Abstract
In Saccharomyces cerevisiae the export of 2-oxoglutarate from the mitochondria, catalyzed by Yhm2p, Odc1p and Odc2p or by at least one of these transporters, has recently been shown to be essential for glutamate biosynthesis in glucose-supplemented minimal synthetic (SM) medium without glutamate, because the triple mutant yhm2∆odc1∆odc2∆ displays a growth defect under these conditions. Surprisingly, in this study it was found that yhm2∆odc1∆odc2∆ cells grow like wild-type (WT) cells in the same medium supplemented with non-fermentable carbon sources. Direct transport assays of 2-oxoglutarate/2-oxoglutarate homoexchange activity in mitochondria from WT and yhm2∆odc1∆odc2∆ cells (solubilized and reconstituted into liposomes) showed that the mitochondrial extract from yhm2∆odc1∆odc2∆ was completely inactive at variance with that from WT cells, showing that S. cerevisiae mitochondria do not contain additional proteins capable of catalyzing 2-oxoglutarate transport efficiently besides Yhm2p, Odc1p and Odc2p. Furthermore, quantitative real-time PCR experiments showed that in both WT and yhm2∆odc1∆odc2∆ cells the expression of GDH1 is low on lactate and high on glucose and, vice versa, the expression of GDH3 is high on lactate and low on glucose. These results may be interpreted to indicate that in S. cerevisiae, grown in glucose-supplemented SM medium, glutamate is synthesized by Gdh1p in the cytosol, whereas in lactate-supplemented SM medium glutamate is synthesized by Gdh3p in the mitochondria; therefore, the pathway of ammonia assimilation under fermentative conditions requires export of 2-oxoglutarate from the mitochondria, whereas the alternative pathway under respiratory conditions does not.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
In Saccharomyces cerevisiae the biosynthesis of glutamate is accomplished via the action of (i) glutamate synthase (Glt1p), which catalyzes the synthesis of two glutamate molecules from glutamine and 2-oxoglutarate (Filetici et al. 1996), and (ii) glutamate dehydrogenases (Gdhp), which synthesize glutamate from 2-oxoglutarate and ammonia (Moye et al. 1985; Avendaño et al. 1997). In S. cerevisiae there are two glutamate dehydrogenase isoforms, Gdh1p and Gdh3p, involved in glutamate biosynthesis. GDH1 is expressed when cells grow on glucose, while GDH3 is expressed when cells grow on non-fermentable carbon sources (DeLuna et al. 2001). These isoenzymes, which are NADPH dependent, probably differ in their subcellular localization; Gdh1p is localized in the cytosol (Perlman and Mahler 1970), whereas Gdh3p probably in the mitochondria (Sickmann et al. 2003). Notably, for ammonium fixation the actions of the cytosolic Gdh1p and Glt1p require the translocation of 2-oxoglutarate across the mitochondrial membrane to connect the mitochondrial matrix with the cytosol. The third isoform of glutamate dehydrogenase, Gdh2p, is involved in glutamate catabolism and is NAD+ dependent (Boles et al. 1993; Miller and Magasanik 1990).
The genome of S. cerevisiae encodes 35 transport proteins belonging to the mitochondrial carrier family (MCF) (Krämer and Palmieri 1992; Palmieri 2013, 2014; Palmieri et al. 2006a; Palmieri and Monné 2016; Palmieri et al. 2000). Among the yeast MCF members, three mitochondrial 2-oxoglutarate transporters have been identified and characterized biochemically (Castegna et al. 2010; Palmieri et al. 2001). Two of them, Odc1p and Odc2p, catalyze the transport of 2-oxoadipate, 2-oxoglutarate and malate (Fiermonte et al. 2001; Palmieri et al. 2001), and their role is probably to provide 2-oxoadipate or 2-oxoglutarate for the cytosolic biosynthesis of lysine or glutamate, respectively (Palmieri et al. 2001). The third yeast MCF member capable of transporting 2-oxoglutarate, Yhm2p, transports citrate, isocitrate and 2-oxoglutarate (Castegna et al. 2010) and its main physiological role is likely to transport citrate into the cytosol to produce NADPH through the action of isocitrate dehydrogenase (Idp2p) (Castegna et al. 2010; Minard and McAlister-Henn 2005). In this way, Yhm2p is able to supply cytosol with reducing equivalents needed to neutralize reactive oxygen species (ROS). Accordingly, the yhm2Δzwf1Δ strain lacking YHM2 and ZWF1, which encodes another cytosolic NADPH source, glucose 6-phosphate dehydrogenase (Nogae and Johnston 1990), is not able to grow under oxidative stress conditions such as presence of H2O2 or high temperatures (37 °C) (Castegna et al. 2010).
