Abstract
Since 1950, more than 620 potato cultivars have been released from Chinese breeding programs, some of which have similar genetic background and phenotype. In this study, 16 parental cultivars widely used in breeding were used to screen 138 simple sequence repeat (SSR) markers. Out of 138 SSR markers, 20 were polymorphic that were used to analyze the genetic diversity of 217 potato cultivars grown in China. In total, 249 alleles were detected with these 20 markers and 244 of them (97.99%) showed polymorphism. The number of alleles ranged from 7 to 22 with an average of 12.45 alleles per primer and polymorphic information content (PIC) ranged from 0.64–0.93 with an average of 0.83. Fragment sizes varied from 80 to 380 bp. Based on PIC values and the clarity of PCR amplification bands, 11 SSR markers were selected and able to differentiate all of the 217 cultivars. The estimated similarity using these polymorphic SSR markers between the cultivars ranged from 0.63 to 0.99, indicating a narrow genetic base. Fingerprinting and genetic diversity analysis in this study provides useful information for the protection of intellectual property, as well as the exploration and utilization of these potato cultivars.
Resumen
Desde 1950, se han liberado más de 620 variedades de papa de los programas chinos de mejoramiento, algunos de los cuales tienen antecedentes genéticos y fenotipo similares. En este estudio, se usaron 16 variedades parentales utilizadas ampliamente en mejoramiento, para hacer un estudio de 138 marcadores de repeticiones simples de secuencia (SSR). De los 138 marcadores SSR, 20 fueron polimórficos que fueron usados para analizar la diversidad genética de 217 variedades de papa cultivadas en China. En total, se detectaron 249 alelos con estos 20 marcadores y 244 de ellos (97.99%) mostraron polimorfismo. El número de alelos tuvo un rango de 7–22 con un promedio de 12.45 alelos por iniciador, y el contenido de la información polimórfica (PIC) varió de 0.64–0.93 con un promedio de 0.83. Los tamaños de los fragmentos variaron de 80–380 pb. Con base en los valores PIC y la claridad de las bandas de amplificación, se seleccionaron 11 marcadores SSR y fue posible diferenciar a la totalidad de las 217 variedades. La similitud estimada usando estos marcadores polimórficos SSR entre las variedades varió de 0.63 a 0.99, indicando una base genética angosta. Los análisis de huellas y diversidad genética en este estudio proporcionan información útil para la protección de propiedad intelectual, así como para la exploración y utilización de estas variedades de papa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potato (Solanum tuberosum L.) is the third most important food crop in the world. It is adapted to a wide range of environmental conditions (Vreugdenhil 2007) and contains high level of nutrients (Gibson and Kurilich 2013). In addition, it also plays an important role in ensuring food security and promoting farmers’ income. In 2016, 162 countries and regions planted 19.25 million hectares of potato and the yield reached 376.83 million tons (http://www.fao.org/faostat/en/#data/QC).
China is the largest producer of potato in the world with a growing area of 5.63 million hectares and total production of 97.4 million tons in 2016 (Qu 2016). Potato cultivars grown in China have contributed immensely to increased yields and acted as germplasm resources for cultivar improvement. In China, breeding for crop improvement in potato started in 1950 and currently more than 620 potato cultivars have been released. For a long period, confusion has existed in cultivar names because of missing or incorrect records. Sometimes the same cultivar might be known by different names, and sometimes different cultivars have the same name. Therefore, a quick and accurate identification of different cultivars has great importance for seed production, protection of intellectual property, and commercialization of cultivars in China.
Cultivar identification in potato is mainly based on traditional morphological traits, such as tuber morphology (shape, skin and flesh color, distribution and depth of the eyes), leaf type, flower color, and sprout appearance. However, collecting trait data is time consuming, difficult and sometimes strongly affected by environmental factors (Chimote et al. 2004). In addition, the repeated use of selected elite parental clones in breeding further narrows down the genetic diversity among the cultivars. Therefore, it is difficult to distinguish similar cultivars morphologically. Isozyme markers developed in the 1970s were widely used to directly detect the gene products, and have proved to be useful in identification of different potato cultivars (Stegemann and Schnick 1985; Oliver and Martinez-Zapater 1985; Douches and Ludlam 1991). However, isozyme analysis is reported to be affected by plant developmental stage (Hahn and Grifo 1996), and is also limited by the number of loci that can be used (Karaagac et al. 2014).
