Abstract
N6-methyladenosine (m6A) is the most abundant and prevalent epigenetic modification of mRNA in mammals. This dynamic modification is regulated by m6A methyltransferases and demethylases, which control the fate of target mRNAs through influencing splicing, translation and decay. Recent studies suggest that m6A modification plays an important role in the progress of cardiac remodeling and cardiomyocyte contractile function. However, the exact roles of m6A in cardiovascular diseases (CVDs) have not been fully explained. In this review, we summarize the current roles of the m6A methylation in the progress of CVDs, such as cardiac remodeling, heart failure, atherosclerosis (AS), and congenital heart disease. Furthermore, we seek to explore the potential risk mechanisms of m6A in CVDs, including obesity, inflammation, adipogenesis, insulin resistance (IR), hypertension, and type 2 diabetes mellitus (T2DM), which may provide novel therapeutic targets for the treatment of CVDs.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
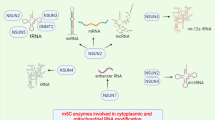
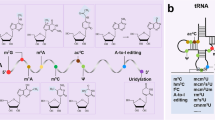
Epigenetic modifications are regulatory mechanisms of gene expression without changing the DNA sequence [1]. These dynamic modifications affect a number of biological processes such as gene expression, protein function, biological senescence, and disease occurrence [2]. Over 100 different types of RNA modifications were found in cellular RNAs [3], in which N6-methyladenosine (m6A) is one of the most common and abundant epigenetic modifications of eukaryotic mRNA in various eukaryotic RNAs since 1974 [4, 5]. Meanwhile, it also widely exists in non-coding RNAs, including lncRNA, miRNA, and circRNAs [6,7,8]. m6A is a reversible methylation modification occurring at the nitrogen-6 position of adenosine. It is catalyzed by a methyltransferase complex, which is composed of Wilms’ tumor 1-associating protein (WTAP), RNA-binding motif protein 15/15B (RBM15/15B), methyltransferase-like 3 (METTL3), and 14 (METTL14), and is removed by demethylases ALKB homolog 5 (ALKBH5) and fat mass and obesity-associated protein (FTO). Moreover, it can be recognized by m6A “reader” YT521-B homology (YTH) domain-containing protein 1/2 (YTHDC1/2) and YTH domain family 1/2/3 (YTHDF1/2/3), and then perform a variety of biological functions. In recent years, there is accumulating evidence that the dynamic regulation of RNA by m6A may play a pivotal role in various biological processes, such as adipogenesis, carcinogenesis, spermatogenesis, circadian rhythms, development, and stem cell renewal [9,10,11,12,13] (Fig. 1).
m6A methylation modification of RNA. m6A methyltransferases and demethylases dynamically and reversibly regulate the m6A modification level of RNA. m6A methyltransferases include METTL3/14, KIAA1429, KC3H13, and RBM15/15B. FTO and ALKBH5 are known as demethylases and perform a demethylation function. The latest research suggested that FTO may have a priority role in m6Am rather than m6A. A series of binding proteins, like YTHDF1/2/3 and YTHDC1/2, recognize m6A-modified mRNA and exert diverse biological functions, such as RNA translation, decay, and splicing
Cardiovascular diseases (CVDs), including cerebrovascular disease, rheumatic heart disease, coronary heart disease, and other conditions, are a common threat to mankind today. Recent studies have revealed that m6A modification may play an important role in the progress of CVDs, such as in the regulation of cardiac remodeling and cardiomyocyte contractile function [14,15,16]. Previous study has investigated that the lack of m6A could lead to cardiac function abnormalities in mice [14], which was associated with impaired cardiomyocyte compensated hypertrophic responses. As the level of m6A in mouse compensated hypertrophic cardiomyocytes increases during serum-mediated hypertrophy stimulation [14]. The study also showed that m6A-deficient mice were more prone to cardiac function damage under serum-induced hypertrophy stimulation [14]. Consistent with this study, Song et al. [17] confirmed that the levels of myocardial m6A were enhanced in hypoxia/deoxygenation-treated neonatal mouse cardiomyocytes and ischemia/reperfusion-treated mouse heart. These results indicated that m6A modification is necessary in maintaining cardiac homeostasis.
In this review, we will focus on the latest progress and potential roles of m6A methylation modification in CVDs, including atherosclerosis (AS), cardiac remodeling, heart failure (HF), and congenital heart disease. Furthermore, the underlying risk factors for CVDs impacted by m6A, including adipogenesis, inflammation, embryonic stem cell (ESC) differentiation, insulin resistance (IR), hypertension, and type 2 diabetes mellitus (T2DM), will also be discussed. Further understanding of the relationship between m6A and CVDs may provide new therapeutic strategies for the treatment of CVDs.
Biological Roles of m6A Modification
It is well known that m6A modification has a significant impact on the process of gene expression, through regulating the degradation, splicing, export, and translation of RNA. Several investigators have demonstrated that m6A modification levels play an important role in numerous biological processes, including adipogenesis, ESC differentiation, circadian rhythms, and cell fate determination [9, 11, 13]. Furthermore, m6A modification deficiency can lead to various diseases, including T2DM, cancer, and developmental arrest.
m6A enzymes include methyltransferases, demethylases, and m6A-binding proteins, and most of which are highly conserved among eukaryotes. Meanwhile, they also involve many important processes, and m6A enzyme deficiency can lead to developmental and functional defects in organisms. A comprehensive evolutionary analysis of the composition of m6A in 64 eukaryotes showed that methyltransferases METTL3 and METTL14 are highly conserved, although not detected in some organisms, such as Giardia intestinalis, Dictyostelium discoideum, and Caenorhabditis elegans [18] (Fig. 2). METTL3 is essential for many processes, including early embryonic development, ESC differentiation, and nervous system development [19,20,21]. Similarly, METTL14 plays a crucial role in maintaining the function of neuronal populations [22]. Its deletion leads to abnormal embryonal and nervous development [21, 23]. Furthermore, it has been confirmed that WTAP is strongly conserved, and its existence was related to METTL3 and METTL14 [18]. RBM15 is present in animals and plants and belongs to a family of RNA-binding motif proteins. It is involved in the process of hematopoietic differentiation [24]. RBM15 depletion has been associated with cardiovascular defects [25]. FTO is conserved in eukaryotes, and its expression has been confirmed in acute megakaryocytic leukemia. In consistence with its role as m6A readers, YTH domain-containing proteins usually exist in organisms with the METTL3-METTL14-WTAP complex. Therefore, m6A is mostly maintained among eukaryotes. Moreover, the broad conservation of m6A enzymes among very different species may suggest that m6A methylation modification is a profound regulatory mechanism that occurred in the last eukaryotic common ancestor.
Conservation of m6A enzymes among eukaryotes. The figure shows detected (colored box) or not detected (dark box) components of the m6A writers, readers, and eraser. On the left are the names of the different organisms being tested. The numbers of YTHDC and YTHDF family proteins are signed in the respective boxes
m6A Methyltransferase
m6A is a reversible modification dynamically regulated by methyltransferases and demethylases, which are also called “writers” and “erasers.” m6A “writers” consist of METTL3/14, KIAA1429, WTAP, zinc finger CCCH domain-containing protein 13 (ZC3H13), and RBM15/15B [26,27,28]. METLL3 and METTL14 are known as core components of the m6A methyltransferase complex [29]. Biochemical characterization has revealed that METTL14 can form heterodimers in a 1:1 ratio with METTL3 in vivo to synergistically enhance the methylation ability [29]. METTL3 is a catalytic subunit, while METTL14 is responsible for substrate recognition. A combination of METTL3 and METTL14 in a 1:1 ratio has been first identified in recombinant proteins isolated from insect cells. RNA probe assays revealed that the METTL3-METTL14 combination displayed significantly higher methyltransferase activity than METTL3 or METTL14 alone [29]. These findings suggest that they have a synergistic effect. It has been shown that METTL14 recognizes histone H3 lysine 36 trimethylation (H3K36me3) modification and mediates selective m6A deposition in mRNA [30, 31]. Although the exact mechanism of this process is unclear, it reveals a possible vital link between H3K36me3 and m6A modification, and indicates the functional importance of METTL14 in the selective binding of m6A and its deposition on mRNA. Besides METTL3 and METTL14, other methyltransferases, such as WTAP, ZC3H13, KIAA1429, and RBM15/RBM15B, play key roles in the localization of the methyltransferase complex in nuclear speckles and U-rich regions adjacent to the m6A sites in mRNAs [27, 28, 32,33,34]. In addition to these, widely studied methylated methyltransferase complexes, zinc finger CCHC-type-containing 4 (ZCCHC4), METTL5, and METTL16 have also been identified as m6A methyltransferases, which can function alone and catalyze m6A on certain structured RNAs, such as 18S rRNA, 28S rRNA, and U6 snRNA [35,36,37,38,39]. Recently, a novel methyltransferase, PCIF1, has been found to be responsible for methylation at the N6-position of adenosine in eukaryotic mRNAs [40, 41]. However, PCIF1 only methylates the first N6-position of adenosine on capped mRNAs but has no effect on the internal m6A formation. PCIF1 regulates cap-dependent mRNA translation through its unique methylation function. Although the specific mechanism has not been fully understood, it explains the influence of m6A on mRNA translation to some extent.