The strain deleted of the ODC1/2 and YHM2 genes (yhm2Δodc1Δodc2Δ) was demonstrated to exhibit a growth defect in a minimal synthetic (SM) medium without lysine or glutamate (Scarcia et al. 2017). The strains lacking only one or two of the above-mentioned three genes did not display a growth defect, showing that all three transporters have an overlapping biochemical function probably consisting in their ability to transport 2-oxoglutarate. Furthermore, the lysine auxotrophy was observed on both fermentable and non-fermentable carbon sources, whereas the glutamate auxotrophy was shown only in glucose-supplemented SM medium (Scarcia et al. 2017).
In this study, we have found that the triple deleted strain yhm2Δodc1Δodc2Δ grows normally in SM medium without glutamate and supplemented with non-fermentable carbon sources; mitochondria from yhm2Δodc1Δodc2Δ are unable to transport 2-oxoglutarate; and in both wild-type (WT) and yhm2Δodc1Δodc2Δ cells glutamate dehydrogenase GDH1 is highly expressed on glucose and little on lactate and, vice versa, GDH3 is little expressed on glucose and highly on lactate. It is concluded that in S. cerevisiae grown on fermentable carbon sources glutamate is synthesized by Gdh1p and on non-fermentable carbon sources by Gdh3p. 2-Oxoglutarate export from the mitochondria is required for the former pathway of ammonium assimilation but not for the latter pathway.
2 Materials and methods
2.1 Strains, media and growth conditions
The strains used in this study are reported in Table 1. The strains were grown at 30 °C in synthetic minimal (SM) medium supplemented with auxotrophic nutrients (Sherman 1991), when required, and 2% glucose, 2% lactate or 3% glycerol as carbon sources. When using solid media, 2% agar was added. In all experiments, the strains were precultured overnight in rich medium (YP) with 2% glucose. For growth studies, the cells were washed in SM medium and serial tenfold dilutions were spotted on solid media. For the preparation of mitochondria the cells were precultured on synthetic complete (SC) medium supplemented with 3% glycerol and 0.1% glucose for 14–16 h, diluted 35-fold in YP supplemented with the same carbon sources, and grown to mid-exponential phase. The mitochondria were isolated as previously described (Daum et al. 1982).
2.2 Reconstitution into liposomes and transport assays
Isolated mitochondria were solubilized with 1% Triton X-100, 50 mM NaCl and 10 mM PIPES, pH 7.0. After incubation for 20 min at 4 °C, the mixture was centrifuged at 138,000×g for 20 min. The mitochondrial extract (30 µg of protein) was reconstituted as previously reported (Palmieri et al. 1999b, 2001). Transport was started by adding [14C]2-oxoglutarate to proteoliposomes preloaded with 10 mM 2-oxoglutarate and terminated by the addition of 30 mM pyridoxal 5′-phosphate and 10 mM bathophenanthroline, which in combination inhibit the activity of many mitochondrial carriers completely and rapidly (Agrimi et al. 2012; Di Noia et al. 2014; Fiermonte et al. 2009; Hoyos et al. 2003; Monné et al. 2015; Palmieri et al. 1999a, 2006b). In controls, the inhibitors were added together with the labeled substrate (the stop-inhibitor method (Indiveri et al. 1994; Marobbio et al. 2006, 2008; Palmieri et al. 1995). The external radioactive substrate was removed, and the radioactivity in the proteoliposomes was measured (Palmieri et al. 1995). The experimental values were corrected by subtracting control values (Marobbio et al. 2002; Marobbio et al. 2003; Porcelli et al. 2014).
2.3 RNA isolation and reverse transcription
Total RNA was isolated from WT or yhm2Δodc1Δodc2Δ cells grown at 30 °C until the early exponential phase was reached (absorbance of 0.8). Aurum Total RNA Kit (Biorad) was used according to the manufacturer’s instructions. The amount of extracted RNA was determined by measuring the absorbance at 260 nm with NanoDrop 1000 (Thermo Scientific), and quality was assessed by the 260/280 absorbance ratio with values of 1.8–2.0 and 260/230 absorbance ratio with values greater than 1.7. The ribosomal RNA band integrity was checked by denaturing agarose/formaldehyde gel electrophoresis and ethidium bromide staining (Sambrook and Russell 2006). The iScript Reverse Transcription Supermix kit (Biorad) with mix of random hexamers and oligo (dT) as primers was used.
