Abstract
Molecular property prediction is an important step in the drug discovery pipeline. Numerous computational methods have been developed to predict a wide range of molecular properties. While recent approaches have shown promising results, no single architecture can comprehensively address all tasks, making this area persistently challenging and requiring substantial time and effort. Beyond traditional machine learning and deep learning architectures for regular data, several deep learning architectures have been designed for graph-structured data to overcome the limitations of conventional methods. Utilizing graph-structured data in quantitative structure–activity relationship (QSAR) modeling allows models to effectively extract unique features, especially where connectivity information is crucial. In our study, we developed residual graph attention networks (ResGAT), a deep learning architecture for molecular graph-structured data. This architecture is a combination of graph attention networks and shortcut connections to address both regression and classification problems. It is also customizable to adapt to various dataset sizes, enhancing the learning process based on molecular patterns. When tested multiple times with both random and scaffold sampling strategies on nine benchmark molecular datasets, QSAR models developed using ResGAT demonstrated stability and competitive performance compared to state-of-the-art methods.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Property prediction is an important step in modern drug discovery, and it continues to capture researchers’ attention [1]. Accurate molecular property determination speeds up screening processes for potential drug candidates, resulting in cost and time savings [2]. Because molecular structures and biological activities (or properties) are closely related, many computational approaches have been developed to predict these properties using structural information. Among these approaches, quantitative structure–activity relationship (QSAR) modeling is a low-cost computational method commonly used to predict a wide range of molecular properties (e.g., lipophilicity, hydrophobicity, solubility) [3]. These QSAR models are flexible in design and optimized for efficient learning of complex structural patterns. Despite initial successes, these modeling tasks remain difficult due to the complexity of chemical structures, class imbalance, high-dimensional data representation, and limited data volume. To address these challenges, robust computational methods and interdisciplinary collaboration are critical.
The graph neural network (GNN) [4], which was specifically designed to handle molecular graphs [5,6,7,8,9], has made a breakthrough over the last two decades. The development of efficient GNN variants allows for the emergence of graph-based representation learning [10,11,12,13]. For years, numerous studies have used GNN and its variants to predict molecular properties. Scarselli et al. [4] proposed the first version of GNN in 2009. Although GNN can handle graph-structured data, their applications have not been widespread due to their relatively low learning efficiency until the graph convolutional network (GCN) was presented [14]. The introduction of GNN has sparked a large number of further research and extensive practice in graph-based deep learning (DL) architectures. Wieder et al. [15] conducted a critical review to summarize DL architectures used for molecular property prediction. Gilmer et al. [16] developed neural fingerprints using CNN customized for graph-structured data. After critically surveying various GNN-based models, Yang et al. [17] introduced and conceptualized the Message Passing Neural Network (MPNN), characterized by two distinct phases: Message passing and Readout. Yang et al. [18] presented the Directed MPNN (D-MPNN) as an upgraded version of MPNN that prioritizes updating information specifically on directed bonds instead of atoms. Xiong et al. [10] introduced AttentiveFP, an attention-based network that achieved robust performance on multiple benchmark datasets. HRGCN+, by Wu et al. [11], combines molecular graphs and descriptors (physicochemical features) to boost prediction efficiency with excellent performance compared to existing methods. Li et al. [8] proposed the Triplet Message Network (TrimNet) for processing molecular graphs, an architecture designed to significantly reduce the number of parameters and enhance the capacity to extract bonding information. The Graph Multiset Transformer (GMT), developed by Baek et al. [19], is a Transformer-based architecture adapting to multiset pooling on graphs. Most recently, the Hierarchical Informative Graph Neural Network (HiGNN), by Zhu et al. [9], is one of the most competitive DL architectures. HiGNN comprises two main blocks: atom–atom interaction and feature-wise attention. The atom–atom interaction block is based on neural tensor networks for knowledge graph reasoning [20], while the feature-wise attention block recalibrates an atom’s representations after the message passing phase, thus enhancing the selective extraction of important features. Initially, the model’s input, the molecular structure, is fragmented into substructures using the BRICS algorithm [21], creating global and hierarchical molecular representations. Since the implementation of HiGNN requires high computational costs to process graph-structured data, it may not be cost- or time-effective for prediction tasks on large datasets. Moreover, the effectiveness of using complex attention mechanisms might not always align with expectations. The performance of models is influenced by various factors, including the complexity of the tasks, data quality, and volume. In such scenarios, architectures with simpler attention mechanisms could offer a more appropriate alternative.