DNA-based molecular markers can overcome the above mentioned limitations and have been successfully used in potato fingerprinting. Moreover, to increase the efficiency of breeding programs, molecular markers have also been used for the analysis of genetic relationships in various potato cultivars, allowing breeders to establish a broad genetic base for breeding purposes (Bisognin and Douches 2002). Such markers include random amplified polymorphic DNA (RAPD) (Demeke et al. 1993; Ford and Taylor 1997; Chakrabarti et al. 2001), amplified fragment length polymorphisms (AFLPs) (Milbourne et al. 1997), inter-simple sequence repeats (ISSRs) (Prevost and Wilkinson 1999; Bornet et al. 2002), simple sequence repeats (SSRs) (Raker and Spooner 2002), single nucleotide polymorphisms (SNPs) (Bali et al. 2017) and various combinations of above markers (McGregor et al. 2000; Gorji et al. 2011).
SSR markers are highly polymorphic, co-dominant and the primer sequences are generally well conserved within and between related species (Karaagac et al. 2014). Additional advantages of SSR markers over dominant marker systems are their high heterozygosity and capacity to reflect ploidy status in potato (Ghislain et al. 2004). Many researchers have therefore developed potato SSR markers from genomic libraries and expressed sequence tags (ESTs) databases (Milbourne et al. 1998; Ashkenazi et al. 2001; Ghislain et al. 2004; Feingold et al. 2005; Ghislain et al. 2009). These markers have been extensively used for DNA fingerprinting (Norero et al. 2002; Coombs et al. 2004; Barandalla et al. 2006; Karaagac et al. 2014; Bali et al. 2017), genetic diversity analysis (Chimote et al. 2004; Ispizúa et al. 2007), germplasm migrations (Ríos et al. 2007), and parental analysis (Spanoghe et al. 2015) in potato.
In the present study, SSR markers were used to fingerprint 217 potato cultivars and to analyze the genetic diversity in Chinese germplasm. The data generated will be used for cultivar identification and future germplasm management programs.
Materials and Methods
Plant Material
Among the 217 cultivars included in the study, eight (Atlantic, Favorita, Mira, Katahdin, Kuannae, Epoka, Schwalbe, and Anemone) were imported from other countries, and the remaining 209 were bred by 44 regional breeding programs in China from 1950 to 2007 (Fig. 1, Table 1).
Map of China showing the geographical distribution of the 44 breeding programs of the cultivars used in the present study. Numbers are cited as map locality in Table 1
Sixteen cultivars that represented the most commonly used parental materials in Chinese potato breeding programs before 2007, were used to screen for polymorphic SSR markers. Katahdin and B76-16 (Xiaoyezi) originated from the United States; Mira, Anemone and Schwalbe originated from Germany; Epoka from Poland; Favorita from the Netherlands; DTO-33 from the International Potato Center (CIP); and, Mengshu 10, Chuanyu 6, Xishu 1, Kexin 2, Gaoyuan 7, Zhongshu 3 and Hutou were domestic cultivars.
DNA Extraction and SSR Analysis
Genomic DNA was extracted from 2 g of fresh young leaves according to the modified CTAB procedure of Doyle and Doyle (1987) and quantified using 1% agarose gel electrophoresis. The DNA samples were diluted to 25 ng μL-1 and stored at – 20 °C until use.
A total of 138 markers located on all 12 potato chromosomes were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). The primer sequences of these markers were obtained from previous studies (Ghislain et al. 2004; Feingold et al. 2005). Marker polymorphism was evaluated using the 16 cultivars mentioned above. Polymorphic markers were used to fingerprint and analyze the genetic diversity of 217 potato cultivars.
PCR amplifications were performed in a 20 μL reaction mixture, containing 2.0 μL of 25 ng μL-1 DNA template, 0.8 μL of 10 pmol μL-1 forward primer, 0.8 μL of 10 pmol μL-1 reverse primer, 1.6 μL of 2.5 mmol L-1 dNTPs, 2.0 μL of 10 × PCR buffer, 0.4 μL of Taq DNA polymerase (2.5 U μL-1) (Tiangen Biotech Co., Ltd., Beijing, China), and 12.4 μL of ddH2O. A modified PCR program was used: 5 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at SSR specific annealing temperature (53 °C to 64 °C) and 45 s at 72 °C; with a final extension step of 7 min at 72 °C. The PCR products were fractioned on 8% denatured polyacrylamide gel electrophoresis, and stained with silver nitrate.