2.4 Quantitative PCR reaction
For quantitative real-time PCR (qPCR), primers based on the cDNA sequences of the investigated genes were designed with Primer Express 3.0 (Applied Biosystems, Life Technologies) and purchased from Invitrogen (Life Technologies). The primer sequences used are reported in Table 2. The qPCR reactions were performed using an ABI Prism 7900 HT (Applied Biosystems, Life Technologies). 20 μL of reaction volume contained 20 ng of template (reverse transcribed first-strand cDNA), 10 μL of SYBR Select Master Mix (Applied Biosystems, Life Technologies), and 300 nM of each primer. The specificity of the PCR amplification was checked with the heat dissociation protocol after the final cycle of PCR. To correct for differences in the amount of starting first-strand cDNAs, the yeast β-actin gene (ACT1) was amplified in parallel as a reference gene. The relative quantification of the investigated genes was performed according to the comparative method (2−ΔΔCt) (Agrimi et al. 2004; Fiermonte et al. 2003, 2004). 2−ΔΔCt = 2−(ΔCt sample−ΔCt calibrator), where ΔCt sample is Ct sample − Ct reference gene and Ct is the threshold cycle. For the calibrator, ΔΔCt = 0 and 2−ΔΔCt = 1. The value of 2−ΔΔCt indicates the fold change in gene expression relative to the calibrator.
3 Results
3.1 Glutamate auxotrophy of the yhm2∆odc1∆odc2∆ strain is carbon source dependent
Yeast Yhm2p (citrate oxoglutarate carrier), Odc1p (oxodicarboxylate carrier isoform 1) and Odc2p (oxodicarboxylate carrier isoform 2) show an overlapping substrate specificity, transporting 2-oxoglutarate and, to different extents, citrate and oxoadipate (Castegna et al. 2010; Palmieri et al. 2001). Recently, the transport of 2-oxoglutarate from the mitochondrial matrix to the cytosol, catalyzed by Yhm2p, Odc1p and Odc2p or by at least one of these transporters, has been shown to be essential for glutamate biosynthesis when yeast cells are grown in glucose-supplemented SM medium in the absence of glutamate (Scarcia et al. 2017). To investigate the role of the above-mentioned three mitochondrial 2-oxoglutarate carriers in glutamate biosynthesis during respiration, the triple deleted strain as well as the WT, yhm2Δ and odc1∆odc2∆ strains were grown in SM medium with lactate as carbon source and without glutamate. Surprisingly, no growth defect of the triple mutant yhm2Δodc1Δodc2Δ strain was observed in lactate-supplemented SM medium lacking glutamate (Fig. 1). Similarly in the absence of glutamate yhm2∆odc1∆odc2∆ cells did not display any growth defect in SM medium supplemented with other non-fermentable carbon sources such as glycerol (data not shown).
Deletion of ODC1, ODC2, and YHM2 results in impaired glutamate biosynthesis in SM medium without glutamate and supplemented with glucose but not with lactate. Tenfold serial dilutions of equally numbered wild-type (WT), yhm2Δ, odc1Δodc2Δ and yhm2Δodc1Δodc2Δ cells were spotted on solid SM medium supplemented with 2% glucose (a), 2% lactate (b) with or without glutamate
3.2 Yhm2p, Odc1p and Odc2p are the only 2-oxoglutarate carriers in S. cerevisiae mitochondria
To investigate whether the lack of glutamate auxotrophy of the yhm2∆odc1∆odc2∆ strain in SM medium supplemented with non-fermentable carbon sources was due to the presence or activation of an unknown mitochondrial 2-oxoglutarate transporter, mitochondria were isolated from WT and mutant cells grown on lactate. They were solubilized using Triton X-100, and the resulting mitochondrial extract was reconstituted into liposomes as previously described (Palmieri et al. 1999b; Punzi et al. 2018). Direct transport measurements showed that proteoliposomes reconstituted with the WT mitochondrial extract were capable of catalyzing the homoexchange between externally added [14C]2-oxoglutarate and intraliposomal 2-oxoglutarate (0.14 ± 0.02 mmol/mg protein/30 min), whereas those reconstituted with the yhm2∆odc1∆odc2∆ mitochondrial extract were completely inactive (Fig. 2). Furthermore, proteoliposomes obtained using the yhm2∆ or odc1∆odc2∆ mitochondrial extract exhibited about 47 and 16% of the WT transport activity, respectively. By contrast, the proteoliposomes prepared using the mitochondrial extracts of all four yeast strains showed an equally efficient [14C]ADP/ADP exchange activity of about 1.9 mmol/g protein/30 min.