Although many models have been developed for predicting molecular properties, there is still a lot of room for improvement. The availability of high-quality data, chemical diversity in datasets, and data curation processes all have a significant impact on prediction efficiency. Insufficient or biased data can limit the model’s ability to learn molecular patterns and make accurate predictions. Furthermore, computational cost is an important consideration when dealing with a large number of molecules. Computationally expensive methods may be impractical for upscaling models. On the other hand, generalizability is one of the issues encountered in most computational methods. Models developed or evaluated on specific sets of molecules for specific properties may perform poorly on other datasets or entirely new data, especially if the training data are not good representatives. Despite being tested against multiple benchmark datasets, all known state-of-the-art methods may fail to show good performance on when applied to a new prediction task. As a result, developing novel methods for molecular property prediction is always one of the key topics in modern drug discovery in order to address various future prediction tasks.
In this study, we introduce the residual graph attention Network (ResGAT), a novel graph-based deep learning architecture for molecular property prediction tasks. This architecture is built on two key insights: (1) the use of regular shortcut connections between blocks, and (2) shortcut connections integrated with a graph attention layer. Incorporating these types of shortcut connections into ResGAT enhances the model’s learning capacity by stabilizing the training process and improving generalization. Our architecture is versatile, capable of handling both regression and classification problems, and allows flexible customization of the number of blocks per block set to accommodate various dataset sizes.
2 Proposed architecture
2.1 Residual graph attention network
We introduce the Residual Graph Attention Network (ResGAT), as described in Algorithm 1, a unique DL architecture designed to process graph-structured data and capable of addressing a variety of molecular property prediction tasks. The ResGAT architecture is constructed using Graph Attention (GAT) layers [22] and two types of shortcut connections [23]. GAT, a masked self-attention layer, is demonstrated to outperform the GCN layer in terms of computing speed and efficiency (see Sect. 2.2). The shortcut connection is a crucial component of the Residual Neural Network (ResNet) [23]. This architecture is designed with three Block Sets, and each of them is specified by L blocks (Fig. 1). A single block consists of two GAT layers activated by the rectified linear unit (ReLU) function. After passing Block Set 3, the outputs are pooled with a global max-pooling layer. Finally, the max-pooled outputs are passed through a fully connected (FC) block comprising three layers. The first two layers are activated by the ReLU function, while the final layer is activated by the Sigmoid function for classification tasks or by the ReLU function for regression tasks. In comparison with ResNet, ResGAT architectures have a smaller number of layers in each block. In ResNet, each block comprises a minimum of four CNN layers, whereas in ResGAT, each block consists of just two GAT layers. Furthermore, we employed another shortcut connection impeded by a graph attention layer, referred to as the ‘graph attention shortcut’. This concept is inspired by a critical analysis conducted by He et al. [23] about the propagation formulations used in residual building blocks with diverse types of shortcut connections. Integrating these two types of shortcut connections into the ResGAT enhances the model’s learning capacity by stabilizing the training process and improving generalization. In addition to the main architecture, ResGAT, we also developed a generic version named ResGCN, which differs only in that all GAT layers [22] are replaced with GCN layers [14]. For model optimization, the number of blocks (num_block) and the feature embedding size (embed_size) in each block set can be tuned. The parameters num_block and embed_size are varied in each block set.