m6A Demethylases
m6A is demethylated by the action of demethylase. In eukaryotes, FTO and ALKBH5 are the two most widely known demethylation enzymes. FTO was shown to have demethylation activity in both single-stranded DNA at neutral pH and cellular mRNAs in vitro [42, 43]. Interestingly, a recent study by Jan et al. found that FTO preferentially demethylates N6, 2’-O-dimethyladenosine (m6Am) instead of m6A and decreases the stability of m6Am mRNAs [44]. The mechanism of ALKBH5 demethylation is to remove methyl groups from m6A-methylated adenosine rather than oxidative demethylation [45]. Zheng et al. [46] reported that ALKBH5 can co-localize with nuclear speckles, which are involved in RNA metabolism, and the assembly of mRNA processing factors in nuclear speckles. This study also demonstrated that ALKBH5 has a selective bias toward m6A within the consensus sequence [46]. Unlike FTO, which can demethylate several different nucleotides, ALKBH5 shows demethylation activity only on m6A [47].
m6A-Binding Proteins
The biological function of m6A modification is regulated by m6A-binding proteins, also called “readers,” including YTHDF1/2/3 and YTHDC1/2. These proteins link methyl-selective RNA to many cellular processes and trigger m6A-dependent regulation of pre-mRNA splicing, translation, initiation, and mRNA decay [3]. Wang et al. [48] revealed that YTHDF1 directly interacts with initiation factors to facilitate the translation of methylated mRNAs. In contrast, YTHDF2-mediated degradation can regulate the lifetime of target transcripts, thereby ensuring the effective production of proteins from dynamic transcripts that are labeled with m6A. With respect to mRNA decay, YTHDF2 can selectively recognize m6A modification sites and promote the degradation of these transcripts through recruiting the CCR4-NOT deadenylase complex [12, 49]. Also, YTHDF3 promotes YTHDF1-mediated cap-dependent translation of target mRNAs and interacts with eukaryotic initiation factor 3 (eIF3), and they collectively affect the distribution of methylated transcripts to YTHDF2, thereby accelerating decay [50, 51]. Although YTHDF1/2 proteins are identified as cytoplasmic m6A readers, YTHDC1 is a nuclear reader. YTHDC1 impacts mRNA splicing, which provides transcriptome-wide insights into the changes in splicing affected by this mRNA methylation reader protein [52]. Moreover, it also exports methylated mRNA from the nucleus to the cytoplasm, and deletion of YTHDC1 leads to a longer residence time of nuclear m6A-containing mRNA, accompanied by accumulation of transcripts in the nucleus and subsequent depletion within the cytoplasm [53]. Except for the YTH family, several RNA-binding proteins such as HNRNPC, HNRNPG, and HNRNPA2B1 also function as m6A readers binding to m6A-modified RNAs [54,55,56]. m6A-dependent RNA structural remodeling facilitates HNRNPC/G binding activities, thereby regulating pre-RNA splicing [54]. HNRNPA2B1, a nuclear m6A reader, enhances microprocessor protein binding with pre-micorRNA, thus promoting micorRNA processing in an m6A/METTL3-dependant manner [56].
m6A and CVDs
Currently, a number of studies focus on m6A and CVDs, such as cardiac hypertrophy, AS, and HF. In different CVDs, the effect of m6A modification is different. Here we explore the relationship between m6A and CVDs, with a view to provide a promising target for the treatment of CVDs.
m6A and Cardiac Remodeling
Cardiac remodeling refers to the compensatory or decompensatory changes of heart genes, proteins, cells, and intercellular materials under the stimulation of pathogenic factors. Emerging evidences reveal that m6A exerts an important role in cardiac remodeling, especially in cardiac hypertrophy and cardiac fibrosis [14, 16]. Cardiac hypertrophy, which occurs at increased workload of heart, has been identified as a common type of cardiac remodeling [57]. Cardiac hypertrophy plays a compensatory role for an increased workload during early stages. However, sustained pathological hypertrophy is an important cause of systolic dysfunction and HF [58]. In the emerging field of transcriptome machinery, mRNA m6A modification plays a key role in cardiac hypertrophy. For instance, m6A methyltransferase METTL3 has been found to regulate kinases and intracellular signaling pathways in the response to cardiac hypertrophy during pressure overload stimulation [14].
METTL3 has a direct effect on cardiac hypertrophy in an m6A-dependent manner. In 3-month-old mice, METTL3-mediated mRNA methylation at the level of N6-adenosine is increased during hypertrophic stimulation, which is necessary for the normal hypertrophic response of cardiomyocytes. Cardiac function (% fractional shortening) and cardiomyocytes size (cross-sectional area, length) measurements showed that increased m6A RNA methylation leads to compensatory myocardial hypertrophy, while reduced m6A drives eccentric cardiomyocyte remodeling and dysfunction, suggesting that this novel stress response mechanism is essential for maintaining normal cardiac function in the heart [14]. In addition, FTO, a widely known m6A demethylase, can improve cardiac contractile dysfunction caused by ischemia. This is achieved through the demethylation activity of FTO, which can selectively demethylate cardiac contractile transcripts, thereby inhibiting their degradation and improving their protein expression under ischemia [16]. FTO also regulates cardiomyocyte response during adipokine stimulation. Gan et al. [59] have demonstrated that nuclear FTO expression in cardiomyocytes as well as the involvement of FTO was significantly increased under adipofactor-induced cardiomyocyte hypertrophic response in cultured neonatal rat cardiomyocytes. Importantly, they also reported that FTO knockdown inhibited the hypertrophic response in neonatal rat cardiomyocytes, suggesting the important regulatory role of FTO in cardiac hypertrophy.
Cardiac fibrosis is a common pathological force in numerous CVDs, characterized by pathological activation of cardiac fibroblasts and excessive accumulation of extracellular matrix (ECM) in the affected tissue [60, 61]. As one of the most important methylation enzymes, METTL3-dependent m6A modification has been shown to regulate the progression of cardiac fibrosis, due to its important influence on the methylation level of fibrosis-related transcripts [62]. The studies indicated that overexpression METTL3 promoted cardiac fibrosis and transdifferentiated fibroblasts into myofibroblasts, while enhance extracellular matrix production in vitro [62]. Similarly, in the cardiac fibrotic mouse model, knockdown of METLL3 can effectively inhibit the progression of cardiac fibrosis [62]. Consistent with these results, a study of the m6A demethylase FTO also demonstrated that decreasing the level of m6A methylation effectively reduced cardiac fibrosis in mouse model of myocardial infarction [16]. These evidences indicated the vital role of m6A in cardiac fibrosis, and reducing the level of m6A methylation is expected to become a new molecular target for controlling fibrosis. We can hypothesize that m6A-dependent regulation of fibrosis, including RNAs splicing, translation, and degradation, maybe the potential novel mechanisms of epigenetics or epitranscriptomics in cardiac fibrosis [16] though the exact mechanisms and signal pathway require to be further investigated.
m6A and AS
AS is characterized by the thickened and hardened lesions, which are composed of lipids and calcifications in the intima and media of arteries [63]. It is a chronic systemic inflammatory disease that causes serious clinical complications and high mortality worldwide. Recently, a large number of researches suggested that m6A has a major effect on the progress and proliferation of AS. The development of AS begins with dysfunction and inflammation of vascular endothelial cells. By detecting the cardiovascular endothelial cells of the AS patients, Zhang et al. [64] found that METTL14, as a methylase, promotes the production of mature miR-19a via increasing the expression of m6A on miR-19a, thereby accelerating the invasion and proliferation of cardiovascular endothelial cells. This study provides new insights into the pathogenesis of AS. Moreover, it has been confirmed that METTL14 knockdown significantly represses TNF-α-induced endothelial cell inflammation by PI3K-Akt and TNF signaling pathways in human endothelial cells [65]. Subsequent in vivo experiments revealed that METTL14 gene knockout can inhibit AS plaque development in an m6A-dependent manner in METTL14 knockout mice [65]. Pyroptosis as a new modality of inflammatory programmed cell death and has been shown to be closely associated with AS. Decreasing m6A modification level on circ_0029589 can increase its expression, thus promoting inflammation and macrophage pyroptosis in peripheral blood mononuclear macrophages of patients with AS [66]. From what has been discussed above, these studies indicate that m6A influences the AS process and development through its regulation of cardiovascular endothelial cell function, endothelial cell inflammation, and macrophage pyroptosis (Fig. 3).
The role of m6A modification in AS. m6A modification facilitates invasion and proliferation of cardiovascular endothelial cells by increasing the production of mature miR-19a. Furthermore, m6A promotes endothelial cell inflammation via accelerating the inflammation-associated signaling pathways, such as the PI3K-Akt and TNF signaling pathways, thereby inhibiting atherosclerotic plaque development. In addition, m6A also represses the expression of circ_0029589, thus promoting macrophage pyroptosis in peripheral blood mononuclear macrophages of AS patients
m6A and HF
HF refers to when the blood supply from the heart is not able to supply the needs of the peripheral tissues [67]. HF contains two major subtypes: HF with preserved ejection fraction (HfpEF, EF≥50%) and HF with reduced ejection fraction (HfrEF, EF≤40%). The pathophysiological basis of HFpEF remains unclear. Recent investigations have shown that systemic low-level inflammation is a key etiology of HfpEF, and it can aggravate microvascular dysfunction and oxidative stress associated with metabolic syndrome [68, 69]. It has been demonstrated that m6A modification can reduce the inflammation level in mouse macrophages and human dental pulp cells (HDPCs) via inhibiting inflammation signaling pathways [70, 71]. This shows that m6A may prevent the occurrence of HfpEF.
