (X × P)) − b] / a, with a = 1, b = 0.4, P = % protein in the sample, X = (g) sample, and V = (L).
Soluble Protein/Peptide Concentrations
The soluble protein/peptide concentration in each supernatant sample was determined in triplicate using the bicinchoninic acid (μBCA) method (BCA Protein Assay kit, Pierce, Fisher, MA, USA) according to the supplier’s guidelines.
Purification of Fractions
Solid-Phase Extraction
The fractionation of supernatant samples was performed by solid-phase extraction (SPE) (Sep-Pak C18 1 g cartridges, Waters, Life Sciences, MA, USA). Briefly, after cartridge equilibration in methanol (100%) followed by deionized water, samples were loaded and sequentially eluted using distilled water containing 50 and 80% (v/v) of acetonitrile (ACN). The samples were dried using a Speed Vac concentrator (Thermo Scientific Savant, Fisher, MA, USA) and stored at − 20 °C until use for antibacterial screening. Dried fractions were reconstituted in 50 mM sodium phosphate buffer at pH 8 and tested for antimicrobial activity as described subsequently. The peptide concentration in each SPE C18 fraction was determined in triplicate using the μBCA method (BCA Protein Assay kit, Pierce, Fisher, MA, USA) according to the supplier’s guidelines.
Antibacterial Screening
Bacterial Strains and Culture Conditions
Based on previous work [24, 32], the three indicative bacterial strains used for antibacterial assays were as follows: Escherichia coli ATCC 25922 (37 °C, Luria Broth), Listeria innocua HPB 13 (30 °C, Tryptic Soy Broth; TSB supplemented with 0.6% yeast extract (YE)), and Vibrio parahaemolyticus ATCC 17802 (30 °C, TSB + 0.6% YE).
Antibacterial Assay
Antibacterial activity against the Gram-positive and Gram-negative bacterial strains noted previously was tested in duplicate. The inhibitory activity of samples was determined by liquid growth inhibition assays using a microdilution method [24]. Briefly, the samples were centrifuged for 10 min at 15,000 ×g, and serial dilutions were made for each sample by transferring 100 μL of the sample into 100 μL of TSB supplemented with 0.6% YE in a 96-well microplate (Falcon, Corning, NY, USA). Into each well, 50 μL of the appropriate bacterial strain, grown overnight, was added [24]. The kinetics of bacterial growth was recorded using Tecan (Nano Quant Infinite F200 Pro, Life Sciences TECAN) with an optical density (OD) of 595 nm, for 22 h. A positive control of bacterial growth inhibition was performed using gentamycin at 200 μg/mL, and a negative control was performed using bacteria in both the culture medium and the sodium phosphate buffer. The bacterial growth curve was established for all fractions in stock concentrations. Once active fractions were identified, the bioassay was repeated at the same concentrations for comparative purposes (0.3 mg/mL for all non-hydrolyzed fractions and 0.5 mg/mL for all hydrolyzed fractions).
The bacterial count was established for active fractions in two replicates. Briefly, after Tecan incubation, a dilution series have been made until 10−6 in a 0.1% (w/v) peptone solution [33]. A volume of 20 μL from the last dilution was incubated overnight in a Petri plate with the appropriate culture medium and temperature. Bacterial count was calculated using the following equation, and obtained values have been converted into log.
The maximum specific growth rates (μmax) for active fractions were calculated from stock concentrations according to the following equation [34]:
Identification of Antibacterial Peptide
Liquid Chromatography Coupled with Mass Spectrometry Analysis
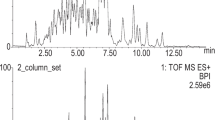
Analyses were performed by the Proteomics Platform of the CHU de Québec-Université Laval Research Center (Quebec, QC, Canada). Briefly, samples were desalted using a C18 Empore filter (Stage-Tip) and then analyzed by LC-MS/MS. Experiments were performed with Ekspert NanoLC425 (Eksigent) coupled to a 5600+ mass spectrometer (Sciex, Framingham, MA, USA) with an ion source. Separation was carried out on picofrit columns (Reprosil 3u, 120A C18, 15 cm × 0.075 mm) followed by elution with a linear gradient (5–35% solvent B (ACN, 0.1% formic acid)) for 35 min at 300 nL/min.
Analyst software version 1.7 was used to establish mass spectra. After the mass spectrum full scanning (400 to 1250 m/z), collision-induced dissociation of the twenty most intense ions was performed with a dynamic exclusion set for 12 s and 50 mDa of tolerance.