2-Oxoglutarate/2-oxoglutarate homoexchange activity in liposomes reconstituted with mitochondrial extracts. The extract (30 µg of protein) of isolated mitochondria from wild-type (lined column), yhm2∆odc1∆odc2∆ (black column), yhm2∆ (gray column) and odc1∆odc2∆ (hatched column) strains were reconstituted into liposomes preloaded with 2-oxoglutarate (20 mM). Transport was started by adding 0.1 mM [14C]2-oxoglutarate to proteoliposomes and terminated after 30 min. The data represent the mean ± SEM for at least three independent experiments
These experiments demonstrate that the triple mutant yhm2∆odc1∆odc2∆ strain is unable to transport 2-oxoglutarate and that S. cerevisiae mitochondria do not contain additional proteins capable of catalyzing 2-oxoglutarate transport efficiently besides Yhm2p, Odc1p and Odc2p.
3.3 Expression of the glutamate dehydrogenase isoforms GDH1 and GDH3 on different carbon sources
Since S. cerevisiae yhm2∆odc1∆odc2∆ cells lacking all three mitochondrial 2-oxoglutarate transporters grow on lactate-supplemented SM medium without glutamate, but not in the same medium supplemented with glucose (Fig. 1) and these cells do not export 2-oxoglutarate from mitochondria to cytosol (Fig. 2), we determined the expression of GDH1 and GDH3 genes in the WT and yhm2∆odc1∆odc2∆ strains, grown in SM medium lacking glutamate and supplemented with either glucose or lactate, by quantitative real-time PCR. In both WT and yhm2∆odc1∆odc2∆ cells GDH1 was expressed considerably more when the cells were grown in glucose-supplemented SM medium than in lactate-supplemented medium (Fig. 3a). In fact, the expression of GDH1 was 2.4-fold higher on glucose than on lactate in the WT cells, and 3.1-fold higher on glucose than on lactate in the triple deleted cells. Notably, although the amount of the GDH1 transcript is higher in yhm2∆odc1∆odc2∆ than in WT cells, the fact that the former cells do not grow in SM medium supplemented with glucose indicates that under these conditions is the lack of 2-oxoglutarate transport from the mitochondria to the cytosol to be limiting the growth and not the amount of Gdh1p. By contrast, in both WT and yhm2∆odc1∆odc2∆ cells GDH3 was remarkably more expressed in lactate-supplemented SM medium than in glucose-supplemented medium (Fig. 3b). Specifically, the increase in GHD3 expression on lactate was 2.3-fold in WT cells and more (3.8-fold) in the triple mutant cells as compared to the expression on glucose. The up-regulation of GHD3 on lactate in WT and much higher in yhm2∆odc1∆odc2∆ most likely reflects a compensatory response to the low glutamate concentration in the cytosol due to the absence of 2-oxoglutarate transport from the mitochondrial matrix to the cytosol.
Expression of GDH1 and GDH3 in wild-type and yhm2Δodc1Δodc2Δ cells on glucose or lactate. qPCR analysis of a GDH1 and b GDH3 mRNAs isolated from wild-type and yhm2∆odc1∆odc2∆ cells grown in SM medium lacking glutamate and supplemented with glucose (white columns) or lactate (black columns). The wild-type cells grown on glucose were used as calibrator. The gene relative quantification was performed according to the comparative method (2−ΔΔCt). Values are mean ± SEM of three independent experiments (*p < 0.05, **p < 0.01 and ***p < 0.001, two-tailed unpaired Student’s t test)
4 Discussion
This study examines the role of three mitochondrial carriers, Yhm2p (citrate oxoglutarate transporter), Odc1p (oxodicarboxylate carrier isoform 1) and Odc2p (oxodicarboxylate carrier isoform 2) and two glutamate dehydrogenase isoforms, Gdh1p and Gdh3p in glutamate biosynthesis of the yeast Saccharomyces cerevisiae.