HfrEF features reduced left ventricular systolic function. Recent research has demonstrated that m6A modification drives eccentric cardiomyocyte remodeling and accelerates the degradation of cardiac contractile transcripts that lead to cardiac dysfunction and HfrEF [14, 17]. Cardiomyocyte-specific deletion of FTO in mice results in a faster progression of HF with marked decrease in ejection fraction and enhanced dilatation [72]. Importantly, Mathiyalagan et al. [16] demonstrated that improving FTO expression in failing mouse hearts can alleviate the ischemia-induced increase in m6A and decrease in cardiac contractility. This is carried out by the demethylation activity of FTO, which can selectively demethylate cardiac contractile transcripts to prevent their degradation and improve protein expression under ischemia [16]. This conclusion is consistent with previous results that m6A affects protein abundance but does not alter mRNA expression levels, indicating a translation regulatory mechanism independent of transcription. These results provide a hypothesis that m6A, as an epigenetic modification, could regulate the progression of HfrEF by affecting the transcription process of the associated myocardial transcript without affecting the translation.
m6A and Congenital Heart Disease
Congenital heart disease is defined as an abnormality in gross cardiac anatomy that occurs in the uterus and represents the most common congenital anomaly groups [73, 74]. It contains left-to-right shunts (e.g., atrial septal defects, ventricular septal defects, and patent ductus arteriosus) and cyanotic congenital heart defects, such as tetralogy of Fallot. These heterogeneous disorders exist separately or as part of a more complex deformity [74]. ESCs are types of stem cells derived from the inner cell mass of the preimplantation embryo. They have the ability of self-renewal and the ability to differentiate into all cell types after injection into the blastocyst, including neural cells, pancreatic beta-cells, and cardiomyocytes [75, 76]. Fuegemann et al. [77] have demonstrated that mouse ESCs can differentiate into different cardiac subtype (nodal-like cells, atrial- and ventricular-) in vitro models. ESCs committed toward the cardiac lineage have been demonstrated to play a crucial role in the early stages of heart development in a mouse model [77]. In addition, stem cell therapy has offered a promising treatment strategy for congenital heart disease patients in recent years. Rupp et al. [78] showed that stem cell transplantation improves left ventricle function and cardiac remodeling. These reports elucidated that ESCs play a crucial role in both the normal heart development and the treatment of congenital heart disease. It has been revealed that as a widespread co-transcriptional modification, m6A can influence ESC directed differentiation [79]. This provides a hypothesis that m6A can regulate the occurrence of congenital heart disease via ESC differentiation.
Atrial and ventricular septal defects are the most common congenital diseases caused by cardiac developmental defects. Notably, cardiomyocytes derived from ESCs are important for maintaining normal heart development and function. A recent study demonstrated that METTL3 and appropriate m6A deposition are necessary for directed differentiation of mouse ESCs into cardiomyocytes [80]. Only 3% of METTL3 knockout colonies of two independent clones generated beating cardiomyocytes, while the wild-type cells could produce beating cardiomyocytes in 50% of colonies in directed differentiation of mouse ESCs in vitro [80]. In addition to METTL3, METTL14 and WTAP also belong to methyltransferases that catalyze m6A RNA modification in ESCs. It has been shown that depletion of METTL14 results in decreased levels of m6A modification in ESCs, increased expression of developmental regulators, and reduced self-renewal capability of ESCs [33]. Similar to METTL14 knockdown, knockdown of Zc3h13 also significantly reduces m6A methylation, impairs self-renewal, and triggers differentiation in ESCs [27]. These studies confirmed that m6A plays a crucial role in the differentiation of ESCs into cardiomyocytes, which is very important for maintaining the normal development and function of the heart. Absence of m6A expression may induce the occurrence of atrial and ventricular septal defects.
In addition, stem cell therapy has offered a promising treatment strategy for congenital heart disease patients in recent years. Numerous studies indicated that ESC transplantation effectively improves the function of damaged cardiomyocytes and left ventricle function [78, 81, 82]. Therefore, m6A modification may be useful for the prevention and treatment of congenital heart disease.
Effects of m6A on the CVD-Related Risk Factors
The CVD-related risk factors, such as obesity, inflammation, adipogenesis, and IR, contribute to the development of CVDs. Therefore, the impact of m6A on these CVD-related risk factors may help us to understand the exact mechanism of the effects of m6A on CVDs (Fig. 4).
Roles of m6A enzymes in the CVD-related risk factors. FTO enhances the occurence of obesity by increasing food intake. METTL3 and YTHDF2 inhibit inflammation response via repressing related signaling ways. METTL3 inhibits NF-κB and MAPK pathways, while YTHDF2 inhibits p65, p38, and ERK pathways. For adipogenesis, m6A methylases and demethylase have opposite effects on it. Methylases METTL3, METTL14, and WTAP promote the protein expression of CCNA2 and then accelerate adipogenesis. On the contrary, demethylase FTO decreases the expression of CCNA2 and CDK2, and then inhibits adipogenesis. In addition, m6A-binding protein YTHDF1 inhibits adipogenesis via increasing the expression of PNPLA2. METTL3 enhances fatty acids and subsequently promotes IR. FTO enhances the level of triglycerides and cholesterol, and subsequently promotes IR.  : Promote
: Promote  : Inhibit
: Inhibit
Obesity
Obesity often is defined as a body mass index (BMI) of ≥30 kg/m2. The association between severity of obesity and cardiometabolic risk factors has been observed even at early stages in life [83]. Obesity is generally regarded as a major and independent risk factor for CVDs, such as hypertension, atrial fibrillation, and HF [84, 85]. Accumulating evidences suggest that the prevalence of hypertension in overweight and obese patients is much higher than that in slim subjects [86,87,88]. The link between hypertension and the incidence of left ventricular hypertrophy has been well established and confirmed by multiple imaging modalities, including echocardiography, electrocardiogram, and cardiac magnetic resonance imaging [89]. Hypertension appears to promote left ventricular hypertrophy in a dose-dependent fashion with even prehypertensive patients at risk for remodeling [89, 90]. Furthermore, the prevalence of cardiac hypertrophy, including left ventricle geometric abnormalities and concentric left ventricular hypertrophy, is markedly increased in obese patients [59, 86].
People who carry FTO risk alleles usually have a high BMI, which may be owing to reduced food satiety and increased food intake, rather than energy expenditure [91, 92]. Several lines of evidence suggest that the strong association between FTO single nucleotide polymorphisms (SNPs) in intron 1 and overweight/obesity may be due to their potential effect on the expression of RPGRIP1L, IRX3, and IRX5, rather than the effect on FTO expression [92,93,94,95]. However, it has also been reported that such FTO SNPs are related to enhanced FTO expression [96, 97]. Moreover, the in vivo experiments showed that mice lacking FTO develop postnatal growth delay and have a decrease in both lean body mass and adipose tissue [98]. In contrast, FTO overexpression in mice can lead to obesity due to enhanced food intake [99], suggesting a crucial role of FTO in obesity. More importantly, as an m6A demethylase, it has been confirmed that the FTO obesity-risk allele (rs9939609 T/A) is related to decreased m6A ghrelin mRNA methylation and increased FTO expression [96]. Liu et al. [100] have demonstrated that inhibition of the expression of FTO increased m6A methylation levels and reduced the occurrence of obesity as well as CVDs. These results suggest that obesity plays a pivotal role in m6A-mediated CVD development.
Inflammation
Inflammation has been regarded as an immune response to tissue injury, harmful stimuli, or infection that protects the body against various pathogens and then restores homeostasis [101]. However, unresolved inflammation due to the inability to accurately control the immune response may result in changes in the expression of cancer-related genes, as well as posttranslational modifications of cellular proteins involved in DNA repair, cell cycle, and apoptosis, thereby promoting the development of cancer [102]. It is well known that prolonged and excessive inflammation triggers immune disorders and causes excessive tissue damage [103]. Inflammation has been reported to be involved in the development of various diseases, including diabetes, arthritis, CVDs, and cancer [104, 105]. In the cardiovascular system, inflammation is a common basis for the pathological and physiological changes in the occurrence and development of AS. Endothelial cells are activated to increase inflammatory cytokines, triggering the inflammatory response that is the mechanism of early AS [106, 107]. In advanced AS, a large number of inflammatory cytokines and macrophages infiltrate the vascular wall, secrete matrix metalloproteinases, and degrade collagen fibers in the extracellular matrix of the plaque, triggering plaque rupture, platelet aggregation, and thrombosis [108]. A number of studies have shown that anti-inflammatory treatments significantly reduced the development of atherosclerotic plaques in different animal models [109,110,111].
The RBP tristetraprolin (TTP) plays a crucial role in the control of inflammatory response by facilitating the degradation of pro-inflammatory cytokine mRNAs [112, 113]. YTHDF2 is an m6A-binding protein and its expression pattern is similar to TTP. YTHDF2 regulates mRNA stability by binding to a G (m6A) C consensus site of m6A, thereby affecting biological processes. Recently, Yu et al. [70] found that in the LPS-stimulated inflammatory response of RAW 264.7 murine macrophages, the YTHDF2 mRNA level was markedly elevated during the first 6h and then decreased. More than that, the authors also demonstrated that YTHDF2 decreases inflammatory cytokine expression via the p38, p65, and ERK signaling pathways in RAW 264.7 murine macrophages. Moreover, the m6A “writer” METTL3 has recently been revealed to play a key role in the regulation of inflammation. Feng et al. [71] reported that METTL3 can inhibit the inflammatory response under LPS stimulation through suppressing the MAPK and NF-κB signaling pathways in HDPCs. Based on these evidences, we can hypothesize that m6A affects CVDs, including AS, cardiac hypertrophy, and HF, by regulating inflammatory response signaling pathways.