Database Searching Mascot
Protein pilot version 5.0 software (Sciex) was used to create MGF peak list files. The files were analyzed by the Mascot (Matrix Science, London, UK; version 2.5.1) using the TAX_Pleocyemata_6692_20150309 database (19,228 entries) with an ion mass tolerance and a parent ion tolerance of 0.1 Da.
Protein Identification
Scaffold (version Scaffold_4.7.3, Proteome Software Inc., Portland, OR) was used to validate the results of MS/MS and protein identification, and peptides with more than 95% probability using the Peptide Prophet algorithm were accepted. Protein identification was accepted if it had greater than 95% homology with other species and contained 2 identified peptides. However, those containing similar peptides, which could not be differentiated using MS/MS analysis, were gathered according to the principles of parsimony.
The peptides obtained were subject to some research on the CAMP R3 Collection Anti-Bacterial peptide online databases to find homology with known active peptides.
Statistical Analysis
Data were analyzed using SAS (University edition, CA). Statistical differences in protein and moisture content, DH, or peptide concentration in samples were determined using Student’s t test. Significant differences were reported at p < 0.001 and p < 0.05. Bacterial growth results were analyzed using the Tukey’s test with significant differences (p < 0.05 and p < 0.001), and bacterial counts results were analyzed by Dunnett’s test with significant differences (p < 0.05).
Results and Discussion
Total Protein Content
The total protein content, expressed as dry weight, varied according to the nature of the snow crab’s component parts. The results (Table 1) indicated a higher protein content in both LE (88.75% ± 2.55) and CL (92.20% ± 1.52), with no significant difference (Student’s t test, p < 0.05). However, the total protein content was lower in HP and SH at 22.36% ± 1.78 and 20.81% ± 0.25, respectively, with no significant difference (Student’s t test, p < 0.05). Of the component parts, CL and LE are known as the richest in muscular proteins [35, 36], while SH is known to have a lower protein content as observed by Shahidi and Synowiecki [37].
The protein content in SH, calculated by subtracting NChitin from Ntot in SH, was 20.81% ± 0.25. The chitin content in C. opilio SH was 13.48% ± 0.79; however, this value is about 8.3% in Atlantic rock crab SH [28, 38], and 12.6–14.5% in green crab SH, using a chemical method, and 5.36% ± 2.1 in snow crab SH, using high-performance liquid chromatography with a refractive index detector [39]. The difference obtained might be due to the difference between crab species and the different method used for snow crab quantification (13.48% ± 0.79 vs 5.36% ± 2.1).
DH and Extraction Efficiency
The peptide/protein concentration in supernatants (soluble peptides/proteins) recovered after extraction was measured for hydrolyzed and non-hydrolyzed component parts to verify the presence of endogenous proteolysis. However, DH was calculated only for the hydrolyzed component parts. The results presented in Table 1 show the highest DH for SH (12.88% ± 1.91) with a small difference of 1.18 mg/mL between peptide/protein concentrations of non-hydrolyzed and hydrolyzed SH. These results suggested that soluble proteinaceous compounds were easily extracted, with or without additional enzymatic treatment (Table 1), with extraction efficiencies of 31.61 and 45.72%, respectively. The highest DH could be explained by protein availability linked to SH rigidity [40]. Proteins are intimately associated with chitin fiber [41], whereas water in SH plays a role in removing protein and salt phases [42]. Indeed, snow crabs were sampled during molting season when the SH was soft, implying an easier hydrolysis after molt [40]. Also, SH rigidity depends on the temperature and food available for crabs [41, 42]. Previous results obtained from crabs fished in previous years exhibited a lower DH (data not shown).
The DH obtained for GI (Table 1) was 3.72% ± 0.09, and the peptide/protein concentration was 1.94 mg/mL after hydrolysis. This low DH could be due to the elastic properties of gill cells and their calcified tissue [43, 44].
Because they are rich in muscular proteins, the highest peptide/protein concentrations (Table 1) occurred in CL (9.84 mg/mL ± 0.23) and LE (8.70 mg/mL ± 0.10), which indicated a noticeable level of enzymatic hydrolysis (3.09% ± 0.13 and 6.93% ± 0.34, respectively). Furthermore, enzymatic hydrolysis using Protamex® increased the extraction efficiency of soluble peptides/proteins. As shown in Table 1, the percentage of extraction for CL, LE, and HE after enzymatic hydrolysis increased at least three-fold. However, the percentage of extraction after hydrolysis for HP, GI, and SH increased only 1.25×, 1.08×, and 1.44×, respectively, consistent with the results obtained for peptide/protein concentrations. These results confirm that peptides obtained in fractions are also due to enzymatic hydrolysis and not just endogenous hydrolysis.
