Abstract
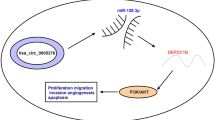
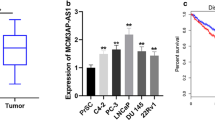
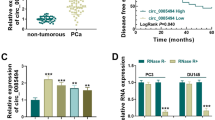
Prostate cancer (PCa) is one of the most common malignant tumors, with a high rate of metastasis and recurrence. Circular RNA_0001165 (circ_0001165) has been shown to be involved in the advance of PCa. However, the interaction between circ_0001165 and microRNA in PCa has not been studied. Quantitative real-time polymerase chain reaction was used to detect the expression of related genes in PCa tissues and cells. The expression of related epithelial-mesenchymal transition proteins was detected by Western blotting. The interaction of miR-654-3p with circ_0001165 or dimethylarginine dimethylaminohydrolase 1 (DDAH1) has been notarized by Dual-luciferase reporter assay and RNA immunoprecipitation assay. Xenotransplantation experiments confirmed the function of circ_0001165 in vivo. Circ_0001165 and DDAH1 are significantly high-expressed in PCa tissues and cells. Silencing circ_0001165 can reduce the proliferation, migration, invasion and tube formation of LNCaP and DU145 cells. MiR-654-3p is a target of circ_0001165. Silencing circ_0001165 can inhibit the malignant behavior of PCa cells by releasing miR-654-3p. In addition, DDAH1 is a target of miR-654-3p. Overexpression of DDAH1 partially restored the inhibitory effect of miR-654-3p on cell proliferation. Animal experiments confirmed the anti-tumor effect of silence circ_0001165 in vivo. Circ_0001165 regulates the expression of DDAH1 by regulating miR-654-3p, thereby mediating the process of PCa, and at least partially promoting the development of PCa cells, providing a novel targeted therapy for PCa.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Xin, L. (2019) Cells of origin for prostate cancer. Adv. Exp. Med. Biol. 1210: 67–86.
Merriel, S. W. D., G. Funston, and W. Hamilton (2018) Prostate cancer in primary care. Adv. Ther. 35: 1285–1294.
Abrams, D. I. (2018) An integrative approach to prostate cancer. J. Altern. Complement. Med. 24: 872–880.
Zhou, C. K., D. P. Check, J. Lortet-Tieulent, M. Laversanne, A. Jemal, J. Ferlay, F. Bray, M. B. Cook, and S. S. Devesa (2016) Prostate cancer incidence in 43 populations worldwide: an analysis of time trends overall and by age group. Int. J. Cancer 138: 1388–1400.
Siegel, R. L., K. D. Miller, and A. Jemal (2016) Cancer statistics, 2016. CA Cancer J. Clin. 66: 7–30.
Sartor, O. and J. S. de Bono (2018) Metastatic prostate cancer. N. Engl. J. Med. 378: 645–657.
Chen, L. L. and L. Yang (2015) Regulation of circRNA biogenesis. RNA Biol. 12: 381–388.
Liu, J., T. Liu, X. Wang, and A. He (2017) Circles reshaping the RNA world: from waste to treasure. Mol. Cancer. 16: 58.
Ding, T., Y. Zhu, H. Jin, P. Zhang, J. Guo, and J. Zheng (2021) Circular RNA circ_0057558 controls prostate cancer cell proliferation through regulating miR-206/USP33/c-Myc axis. Front. Cell Dev. Biol. 9: 644397.
Li, X., B. Azhati, W. Wang, M. Rexiati, C. Xing, and Y. Wang (2021) Circular RNA UBAP2 promotes the proliferation of prostate cancer cells via the miR-1244/MAP3K2 axis. Oncol. Lett. 21: 486.
Mao, Y., W. Li, B. Hua, X. Gu, W. Pan, Q. Chen, B. Xu, C. Lu, and Z. Wang (2021) Circular RNA_PDHX promotes the proliferation and invasion of prostate cancer by sponging MiR-378a-3p. Front. Cell Dev. Biol. 8: 602707.
Yan, Z., Y. Xiao, Y. Chen, and G. Luo (2020) Screening and identification of epithelial-to-mesenchymal transition-related circRNA and miRNA in prostate cancer. Pathol. Res. Pract. 216: 152784.
Hansen, T. B., T. I. Jensen, B. H. Clausen, J. B. Bramsen, B. Finsen, C. K. Damgaard, and J. Kjems (2013) Natural RNA circles function as efficient microRNA sponges. Nature. 495: 384–388.
Gregory, R. I. and R. Shiekhattar (2005) MicroRNA biogenesis and cancer. Cancer Res. 65: 3509–3512.