Adipogenesis
Adipogenesis is an important factor that decides the adipose content inside the body. Effective isolation of lipids to prevent lipotoxicity in other tissues, such as liver, muscle, and heart, is crucial for maintaining metabolic homeostasis. Adipose tissue plays a crucial role in the innate immune system, far more than just an inert mass of energy storage [114]. However, when there is a disruption of adipogenesis, hypertrophic adipocytes lose their functional activities and adiponectin production [115], which increases the risk of AS.
Adipogenesis is a highly coordinated process regulated by extracellular signals and transcriptional cascades [116]. Numerous studies have shown that m6A is widely involved in regulating various aspects of adipogenesis. Kobayashi et al. [117] showed that the RNA N6-adenosine methyltransferase complex, which is composed of WTAP, METTL14, and METTL3, positively regulates adipogenesis by facilitating cell cycle transition in mitotic clonal expansion (MEC) during adipogenesis in 3T3-L1 cells. Deletion of each of these three proteins results in impaired adipogenesis and cell cycle arrest associated with inhibition of cyclin A2 (CCNA2) upregulation during MEC [117]. Furthermore, FTO has also been shown to have a negative effect on adipogenesis by relying on m6A. FTO can control the splicing of adipogenic regulatory factor RNA by modulating the level of m6A around the splicing site, thus affecting the differentiation of preadipocytes [118]. More than that, Wu et al. [119] showed that FTO knockdown suppressed adipogenesis by decreasing protein expression of cyclin-dependent kinase 2 (CDK2) and key cell cycle regulators, in an m6A-YTHDF2-dependent manner. This paper demonstrated a key role of FTO/m6A/YTHDF2 signaling in adipogenesis in 3T3-L1 cells [119]. As a homology of YTHDF2, Wang et al. [120] showed that m6A promotes the translation of patatin-like phospholipase domain containing 2 (PNPLA2) and increases protein expression through YTHDF1, thus suppressing adipogenesis in Landrace and Jinhua pigs.
In addition to the effect of m6A itself on adipogenesis, many RNA or protein effects on lipid formation also need to be realized through m6A. Mitochondrial carrier homology 2 (MTCH2) protein expression was positively correlated with m6A levels. It promotes adipogenesis in porcine intramuscular preadipocytes by an m6A-YTHDF1-dependent mechanism [121]. Epigallocatechin gallate (EGCG) enhances the induction of CCNA2 and CDK2 through YTHDF2 and FTO-induced adipogenesis inhibition in 3T3-L1 cells [122]. Zinc finger protein 217 (Zfp217) regulates m6A mRNA methylation through transcriptional activation of the m6A demethylase FTO, thus facilitating adipogenesis [123]. But interestingly, a subsequent study showed that ZFP217 increases the expression of the m6A methyltransferase METTL3, which upregulates the level of m6A and accelerates adipogenesis in 3T3-L1 cells [124]. These data suggest a new mechanism by which m6A influences AS by regulating adipogenesis.
IR
IR can be defined as decreased tissue responsiveness to insulin with increased production of insulin to provide a normal biological response [125]. A clinical investigation showed an increased prevalence of CVDs in the IR subset of patients with prediabetes mellitus despite fasting plasma glucose concentrations that are not different from the non-IR patients, and revealed the important role that differences in IR play in modulation of CVD risk in nondiabetic subjects [126]. A meta-analysis revealed that IR, evaluated by Homeostasis Model Assessment (HOMA), was a better predictor of CVD events than fasting levels of insulin or glucose in adults without diabetes mellitus [127]. Based on HOMA, a 1-unit increase in IR is associated with a 5.4% increase in CVD risk [128]. In addition, IR causes pathophysiological abnormalities, which adversely affect the structure and function of the heart [129]. IR-induced reactive oxygen species plays a causal role in left ventricular remodeling and myocardial dysfunction [129]. Moreover, 50% of normotensive, asymptomatic IR patients have diastolic dysfunction, which may lead to a 4- to 8-fold increase in the risk of HF and other myocardial dysfunctions that usually progress to sudden death [130]. In addition, IR is highly prevalent among nondiabetic patients with HF in comparison with healthy patients and is related to reduced exercise capacity in patients with HF [131].
Previous studies have elucidated the important roles of m6A methylase METTL3 and demethylase FTO in the regulation of IR [132,133,134]. Xie et al. [132] revealed that the IR index is positively correlated with the m6A level and METTL3 expression in the liver tissue of patients with type 2 diabetes, compared with nondiabetes subjects. Moreover, the authors also found that hepatocyte-specific deletion of METTL3 in mice decreased the m6A methylation and improved insulin sensitivity, which revealed the key roles for METTL3-mediated m6A methylation in IR [132]. Iskandar et al. [133] demonstrated that FTO rs9939609 gene contributed to a higher HOMA-IR index in type 2 diabetes, and it was significantly positively correlated to the familial history of diabetes. These results are novel and support a key role of m6A RNA methylation in the development of IR and HF, and point that m6A can be an effective therapeutic strategy for HF.
Hypertension
Hypertension is a long-term increase in blood pressure (BP) in the arteries. It is a serious chronic disease since persistent high BP negatively affects target organs such as the heart and kidney. It is known that hypertension is an important risk factor closely related to CVDs [135]. Mo et al. showed that m6A plays a crucial role in the regulation of BP [136]. Genetic variation influences m6A expression by altering the RNA sequence of target sites, which is called m6A-associated SNP [137]. Many m6A-associated SNPs, including rs9847953 and rs197922, affect related gene expression (e.g., C1orf167, DOT1L), resulting in effect BP [136].
In spontaneously hypertensive rat pericytes, m6A expression levels and distribution sites are different from those in Wistar Kyoto pericytes [138]. The study revealed that compared with Wistar Kyoto rat pericytes, the m6A methylation level of spontaneously hypertensive rat pericytes reduced, meaning that m6A meth ylation was altered when hypertension occurred. In addition, m6A sequencing and gene ontology enrichment analysis found that the increase m6A peaks in hypertensive rat pericytes were mainly associated with inflammatory response, RNA methyltransferase activity, and proximal tubule development. These findings may allow an illustration of the underlying mechanism of hypertension in the perspective of m6A modification.
T2DM
T2DM is a common metabolic disease, characterized by disorder of glucose metabolism and hyperglycemia. As an independent risk factor for CVDs, T2DM causes a variety of cardiovascular complications, and more importantly, increases mortality in people with cardiovascular disease [139]. In view of the critical role of environment and lifestyle in T2DM, epigenetic modifications that change under environmental stimuli are likely to have special significance [140]. As a widespread epigenetic modification in eukaryotic RNAs, the lower level of m6A has been found and associated with increased FTO expression but not ALKBH5 expression in T2DM patients [141]. FTO mRNA expression was significantly enhanced in response to high glucose stimulation, thus reducing the m6A content [142]. These results suggest that m6A is vital for blood glucose regulation and T2DM, and we note that this may be related to the regulation of m6A on the insulin secretion, IR, and liver gluconeogenesis [142, 143].
A study of islet cells in people with T2DM showed that m6A was significantly reduced in β-cells, rather than α-cells, which provided an evidence for m6A to control cell insulin secretion [143]. Subsequent gene ontology revealed that m6A was widely present in the insulin secretion-related genes [143]. More importantly, further pathway analyses indicated that low m6A downregulated insulin/IGF1–AKT–PDX1 pathway, which is crucial in insulin secretion, thus impaired insulin secretion [143]. Moreover, FTO positively regulates gluconeogenic-related genes, such as glucose-6-phosphatase catalytic subunit (G6Pc) and forkhead box O1 (FOXO1), in m6A-dependent manner. In T2DM human, decreased m6A promoted hepatic gluconeogenesis, leading to elevated blood glucose through decreasing the expression of gluconeogenic-related genes [142]. These suggest that m6A act as an important role in T2DM, and low m6A may serve as a new potential biomarker for T2DM, which needs further confirmation.
Effects of N6-methyladenine DNA modification on CVDs
DNA methylation is an important component of epigenetic modifications involved in regulation of many disease processes. The most well-characterized type of DNA methylation in eukaryotes is 5-methylcytosine (5mC). Contrarily, N6-methyladenine DNA (6mA) modification is identified as the most common DNA modification in prokaryotes. Importantly, several lines of evidence have demonstrated that 6mA modification is widely present in the human genome, particularly in the mitochondria, in which (G/C) AGG (C/T) is the most significantly associated motif with 6mA modification [144, 145]. ALKBH1 and N-6 adenine-specific DNA methyltransferase 1 (N6AMT1) have been identified as the demethylase and methyltransferase for 6mA modification in human cells, respectively [144].
Zhang et al. [146] explored epigenetic modifications in the cardiovascular system and found that DNA methylation and demethylation are important factors involved in cardiovascular aging and CVDs. Numerous studies have shown that the risk factors of CVDs, such as obesity, inflammation, smoking, and IR, can lead to dysregulation of DNA methylation [147,148,149,150]. A longitudinal study of 11461 participants from population-based cohorts has shown that the methylation quantitative trait loci (meQTL) promote gene expression, demonstrating a potential causal relationship between DNA methylation and the occurrence of coronary heart disease/myocardial infarction [151]. A recent study by Guo et al. found that decreased 6mA modification is implicated in the development of hypertension [152]. Elevated ALKBH1 expression is responsible for reduced 6mA DNA level in VSMCs both in vitro and in vivo hypertension models [152], suggesting a potential epigenetic role for ALKBH1-6mA regulation in the development of CVDs. In addition, several investigators have investigated that levels of global NA methylation are significantly elevated in human AS plaques (Methylation of the estrogen receptor gene is associated with aging and atherosclerosis in the cardiovascular system), indicating a close relationship between DNA methylation and AS. Taken together, these findings suggest that methylation modification of DNA in blood vessel cells and mitochondria plays an important role in CVDs, which may be due to the influence of 6mA modification on the generation of related lncRNAs and miRNAs.