Peptide Purification and Antibacterial Activity
All component parts from C. opilio were hydrolyzed and fractionated by SPE C18 prior to antibacterial activity assays. Antibacterial activity was only detected in non-hydrolyzed HP eluted with 50 or 80% ACN (HP− 50%, HP− 80%) and in hydrolyzed HP eluted with 50 or 80% ACN (HP+ 50%, HP+ 80%), and different concentrations were active against only L. innocua HPB13.
The four bioactive fractions delayed the L. innocua HPB 13 exponential phase (Fig. 1). This delay was more significant in the presence of HP− 50%, HP+ 50%, and HP− 80% at 20.70, 43.05, and 0.30 mg/mL, respectively (Fig. 1), than in the presence of HP+ 80% at 0.50 mg/mL. However, in comparing fraction concentrations and their activities against L. innocua HPB13, it is worthwhile to note that HP− 80% and HP+ 80% fractions had better antibacterial activity, even if the total peptide/protein concentrations were lower. Thus, the HP− 80% fraction may contain native bioactive peptides that are partially degraded after hydrolysis. Above all, other studies have shown that increased DH increases the amount of small peptide below 500 Da, which influences bioactivity [45].
Growth response of Listeria innocua HPB13 in the presence of four active fractions from hepatopancreas (HP) at stock concentrations (HP− 50% (white triangle), HP+ 50% (black triangle), HP− 80% (white circle), and HP+ 80% (black circle)), L. innocua HPB13 (black square), and gentamycin 200 μg/mL (white square)
To normalize the antibacterial results, concentrations of non-hydrolyzed and hydrolyzed fractions were established at 0.30 and 0.50 mg/mL, respectively. At these concentrations, HP− 50% and HP+ 50% fractions lost their antibacterial activities (data not shown).
These results showed that C. opilio HP is a source of antibacterial peptides, as with Homarus americanus HP [46] or Portunus segnis crab HP [47]. Also, it has been suggested that this activity corroborates the presence of highly hydrophobic peptides (such as tachyplesin I from the horseshoe crab) rich in tryptophan residues that allow binding to the hydrophobic, lipopolysaccharide-rich bacterial membrane [48]. Conversely, non-hydrolyzed fractions HP− 50% and HP− 80% have shown antibacterial activity, which supposes the possible presence of native antibacterial peptides. As described by Destoumieux et al., the native peptide, panaeidin, was identified from hemocytes of the marine crustacean, Penaeus vannamei [49]. Other native peptides, rich in hydrophobic amino acids and exhibiting antimicrobial activity, have been found in different crab species, such as tachyplesin in the horseshoe crab Tachypleus tridentatus, hyastatin and arasin 1 in the spider crab Hyas araneus, and callinectin found in the blue crab Callinectes sapidus [20, 21, 23, 48]. Additionally, peptides can be generated by endogenous proteolysis, for example, a proline-rich antibacterial peptide from the shore crab C. maenas [50].
The μmax for four active fractions has been calculated from the same stock concentrations. As seen in Fig. 2, μmax increased at lower concentrations and decreased at higher concentrations. At low concentrations, fractions have a slightly increased L. innocua growth rate that could be due to the presence of nutrients, such as free amino acids, as a carbon source. However, by increasing the concentrations of HP− 50%, HP− 80%, HP+ 50%, and HP+ 80%, from 5.170, 0.075, 10.760, and 0.125 mg/mL, respectively, to reach their stock concentrations, the μmax decreased due to the active peptide concentration. Furthermore, the non-hydrolyzed fractions (HP− 50% and HP− 80%) needed a lower concentration than the hydrolyzed fractions, meaning that the hydrophobic fractions composed of native peptides are possibly more active.
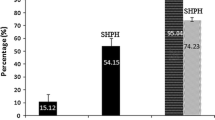
In the presence of these four fractions at stock concentrations, the concentration of L. innocua HPB 13 was significantly decreased for HP− 50%, HP− 80%, and HP+ 80% at 0.28, 0.36, and 0.21 Log (Fig. 3), respectively, compared to the control (Dunnett’s test, p < 0.05). The results obtained suggested that these fractions had a bacteriostatic effect [51] on L. innocua HPB 13 because bacterial colonies could be observed on solid media after incubation.
After separation by RP-HPLC, the HP+ 50% fraction did not show any bioactivity. This suggested that activity probably results from the synergistic action of peptides, which could not have been detected after purification using RP-HPLC [30]. Thus, all active fractions were analyzed directly using LC-MS/MS.