2.2 Graph attention layer
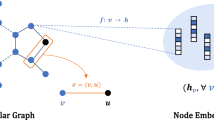
The graph attention (GAT) layer was completely formulated by Veličković et al. [22] based on a previously published work done by Bahdanau et al. [24]. A graph \(\mathcal {G(V, E)}\) with N nodes (vertices) is defined by a vector of node features \(h = \{\ h_1, h_2, h_3,..., h_N \}\) with \( h \in \mathbb {R}^F\). The vector h is operated by the GAT layer to return \( h' = \{ h'_1, h'_2, h'_3,..., h'_N \}\) with \( h' \in \mathbb {R}^{F'}\). The weight matrix \({\textbf {W}} \in \mathbb {R}^{F' \times F} \) is multiplied to every node; and F and \(F'\) are the numbers of input and output features, respectively. The attention output \(e_{uv}\) of node\(_u\) directed from node\(_v\) is computed as:
where a is the self-attention feedforward layer parameterized by the learnable vector of parameters \(a^T\) and || denotes the concatenation operation. The Softmax function is applied to normalize the attention output. Each normalized attention output \(\alpha _{uv}\) is computed as:
The equation for \(\alpha _{uv}\) is rewritten as:
Since the layer is designed to force each node to attend to all other nodes in the network, the output vector of node\(_u\) (\(\mathbf {h'}_u\)) is finally obtained by the summation of all products of the normalized attention outputs (\(\alpha _{uv}\)) and the weighted node feature vectors (\(h_v\)) of other neighboring nodes.
where \(\sigma \) is the nonlinearity activation function. Algorithm 2 describes the operation mechanism of the GAT layer.
3 Experiments
3.1 Overview
The major steps in our experiments are presented in Fig. 2. First, the original benchmark datasets were downloaded from the MoleculeNet website [25]. To qualify the data for the modeling experiment, all the benchmark datasets were curated (see Sect. 3.3) before being encoded (see Sect. 3.4). Then, each refined dataset was divided into two parts: a train-val set and a test set with a ratio of 90:10. The train-val data was then split into a new training set and a validation set with a ratio of 90:10. The validation set was used for hyperparameter tuning. Once hyperparameter tuning was completed, the model was retrained using the best hyperparameters, and then it was evaluated using the test set for benchmarking (see Sect. 3.6).
3.2 Benchmark datasets
To investigate the performance of ResGAT, we conducted a large number of modeling experiments on nine benchmark molecular datasets, including ESOL, FreeSolv, Lipo, BACE, BBBP, HIV, ClinTox, SIDER, and Tox21. These datasets were collected from MoleculeNet [25], an online source containing molecular datasets specially designed to benchmark machine learning methods on property prediction tasks. Table 1 gives information on the datasets used in the study. After collecting these datasets, we performed data curation to remove unqualified samples. Generally, the number of samples from all refined datasets decreased after the data curation was completed (Table 2). The details of data curation are provided in Sect. 3.3.
3.3 Data curation
Before conducting experiments, we performed data curation to qualify the chemical data for model development and evaluation. Our data curation pipeline [26,27,28] includes four phases: (1) Validation, (2) Cleaning, (3) Normalization, and (4) Final verification. Before entering the pipeline, all chemical data (in the SMILES format) were converted into their corresponding canonical forms. In phase (1), molecules whose chemical types belong to one of three classes, including inorganics, mixtures, and organometallics, are removed. In phase (2), salts and manipulating charged molecules are eliminated. Charged molecules may be formed by metal-containing structures or polar organic groups. While metal-containing charged molecules are rejected, organic charged molecules are converted to non-charged forms. The neutralization of charged organic molecules, however, remains a controversial topic among scientists, as they face challenges in precisely determining the experimental conditions under which these molecules exhibit activities. In phase (3), detautomerization, destereoisomerization, and removal of chemotypes are executed to unify tautomers, stereoisomers, or chemotypes of the same molecules into canonical forms. A molecule possessing unstable substructures often undergoes interchange among multiple intermediate forms. When considering a group of tautomers (or chemotypes) for the same molecule, the intermediate form that exhibits the highest degree of structural equivalence compared to other forms is selected as the canonical tautomer (or chemotype). At the end of these three first phases, any duplicates found are discarded. In phase (4), samples (molecules) whose labels conflict with each other are manually processed. Conflicting samples can arise in any of the three situations outlined below: (a) a group of identical molecules with different labels; (b) a group of identical molecules with duplicated labels; and (c) a group of different molecules identified by the same CAS registry number. Samples in situations (a) or (c) are excluded, whereas those in situations (b) are retained and unified. Finally, structural verification is accomplished to confirm identity and validity using the two largest chemical databases: PubChem and ChEMBL. Table 2 compares the number of samples in refined datasets with the original ones.