Conclusion and Future Perspectives
m6A research has revealed potential links between this epigenetic modification and CVDs. m6A has a wild range of effects on obesity, adipogenesis, inflammation, ESC differentiation, IR, hypertension, and T2DM. These may explain the mechanisms via which m6A affects the development of CVDs, including cardiac remodeling, AS, HF, and congenital heart disease.
However, m6A research is a relatively new field and many problems are still unexplained. At present, studies of the effects of m6A methylases on CVDs mainly focus on METTL3. The role of other enzymes that also have methylation activity and are even homologs of METTL3, such as METTL14 and WTAP, in CVDs is still not clear. Similarly, the role of m6A demethylases belonging to the AlkB family in the development of CVDs is unknown, besides FTO. In addition to this, m6A has been reported to enhance the endothelial cell-induced angiogenic response in an animal model [16]. When endothelial cells are dysfunctional, their mediated angiogenic response is reduced, thereby contributing to HF [153, 154]. Whether endothelial cell dysfunction is a potential pathological mechanism linking m6A and CVDs is still unknown. Our paper has provided a hypothesis that m6A may influence the development of congenital heart disease by affecting stem cell differentiation, but there is still no direct evidence for the role of m6A in congenital heart disease. Moreover, a recent study confirmed that transcripts of Ca+2-handling SERCA2a and RYR2 were hypermethylated in the failing human hearts’ left ventricle tissue [16]. However, it is not completely clear whether m6A can regulate the cardiac calcium pathway or even cause arrhythmia, and more studies are needed to prove it. In the future, a more in-depth exploration of m6A methylation modification may provide a novel therapeutic strategy for CVDs.
Abbreviations
- m6A:
-
N6-methyladenosine
- AS:
-
Atherosclerosis
- IR:
-
Insulin resistance
- T2DM:
-
Type 2 diabetes mellitus
- WTAP:
-
Wilms’ tumor 1-associating protein
- RBM15:
-
RNA-binding motif protein 15
- METTL3:
-
Methyltransferase-like 3
- METTL14:
-
Methyltransferase-like 14
- ALKBH5:
-
ALKB homolog 5
- FTO:
-
Fat mass and obesity-associated
- YTH:
-
YT521-B homology
- CVDs:
-
Cardiovascular diseases
- HF:
-
Heart failure
- ESC:
-
Embryonic stem cell
- ZC3H13:
-
Zinc finger CCCH domain-containing protein 13
- RBM:
-
RNA-binding motif protein
- H3K36me3:
-
Histone H3 lysine 36 trimethylation
- ZCCHC4:
-
Zinc finger CCHC-type-containing 4
- m6Am:
-
N6, 2’-O-dimethyladenosine
- eIF3:
-
Eukaryotic initiation factor 3
- HfpEF:
-
HF with preserved ejection fraction
- HfrEF:
-
HF with reduced ejection fraction
- HDPCs:
-
Human dental pulp cells
- BMI:
-
Body mass index
- SNPs:
-
Single nucleotide polymorphisins
- TTP:
-
The RBP tristetraprolin
- MEC:
-
Mitotic clonal expansion
- CCNA2:
-
Cyclin A2
- CDK2:
-
Cyclin-dependent kinase 2
- PNPLA2:
-
Patatin-like phospholipase domain containing 2
- MTCH2:
-
Mitochondrial carrier homology 2
- EGCG:
-
Epigallocatechin gallate
- Zfp217:
-
Zinc finger protein 217
- HOMA:
-
Homeostasis Model Assessment
- BP:
-
Blood pressure
- G6Pc:
-
Glucose-6-phosphatase catalytic subunit
- FOXO1:
-
Forkhead box O1
- 6mA:
-
N6-methyladenine DNA
- 5mC:
-
5-methylcytosine
- N6AMT1:
-
N-6 adenine-specific DNA methyltransferase 1
- meQTL:
-
Methylation quantitative trait loci
References
Kim, T. K., Gore, S. D., & Zeidan, A. M. (2015). Epigenetic therapy in acute myeloid leukemia: current and future directions. Seminars in Hematology, 52, 172–183.
Wu, X., Sang, L., & Gong, Y. (2018). N6-methyladenine RNA modification and cancers. American Journal of Cancer Research, 8, 1957–1966.
Roundtree, I. A., Evans, M. E., Pan, T., & He, C. (2017). Dynamic RNA modifications in gene expression regulation. Cell, 169, 1187–1200.
Adams, J. M., & Cory, S. (1975). Modified nucleosides and bizarre 5’-termini in mouse myeloma mRNA. Nature, 255, 28–33.
Desrosiers, R., Friderici, K., & Rottman, F. (1974). Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proceedings of the National Academy of Sciences of the United States of America, 71, 3971–3975.
Yang, D., Qiao, J., Wang, G., Lan, Y., Li, G., Guo, X., et al. (2018). N6-Methyladenosine modification of lincRNA 1281 is critically required for mESC differentiation potential. Nucleic Acids Research, 46(8), 3906–3920.
Alarcón, C. R., Lee, H., Goodarzi, H., Halberg, N., & Tavazoie, S. F. (2015). N6-methyladenosine marks primary microRNAs for processing. Nature, 519(7544), 482–485.
Chen, Y. G., Chen, R., Ahmad, S., Verma, R., Kasturi, S. P., Amaya, L., et al. (2019). N6-methyladenosine modification controls circular rna immunity. Molecular Cell, 76(1), 96–109.
Jia, G., Fu, Y., & He, C. (2013). Reversible RNA adenosine methylation in biological regulation. Trends in Genetics, 29, 108–115.
Niu, Y., Zhao, X., Wu, Y. S., & Li, M. M. (2013). N6-methyl-adenosine (m6A) in RNA: an old modificatio with a novel epigenetic function. Genomics, Proteomics & Bioinformatics, 11, 8–17.
Fustin, J. M., Doi, M., Yamaguchi, Y., & Hida, H. (2013). RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell, 155, 793–806.
Wang, X., Lu, Z., Gomez, A., & Hon, C. G. (2014). N6-methyladenosine-dependent regulation of messenger RNA stability. Nature, 505, 117–120.
Schwartz, S., Agarwala, S. D., Mumbach, M. R., & Jovanovic, M. (2013). High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell, 155, 1409–1421.
Dorn, L. E., Lasman, L., Chen, J., & Xu, X. (2019). The N(6)-methyladenosine mRNA methylase METTL3 controls cardiac homeostasis and hypertrophy. Circulation, 139, 533–545.
Kmietczyk, V., Riechert, E., Kalinski, L., & Boileau, E. (2019). m(6)A-mRNA methylation regulates cardiac gene expression and cellular growth. Life Science Alliance, 2, 2.
Mathiyalagan, P., Adamiak, M., Mayourian, J., & Sassi, Y. (2019). FTO-dependent N(6)-methyladenosine regulates cardiac function during remodeling and repair. Circulation, 139, 518–532.
Song, H., Feng, X., Zhang, H., Luo, Y., Huang, J., Lin, M., et al. (2019). METTL3 and ALKBH5 oppositely regulate m6A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy, 15(8), 1419–1437.
Balacco, D. L., & Soller, M. (2019). The m6A writer: rise of a machine for growing tasks. Biochemistry, 58(5), 363–378.
Frye, M., Harada, B. T., Behm, M., & He, C. (2018). RNA modifications modulate gene expression during development. Science, 361(6409), 1346–1349.
Geula, S., Moshitch-Moshkovitz, S., Dominissini, D., Mansour, A. A., Kol, N., Salmon-Divon, M., et al. (2015). m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science, 347(6225), 1002–1006.
Ma, C., Chang, M., Lv, H., Zhang, Z. W., Zhang, W., He, X., et al. (2018). RNA m6A methylation participates in regulation of postnatal development of the mouse cerebellum. Genome Biology, 19(1), 68.
Yoon, K. J., Ringeling, F. R., Vissers, C., Jacob, F., Pokrass, M., Jimenez-Cyrus, D., et al. (2017). Temporal control of mammalian cortical neurogenesis by m6A methylation. Cell, 171(4), 877–889.
Meng, T. G., Lu, X., Guo, L., Hou, G. M., Ma, X. S., Li, Q. N., et al. (2019). Mettl14 is required for mouse postimplantation development by facilitating epiblast maturation. The FASEB Journal, 33(1), 1179–1187.
Tran, N. T., Su, H., Khodadadi-Jamayran, A., Lin, S., Zhang, L., Zhou, D., et al. (2016). The AS-RBM15 lncRNA enhances RBM15 protein translation during megakaryocyte differentiation. EMBO Reports, 17(6), 887–900.
Raffel, G. D., Chu, G. C., Jesneck, J. L., Cullen, D. E., Bronson, R. T., Bernard, O. A., et al. (2016). Ott1 (Rbm15) is essential for placental vascular branching morphogenesis and embryonic development of the heart and spleen. Molecular and Cellular Biology, 29(2), 333–341.
Yue, Y., Liu, J., Cui, X., & Cao, J. (2018). VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discovery, 4, 10.
Wen, J., Lv, R., Ma, H., & Shen, H. (2018). Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Molecular Cell, 69, 1028–1038.
Patil, D. P., Chen, C. K., & Pickering, B. F. A. (2016). Chow, m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature, 537, 369–373.