Macfarlane, L. A. and P. R. Murphy (2010) MicroRNA: biogenesis, function and role in cancer. Curr. Genomics. 11: 537–561.
Pu, J. T., Z. Hu, D. G. Zhang, T. Zhang, K. M. He, and T. Y. Dai (2020) MiR-654-3p suppresses non-small cell lung cancer tumourigenesis by inhibiting PLK4. Onco Targets Ther. 13: 7997–8008.
Yang, J., Z. Zhang, S. Chen, W. Dou, R. Xie, and J. Gao (2020) miR-654-3p predicts the prognosis of hepatocellular carcinoma and inhibits the proliferation, migration, and invasion of cancer cells. Cancer Biomark. 28: 73–79.
Formosa, A., E. K. Markert, A. M. Lena, D. Italiano, E. Finazzi-Agro’, A. J. Levine, S. Bernardini, A. V. Garabadgiu, G. Melino, and E. Candi (2014) MicroRNAs, miR-154, miR-299-5p, miR-376a, miR-376c, miR-377, miR-381, miR-487b, miR-485-3p, miR-495 and miR-654-3p, mapped to the 14q32.31 locus, regulate proliferation, apoptosis, migration and invasion in metastatic prostate cancer cells. Oncogene. 33: 5173–5182.
Frey, D., O. Braun, C. Briand, M. Vasák, and M. G. Grütter (2006) Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 14: 901–911.
Leiper, J. M., J. Santa Maria, A. Chubb, R. J. MacAllister, I. G. Charles, G. S. Whitley, and P. Vallance (1999) Identification of two human dimethylarginine dimethylaminohydrolases with distinct tissue distributions and homology with microbial arginine deiminases. Biochem. J. 343: 209–214.
Dayal, S., R. N. Rodionov, E. Arning, T. Bottiglieri, M. Kimoto, D. J. Murry, J. P. Cooke, F. M. Faraci, and S. R. Lentz (2008) Tissue-specific downregulation of dimethylarginine dimethylaminohydrolase in hyperhomocysteinemia. Am. J. Physiol. Heart Circ. Physiol. 295: H816–H825.
Ye, J., J. Xu, Y. Li, Q. Huang, J. Huang, J. Wang, W. Zhong, X. Lin, W. Chen, and X. Lin (2017) DDAH1 mediates gastric cancer cell invasion and metastasis via Wnt/β-catenin signaling pathway. Mol. Oncol. 11: 1208–1224.
Ding, X., J. Zheng, and M. Cao (2021) Circ_0004771 accelerates cell carcinogenic phenotypes via suppressing miR-1253-mediated DDAH1 inhibition in breast cancer. Cancer Manag. Res. 13: 1–11.
Reddy, K. R. K., C. Dasari, D. Duscharla, B. Supriya, N. S. Ram, M. V. Surekha, J. M. Kumar, and R. Ummanni (2018) Dimethylarginine dimethylaminohydrolase-1 (DDAH1) is frequently upregulated in prostate cancer, and its overexpression conveys tumor growth and angiogenesis by metabolizing asymmetric dimethylarginine (ADMA). Angiogenesis. 21: 79–94.
Arnaiz, E., C. Sole, L. Manterola, L. Iparraguirre, D. Otaegui, and C. H. Lawrie (2019) CircRNAs and cancer: biomarkers and master regulators. Semin. Cancer Biol. 58: 90–99.
Ricciuti, B., J. Foglietta, V. Bianconi, A. Sahebkar, and M. Pirro (2019) Enzymes involved in tumor-driven angiogenesis: a valuable target for anticancer therapy. Semin. Cancer Biol. 56: 87–99.
Yu, Q., P. Li, M. Weng, S. Wu, Y. Zhang, X. Chen, Q. Zhang, G. Shen, X. Ding, and S. Fu (2018) Nano-vesicles are a potential tool to monitor therapeutic efficacy of carbon ion radiotherapy in prostate cancer. J. Biomed. Nanotechnol. 14: 168–178.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest. The design of this protocol follows the tenets of the Declaration of Helsinki, approved by the Ethics Committee of Wuhan Hankou Hospital.
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Table S1
. Primer sequences for quantitative real-time polymerase chain reaction
Rights and permissions
About this article
Cite this article
Ji, S., Zhang, C., Xiang, Y. et al. Hsa_circ_0001165 Regulates the Malignant Phenotype and Angiogenesis of Prostate Cancer Cells through miR-654-3p/DDAH1. Biotechnol Bioproc E 28, 125–136 (2023). https://doi.org/10.1007/s12257-021-0229-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-021-0229-4