Molecular encoding scheme. A N-atom molecule with M bonds is transformed into an atom matrix of size \(N \times 41\) and a bond index matrix of size \(M \times 2\). The atom matrix is a column-wise combination of N vectors of size of \(41 \times 1\). The bond index matrix is a combination of M vectors of size \(2 \times 1\) indicating connectivity between atom ith and atom jth
3.4 Molecular encoding scheme
Figure 3 explains the molecular encoding scheme used in our study. For each molecule constituted by N heavy atoms (excluding hydrogen) and M bonds connecting these atoms, its representations are defined with two matrices: an atom matrix with a dimension of \(N\times 41\) and a bonding matrix with a dimension of \(M\times 2\). The values of N and M vary across molecules.
The atom matrix is created in several steps. The molecular structure is first analyzed to determine the appearance order of heavy atoms, and these atoms are then assigned indices. For each atom, a set of 41 features is computed using the RDKit library [29]. The details of these features are described in Table 3. For a heavy atom, a 41-dimensional feature vector is organized as a binary vector with a size of 1\(\times \)41. The feature vector consists of 16 Atomic features, 9 Degree features, 6 Orbital hybridization features, 5 Number of hydrogens features, 2 Cahn-Ingold-Prelog (CIP) priority features, 2 IsCharge features, 1 IsAromatic feature, and 1 Chirality feature. The Atomic features determine the atom based on the atom list. The Degree features indicate the number of bonds formed by the atom with neighboring heavy atoms, ranging from 0 to \(>=7\). The Orbital Hybridization features describe the specific type of orbital hybridization of a heavy atom uses to form its bonds. The Number of Hydrogens feature counts the number of hydrogen atoms that have bonds with a heavy atom. The CIP Priority features identify the spatial orientation of a chiral center (atom bonding to four different groups): clockwise (R) or counterclockwise (S). The IsCharge features present the charge state of an atom to assign it either the ‘formal charge (FC)’ or ‘radical electron (RE)’ state. The IsAromatic feature defines whether an atom is a member of any ring or cyclic structure. The Chirality feature identifies whether a heavy atom has a chiral center. To create the bond matrix, the connectivity map of all heavy atoms is computed (as shown in Fig. 3). The atom matrix carries information on the node features, while the bond matrix stores information on the edge indices.
3.5 Model development
While GAT is flexible in learning graph structures with highly varying neighbor relationships, it requires a higher number of parameters to learn attention coefficients. In contrast, GCN is parameter-efficient due to shared weights across the graph but may struggle with generalizability when faced with new graph patterns. GCN is well-suited for relatively uniform or well-defined graph patterns, while GAT is more appropriate for handling complex graph structures. To address diverse scenarios, we introduce another variant of the ResGAT architecture named ResGCN, wherein all GAT layers are replaced with GCN layers.
All models constructed with these two architectures were tuned, trained, and tested under the same conditions and settings. For each dataset, the training and validation sets were used for model tuning and development, while the test set was used for model evaluation. Test sets were not involved in any stage of model selection. Initially, the number of blocks (num_block) in each block set was fixed at 1 to tune the feature embedding size (embed_size) of the graph layer with three values of 64, 128, and 256. After tuning the parameter embed_size for a block in each block set, we tuned the parameter num_block for each block set with three values of 1, 2, and 3. Finally, the learning rate was tuned with three values of \(1\times 10^{-4}\), \(5\times 10 ^{-4}\), and \(1\times 10^{-3}\). Models implemented for different prediction tasks have different hyperparameters. The loss functions for regression and classification tasks are mean squared error (MSE) and binary cross entropy (BCE), respectively, and are computed as:
where n is the number of samples; y is the ground truth (label), and \(\hat{y}\) is the predicted value or probability of the regression or classification task, respectively.