Data Analysis
A data file was constructed by scoring 0 for absence and 1 for presence of specific amplification product for each sample. The following related genetic parameters were then calculated:
-
(1).
Percentage of polymorphic loci p = (k/n) × 100%, where k is the number of polymorphic loci, and n is the total number of measured loci.
-
(2).
Polymorphic information content (PIC) =\( 1-\sum \limits_1^i{f}_i^2 \), where: fi is the frequency of the ith allele in a locus.
Cluster analysis was performed with NTSYS Pc2.11 using the Dice co-efficient (Dice 1945) for calculating similarities and unweighted pair-group arithmetic average method (UPGMA) for constructing dendrograms.
Results
SSR Marker Screening
Among the 138 SSR markers screened on 16 potato cultivars (data not shown), 20 SSR markers generated polymorphism. These 20 markers were used to fingerprint 217 cultivars: a total of 249 alleles were obtained, of which 244 alleles were polymorphic (97.99%). The number of alleles in the 20 loci ranged from seven (primers S7 and S122) to 22 (primer S189) with an average of 12.45. The PIC values for the markers varied from 0.64 (primer S122) to 0.93 (primer S189) with an average of 0.83. The size of amplifications ranged from 80 to 380 bp (Table 2).
Construction of DNA Fingerprinting
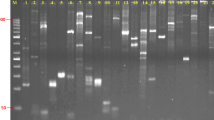
The 20 polymorphic SSR markers were used to fingerprint 217 cultivars. Eleven markers (S122, S168, S151, S184, S7, S170, S118, S192, S180, S174, and S25) demonstrated high PIC values, high quality amplifications, and were able to differentiate all 217 cultivars (Supplemental Material 1). These markers produced 132 alleles, of which 129 (97.73%) were polymorphic (Table 2). The DNA fingerprint for each cultivar was unique. Figure 2 shows the amplification of 17 potato cultivars by marker S122. The smallest allele amplified by S122 was designated as 1 and the largest allele amplified was designated as 7.
Thirty-one cultivars produced unique alleles with certain markers (Table 3). For Cooperation 001 and Qinyu 31, three markers generated unique alleles. Kexin 20, Cooperation 002, Cooperation 003, Cooperation 88, Jinshu 14 and Zhongdianhong amplified unique alleles with two markers. Only one marker in the remaining 23 cultivars produced unique alleles.
Genetic Diversity Analysis
Based on the results from 20 polymorphic SSR markers, a similarity matrix was used to generate a UPGMA dendrogram (Supplemental Material 2). The dendrogram showed that all of the 217 cultivars were closely related and lacked the formation of distinct clusters.
In most cases, the dendrogram grouping fits well with the recorded pedigree information. Cultivars from the same cross usually clustered together, for example, Kexin 4 and Kexin 5 (Anemone X Katahdin); Zhengshu 5 and Zhengshu 6 (Gaoyuan 7 X Zheng 762-93); Zhongshu 8 and Zhongshu 12 (W953 X FL475); and, Yunshu 101, Yunshu 102 and Yunshu 201 (S95-105 X Neishu 7). Cultivars sharing one of the parents also clustered together, such as Kangyi 1, Kangbingchi and Jinkengbai which had Epoka as one of their parents; Wannong 4, Wanshu 8 and Wanyu 9 which had Duozibai as the male parent; and, Chuanyu 39 and Chuanyu 4 which had 7XY.1 as the male parent. Cultivars usually clustered together with their parent(s). As examples, Jinshu 5 and Jinshu 8 clustered with their female parent, Junshu 2; and, Liangshu 8 clustered together with its female parent, Liangshu 97. Most of these 217 cultivars were progeny of cultivars or accessions from the United States, Germany, Poland, the Netherlands, and CIP, and they were distributed in almost all the clusters. There was no obvious clustering trend by their country of origin: perhaps because a few foreign elite lines were used widely as parental clones in most breeding programs.