Liu, J., Yue, Y., Han, D., & Wang, X. (2014). A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nature Chemical Biology, 10, 93–105.
Huang, H., Weng, H., & Chen, J. (2020). The biogenesis and precise control of RNA m(6)A methylation. Trends in Genetics, 36, 44–52.
Huang, H., Weng, H., Zhou, K., & Wu, T. (2019). Histone H3 trimethylation at lysine 36 guides m(6)A RNA modification co-transcriptionally. Nature, 567, 414–419.
Ping, X. L., Sun, B. F., Wang, L., & Xiao, W. (2014). Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Research, 24, 177–189.
Wang, Y., Li, Y., Toth, J. I., & Petroski, M. D. (2014). N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biology, 16, 191–198.
Schwartz, S., Mumbach, M. R., Jovanovic, M., & Wang, T. (2014). Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Reports, 8, 284–296.
Mendel, M., Chen, K. M., Homolka, D., & Gos, P. (2018). Methylation of structured RNA by the m(6)A writer METTL16 is essential for mouse embryonic development. Molecular Cell, 71, 986–1000.
Pendleton, K. E., Chen, B., Liu, K., & Hunter, O. V. (2017). The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell, 169, 824–835.
Ma, H., & Wang, X. (2019). N(6-)Methyladenosine methyltransferase ZCCHC4 mediates ribosomal RNA methylation. Nature Chemical Biology, 15, 88–94.
Nvan, T., Ernst, F. G. M., Hawley, B. R., & Zorbas, C. (2019). The human 18S rRNA m6A methyltransferase METTL5 is stabilized by TRMT112. Nucleic Acids Research, 47, 7719–7733.
Warda, A. S., Kretschmer, J., Hackert, P., & Lenz, C. (2017). Human METTL16 is a N(6)-methyladenosine (m(6)A) methyltransferase that targets pre-mRNAs and various non-coding RNAs. EMBO Reports, 18, 2004–2014.
Akichika, S., Hirano, S., & Shichino, Y. (2019). Cap-specific terminal N (6)-methylation of RNA by an RNA polymerase II-associated methyltransferase. Science, 363(64213).
Sun, H., Zhang, M., Li, K., & Bai, D. (2019). Cap-specific, terminal N(6)-methylation by a mammalian m(6)Am methyltransferase. Cell Research, 29, 80–82.
Jia, G., Fu, Y., Zhao, X., & Dai, Q. (2011). N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chemical Biology, 7, 885–887.
Gerken, T., Girard, C. A., Tung, Y. C., & Webby, C. J. (2007). The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science, 318, 1469–1472.
Mauer, J., Luo, X., Blanjoie, A., & Jiao, X. (2017). Reversible methylation of m(6)Am in the 5’ cap controls mRNA stability. Nature, 541, 371–375.
Chen, W., Zhang, L., Zheng, G., & Fu, Y. (2014). Crystal structure of the RNA demethylase ALKBH5 from zebrafish. FEBS Letters, 588, 892–898.
Zheng, G., Dahl, J. A., Niu, Y., & Fedorcsak, P. (2013). ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Molecular Cell, 49, 18–29.
Feng, C., Liu, Y., Wang, G., & Deng, Z. (2014). Crystal structures of the human RNA demethylase Alkbh5 reveal basis for substrate recognition. The Journal of Biological Chemistry, 289, 11571–11583.
Wang, X., Zhao, B. S., Roundtree, I. A., & Lu, Z. (2015). N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell, 161, 1388–1399.
Du, H., Zhao, Y., He, J., & Zhang, Y. (2016). YTHDF2 destabilizes m(6)A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nature Communications, 7, 12626.
Shi, H., Wang, X., Lu, Z., & Zhao, B. S. (2017). YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Research, 27, 315–328.
Meyer, K. D., Patil, D. P., Zhou, J., & Zinoviev, A. (2015). 5’ UTR m(6)A promotes Cap-independent translation. Cell, 163, 999–1010.
Roundtree, I. A., & He, C. (2016). Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Trends in Genetics, 32, 320–321.
Roundtree, I. A., Luo, G. Z., Zhang, Z., & Wang, X. (2017). YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife, 6.
Liu, N., Dai, Q., Zheng, G., He, C., Parisien, M., & Pan, T. (2015). N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature, 518(7540), 560–564.
Zhou, K. I., Shi, H., Lyu, R., Wylder, A. C., Matuszek, Ż., Pan, J. N., et al. (2019). Regulation of co-transcriptional pre-mRNA splicing by m6A through the low-complexity protein hnRNPG. Molecular Cell, 76(1), 70–81.
Alarcón, C. R., Goodarzi, H., Lee, H., Liu, X., Tavazoie, S., & Tavazoie, S. F. (2015). HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell., 162(6), 1299–1308.
Kehat, I., & Molkentin, J. D. (2010). Molecular pathways underlying cardiac remodeling during pathophysiological stimulation. Circulation, 122, 2727–2735.
Li, L., Xu, J., He, L., & Peng, L. (2016). The role of autophagy in cardiac hypertrophy. Acta Biochimica et Biophysica Sinica Shanghai, 48, 491–500.
Gan, X. T., Zhao, G., Huang, C. X., & Rowe, A. C. (2013). Identification of fat mass and obesity associated (FTO) protein expression in cardiomyocytes: regulation by leptin and its contribution to leptin-induced hypertrophy. PLoS One, 8.
Frangogiannis, N. G. (2019). Cardiac fibrosis: cell biological mechanisms, molecular pathways and therapeutic opportunities. Molecular Aspects of Medicine, 65, 70–99.
González, A., Schelbert, E. B., Díez, J., & Butler, J. (2018). Myocardial interstitial fibrosis in heart failure: biological and translational perspectives. Journal of the American College of Cardiology, 71(15), 1696–1706.
Li, T., Zhuang, Y., Yang, W., Xie, Y., Shang, W., Su, S., et al. (2021). Silencing of METTL3 attenuates cardiac fibrosis induced by myocardial infarction via inhibiting the activation of cardiac fibroblasts. The FASEB Journal, 35(2), e21162.
Chen, P. Y., Qin, L., Baeyens, N., & Li, G. (2015). Endothelial-to-mesenchymal transition drives atherosclerosis progression. The Journal of Clinical Investigation, 125, 4514–4528.
Zhang, B. Y., Han, L., Tang, Y. F., Zhang, G. X., Fan, X. L., Zhang, J. J., et al. (2020). METTL14 regulates M6A methylation-modified primary miR-19a to promote cardiovascular endothelial cell proliferation and invasion. European Review for Medical and Pharmacological Sciences, 24(12), 7015–7023.
Jian, D., Wang, Y., Jian, L., Tang, H., Rao, L., Chen, K., et al. (2020). METTL14 aggravates endothelial inflammation and atherosclerosis by increasing FOXO1 N6-methyladeosine modifications. Theranostics, 10(20), 8939–8956.
Guo, M., Yan, R., Ji, Q., Yao, H., Sun, M., Duan, L., et al. (2020). IFN regulatory Factor-1 induced macrophage pyroptosis by modulating m6A modification of circ_0029589 in patients with acute coronary syndrome. International Immunopharmacology, 86, 106800.
Savarese, G., & Lund, L. H. (2017). Global public health burden of heart failure. Cardiac Failure Review, 3, 7–11.
Azevedo, P. S., Polegato, B. F., Minicucci, M. F., & Paiva, S. A. (2016). Cardiac remodeling: concepts, clinical impact, pathophysiological mechanisms and pharmacologic treatment. Arquivos Brasileiros de Cardiologia, 106, 62–69.
Toischer, K., Rokita, A. G., Unsold, B., & Zhu, W. (2010). Differential cardiac remodeling in preload versus afterload. Circulation, 122, 993–1003.
Yu, R., Li, Q., Feng, Z., & Cai, L. (2019). m6A reader YTHDF2 regulates LPS-induced inflammatory response. International Journal of Molecular Sciences, 20.
Feng, Z., Li, Q., Meng, R., & Yi, B. (2018). METTL3 regulates alternative splicing of MyD88 upon the lipopolysaccharide-induced inflammatory response in human dental pulp cells. Journal of Cellular and Molecular Medicine, 22, 2558–2568.
Berulava T, Buchholz Es, V. Elerdashvili, Pena T. (2020). Changes in m6A RNA methylation contribute to heart failure progression by modulating translation. European Journal of Heart Failure, 22(1), 54-66.
Sun, R., Liu, M., Lu, L., & Zheng, Y. (2015). Congenital heart disease: causes, diagnosis, symptoms, and treatments. Cell Biochemistry and Biophysics, 72, 857–860.
Khoshnood, B., Lelong, N., Houyel, L., & Thieulin, A. C. (2012). Prevalence, timing of diagnosis and mortality of newborns with congenital heart defects: a population-based study. Heart, 98, 1667–1673.
Lorzadeh, N., & Kazemirad, N. (2018). Embryonic stem cells and infertility. American Journal of Perinatology, 35, 925–930.
Stubbs, S. L., Crook, J. M., Morrison, W. A., & Newcomb, A. E. (2011). Toward clinical application of stem cells for cardiac regeneration. Heart, Lung & Circulation, 20, 173–179.
Fuegemann, C. J., Samraj, A. K., Walsh, & Fleischmann, B. K. (2010). Differentiation of mouse embryonic stem cells into cardiomyocytes via the hanging-drop and mass culture methods. Current Protocols in Stem Cell Biology, 1.