Peptide Characterization
Mass spectrometry results show a higher percentage of potential active peptide in HP− 80% and HP+ 80% (32 and 19%) than in HP− 50% or HP+ 50% (14 and 15%). These results were consistent with those obtained from the antibacterial assays where more antibacterial activity was found in fractions eluted with 80% ACN.
A total of 45 protein precursors identified from peptide sequences corresponded to ribosomal proteins, tissue proteins, and enzymes (Fig. 4). Eleven peptide sequences have a homology percentage greater than 80% with related AMPs (Table 2). The peptides identified in non-hydrolyzed fractions mostly originated from enzymes such as ATP synthase subunit β, superoxide dismutase, and adenosylhomocysteinase (41 and 63% for HP− 50% and HP− 80%, respectively). Peptides identified in hydrolyzed fractions were related to proteins from tissues such as tubulin β, thymosin, β-actin, and myosin heavy chains (46 and 79% for HP+ 50% and HP+ 80%, respectively).
Mascot analysis and scaffold results showed that peptides related to precursor proteins were mostly found in various marine invertebrates, such as the mud crab, Scylla paramamosain, and the giant tiger prawn, P. monodon (Table 2). Peptide sequences were analyzed using Blast on CAMP Collection Anti-Bacterial Peptides online databases and AMP databases to highlight AMPs. No matches with AMPs from the AMP database were found. CAMP R3 Blast tools showed that 11 peptide sequences possessed homology (> 80%) with eight known AMPs. Of these AMPs, five were well-known and related to cationic antimicrobial proteins. On the other hand, the results obtained with a fixed homology of 80% or more did not find a common peptide for all four fractions. However, the crustin-like peptide was common in both HP− 50% and HP− 80% fractions. This antimicrobial peptide has been isolated for the first time from C. maenas (shore crab) hemocytes [22]. Previous studies have shown that crustins are cationic AMPs rich in hydrophobic amino acid, cysteine [53]. This peptide, with an isoelectric point between 8.03 and 8.64 (https: //web.expasy.org/protparam/), has an effect against Gram-positive bacteria and active fractions obtained from HP of snow crab [16]. Several isoforms have been described in many crustaceans and marine organisms [16], such as Crustin1 and 2 that were transcribed in the crayfish, Pacifastacus leniusculus, in the presence of bacteria in HP [54]. It has also been reported that a crustin identified in the blue crab, Callinectes sapidus HP, was induced after cells were stressed by tributyltin [55].
Moreover, in HP− 50% and HP+ 50% fractions, peptide sequences exhibited homologies with two AMPs known as Odorranain-C7 and Yellowfin tuna GAPDH-related antimicrobial peptide (YFGAP). Odorranain-C7 was identified in both non-hydrolyzed and hydrolyzed fractions whereas YFGAP was only found in hydrolyzed fractions. These peptides were identified from the skins of Graham’s frog, Harmonia axyridis, and the yellowfin tuna, Thunnus albacares, respectively [56, 57]. In addition, using a cDNA microarray, it has been shown that Odorranain, is present in Penaeus stylirostris (shrimp) HP and its expression is increased after infection by White Spot Virus (WSV) [58]. Also, glyceraldehyde-3-phosphate dehydrogenase is moderately expressed in P. stylirostris HP after WSV infection [59]. All these AMPs act against Gram-negative and Gram-positive bacteria; however, fractions tested in this study had antibacterial activity against only Gram-positive bacteria. This bioactivity was absent after HPLC purification of the active fractions, suggesting that the peptides may act in synergy. The same suggestion has been reported from Lüders et al. noting the importance of the synergy between cationic and non-cationic peptides in the immune responses of bacterial, invertebrate, and vertebrate species [60]. For instance, the synergy between big defensin and tachycitin in horseshoe crabs increases the antimicrobial activity of big defensin against Gram-negative bacteria.
Another predicted antibacterial peptide was identified from Harmonia axyridis (Table 2) and found in HP+ 50% and HP+ 80% fractions. This peptide is known to be active against both Gram-positive and Gram-negative bacteria and its protein precursor, the elongation factor 1-alpha, known as an apoptotic-related protein in shrimp Litopenaeus vannamei HP, which is up-regulated after WSV infection.
The identification of antibacterial peptides in marine organisms (tuna and giant tiger prawn) with high homology to peptides in our active fractions is promising. Protein precursors of these antibacterial peptides are involved in the initialization of humoral defense in HP of marine crustaceans.