Nevertheless, the relationship between domestic cultivar distribution and agro-ecological zone could be observed in the UPGMA dendrogram. There are four main potato agro-ecological zones in China, including the northern single cropping zone, the southwest mixed cropping zone, the central double cropping zone and the southern winter cropping zone. Most of the cultivars that were developed in the same cropping zone clustered together. Most of the cultivars from the northern single cropping zone clustered in a large group. Among them, Kexin 19, Dongnong 304, Hushu 5, Kexin 2, Kexin 3, Kexin 13, Kexin 9, Chunshu 5, Hushu 1, Chaobai, Kexin 4, and Kexin 5, all from Northeast China, clustered in one subgroup. Surprisingly, pedigree information revealed that, most of these cultivars were descendants of Mira (one to two generations). Similarly, Qingshu 8, Longshu 1, Wensheng 4, Linshu 7, Xindaping, Ningshu 9, Ningshu 1, Shengli 1, Gaoyuan 7, Weishu 1, Yushu CA, Linshu 3, Weishu 8, Annong 5, Ningshu 6, Kangyi 1, Longshu 3, and Ningshu 7, all from Northwest China, clustered in another subgroup, and they were mostly progeny of the Polish cultivar Epoka and German cultivars Schwalbe and Industria. Cultivars from the southwest mixed cropping zone also clustered in one group, for example, Yunshu 201, Yunshu 102, Yunshu 301, Yunshu 501, Yunshu 101, Chuanyu 6 and Weiyu 3. These cultivars were mainly derived from Mira and neo-tuberosum. Clustering analysis also showed that most of the cultivars that were bred by the same program clustered within the same group, for example, Kexin 2, Kexin 3, Kexin 4, Kexin 5, Kexin 9, Kexin 13, Kexin 14 and Kexin 19 from one breeding program clustered together; Yunshu 201, Yunshu 102, Yunshu 301, Yunshu 501 and Yunshu 101 from another breeding program clustered together.
Discussion
Potato cultivar identification is of great importance for seed production, germplasm management and breeders’ right protection. In this study, we successfully identified 20 polymorphic SSR markers to analyze the fingerprints of 217 potato cultivars grown in China, and 11 of these markers enabled complete differentiation among all the cultivars. Our results indicate that SSR markers have enough power to differentiate the large number of potato cultivars in China. Moreover, genetic diversity analysis of 217 potato cultivars based on 20 polymorphic SSR markers indicates a narrow genetic background in these cultivars.
In our study, the number of alleles per locus (7–22) and PIC values (0.64–0.93) are comparable to the values reported by Ghislain et al. (2004); however, slight differences in these two studies could be attributed to the primer combinations used and the genetic background of the potato cultivars. The study by Ghislain et al. was done on 931 accessions in eight taxonomic groups of cultivated potato ranging from diploids to pentaploids, whereas, our study was done on 217 potato cultivars which were autotetraploid genotypes of different origin (the United States, Germany, Poland, the Netherlands, CIP, Canada, Czech Republic and domestic cultivars).
Previous studies have shown that a small number of SSR markers with high polymorphism could be used to distinguish large numbers of potato varieties. In this study, a subset of 11 SSR markers was sufficient to distinguish all 217 potato cultivars grown in China. Moisan-Thiery et al. (2005) successfully differentiated 286 potato cultivars produced in France with five SSR markers. Reid and Kerr (2007) developed six high polymorphic SSR markers that could differentiate more than 400 cultivars of European germplasm. Ghislain et al. (2009) also developed a potato genetic identification kit to differentiate 93.5% and 98.8% of the 742 landraces using 24 and 51 SSR markers, respectively. Karaagac et al. (2014) reported that 50 tetraploid genotypes could be differentiated with six SSR markers. These studies all indicated that SSR markers can efficiently identify potato germplasm materials at the molecular level. Further, we found that the number of unique alleles amplified by a SSR marker was not directly associated with the number of alleles detected or PIC values. For example, a maximum of six different unique alleles were amplified from six cultivars by marker S118 (Table 3). However, only 11 alleles for this marker were found in the 217 cultivars investigated and its PIC value was 0.87; these were moderate values for both number of alleles and PIC from among the 11 markers used. This finding was in agreement with results of Wang et al. (2003) on maize.