3.6 Evaluation metrics
To evaluate all models, we used the Root Mean Squared Error (RMSE) and the Area Under the Receiver Operating Characteristic (ROC) curve (AUCROC) for regression tasks and classification tasks, respectively. For multitask classification tasks, the average AUCROC was computed based on the number of tasks. Our code is made publicly available in our GitHub repository.Footnote 1
4 Results and discussion
4.1 Model evaluation
This experiment was conducted to evaluate the performance of ResGAT (our proposed method) and ResGCN (a generic version of ResGAT). The only difference between ResGAT and ResGCN is the type of graph neural network used. In ResGAT, the graph layer is a GAT layer, while in ResGCN, the graph layer is a GCN layer. Both architectures were used to develop nine prediction models across three types of tasks: regression, binary classification, and multi-task classification. All models were implemented under the same conditions for a fair assessment.
Experimental results show that ResGAT and ResGCN have equivalent performance in all classification tasks under both sampling strategies (Fig. 4). Under the random sampling strategy, ResGCN has slightly lower performance on the BACE, BBBP, and HIV datasets but higher performance on the ClinTox dataset compared to ResGAT. The performances of both models on the SIDER and Tox21 datasets are almost similar. For the regression tasks, ResGCN obtains higher performance on all datasets compared to ResGAT. Under the scaffold sampling strategy, ResGCN shows greater efficiency on the ESOL dataset, whereas ResGAT obtains better performance on the FreeSolve dataset. Their effectiveness on the Lipo dataset is comparable. For classification tasks, ResGCN achieves higher performance on the BBBP and ClinTox datasets. Meanwhile, ResGAT works effectively on the BACE dataset only. Both models demonstrate similar levels of prediction power on the rest of the classification datasets.
4.2 Model benchmarking
To examine the efficiency of ResGAT, we developed a series of prediction models using five other state-of-the-art architectures, including AttentiveFP [10], GMT [19], TrimNet [8], D-MPNN [18], and HiGNN [9]. The GCN [14] and GAT [22] architectures were also used to develop two baseline graph models. Models of state-of-the-art architectures were reimplemented using source codes provided by their authors. The parameters of all the reimplemented models were also fairly tuned. Besides, two sampling methods, random and scaffold, were employed. For each dataset, the modeling experiment for a particular architecture was repeated ten times to avoid sampling bias.
Tables 4, 5, and 6 provide detailed results of modeling experiments on nine benchmark datasets of regression, binary classification, and multitask classification tasks, respectively. These tables compare the performance of the models constructed using our proposed architecture with those constructed using state-of-the-art architectures. The experimental results show that our models (developed using ResGAT or ResGCN) are ranked in the top 3 in five out of the nine datasets and in seven out of the nine datasets under the random sampling and scaffold sampling strategies, respectively. Under the random sampling strategy, our models obtain three 1st-ranks on the BACE, HIV, and ClinTox datasets; two 2nd-ranks on the ClinTox and SIDER datasets; and three 3rd-ranks on the FreeSolve, BACE, and SIDER datasets. Under the scaffold sampling strategy, our models achieve two 2nd-ranks (on the FreeSolv and BACE datasets) and three 3rd-ranks (on the Lipo, BBBP, and HIV datasets). D-MPNN is a very robust architecture when 13 D-MPNN-based models are ranked in the top-3 of both sampling strategies. However, most of them obtain only 2nd-ranks and 3rd-ranks. Also, HiGNN is an efficient architecture compared to others. Although there are only nine out of eighteen HiGNN-based models present in the top 3, they had seven 1st-ranks on the ESOL, Lipo, BACE, BBBP, HIV, and ClinTox datasets. The TrimNet and GMT architectures work better on regression tasks while showing low predictive efficiency in classification tasks. Models implemented using the AttentiveFP architecture achieve competitive performance on classification tasks, especially for those with a large number of tasks. To rank the overall performance of all implemented architectures, we create a summary table describing the performance ranking of the models on the test sets. For each dataset, the performance of models is ranked from 1 (highest) to 9 (smallest) scores. Every architecture is assigned scores from nine datasets. The maximum score is 81, and the minimum score is 9 (Table 7). Based on average ranking scores, the ResGAT and ResGCN are in the top 3. Under the random sampling strategy, D-MPNN and HiGNN have the smallest average ranking scores of 3.22, followed by ResGAT (3.33), ResGCN (3.89), and others. Under the scaffold sampling strategy, D-MPNN achieves average ranking scores of 3.22, followed by ResGAT (3.44), ResGCN (3.89), and others.