Conclusion
The aim of this study was to demonstrate the potential of C. opilio by-products for increased valorization through the detection of AMPs. We have demonstrated, using Protamex® with DH at 4.89%, that C. opilio HP− and HP+ are sources of antibacterial peptides. SPE fractionation and concentration allowed us to detect antibacterial peptides from HP− 50%, HP− 80%, HP+ 50%, and HP+ 80% fractions that exhibit activity against L. innocua HPB 13. Analysis of SPE fractions using LC-MS/MS and the CAMP R3 database identified 11 sequences with > 80% amino acid homology to antibacterial peptides. These sequences corresponded to crustin-like, YFGAP, Odorranain-C7, and other uncharacterized antibacterial peptides. These results require further investigation such as purification of specific active peptides from the fractions using other biochemical methods or immunological tools. Molecular methods, such as PCR, could be used to amplify the gene coding for crustin-like and YFGAP, followed by sequencing to identify the nucleic acid sequence of precursor genes. Chemical synthesis could also be used to confirm antibacterial activities. The active peptides we obtained after enzymatic hydrolysis could be used as natural preservatives to replace conventional food preservative agents, especially in the context of bioresource valorizations. Known to be caught in the Atlantic and Pacific Oceans as well as in the Arctic Sea, C. opilio’s by-products might also be a source of free amino acids and short peptides in protein hydrolysates that could be used in microbiology, food industry, cosmetics, pharmaceutical, and nutritional products due to their multiple functional properties.
References
MAPAQ (2015) Monographie de l’industrie du crabe des neiges
DFO (2015) Snow crab. http://www.dfo-mpo.gc.ca/fm-gp/sustainable-durable/fisheries-peches/snow-crab-eng.htm. [Accessed: 01-Apr-2017]
Carbonneau ME (2013) Coproduits de crabe des neiges. Merinov. https://www.google.ca/url?sa=t&rct=j&q=&esrc=s&source=web&cd=1&ved=0ahUKEwievZWn08vZAhUL0IMKHSi0CNoQFggpMAA&url=https%253A%252F%252Fwww.merinov.ca%252Ffr%252Fpubli%253Ftask%253Dcallelement%2526format%253Draw%2526item_id%253D739%2526element%253D5683814e-953d-40a2-8d2a-15d5ece9d966%25. [Accessed: 01-Feb-2018]
Bouazza A (2009) Purification, caractérisation et etude des proprieties immunostumulantes de l’hémocyanine de crabe des neiges, Chinoecetes Opilio. Dissertation, Laval University
Lafarga T, Hayes M (2017) Bioactive protein hydrolysates in the functional food ingredient industry: overcoming current challenges. Food Rev Int 33:217–246. https://doi.org/10.1080/87559129.2016.1175013
Beaulieu L, Thibodeau J, Bryl P, Carbonneau ME (2009) Characterization of enzymatic hydrolyzed snow crab (Chionoecetes opilio) by-product fractions: a source of high-valued biomolecules. Bioresour Technol 100:3332–3342. https://doi.org/10.1016/j.biortech.2009.01.073
Jiao G, Yu G, Zhang J, Ewart HS (2011) Chemical structures and bioactivities of sulfated polysaccharides from marine algae. Mar Drugs 9:196–233. https://doi.org/10.3390/md9020196
Sanina NM, Velansky PV, Kostetsky EY (2016) The thermotropic behavior and fatty radical composition of major phospholipids of the tanner crab Chionoecetes bairdi Rathbun, 1924. Russ J Mar Biol 42:81–86. https://doi.org/10.1134/S1063074016010156
Matos J, Cardoso C, Bandarra NM, Afonso C (2017) Microalgae as a healthy ingredient for functional food: a review. Food Funct 8:2672–2685. https://doi.org/10.1039/C7FO00409E
Ng TB, Cheung RCF, Wong JH, Bekhit AA, Bekhit AED (2015) Antibacterial products of marine organisms. Appl Microbiol Biotechnol 99:4145–4173. https://doi.org/10.1007/s00253-015-6553-x
Tincu JA, Taylor SW (2004) Antimicrobial peptides from marine invertebrates. Antimicrob Agents Chemother 48:3645–3654. https://doi.org/10.1128/AAC.48.10.3645-3654.2004
Otero-González AJ, Magalhães BS, Garcia-Villarino M, López-Abarrategui C, Sousa DA, Dias SC, Franco OL (2010) Antimicrobial peptides from marine invertebrates as a new frontier for microbial infection control. FASEB J 24:1320–1334. https://doi.org/10.1096/fj.09-143388
Lahl WJ (1994) Enzymatic production of protein by hydrolysates for food use. Food Sci 48:68–71
Tam JP, Lu YA, Yang JL (2000) Marked increase in membranolytic selectivity of novel cyclic tachyplesins constrained with an antiparallel two-strand cystine knot framework. Biochem Biophys Res Commun 267:783–790. https://doi.org/10.1006/bbrc.1999.2035