The analysis of genetic diversity of 217 potato cultivars indicated that the genetic base of potato germplasm in China was narrow. The vast majority of potato germplasm resources in China originated from abroad. There’re 288 cultivars released in China during the period from 1950 to 2007, which can be mainly derived from cultivars or accessions from the United States, Germany, Poland, the Netherlands, and CIP. Among them, 100 cultivars were progeny of American cultivars including Katahdin, Early Rose, Houma, Wauseon, Pontiac, Triumph, Atlantic, and B76-16, which account for 34.7% of all the released cultivars in China during this time. 93 cultivars were progeny of German cultivars including Mira, Anemone, Schwalbe, Industria, Jubel, Merkur, Flava, Pepo, Amsel, Apta, and Mitterfrühe, accounting for 32.3%. 53 cultivars were progeny of Polish cultivar Epoka, accounting for 18.4%. Moreover, there’re 20 and 4 cultivars that can be derived from the CIP resources and Netherlands cultivar Favorita, respectively. The other few cultivars were derived from other countries, domestic cultivars, or have unknown pedigree information. Sixteen cultivars we originally selected to screen for SSR marker polymorphism represented most of the germplasm of potato cultivars in China bred from 1950 to 2007.
In China, the use of a limited number of parental lines in the breeding programs may contribute to the narrow genetic base of improved cultivars. For example, six cultivars or accessions of Mira, Katahdin, Epoka, Anemone, Duozibai, and B76-16 (Xiaoyezi) were found repeatedly in the parentage of 74 cultivars released before 1983, accounting for 68.8% of the total cultivars released in that period of time. In this study, cultivars released before 1983 were mostly clustered in one single group. The clustering together of most of the cultivars that came from the same cropping zone or the cultivars that were bred by the same program, strengthens the fact that few elite parental lines have been commonly and repeatedly used in potato breeding programs in China. In contrary, there were some closely related cultivars that did not cluster together in the same group. For example, Cultivar Linshu 2 and Linshu 3 were both bred from the same parents (B76-16 X Everest) but were placed in different groups. This may be attributed to non-specific fingerprints amplified during PCR or incorrect pedigree records for these cultivars.
The cultivated potato is a heterozygous autotetraploid species with four homologous sets of chromosomes (2n=4x=48) (Gebhardt and Valkonen 2001). The selection of parental combinations is mainly dependent on morphological traits like growth habit, yield, resistance and processing quality, and less on pedigree information since the data is often scanty and sometimes incorrect (Braun and Wenzel 2005). However, morphological traits do not always provide a good measure of genetic makeup and may not accurately reveal genetic variation (Tanksley and McCouch 1997). Genetic relatedness measures based on molecular markers may not predict similarity in trait values or parental performance (Jansky et al. 2015). However, in order to improve heterotic effects, better knowledge about the degree of relationship between parents is required. In the present study, SSR markers were used to estimate genetic relationships in tetraploid potato cultivars with known pedigrees. The results showed that the cultivars with similar pedigrees grouped together in most cases. This is in agreement with the results of Braun and Wenzel (2005), who used AFLP and SSR markers to evaluate genetic diversity in the tetraploid potato and to compare the findings with known pedigree information, and they found that in most cases the grouping of related genotypes and the known pedigree information was reflected well in dendrograms. Although our study found a narrow genetic base in the cultivars, breeders could try to choose distinct cultivars grouped in different subclasses as parents to introduce more diversity in the hybrids. In order to further improve the yield and quality of Chinese potato cultivars, it is necessary to extensively increase the genetic basis of the hybrid parents through the introduction and utilization of new germplasm resources.
References
Ashkenazi, V., E. Chani, U. Lavi, D. Levy, J. Hillel, and R.E. Veilleux. 2001. Development of microsatellite markers in potato and their use in phylogenetic and fingerprinting analyses. Genome 44 (1): 50–62.
Bali, S., V. Sathuvalli, C. Brown, R. Novy, L. Ewing, J. Debons, D. Douches, J. Coombs, D. Navarre, J. Whitworth, B. Charlton, S. Yilma, C. Shock, J. Stark, M. Pavek, and N.R. Knowles. 2017. Genetic fingerprinting of potato varieties from the northwest potato variety development program. American Journal of Potato Research 94 (1): 54–63.
Barandalla, L., J.I. Ruiz de Galarreta, D. Rios, and E. Ritter. 2006. Molecular analysis of local potato cultivars from Tenerife Island using microsatellite markers. Euphytica 152: 283–291.
Bisognin, D.A., and D.S. Douches. 2002. Genetic diversity in diploid and tetraploid late blight resistant potato germplasm. Horticultural Science 37 (1): 178–183.