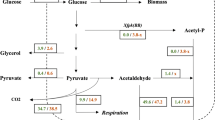
The results reported above can be interpreted to indicate that the synthesis of glutamate in S. cerevisiae is accomplished by different pathways on fermentable and non-fermentable carbon sources (Fig. 4). When S. cerevisiae cells grow on glucose glutamate is synthesized in the cytosol by Gdh1p from 2-oxoglutarate and ammonia, and 2-oxoglutarate is exported from the mitochondrial matrix, where it is produced, to the cytosol by Odc1p, Odc2p or Yhm2p. These three mitochondrial transporters are obligate exchangers. In view of their substrate specificity, we can infer that Odc1p and Odc2p transport 2-oxoglutarate from the mitochondrial matrix to the cytosol in exchange for a cytosolic dicarboxylate (most likely oxaloacetate or malate), and Yhm2p exports 2-oxoglutarate from the mitochondria in exchange for succinate or oxaloacetate. Alternatively, Yhm2p can export citrate, instead of 2-oxoglutarate, from the mitochondria to the cytosol where it is converted into 2-oxoglutarate by Idp2p (NADP-dependent isocitrate dehydrogenase) (Fig. 4). This reaction also catalyzes the reduction of NADP+ to NADPH which can be used by Gdh1p to produce glutamate. The conclusion that S. cerevisiae when grown on glucose synthesizes glutamate by the action of Gdh1p, a pathway that requires the essential intervention of Odc1p, Odc2p and Yhm2p or at least one of them for the export of 2-oxoglutarate (or citrate), is substantiated by the fact that (i) the triple mutant yhm2∆odc1∆odc2∆ does not grow on glucose-supplemented SM medium lacking glutamate [(Scarcia et al. 2017) and Fig. 1 of this study]; (ii) Odc1p, Odc2p and Yhm2p are individually capable of transporting 2-oxoglutarate (Castegna et al. 2010; Palmieri et al. 2001); (iii) S. cerevisiae mitochondria do not contain other efficient transporters of 2-oxoglutarate (Fig. 2), and (iv) the expression of GDH3 is very low on glucose as fermentable carbon source (Fig. 3), suggesting that its contribution to glutamate synthesis is negligible under these conditions. On the contrary, when S. cerevisiae cells grow on non-fermentable carbon sources glutamate is synthesized from 2-oxoglutarate and ammonia by the action of GDH3 in the mitochondrial matrix. Consistently, (a) the triple mutant yhm2∆odc1∆odc2∆ does not exhibit any growth defect on lactate-supplemented SM medium without glutamate (Fig. 1); (b) GDH3 is strongly upregulated on lactate, as compared to on glucose, in WT and much more in yhm2∆odc1∆odc2∆ cells, and (c) the gdh3∆ cells grow remarkably less than WT cells on ethanol as non-fermentable carbon source (DeLuna et al. 2001).
Pathways of glutamate synthesis in S. cerevisiae cells growing in SM medium without glutamate and supplemented with glucose or lactate. The pathway of glutamate synthesis occurring on glucose is indicated by dashed lines, whereas that occurring on lactate by continuous bold lines. In the pathway on sucrose the dashed bold lines were used to indicate the Odc1p-, Odc2p- and Yhm2p-mediated transport of 2-oxoglutarate from the mitochondrial matrix to the cytosol and the Gdh1p-mediated 2-oxoglutarate amination to glutamate, and the dashed unbold lines to indicate the Yhm2p-mediated citrate transport from the mitochondrial matrix to the cytosol, isocitrate formation from citrate and the Idp2p-mediated, NADP+-dependent production of 2-oxoglutarate from isocitrate. TCA cycle tricarboxylic acid cycle, OAA oxaloacetate, Yhm2p citrate oxoglutarate carrier, Odc1p oxodicarboxylate carrier isoform 1, Odc2p oxodicarboxylate carrier isoform 2, Agc1p aspartate glutamate carrier 1, Gdh1p NADP+-dependent glutamate dehydrogenase 1 (cytosolic isoform), Gdh3p NADP+-dependent glutamate dehydrogenase 3 (mitochondrial isoform), Idp2p NADP+-dependent isocitrate dehydrogenase 2 (cytosolic isoform)
While the subcellular localization of Gdh1p has been clearly demonstrated to be cytosolic (Perlman and Mahler 1970), the Gdh3p subcellular localization has not been definitively established. Using GFP-tagged proteins, Huh et al. (2003) localized Gdh1p and Gdh2p in the cytosol, but did not detect Gdh3p probably because yeast cells were grown on glucose. Conversely, glutamate dehydrogenase isoform 3 and 2 were localized in mitochondria in a large-scale proteomic study (Sickmann et al. 2003). Our data strongly support the contention that Gdh3p is localized to mitochondria because ammonia fixation in the presence of a non-fermentable substrate does not require the presence of Odc1p, Odc2p or Yhm2p, i.e., the export of 2-oxoglutarate, as demonstrated by the glutamate prototrophy of the yhm2∆odc1∆odc2∆ strain on lactate (Fig. 1). Obviously in the Gdh3p-mediated pathway of glutamate synthesis this intramitochondrially produced amino acid has to be exported to the cytosol (Fig. 4), a transport step that can be catalyzed by the aspartate glutamate carrier Agc1p (Cavero et al. 2003) and/or by a not yet identified glutamate transport system localized in the mitochondrial membrane.