Miyata T, Tokunaga F, Yoneya T, Yoshikawa K, Iwanaga S, Niwa M, Takao T, Shimonishi Y (1989) Antimicrobial peptides, isolated from horseshoe crab hemocytes, tachyplesin II, and polyphemusins I and II: chemical structures and biological activity. J Biochem 106:663–668. https://doi.org/10.1093/oxfordjournals.jbchem.a122913
Antony SP, Singh ISB, Sudheer NS, Vrinda S, Priyaja P, Philip R (2011) Molecular characterization of a crustin-like antimicrobial peptide in the giant tiger shrimp, Penaeus monodon, and its expression profile in response to various immunostimulants and challenge with WSSV. Immunobiology 216:184–194. https://doi.org/10.1016/j.imbio.2010.05.030
Sruthy KS, Chaithanya ER, Sathyan N, Nair A, Antony SP, Bright Singh IS, Philip R (2015) Molecular characterization and phylogenetic analysis of novel isoform of anti-lipopolysaccharide factor from the Mantis shrimp, Miyakea nepa. Probiotics Antimicrob Proteins 7:275–283. https://doi.org/10.1007/s12602-015-9198-2
Lee SY, Lee BL, Soderhall K (2003) Processing of an antibacterial peptide from hemocyanin of the freshwater crayfish Pacifastacus leniusculus. J Biol Chem 278:7927–7933. https://doi.org/10.1074/jbc.M209239200
Munoz M, Vandenbulcke F, Garnier J, Gueguen Y, Bulet P, Saulnier D, Bachère E (2004) Involvement of penaeidins in defense reactions of the shrimp Litopenaeus stylirostris to a pathogenic vibrio. Cell Mol Life Sci 61:961–972. https://doi.org/10.1007/s00018-003-3441-9
Stensvag K, Haug T, Sperstad SV et al (2008) Arasin 1, a proline-arginine-rich antimicrobial peptide isolated from the spider crab, Hyas araneus. Dev Comp Immunol 32:275–285. https://doi.org/10.1016/j.dci.2007.06.002
Sperstad SV, Haug T, Vasskog T, Stensvag K (2009) Hyastatin, a glycine-rich multi-domain antimicrobial peptide isolated from the spider crab (Hyas araneus) hemocytes. Mol Immunol 46:2604–2612. https://doi.org/10.1016/j.molimm.2009.05.002
Relf JM, Chisholm JRS, Kemp GD, Smith VJ (1999) Purification and characterization of a cysteine-rich 11.5-kDa antibacterial protein from the granular haemocytes of the shore crab, Carcinus maenas. Eur J Biochem 264:350–357. https://doi.org/10.1046/j.1432-1327.1999.00607.x
Noga EJ, Stone KL, Wood A, Gordon WL, Robinette D (2011) Primary structure and cellular localization of callinectin, an antimicrobial peptide from the blue crab. Dev Comp Immunol 35:409–415. https://doi.org/10.1016/j.dci.2010.11.015
Beaulieu L, Thibodeau J, Desbiens M, Saint-Louis R, Zatylny-Gaudin C, Thibault S (2010) Evidence of antibacterial activities in peptide fractions originating from snow crab (Chionoecetes opilio) by-products. Probiotics Antimicrob Proteins 2:197–209. https://doi.org/10.1007/s12602-010-9043-6
Gaillard M, Bernatchez L, Tremblay R, Audet C (2015) Regional variation in energy storage strategies in American glass eels from Eastern Canada. Comp Biochem Physiol A Mol Integr Physiol 188:87–95. https://doi.org/10.1016/j.cbpa.2015.06.019
Ebeling ME (1968) The Dumas method for nitrogen in feeds. J Assoc Off Anal Chem 51:766–770
Spinelli J, Lehman L, Wieg D (1974) Composition, processing, and utilization of red crab (Pleuroncodes planipes) as an aquacultural feed ingredient. J Fish Res Board Can 31:1025–1029. https://doi.org/10.1139/f74-115
Hamed I, Ozogul F, Regenstein JM (2016) Industrial applications of crustacean by-products (chitin, chitosan, and chitooligosaccharides): a review. Trends Food Sci Technol 48:40–50. https://doi.org/10.1016/j.tifs.2015.11.007
AOAC, Official Methods of Analysis (1990) Arlington, Virginia, USA
Robert M, Zatylny-Gaudin C, Fournier V, Corre E, Le Corguillé G, Bernay B, Henry J (2014) Transcriptomic and peptidomic analysis of protein hydrolysates from the white shrimp (L. vannamei). J Biotechnol 186:30–37. https://doi.org/10.1016/j.jbiotec.2014.06.020
Church FC, Swaisgood HE, Porter DH, Catignani GL (1983) Spectrophotometric assay using o-phthaldialdehyde for determination of proteolysis in milk and isolated milk proteins. J Dairy Sci 66:1219–1227. https://doi.org/10.3168/jds.S0022-0302(83)81926-2
Doyen A, Saucier L, Beaulieu L, Pouliot Y, Bazinet L (2012) Electroseparation of an antibacterial peptide fraction from snow crab by-products hydrolysate by electrodialysis with ultrafiltration membranes. Food Chem 132:1177–1184. https://doi.org/10.1016/j.foodchem.2011.11.059