Bornet, B., G. Goraguer, G. Joly, and M. Branchard. 2002. Genetic diversity in European and Argentinean cultivated potatoes (Solanum tuberosum subsp. tuberosum) detected by inter-simple sequence repeats (ISSRs). Genome 45: 481–484.
Braun, A., and G. Wenzel. 2005. Molecular analysis of genetic variation in potato (Solanum tuberosum L.). I. German cultivars and advanced clones. Potato Research 47: 81–92.
Chakrabarti, S.K., D. Pattanayak, and P.S. Naik. 2001. Fingerprinting Indian potato cultivars by random amplified polymorphic DNA (RAPD) markers. Potato Research 44: 375–387.
Chimote, V.P., S.K. Chakrabarti, D. Pattanayak, and P.S. Naik. 2004. Semi-automated simple sequence repeat analysis reveals narrow genetic base in Indian potato cultivars. Biologia Plantarum 48 (4): 517–522.
Coombs, J.J., L.M. Frank, and D.S. Douches. 2004. An applied fingerprinting system for cultivated potato using simple sequence repeats. American Journal of Potato Research 81 (4): 243–250.
Demeke, T., L.M. Kawchuk, and D.R. Lynch. 1993. Identification of potato cultivars and clonal variants by random amplified polymorphic DNA analysis. American Potato Journal 70: 561–570.
Dice, L.R. 1945. Measures of the amount of ecologic association between species. Ecology 26: 297–302.
Douches, D.S., and K. Ludlam. 1991. Electrophoretic characterization of north American potato cultivars. American Potato Journal 68: 767–780.
Doyle, J.J., and J.L. Doyle. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19: 11–15.
Feingold, S., J. Lloyd, N. Norero, M. Bonierbale, and J. Lorenzen. 2005. Mapping and characterization of new EST-derived microsatellites for potato (Solanum tuberosum L.). Theoretical and Applied Genetics 111 (3): 456–466.
Ford, R., and P.W.J. Taylor. 1997. The application of RAPD markers for potato (Solanum tuberosum L.) cultivar identification in the Australian potato industry. Australian Journal of Agricultural Research 48: 1213–1217.
Gebhardt, C., and J.P.T. Valkonen. 2001. Organization of genes controlling disease resistance in the potato genome. Annual Review of Phytopathology 39: 79–102.
Ghislain, M., D.M. Spooner, F. Rodriguez, F. Villamon, J. Nunez, C. Vasquez, R. Waugh, and M. Bonierbale. 2004 Selection of highly informative and user-friendly microsatellites (SSRs) for genotyping of cultivated potato. Theoretical and Applied Genetics 108 (5): 881–890.
Ghislain, M., J. Nunez, M. del Rosario Herrera, J. Pignataro, F. Guzman, M. Bonierbale, and D.M. Spooner. 2009. Robust and highly informative microsatellite-based genetic identity kit for potato. Molecular Breeding 23 (3): 377–388.
Gibson, S., and A.C. Kurilich. 2013. The nutritional value of potatoes and potato products in the UK diet. Nutrition Bulletin 38 (4): 389–399.
Gorji, A.M., P. Poczai, Z. Polgar, and J. Taller. 2011. Efficiency of arbitrarily amplified dominant markers (SCOT, ISSR and RAPD) for diagnostic fingerprinting in tetraploid potato. American Journal of Potato Research 88: 226–237.
Hahn, W.J., and F.T. Grifo. 1996. Molecular markers in plant conservation genetics. In: B.W.S. Sobral (Ed.), The impact of plant molecular genetics. Birkh/iuser, Boston, Chapter 7, 114–136.
Ispizúa, V.N., I.R. Guma, S. Feingold, and A.M. Clausen. 2007. Genetic diversity of potato landraces from northwestern Argentina assessed with simple sequence repeats (SSRs). Genetic Resources and Crop Evolution 54: 1833–1848.
Jansky, S.H., J. Dawson, and D.M. Spooner. 2015. How do we address the disconnect between genetic and morphological diversity in germplasm collections? American Journal of Botany 102: 1213–1215.
Karaagac, E., S. Yilma, A. Cuesta-Marcos, and M.I. Vales. 2014. Molecular analysis of potatoes from the pacific northwest tri-state variety development program and selection of markers for practical DNA fingerprinting applications. American Journal of Potato Research 91 (2): 195–203.