References
Agrimi G, Di Noia MA, Marobbio CM, Fiermonte G, Lasorsa FM, Palmieri F (2004) Identification of the human mitochondrial S-adenosylmethionine transporter: bacterial expression, reconstitution, functional characterization and tissue distribution. Biochem J 379:183–190. https://doi.org/10.1042/BJ20031664
Agrimi G, Russo A, Scarcia P, Palmieri F (2012) The human gene SLC25A17 encodes a peroxisomal transporter of coenzyme A, FAD and NAD(+). Biochem J 443:241–247. https://doi.org/10.1042/BJ20111420
Avendaño A, Deluna A, Olivera H, Valenzuela L, Gonzalez A (1997) GDH3 encodes a glutamate dehydrogenase isozyme, a previously unrecognized route for glutamate biosynthesis in Saccharomyces cerevisiae. J Bacteriol 179:5594–5597. https://doi.org/10.1128/jb.179.17.5594-5597.1997
Boles E, Lehnert W, Zimmermann FK (1993) The role of the NAD-dependent glutamate dehydrogenase in restoring growth on glucose of a Saccharomyces cerevisiae phosphoglucose isomerase mutant. Eur J Biochem 217:469–477. https://doi.org/10.1111/j.1432-1033.1993.tb18266.x
Castegna A, Scarcia P, Agrimi G, Palmieri L, Rottensteiner H, Spera I, Germinario L, Palmieri F (2010) Identification and functional characterization of a novel mitochondrial carrier for citrate and oxoglutarate in Saccharomyces cerevisiae. J Biol Chem 285:17359–17370. https://doi.org/10.1074/jbc.m109.097188
Cavero S, Vozza A, del Arco A, Palmieri L, Villa A, Blanco E, Runswick MJ, Walker JE, Cerdán S, Palmieri F, Satrústegui J (2003) Identification and metabolic role of the mitochondrial aspartate-glutamate transporter in Saccharomyces cerevisiae. Mol Microbiol 50:1257–1269. https://doi.org/10.1046/j.1365-2958.2003.03742.x
Daum G, Gasser SM, Schatz G (1982) Import of proteins into mitochondria. Energy-dependent, two-step processing of the intermembrane space enzyme cytochrome b2 by isolated yeast mitochondria. J Biol Chem 257:13075–13080
DeLuna A, Avendaño A, Riego L, González A (2001) NADP-glutamate dehydrogenase isoenzymes of Saccharomyces cerevisiae. J Biol Chem 276:43775–43783. https://doi.org/10.1074/jbc.m107986200
Di Noia MA, Todisco S, Cirigliano A, Rinaldi T, Agrimi G, Iacobazzi V, Palmieri F (2014) The human SLC25A33 and SLC25A36 genes of solute carrier family 25 encode two mitochondrial pyrimidine nucleotide transporters. J Biol Chem 289:33137–33148. https://doi.org/10.1074/jbc.M114.610808
Fiermonte G, Dolce V, Palmieri L, Ventura M, Runswick MJ, Palmieri F, Walker JE (2001) Identification of the human mitochondrial oxodicarboxylate carrier. Bacterial expression, reconstitution, functional characterization, tissue distribution, and chromosomal location. J Biol Chem 276:8225–8230. https://doi.org/10.1074/jbc.M009607200
Fiermonte G, Dolce V, David L, Santorelli FM, Dionisi-Vici C, Palmieri F, Walker JE (2003) The mitochondrial ornithine transporter. Bacterial expression, reconstitution, functional characterization, and tissue distribution of two human isoforms. J Biol Chem 278:32778–32783. https://doi.org/10.1074/jbc.M302317200
Fiermonte G, De Leonardis F, Todisco S, Palmieri L, Lasorsa FM, Palmieri F (2004) Identification of the mitochondrial ATP-Mg/Pi transporter. Bacterial expression, reconstitution, functional characterization, and tissue distribution. J Biol Chem 279:30722–30730. https://doi.org/10.1074/jbc.M400445200
Fiermonte G, Paradies E, Todisco S, Marobbio CM, Palmieri F (2009) A novel member of solute carrier family 25 (SLC25A42) is a transporter of coenzyme A and adenosine 3′,5′-diphosphate in human mitochondria. J Biol Chem 284:18152–18159. https://doi.org/10.1074/jbc.M109.014118
Filetici P, Martegani MP, Valenzuela L, González A, Ballario P (1996) Sequence of the GLT1 gene from Saccharomyces cerevisiae reveals the domain structure of yeast glutamate synthase Yeast 12:1359–1366. https://doi.org/10.1002/(sici)1097-0061(199610)12:13<1359::aid-yea3>3.0.co;2-5