Rupp, S., Zeiher, A. M., Dimmeler, S., & Tonn, T. (2010). A regenerative strategy for heart failure in hypoplastic left heart syndrome: intracoronary administration of autologous bone marrow-derived progenitor cells. The Journal of Heart and Lung Transplantation, 29, 574–577.
Slobodin, B., Han, R., Calderone, V., Vrielink, J. A. F. O., Loayza-Puch, F., Elkon, R., et al. (2017). Transcription impacts the efficiency of mRNA translation via co-transcriptional N6-adenosine methylation. Cell, 169(2), 326–337.
Batista, P. J., Molinie, B., Wang, J., & Qu, K. (2014). m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell, 15, 707–719.
Kadota, S., Pabon, L., Reinecke, H., & Murry, C. E. (2017). In vivo maturation of human induced pluripotent stem cell-derived cardiomyocytes in neonatal and adult rat hearts. Stem Cell Reports, 8, 278–289.
Cho, G. S., Lee, D. I., Tampakakis, E., & Murphy, S. (2017). Neonatal transplantation confers maturation of PSC-derived cardiomyocytes conducive to modeling cardiomyopathy. Cell Reports, 18, 571–582.
Skinner, A. C., Perrin, E. M., Moss, L. A., & Skelton, J. A. (2015). Cardiometabolic risks and severity of obesity in children and young adults. The New England Journal of Medicine, 373, 1307–1317.
Lavie, C. J., De Schutter, A., Parto, P., & Jahangir, E. (2016). Obesity and prevalence of cardiovascular diseases and prognosis-the obesity paradox updated. Progress in Cardiovascular Diseases, 58, 537–547.
Lavie, C. J., Sharma, A., Alpert, M. A., & De Schutter, A. (2016). Update on obesity and obesity paradox in heart failure. Progress in Cardiovascular Diseases, 58, 393–400.
Lavie, C. J., Patel, D. A., Milani, R. V., & Ventura, H. O. (2014). Impact of echocardiographic left ventricular geometry on clinical prognosis. Progress in Cardiovascular Diseases, 57, 3–9.
Bastien, M., Poirier, P., Lemieux, I., & Despres, J. P. (2014). Overview of epidemiology and contribution of obesity to cardiovascular disease. Progress in Cardiovascular Diseases, 56, 369–381.
Ng, M., Fleming, T., Robinson, M., & Thomson, B. (2014). Global, regional, and national prevalence of overweight and obesity in children and adults during 1980-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet, 384, 766–781.
Heckbert, S. R., Post, W., Pearson, G. D., & Arnett, D. K. (2006). Traditional cardiovascular risk factors in relation to left ventricular mass, volume, and systolic function by cardiac magnetic resonance imaging: the Multiethnic Study of Atherosclerosis. Journal of the American College of Cardiology, 48, 2285–2292.
Santos, A. B., Gupta, D. K., Bello, N. A., & Gori, M. (2016). Prehypertension is associated with abnormalities of cardiac structure and function in the atherosclerosis risk in communities study. American Journal of Hypertension, 29, 568–574.
Haupt, A., Thamer, C., Staiger, H., & Tschritter, O. (2009). Variation in the FTO gene influences food intake but not energy expenditure. Experimental and Clinical Endocrinology & Diabetes, 117, 194–197.
Cecil, J. E., Tavendale, R., Watt, P., & Hetherington, M. M. (2008). An obesity-associated FTO gene variant and increased energy intake in children. The New England Journal of Medicine, 359, 2558–2566.
Smemo, S., Tena, J. J., Kim, K. H., & Gamazon, E. R. (2014). Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature, 507, 371–375.
Claussnitzer, M., Dankel, S. N., Kim, K. H., & Quon, G. (2015). FTO obesity variant circuitry and adipocyte browning in humans. The New England Journal of Medicine, 373, 895–907.
Stratigopoulos, G., Martin Carli, J. F., O’Day, D. R., & Wang, L. (2014). Hypomorphism for RPGRIP1L, a ciliary gene vicinal to the FTO locus, causes increased adiposity in mice. Cell Metabolism, 19, 767–779.
Karra, E., O’Daly, O. G., Choudhury, A. I., & Yousseif, A. (2013). A link between FTO, ghrelin, and impaired brain food-cue responsivity. The Journal of Clinical Investigation, 123, 3539–3551.
Melnik, B. C. (2015). Milk: an epigenetic amplifier of FTO-mediated transcription? Implications for Western diseases. Journal of Translational Medicine, 13, 385.
Fischer, J., Koch, L., Emmerling, C., & Vierkotten, J. (2009). Inactivation of the Fto gene protects from obesity. Nature, 458, 894–898.
Church, C., Moir, L., McMurray, F., & Girard, C. (2010). Overexpression of Fto leads to increased food intake and results in obesity. Nature Genetics, 42, 1086–1092.
Liu, C., Mou, S., & Pan, C. (2013). The FTO gene rs9939609 polymorphism predicts risk of cardiovascular disease: a systematic review and meta-analysis. PLoS One, 8.
Fernandes, J. V., Cobucci, R. N., Jatoba, C. A., & Fernandes, T. A. (2015). The role of the mediators of inflammation in cancer development. Pathology Oncology Research, 21, 527–534.
Murata, M. (2018). Inflammation and cancer. Environmental Health and Preventive Medicine, 23, 50.
Park, S. B., Park, G. H., Um, Y., & Kim, H. N. (2018). Wood-cultivated ginseng exerts anti-inflammatory effect in LPS-stimulated RAW264.7 cells. International Journal of Biological Macromolecules, 116, 327–334.
Zou, Y. H., Zhao, L., Xu, Y. K., & Bao, J. M. (2018). Anti-inflammatory sesquiterpenoids from the Traditional Chinese Medicine Salvia plebeia: regulates pro-inflammatory mediators through inhibition of NF-kappaB and Erk1/2 signaling pathways in LPS-induced Raw264.7 cells. Journal of Ethnopharmacology, 210, 95–106.
Zou, J., Guo, P., Lv, N., & Huang, D. (2015). Lipopolysaccharide-induced tumor necrosis factor-alpha factor enhances inflammation and is associated with cancer (Review). Molecular Medicine Reports, 12, 6399–6404.
Tabas, I., Garcia-Cardena, G., & Owens, G. K. (2015). Recent insights into the cellular biology of atherosclerosis. The Journal of Cell Biology, 209, 13–22.
Chistiakov, D. A., Melnichenko, A. A., Grechko, A. V., & Myasoedova, V. A. (2018). Potential of anti-inflammatory agents for treatment of atherosclerosis. Experimental and Molecular Pathology, 104, 114–124.
Liu, Y., Yu, H., Zhang, Y., & Zhao, Y. (2008). TLRs are important inflammatory factors in atherosclerosis and may be a therapeutic target. Medical Hypotheses, 70, 314–316.
Rosenson, R. S., Hislop, C., Elliott, M., & Stasiv, Y. (2010). Effects of varespladib methyl on biomarkers and major cardiovascular events in acute coronary syndrome patients. Journal of the American College of Cardiology, 56, 1079–1088.
Alaarg, A., Senders, M. L., Varela-Moreira, A., & Perez-Medina, C. (2017). A systematic comparison of clinically viable nanomedicines targeting HMG-CoA reductase in inflammatory atherosclerosis. Journal of Controlled Release, 262, 47–57.
Alaarg, A., Zheng, K. H., van der Valk, F. M., & da Silva, A. E. (2016). Multiple pathway assessment to predict anti-atherogenic efficacy of drugs targeting macrophages in atherosclerotic plaques. Vascular Pharmacology, 82, 51–59.
Tiedje, C., Diaz-Munoz, M. D., Trulley, P., Ahlfors, H., Laass, K., Blackshear, P. J., et al. (2016). The RNAbinding protein TTP is a global post-transcriptional regulator of feedback control in inflammation. Nucleic Acids Research, 44, 7418–7440.
Bulbrook, D., Brazier, H., Mahajan, P., Kliszczak, M., Fedorov, O., Marchese, F. P., et al. (2018). Tryptophan-mediated interactions between tristetraprolin and the CNOT9 subunit are required for CCR4-NOT deadenylase complex recruitment. Journal of Molecular Biology, 430(5), 722–736.
Hafidi, M. E., & Buelna-Chontal, M. (2019). Adipogenesis: a necessary but harmful strategy. International Journal of Molecular Sciences, 20.
Woo, C. Y., Jang, J. E., Lee, S. E., & Koh, E. H. (2019). Mitochondrial dysfunction in adipocytes as a primary cause of adipose tissue inflammation. Diabetes and Metabolism Journal, 43, 247–256.
Rosen, E. D., & MacDougald, O. A. (2006). Adipocyte differentiation from the inside out. Nature Reviews. Molecular Cell Biology, 7, 885–896.
Kobayashi, M., Ohsugi, M., Sasako, T., & Awazawa, M. (2018). The RNA methyltransferase complex of WTAP, METTL3, and METTL14 regulates mitotic clonal expansion in adipogenesis. Molecular and Cellular Biology, 38.
Zhao, X., Yang, Y., Sun, B. F., & Shi, Y. (2014). FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Research, 24, 1403–1419.
Wu, R., Liu, Y., Yao, Y., & Zhao, Y. (2018). FTO regulates adipogenesis by controlling cell cycle progression via m(6)A-YTHDF2 dependent mechanism. Biochimica et Biophysica Acta - Molecular and Cell Biology of Lipids, 1863, 1323–1330.
Wang, X., Sun, B., Jiang, Q., & Wu, R. (2018). mRNA m(6)A plays opposite role in regulating UCP2 and PNPLA2 protein expression in adipocytes. International Journal of Obesity, 42, 1912–1924.