In our experiments, we implemented all DL models using PyTorch 1.12.0 and PyTorch Geometric 2.0.4, training them on an i9-13900K with 64 GB of RAM and one NVIDIA GTX 3060. The training process time (seconds per epoch), recorded in Table 8, reflects the computing resources required. Our models demonstrate superior time and cost efficiency compared to state-of-the-art models, notably outperforming D-MPNN- and HiGNN-based models with up to a 50% reduction in training time. While only models developed with two baseline architectures (GCN and GAT) have shorter training times, for the ClinTox dataset, ResGCN-based models exhibit slightly higher training times than other models, whereas ResGAT-based models still require less time. Testing completion for our models ranged from 0.02 to 0.48 s, depending on the dataset, comparable to the two baseline models and faster than all other models. In summary, the results confirm that our proposed architectures are not only robust but also time-effective.
4.3 Limitations and future work
Besides achieving goals, our proposed architecture still has limitations that need to be improved in the future. Overall, compared to D-MPNN and HiGNN, ResGAT (and ResGCN) show less efficiency in regression problems. In binary classification tasks, ResGAT obtains better performance under the random sampling strategy, while HiGNN demonstrates its powerful architecture under the scaffold sampling strategy. In multitask classification tasks, although ResGAT works more effectively than HiGNN under the random sampling strategy, its performance under the scaffold sampling strategy needs to be enhanced. From our experimental results, it can be observed that our proposed architecture is more efficient when dealing with classification tasks than regression tasks. It can work competently on large datasets, especially for multitask classification problems.
5 Conclusion
In this study, we presented ResGAT, an innovative DL architecture designed for predicting molecular properties from graph-structured data. ResGAT is versatile, capable of handling both regression and classification tasks, and it offers a flexible tuning mechanism to accommodate various dataset sizes. The depth of the architecture can be adjusted to specific needs, and our experimental findings validate its robustness and efficiency. Our results indicate that ResGAT, along with ResGCN, are competitive with other state-of-the-art architectures. Further investigation and improvements are anticipated to enhance their predictive power.