Hoyos ME, Palmieri L, Wertin T, Arrigoni R, Polacco JC, Palmieri F (2003) Identification of a mitochondrial transporter for basic amino acids in Arabidopsis thaliana by functional reconstitution into liposomes and complementation in yeast. Plant J 33:1027–1035
Huh W-K, Falvo JV, Gerke LC, Carroll AS, Howson RW, Weissman JS, O’Shea EK (2003) Global analysis of protein localization in budding yeast. Nature 425:686–691. https://doi.org/10.1038/nature02026
Indiveri C, Tonazzi A, Palmieri F (1994) The reconstituted carnitine carrier from rat liver mitochondria: evidence for a transport mechanism different from that of the other mitochondrial translocators. Biochim Biophys Acta 1189:65–73. https://doi.org/10.1016/0005-2736(94)90281-x
Krämer R, Palmieri F (1992) Chapter 16 Metabolite carriers in mitochondria. In: Ernster L (ed) Molecular mechanisms in bioenergetics. Elsevier Science Publishers B. V., Amsterdam. https://doi.org/10.1016/s0167-7306(08)60184-2
Marobbio CMT, Vozza A, Harding M, Bisaccia F, Palmieri F, Walker JE (2002) Identification and reconstitution of the yeast mitochondrial transporter for thiamine pyrophosphate. EMBO J 21:5653–5661. https://doi.org/10.1093/emboj/cdf583
Marobbio CMT, Agrimi G, Lasorsa FM, Palmieri F (2003) Identification and functional reconstitution of yeast mitochondrial carrier for S-adenosylmethionine The. EMBO J 22:5975–5982. https://doi.org/10.1093/emboj/cdg574
Marobbio CMT, Di Noia MA, Palmieri F (2006) Identification of a mitochondrial transporter for pyrimidine nucleotides in Saccharomyces cerevisiae: bacterial expression, reconstitution and functional characterization. Biochem J 393:441–446. https://doi.org/10.1042/bj20051284
Marobbio CMT, Giannuzzi G, Paradies E, Pierri CL, Palmieri F (2008) alpha-Isopropylmalate, a leucine biosynthesis intermediate in yeast, is transported by the mitochondrial oxalacetate carrier. J Biol Chem 283:28445–28453. https://doi.org/10.1074/jbc.M804637200
Miller SM, Magasanik B (1990) Role of NAD-linked glutamate dehydrogenase in nitrogen metabolism in Saccharomyces cerevisiae. J Bacteriol 172:4927–4935. https://doi.org/10.1128/jb.172.9.4927-4935.1990
Minard KI, McAlister-Henn L (2005) Sources of NADPH in yeast vary with carbon source. J Biol Chem 280:39890–39896. https://doi.org/10.1074/jbc.m509461200
Monné M, Miniero DV, Obata T, Daddabbo L, Palmieri L, Vozza A, Nicolardi MC, Fernie AR, Palmieri F (2015) Functional characterization and organ distribution of three mitochondrial ATP–Mg/Pi carriers in Arabidopsis thaliana. Biochim Biophys Acta 1847:1220–1230. https://doi.org/10.1016/j.bbabio.2015.06.015
Moye WS, Amuro N, Rao JK, Zalkin H (1985) Nucleotide sequence of yeast GDH1 encoding nicotinamide adenine dinucleotide phosphate-dependent glutamate dehydrogenase. J Biol Chem 260:8502–8508
Nogae I, Johnston M (1990) Isolation and characterization of the ZWF1 gene of Saccharomyces cerevisiae, encoding glucose-6-phosphate dehydrogenase. Gene 96:161–169. https://doi.org/10.1016/0378-1119(90)90248-p
Palmieri F (2013) The mitochondrial transporter family SLC25: identification, properties and physiopathology. Mol Aspects Med 34:465–484. https://doi.org/10.1016/j.mam.2012.05.005
Palmieri F (2014) Mitochondrial transporters of the SLC25 family and associated diseases: a review. J Inherit Metab Dis 37:565–575. https://doi.org/10.1007/s10545-014-9708-5
Palmieri F, Monné M (2016) Discoveries, metabolic roles and diseases of mitochondrial carriers: a review. Biochim Biophys Acta 1863:2362–2378. https://doi.org/10.1016/j.bbamcr.2016.03.007
Palmieri F, Indiveri C, Bisaccia F, Iacobazzi V (1995) Mitochondrial metabolite carrier proteins: purification, reconstitution, and transport studies. Methods Enzymol 260:349–369. https://doi.org/10.1016/0076-6879(95)60150-3
Palmieri L, Lasorsa FM, Iacobazzi V, Runswick MJ, Palmieri F, Walker JE (1999a) Identification of the mitochondrial carnitine carrier in Saccharomyces cerevisiae. FEBS Lett 462:472–476