Sharpe AN, Kilsby DC (1971) A rapid, inexpensive bacterial count technique using agar droplets. J Appl Bacteriol 34:435–440. https://doi.org/10.1111/j.1365-2672.1971.tb02303.x
Beaulieu L, Bondu S, Doiron K, Rioux LE, Turgeon SL (2015) Characterization of antibacterial activity from protein hydrolysates of the macroalga Saccharina longicruris and identification of peptides implied in bioactivity. J Funct Foods 17:685–697. https://doi.org/10.1016/j.jff.2015.06.026
Adeyeye EI (2002) Determination of the chemical composition of the nutritionally valuable parts of male and female common west African fresh water crab Sudananautes africanus africanus. Int J Food Sci Nutr 53:189–196. https://doi.org/10.1080/09637480220132805
Skonberg DI, Perkins BL (2002) Nutrient composition of green crab (Carcinus maenus) leg meat and claw meat. Food Chem 77:401–404. https://doi.org/10.1016/S0308-8146(01)00364-8
Shahidi F, Synowiecki J (1991) Isolation and characterization of nutrients and value-added products from snow crab (Chionoecetes opilio) and shrimp (Pandalus borealis) processing discards. J Agric Food Chem 39:1527–1532. https://doi.org/10.1021/jf00008a032
Carbonneau ME (2013) Coproduits du crabe commun, carapaces. Merinov. http://www.merinov.ca/fr/app-publication/item/fiche-biomasse-marine-sous-valorisee-crabe-commun-carapace. [Accessed: 01-Feb-2018]
Crespo MOP, Martínez MV, Hernández JL, Lage Yusty MA (2006) High-performance liquid chromatographic determination of chitin in the snow crab, Chionoecetes opilio. J Chromatogr 1116:189–192. https://doi.org/10.1016/j.chroma.2006.03.068
DFO (2016) Snow crab. http://www.dfo-mpo.gc.ca/species-especes/profiles-profils/snow-crab-crabe-neiges-atl-eng.html. [Accessed: 24-May-2017]
Joffe I, Hepburn HR, Nelson KJ, Green N (1975) Mechanical properties of a crustacean exoskeleton. Comp Biochem Physiol Physiol 50:545–549. https://doi.org/10.1016/0300-9629(75)90312-6
Hepburn HR, Joffe I, Green N, Nelson KJ (1975) Mechanical properties of a crab shell. Comp Biochem Physiol 50:551–554
Taylor HH (1990) Pressure-flow characteristics of crab gills: implications for regulation of hemolymph pressure. Physiol Biochem Zool 63:72–89. https://doi.org/10.1086/physzool.63.1.30158154
Wheeler K, Shields JD, Taylor DM (2007) Pathology of hematodinium infections in snow crabs (Chionoecetes opilio) from Newfoundland, Canada. J Invertebr Pathol 95:93–100. https://doi.org/10.1016/j.jip.2007.01.002
You L, Zhao M, Cui C, Zhao H, Yang B (2009) Effect of degree of hydrolysis on the antioxidant activity of loach (Misgurnus anguillicaudatus) protein hydrolysates. Innov Food Sci Emerg Technol 10:235–240. https://doi.org/10.1016/j.ifset.2008.08.007
Mori K, Stewart JE (1978) The hemolymph bactericidin of American lobster (Homarus americanus): adsorption and activation. J Fish Res Board Canada 35:1504–1507. https://doi.org/10.1139/f78-238
Hajirasouli M, Pizooki J (2014) Antimicrobial potential of hemolymph and hepatopancreas of Portunus segnis crabs. Int J Pharm Pharm Sci 6:6–8
Kushibiki T, Kamiya M, Aizawa T, Kumaki Y, Kikukawa T, Mizuguchi M, Demura M, Kawabata SI, Kawano K (2014) Interaction between tachyplesin I, an antimicrobial peptide derived from horseshoe crab, and lipopolysaccharide. BBA—Proteins Proteomics 1844:527–534. https://doi.org/10.1016/j.bbapap.2013.12.017
Destoumieux D, Bulet P, Loew D, van Dorsselaer A, Rodriguez J, Bachère E (1997) Penaeidins, a new family of antimicrobial peptides isolated from the shrimp Penaeus vannamei (Decapoda). J Biol Chem 272:28398–28406. https://doi.org/10.1074/jbc.272.45.28398
Schnapp D, Kemp GD, Smith VJ (1996) Purification and characterization of a proline-rich antibacterial peptide, with sequence similarity to bactenecin-7, from the haemocytes of the shore crab, Carcinus maenas. Eur J Biochem 240:532–539. https://doi.org/10.1111/j.1432-1033.1996.0532h.x
Pankey GA, Sabath LD (2004) Clinical relevance of bacteriostatic versus bactericidal mechanisms of action in the treatment of Gram-positive bacterial infections. Clin Infect Dis 38:864–870. https://doi.org/10.1086/381972
Vilcinskas A, Mukherjee K, Vogel H (2013) Expansion of the antimicrobial peptide repertoire in the invasive ladybird Harmonia axyridis. Proc Biol Sci 280:2012–2113. https://doi.org/10.1098/rspb.2012.2113.