References
Tang Y, Zhu W, Chen K, Jiang H (2006) New technologies in computer-aided drug design: toward target identification and new chemical entity discovery. Drug Discov Today Technol 3(3):307–313. https://doi.org/10.1016/j.ddtec.2006.09.004
Shen J, Nicolaou CA (2019) Molecular property prediction: recent trends in the era of artificial intelligence. Drug Discov Today Technol 32–33:29–36. https://doi.org/10.1016/j.ddtec.2020.05.001
Muratov EN, Bajorath J, Sheridan RP, Tetko IV, Filimonov D, Poroikov V, Oprea TI, Baskin II, Varnek A, Roitberg A, Isayev O, Curtalolo S, Fourches D, Cohen Y, Aspuru-Guzik A, Winkler DA, Agrafiotis D, Cherkasov A, Tropsha A (2020) QSAR without borders. Chem Soc Rev 49(11):3525–3564. https://doi.org/10.1039/d0cs00098a
Scarselli F, Gori M, Tsoi AC, Hagenbuchner M, Monfardini G (2009) The graph neural network model. IEEE Trans Neural Netw 20(1):61–80. https://doi.org/10.1109/tnn.2008.2005605
Baskin II, Palyulin VA, Zefirov NS (1997) A neural device for searching direct correlations between structures and properties of chemical compounds. J Chem Inf Comput Sci 37(4):715–721. https://doi.org/10.1021/ci940128y
Micheli A, Sperduti A, Starita A, Bianucci AM (2000) Analysis of the internal representations developed by neural networks for structures applied to quantitative structure–activity relationship studies of benzodiazepines. J Chem Inf Comput Sci 41(1):202–218. https://doi.org/10.1021/ci9903399
Goulon A, Picot T, Duprat A, Dreyfus G (2007) Predicting activities without computing descriptors: graph machines for QSAR. SAR QSAR Environ Res 18(1–2):141–153. https://doi.org/10.1080/10629360601054313
Li P, Li Y, Hsieh C-Y, Zhang S, Liu X, Liu H, Song S, Yao X (2020) TrimNet: learning molecular representation from triplet messages for biomedicine. Brief Bioinform 22(4):bbaa266. https://doi.org/10.1093/bib/bbaa266
Zhu W, Zhang Y, Zhao D, Xu J, Wang L (2022) HiGNN: a hierarchical informative graph neural network for molecular property prediction equipped with feature-wise attention. J Chem Inf Model 63(1):43–55. https://doi.org/10.1021/acs.jcim.2c01099
Xiong Z, Wang D, Liu X, Zhong F, Wan X, Li X, Li Z, Luo X, Chen K, Jiang H, Zheng M (2019) Pushing the boundaries of molecular representation for drug discovery with the graph attention mechanism. J Med Chem 63(16):8749–8760. https://doi.org/10.1021/acs.jmedchem.9b00959
Wu Z, Jiang D, Hsieh C-Y, Chen G, Liao B, Cao D, Hou T (2021) Hyperbolic relational graph convolution networks plus: a simple but highly efficient QSAR-modeling method. Brief Bioinform 22(5):bbab112. https://doi.org/10.1093/bib/bbab112
Li P, Wang J, Qiao Y, Chen H, Yu Y, Yao X, Gao P, Xie G, Song S (2021) An effective self-supervised framework for learning expressive molecular global representations to drug discovery. Brief Bioinform 22(6):bbab109. https://doi.org/10.1093/bib/bbab109
Cai H, Zhang H, Zhao D, Wu J, Wang L (2022) FP-GNN: a versatile deep learning architecture for enhanced molecular property prediction. Brief Bioinform 23(6):bbac408. https://doi.org/10.1093/bib/bbac408
Kipf TN, Welling M (2016) Semi-supervised classification with graph convolutional networks. arXiv. https://doi.org/10.48550/arxiv.1609.02907
Wieder O, Kohlbacher S, Kuenemann M, Garon A, Ducrot P, Seidel T, Langer T (2020) A compact review of molecular property prediction with graph neural networks. Drug Discov Today Technol 37:1–12. https://doi.org/10.1016/j.ddtec.2020.11.009
Duvenaud D, Maclaurin D, Aguilera-Iparraguirre J, Gómez-Bombarelli R, Hirzel T, Aspuru-Guzik A, Adams RP (2015) Convolutional networks on graphs for learning molecular fingerprints. arXiv. https://doi.org/10.48550/arxiv.1509.09292