Jiang, Q., Sun, B., Liu, Q., & Cai, M. (2019). MTCH2 promotes adipogenesis in intramuscular preadipocytes via an m(6)A-YTHDF1-dependent mechanism. The FASEB Journal, 33, 2971–2981.
Wu, R., Yao, Y., Jiang, Q., & Cai, M. (2018). Epigallocatechin gallate targets FTO and inhibits adipogenesis in an mRNA m(6)A-YTHDF2-dependent manner. International Journal of Obesity, 42, 1378.
Song, T., Yang, Y., Wei, H., & Xie, X. (2019). Zfp217 mediates m6A mRNA methylation to orchestrate transcriptional and post-transcriptional regulation to promote adipogenic differentiation. Nucleic Acids Research, 47, 6130–6144.
Liu, Q., Zhao, Y., Wu, R., & Jiang, Q. (2019). ZFP217 regulates adipogenesis by controlling mitotic clonal expansion in a METTL3-m(6)A dependent manner. RNA Biology, 16, 1785–1793.
Budiyani, L., Purnamasari, D., Simadibrata, M., & Abdullah, M. (2018). Insulin resistance in gastroesophageal reflux disease. Acta Medica Indonesiana, 50, 336–342.
Salazar, M. R., Carbajal, H. A., Espeche, W. G., & Aizpurua, M. (2016). Insulin resistance: the linchpin between prediabetes and cardiovascular disease. Diabetes & Vascular Disease Research, 131, 157–163.
Gast, K. B., Tjeerdema, N., Stijnen, T., & Smit, J. W. (2012). Insulin resistance and risk of incident cardiovascular events in adults without diabetes: meta-analysis. PLoS One, 7.
Sandeep, S., Gokulakrishnan, K., Deepa, M., & Mohan, V. (2011). Insulin resistance is associated with increased cardiovascular risk in Asian Indians with normal glucose tolerance--the Chennai Urban Rural Epidemiology Study (CURES-66). The Journal of the Association of Physicians of India, 59, 480–484.
Reddy, K. J., Singh, M., Bangit, J. R., & Batsell, R. R. (2010). The role of insulin resistance in the pathogenesis of atherosclerotic cardiovascular disease: an updated review. Journal of Cardiovascular Medicine (Hagerstown, Md.), 11, 633–647.
Ritchie, R. H. (2009). Evidence for a causal role of oxidative stress in the myocardial complications of insulin resistance. Heart, Lung & Circulation, 18, 11–18.
AlZadjali, M. A., Godfrey, V., Khan, F., & Choy, A. (2009). Insulin resistance is highly prevalent and is associated with reduced exercise tolerance in nondiabetic patients with heart failure. Journal of the American College of Cardiology, 53, 747–753.
Xie, W., Ma, L. L., Xu, Y. Q., & Wang, B. H. (2019). METTL3 inhibits hepatic insulin sensitivity via N6-methyladenosine modification of Fasn mRNA and promoting fatty acid metabolism. Biochemical and Biophysical Research Communications, 518, 120–126.
Iskandar, K., Patria, S. Y., Huriyati, E., & Luglio, H. F. (2018). Effect of FTO rs9939609 variant on insulin resistance in obese female adolescents. BMC Research Notes, 11, 300.
Khoshi, A., Bajestani, M. K., Shakeri, H., & Goodarzi, G. (2019). Association of Omentin rs2274907 and FTO rs9939609 gene polymorphisms with insulin resistance in Iranian individuals with newly diagnosed type 2 diabetes. Lipids in Health and Disease, 18, 142.
Han, M., Li, Q., Liu, L., Zhang, D., Ren, Y., Zhao, Y., et al. (2019). Prehypertension and risk of cardiovascular diseases: a meta-analysis of 47 cohort studies. Journal of Hypertension, 37(12), 2325–2332.
Mo, X. B., Lei, S. F., Zhang, Y. H., & Zhang, H. (2019). Examination of the associations between m6A-associated single-nucleotide polymorphisms and blood pressure. Hypertension Research, 42(10), 1582–1589.
Zheng, Y., Nie, P., Peng, D., He, Z., Liu, M., Xie, Y., et al. (2018). m6AVar: a database of functional variants involved in m6A modification. Nucleic Acids Research, 46, D139–D145.
Wu, Q., Yuan, X., Han, R., Zhang, H., & Xiu, R. (2019). Epitranscriptomic mechanisms of N6-methyladenosine methylation regulating mammalian hypertension development by determined spontaneously hypertensive rats pericytes. Epigenomics, 11(12), 1359–1370.
Fox, C. S., Golden, S. H., Anderson, C., Bray, G. A., Burke, L. E., de Boer, I. H., et al. (2015). Update on prevention of cardiovascular disease in adults with type 2 diabetes mellitus in light of recent evidence: a scientific statement from the American Heart Association and the American Diabetes Association. Diabetes Care, 38(9), 1777–1803.
Gilbert, E. R., & Liu, D. (2012). Epigenetics: the missing link to understanding β-cell dysfunction in the pathogenesis of type 2 diabetes. Epigenetics, 7(8), 841–852.
Shen, F., Huang, W., Huang, J. T., Xiong, J., Yang, Y., Wu, K., et al. (2015). Decreased N(6)-methyladenosine in peripheral blood RNA from diabetic patients is associated with FTO expression rather than ALKBH5. The Journal of Clinical Endocrinology and Metabolism, 100(1), 148–154.
Yang, Y., Shen, F., Huang, W., Qin, S., Huang, J. T., Sergi, C., et al. (2019). Glucose is involved in the dynamic regulation of m6A in patients with type 2 diabetes. The Journal of Clinical Endocrinology and Metabolism, 104(3), 665–673.
De Jesus, D. F., Zhang, Z., Kahraman, S., Brown, N. K., Chen, M., Hu, J., et al. (2019). m6A mRNA methylation regulates human β-cell biology in physiological states and in type 2 diabetes. Nature Metabolism, 1(8), 765–774.
Xiao, C. L., Zhu, S., He, M., & Chen, D. (2018). N(6)-methyladenine DNA modification in the human genome. Molecular Cell, 71, 306–318.
Koh, C. W. Q., Goh, Y. T., Toh, J. D. W., & Neo, S. P. (2018). Single-nucleotide-resolution sequencing of human N6-methyldeoxyadenosine reveals strand-asymmetric clusters associated with SSBP1 on the mitochondrial genome. Nucleic Acids Research, 46, 11659–11670.
Zhang, W., Song, M., Qu, J., & Liu, G. H. (2018). Epigenetic modifications in cardiovascular aging and diseases. Circulation Research, 123, 773–786.
Kim, A. Y., Park, Y. J., Pan, X., & Shin, K. C. (2015). Obesity-induced DNA hypermethylation of the adiponectin gene mediates insulin resistance. Nature Communications, 6, 7585.
Zhao, J., Goldberg, J., Bremner, J. D., & Vaccarino, V. (2012). Global DNA methylation is associated with insulin resistance: a monozygotic twin study. Diabetes, 61, 542–546.
Stenvinkel, P., Karimi, M., Johansson, S., & Axelsson, J. (2007). Impact of inflammation on epigenetic DNA methylation - a novel risk factor for cardiovascular disease? Journal of Internal Medicine, 261, 488–499.
Breton, C. V., Byun, H. M., Wenten, M., & Pan, F. (2009). Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. American Journal of Respiratory and Critical Care Medicine, 180, 462–467.
Agha, G., Mendelson, M. M., Ward-Caviness, C. K., & Joehanes, R. (2019). Blood leukocyte DNA methylation predicts risk of future myocardial infarction and coronary heart disease. Circulation, 140, 645–657.
Guo, Y., Pei, Y., & Li, C. W. (2020). DNA N(6)-methyladenine modification in hypertension. Aging (Albany NY), 12, 6276–6291.
Yan, X. C., Cao, J., Liang, L., & Wang, L. (2016). miR-342-5p is a notch downstream molecule and regulates multiple angiogenic pathways including notch, vascular endothelial growth factor and transforming growth factor beta signaling. Journal of the American Heart Association, 5.
Good, R. B., Gilbane, A. J., Trinder, S. L., & Denton, C. P. (2015). Endothelial to mesenchymal transition contributes to endothelial dysfunction in pulmonary arterial hypertension. The American Journal of Pathology, 185, 1850–1858.
Funding
Key Project of Hunan Provincial Department of Education (20A427). This work was supported by the grants from the National Natural Sciences Foundation of China (Nos. 81970390, 82060065), the Natural Science Foundation of Hunan Province (Nos. 2018JJ3455, 2018JJ2341, 2019JJ40249), the Key Project of the Natural Science Foundation of Guangxi Zhuang Autonomous Region, China(No. 2020GXNSFDA297011), the Foundation for Guangxi Key Laboratory of Diabetic Systems Medicine (No. 20-065-77), the Outstanding Young Aid Program for Education Department of Hunan Province (No. 18B274), the Major Project of social science achievement review committee in Hunan Province (No. XSP20ZDI013), and the Hunan Provincial Innovation Foundation For Postgraduate (No. CX20200965).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Associate Editor Junjie Xiao oversaw the review of this article
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, Ys., Ouyang, Xp., Yu, Xh. et al. N6-Adenosine Methylation (m6A) RNA Modification: an Emerging Role in Cardiovascular Diseases. J. of Cardiovasc. Trans. Res. 14, 857–872 (2021). https://doi.org/10.1007/s12265-021-10108-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-021-10108-w