Smith VJ, Fernandes JMO, Kemp GD, Hauton C (2008) Crustins: enigmatic WAP domain-containing antibacterial proteins from crustaceans. Dev Comp Immunol 32:758–772. https://doi.org/10.1016/j.dci.2007.12.002
Jiravanichpaisal P, Lee SY, Kim YA, Andrén T, Soderhall I (2007) Antibacterial peptides in hemocytes and hematopoietic tissue from freshwater crayfish Pacifastacus leniusculus: characterization and expression pattern. Dev Comp Immunol 31:441–455. https://doi.org/10.1016/j.dci.2006.08.002
Oberdorster E, Rittschof D, Mcclellan-Green P (1998) Induction of cytochrome P450 3A and heat shock protein by tributyltin in blue crab, Callinectes sapidus. Aquat Toxicol 41:83–100. https://doi.org/10.1016/S0166-445X(97)00067-2
Li J, Xueqing X, Chunhua X et al (2007) Anti-infection peptidomics of amphibian skin. Mol Cell Proteomics 6:882–894. https://doi.org/10.1074/mcp.M600334-MCP200
Seo JK, Lee MJ, Go HJ, Park TH, Park NG (2012) Purification and characterization of YFGAP, a GAPDH-related novel antimicrobial peptide, from the skin of yellowfin tuna, Thunnus albacares. Fish Shellfish Immunol 33:743–752. https://doi.org/10.1016/j.fsi.2012.06.023
Dhar AK, Dettori A, Roux MM, Klimpel AR, Reid B (2003) Identification of differentially expressed genes in shrimp (Penaeus stylirostris) infected with White spot syndrome virus by cDNA microarrays. Arch Virol 148:2381–2396. https://doi.org/10.1007/s00705-003-0172-z
Dhar AK, Bowers RM, Licon KS, Veazey G, Read B (2009) Validation of reference genes for quantitative measurement of immune gene expression in shrimp. Mol Immunol 46:1688–1695. https://doi.org/10.1016/j.molimm.2009.02.020
Lüders T, Birkemo GA, Fimland G, Nissen-Meyer J, Nes IF (2003) Strong synergy between a eukaryotic antimicrobial peptide and bacteriocins from lactic acid bacteria. Appl Environ Microbiol 69:1797–1799. https://doi.org/10.1128/AEM.69.3.1797-1799.2003
Acknowledgments
The authors wish to thank Réjean Tremblay for welcoming us in his laboratory (ISMER-UQAR, QC, Canada) and for snow crab dissection; Jean-Bruno Nadalini (ISMER-UQAR, QC, Canada), Diane Gagnon (Université Laval, QC, Canada), and Marine Béguin (Université Laval, QC, Canada) for their technical expertise. The authors would also like to thank the Natural Sciences and Engineering Research Council of Canada (NSERC) for their financial support (NSERC discovery grant; RGPIN/327203-2011).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
El Menif, E., Offret, C., Labrie, S. et al. Identification of Peptides Implicated in Antibacterial Activity of Snow Crab Hepatopancreas Hydrolysates by a Bioassay-Guided Fractionation Approach Combined with Mass Spectrometry. Probiotics & Antimicro. Prot. 11, 1023–1033 (2019). https://doi.org/10.1007/s12602-018-9484-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-018-9484-x