McGregor, C.E., M.M. Greyling, and L. Warnich. 2000. The use of simple sequence repeats (SSRs) to identify commercially important potato (Solanum tuberosum L.) cultivars in South Africa. South African Journal of Plant and Soil 17 (4): 177–179.
Milbourne, D., R. Meyer, J.E. Bradshaw, E. Baird, N. Bonar, J. Provan, W. Powell, and R. Waugh. 1997. Comparison of PCR-based marker system for the analysis of genetic relationships in cultivated potato. Molecular Breeding 3: 127–136.
Milbourne, D., R.C. Meyer, A.J. Collins, L.D. Ramsay, C. Gebhardt, and R. Waugh. 1998. Isolation, characterization and mapping of simple sequence repeat loci in potato. Molecular and General Genetics 259: 233–245.
Moisan-Thiery, M., S. Marhadour, M.C. Kerlan, N. Dessenne, M. Perramant, T. Gokelaere, and Y. Le Hingrat. 2005. Potato cultivar identification using simple sequence repeats markers (SSR). Potato Research 48 (3–4): 191–200.
Norero, N., J. Malleville, M. Huarte, and S. Feingold. 2002. Cost efficient potato (Solanum tuberosum L.) cultivar identification by microsatellite amplification. Potato Research 45: 131–138.
Oliver, J.L., and J.M. Martinez-Zapater. 1985. A genetic classification of potato cultivars based on allozyme patterns. Theoretical and Applied Genetics 69: 305–311.
Prevost, A., and M.J. Wilkinson. 1999. A new system of comparing PCR primers applied to ISSR fingerprinting. Theoretical and Applied Genetics 98: 107–112.
Qu, D.Y. 2016. China Agriculture Statistical Report. Beijing: China Agriculture Press.
Raker, C.M., and D.M. Spooner. 2002. Chilean tetraploid cultivated potato, Solanum tuberosum, is distinct from the Andean populations: Microsatellite data. Crop Science 42: 1451–1458.
Reid, A., and E.M. Kerr. 2007. A rapid simple sequence repeat (SSR)-based identification method for potato cultivars. Plant Genetic Resources: Characterization and Utilization 5 (1): 7–13.
Ríos, D., M. Ghislain, F. Rodriguez, and D.M. Spooner. 2007. What is the origin of the European potato? Evidence from Canary Island landraces. Crop Science 47 (3): 1271–1280.
Spanoghe, M., T. Marique, J. Rivière, D. Lanterbecq, and M. Gadenne. 2015. Investigation and development of potato parentage analysis methods using multiplexed SSR fingerprinting. Potato Research 58: 43–65.
Stegemann, H., and D. Schnick. 1985. Index 1985 of European potato varieties. Mitteilungen der Biologische Bundesanstalt 227: 1–128.
Tanksley, S.D., and S.R. McCouch. 1997. Seed banks and molecular maps: Unlocking genetic potential from the wild. Science 277: 1063–1066.
Vreugdenhil, D. 2007. Potato biology and biotechnology. Amsterdam: Elsevier press.
Wang, F.G., J.R. Zhao, J.L. Guo, H.D. She, and G. Chen. 2003. Comparison of three DNA fingerprint analyzing methods for maize cultivars' identification. Molecular Plant Breeding 1 (5/6): 655–661.
Acknowledgments
We thank the 22 breeding programs in China for providing the test materials. This study was supported by National Key R&D Program of China (2017YFD0101905) and China Agriculture Research System (CARS-9).
Author information
Authors and Affiliations
Contributions
LJ conceived the project; YD performed the molecular marker analysis; JL and JX conducted part of the experiment; SD collected the materials tested; CB and WP managed the field experiment; GL and JH contributed to the critical revision of the artwork and illustrations. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that there are no conflicts of interest.
Additional information
Key message
Out of 138 SSR markers tested, 20 were polymorphic and they were used to fingerprint and analyze the genetic diversity of 217 potato cultivars grown in China. A set of 11 SSR markers was sufficient to differentiate all the cultivars. The genetic diversity analysis suggested narrow genetic background of these cultivars.
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Duan, Y., Liu, J., Xu, J. et al. DNA Fingerprinting and Genetic Diversity Analysis with Simple Sequence Repeat Markers of 217 Potato Cultivars (Solanum tuberosum L.) in China. Am. J. Potato Res. 96, 21–32 (2019). https://doi.org/10.1007/s12230-018-9685-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12230-018-9685-6