Gilmer J, Schoenholz, SS, Riley PF, Vinyals O, Dahl GE (2017) Neural message passing for quantum chemistry. In: Precup D, Teh YW (eds) Proceedings of the 34th international conference on machine learning. Proceedings of machine learning research, vol 70. PMLR, Sydney, NSW, Australia, pp 1263–1272. https://doi.org/10.5555/3305381.3305512
Yang K, Swanson K, Jin W, Coley C, Eiden P, Gao H, Guzman-Perez A, Hopper T, Kelley B, Mathea M, Palmer A, Settels V, Jaakkola T, Jensen K, Barzilay R (2019) Analyzing learned molecular representations for property prediction. J Chem Inf Model 59(8):3370–3388. https://doi.org/10.1021/acs.jcim.9b00237
Baek J, Kang M, Hwang SJ (2021) Accurate learning of graph representations with graph multiset pooling. arXiv. https://doi.org/10.48550/arxiv.2102.11533
Socher R, Chen D, Manning CD, Ng AY (2013) Reasoning with neural tensor networks for knowledge base completion. In: Burges CJ, Bottou L, Welling M, Ghahramani Z, Weinberger KQ (eds) NIPS’13: proceedings of the 26th international conference on neural information processing systems, vol 1. Curran Associates, Inc., Lake Tahoe, Nevada, United States, pp 926–934. https://doi.org/10.5555/2999611.2999715
Degen J, Wegscheid-Gerlach C, Zaliani A, Rarey M (2008) On the art of compiling and using ‘drug-like’ chemical fragment spaces. ChemMedChem 3(10):1503–1507. https://doi.org/10.1002/cmdc.200800178
Veličković P, Cucurull G, Casanova A, Romero A, Liò P, Bengio Y (2017) Graph attention networks. arXiv. https://doi.org/10.48550/arXiv.1710.10903
He K, Zhang X, Ren S, Sun J (2016) Identity mappings in deep residual networks. In: 14th European conference on computer vision (ECCV), vol 4. Springer, Amsterdam, pp 630–645. https://doi.org/10.1007/978-3-319-46493_038
Bahdanau D, Cho K, Bengio Y (2014) Neural machine translation by jointly learning to align and translate. arXiv. https://doi.org/10.48550/arXiv.1409.0473
Wu Z, Ramsundar B, Feinberg EN, Gomes J, Geniesse C, Pappu AS, Leswing K, Pande V (2018) MoleculeNet: a benchmark for molecular machine learning. Chem Sci 9(2):513–530. https://doi.org/10.1039/c7sc02664a
Nguyen-Vo T-H, Trinh QH, Nguyen L, Nguyen-Hoang P-U, Nguyen T-N, Nguyen DT, Nguyen BP, Le L (2021) iCYP-MFE: identifying human cytochrome P450 inhibitors using multitask learning and molecular fingerprint-embedded encoding. J Chem Inf Model 62(21):5059–5068. https://doi.org/10.1021/acs.jcim.1c00628
Nguyen L, Nguyen Vo T-H, Trinh QH, Nguyen BH, Nguyen-Hoang P-U, Le L, Nguyen BP (2022) iANP-EC: identifying anticancer natural products using ensemble learning incorporated with evolutionary computation. J Chem Inf Model 62(21):5080–5089. https://doi.org/10.1021/acs.jcim.1c00920
Vinh T, Trinh QH, Nguyen L, Nguyen-Vo T-H, Nguyen BP (2024) Predicting cardiotoxicity of molecules using attention-based graph neural network. J Chem Inf Model 64(6):1816–1827. https://doi.org/10.1021/acs.jcim.3c01286
Landrum G et al (2022) RDKit: open-source cheminformatics software (Release 2022.03.2). https://doi.org/10.5281/zenodo.591637
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions The work of B.P.N was supported in part by the Faculty Strategic Research Grant (FSRG) Number 411494 at Victoria University of Wellington (VUW) and the Endeavour Fund (Smart Ideas) from the New Zealand Ministry of Business, Innovation and Employment (MBIE) under contract RTVU2301.
Author information
Authors and Affiliations
Contributions
T.-H.N.-V.: Conceptualization, software, formal analysis, visualization, writing—original draft. T.T.T.D: Validation, writing—review and editing, supervision. B.P.N.: Conceptualization, funding acquisition, writing—review and editing, supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nguyen-Vo, TH., Do, T.T.T. & Nguyen, B.P. ResGAT: Residual Graph Attention Networks for molecular property prediction. Memetic Comp. 16, 491–503 (2024). https://doi.org/10.1007/s12293-024-00423-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12293-024-00423-5