Palmieri L, Vozza A, Honlinger A, Dietmeier K, Palmisano A, Zara V, Palmieri F (1999b) The mitochondrial dicarboxylate carrier is essential for the growth of Saccharomyces cerevisiae on ethanol or acetate as the sole carbon source. Mol Microbiol 31:569–577. https://doi.org/10.1046/j.1365-2958.1999.01197.x
Palmieri L, Lasorsa FM, Vozza A, Agrimi G, Fiermonte G, Runswick MJ, Walker JE, Palmieri F (2000) Identification and functions of new transporters in yeast mitochondria. Biochim Biophys Acta 1459:363–369. https://doi.org/10.1016/S0005-2728(00)00173-0
Palmieri L, Agrimi G, Runswick MJ, Fearnley IM, Palmieri F, Walker JE (2001) Identification in Saccharomyces cerevisiae of two isoforms of a novel mitochondrial transporter for 2-oxoadipate and 2-oxoglutarate. J Biol Chem 276:1916–1922. https://doi.org/10.1074/jbc.M004332200
Palmieri F, Agrimi G, Blanco E, Castegna A, Di Noia MA, Iacobazzi V, Lasorsa FM, Marobbio CM, Palmieri L, Scarcia P, Todisco S, Vozza A, Walker J (2006a) Identification of mitochondrial carriers in Saccharomyces cerevisiae by transport assay of reconstituted recombinant proteins. Biochim Biophys Acta 1757:1249–1262. https://doi.org/10.1016/j.bbabio.2006.05.023
Palmieri L, Arrigoni R, Blanco E, Carrari F, Zanor MI, Studart-Guimaraes C, Fernie AR, Palmieri F (2006b) Molecular identification of an Arabidopsis S-adenosylmethionine transporter. Analysis of organ distribution, bacterial expression, reconstitution into liposomes, and functional characterization. Plant Physiol 142:855–865. https://doi.org/10.1104/pp.106.086975
Perlman PS, Mahler HR (1970) Intracellular localization of enzymes in yeast. Arch Biochem Biophys 136:245–259. https://doi.org/10.1016/0003-9861(70)90348-6
Porcelli V, Fiermonte G, Longo A, Palmieri F (2014) The human gene SLC25A29, of solute carrier family 25, encodes a mitochondrial transporter of basic amino acids. J Biol Chem 289:13374–13384. https://doi.org/10.1074/jbc.M114.547448
Punzi G, Porcelli V, Ruggiu M, Hossain MF, Menga A, Scarcia P, Castegna A, Gorgoglione R, Pierri CL, Laera L, Lasorsa FM, Paradies E, Pisano I, Marobbio CMT, Lamantea E, Ghezzi D, Tiranti V, Giannattasio S, Donati MA, Guerrini R, Palmieri L, Palmieri F, De Grassi A (2018) SLC25A10 biallelic mutations in intractable epileptic encephalopathy with complex I deficiency. Hum Mol Genet 27:499–504. https://doi.org/10.1093/hmg/ddx419
Sambrook J, Russell DW (2006) Separation of RNA according to size: electrophoresis of RNA through agarose gels containing formaldehyde. Cold Spring Harbor Protoc. https://doi.org/10.1101/pdb.prot4050
Scarcia P, Palmieri L, Agrimi G, Palmieri F, Rottensteiner H (2017) Three mitochondrial transporters of Saccharomyces cerevisiae are essential for ammonium fixation and lysine biosynthesis in synthetic minimal medium. Mol Genet Metab 122:54–60. https://doi.org/10.1016/j.ymgme.2017.07.004
Sherman F (1991) Getting started with yeast. Methods Enzymol 194:3–21. https://doi.org/10.1016/0076-6879(91)94004-v
Sickmann A, Reinders J, Wagner Y, Joppich C, Zahedi R, Meyer HE, Schönfisch B, Perschil I, Chacinska A, Guiard B, Rehling P, Pfanner N, Meisinger C (2003) The proteome of Saccharomyces cerevisiae mitochondria. Proc Natl Acad Sci USA 100:13207–13212. https://doi.org/10.1073/pnas.2135385100
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Acknowledgements
This work was supported by Grants from the Ministero dell’Istruzione, dell’Università e della Ricerca (MIUR) and the Center of Excellence on Comparative Genomics (CEGBA).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Scarcia, P., Agrimi, G., Germinario, L. et al. In Saccharomyces cerevisiae grown in synthetic minimal medium supplemented with non-fermentable carbon sources glutamate is synthesized within mitochondria. Rend. Fis. Acc. Lincei 29, 483–490 (2018). https://doi.org/10.1007/s12210-018-0687-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12210-018-0687-6