Abstract
Background
Oral squamous cell carcinoma (OSCC) is a commonly occurring malignancy with complex genetic alterations contributing to its development. The H-Ras, a proto-oncogene, becomes an oncogene when mutated and has been implicated in various cancers. This systematic review aims to research to what extent H-Ras expression and mutation contribute to the development and progression of OSCC, and how does this molecular alteration impacts the clinical characteristics and prognosis in patients with OSCC.
Methods
A thorough electronic scientific literature search was carried out in PUBMED, SCOPUS, and GOOGLE SCHOLAR databases from 2007 to 2021. The search strategy yielded 120 articles. Following aggregation and filtering all results through our inclusion and exclusion criteria total 9 articles were included in our literature review. It has also been registered with PROSPERO (CRD42023485202).
Results
It was found that mutations in the Ras gene commonly reported in hotspots at codons 12, 13, and 61 resulting in the activation of downstream signaling pathways causing abnormal and uncontrolled cell growth. This systematic review has shown an increased prevalence of H-Ras mutation in well-differentiated OSCC and also the prevalence of H-Ras mutation in individuals engaging in multiple risk behaviors, particularly chewing tobacco, demonstrated a significant association with a higher prevalence of H-Ras positivity.
Conclusion
This review sheds light on the prevalence of H-Ras mutations, their association with clinical characteristics, and their potential implications for OSCC prognosis. It also enhances our comprehension of the molecular mechanisms that underlie OSCC and paves the way for further research into targeted treatments based on H-Ras alterations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oral squamous cell carcinoma (OSCC) stands out as the most common malignant tumor affecting the oral cavity, representing 80 to 90% of all malignancies in this area [1]. It is a medical condition associated with high morbidity and mortality along with a compromised quality of life. OSCC accounts for the majority of oral malignancies with only 50–60% of cases exhibiting a 5 year survival rate [2]. The habits, environment, and genetic factors interplay to form the risk factors for OSCC. The tongue is the anatomical location that is most commonly affected [3, 4]. Moreover, the primary etiological factor for squamous cell carcinoma affecting the lower lip is intense exposure to sunlight [5]. While smoking and alcohol consumption are recognized as significant risk factors for OSCC, it’s essential to highlight that only a fraction of individuals who engage in these habits ultimately develop oral cancer [6]. Due to the limited duration of exposure to significant risk factors, such as prolonged sunlight exposure and the use of tobacco and/or alcohol, young individuals face a distinct set of circumstances. Studies propose variations in the etiology of Oral Squamous Cell Carcinoma (OSCC) between younger and older patients [7,8,9,10,11]. Furthermore, some research indicates that younger individuals with OSCC are more likely to be nonsmokers and nondrinkers [9, 10, 12]. This observation implies that there are likely other genetic factors at play in the onset and advancement of the disease.
Although multiple cancer-related genes have been discovered as possible therapeutic targets, there are only a few molecular treatment options available for OSCC. Therefore, surgery is still the first line of treatment [13]. Numerous genes have been reported to play a crucial role in the etiology of OSCC in recent literature [14, 15]. Literature search has revealed that approximately 30% of human tumors harbor mutations in either K-Ras, N-Ras, or H-Ras genes that are vital components of the Ras-Raf-MEK-ERK-MAP kinase signaling pathway [16]. The Ras-Raf-MEK-ERK-MAP kinase signaling pathway regulates cell cycle progression and apoptosis in various cell types. Therefore, key genes and their subsequent proteins can be significant targets for cancer therapeutics.
Harvey-Ras (H-Ras) is the first discovered human proto-oncogene [17]. The gene has received a lot of interest, particularly in oral cancer patients. Over the past two decades, there has been evidence indicating frequent mutations in Ras genes across various tumor types, highlighting their involvement in tumor proliferation and maintenance [18]. This is attributed not just to the challenging nature of directly inhibiting Ras proteins but also to the remarkable capability of Ras-mutant cancers in rendering therapeutic agents ineffective, thereby enhancing tumor fitness [19, 20].
The H-RAS is a small G-protein in the Ras superfamily that possesses inherent guanosine triphosphatase (GTPase) activity, facilitating the transmission of growth signals from the cell surface to intracellular effectors via mitogenic activating protein kinase (MAPK), c‑Jun N‑terminal kinase (JNK), and p38‑kinase pathways. These pathways play a crucial role in regulating normal cell proliferation functions [21, 22]. The HRAS protein functions as a molecular toggle, alternating between the active Guanosine triphosphate (GTP)-bound state and the inactive Guanosine diphosphate (GDP)-bound state. When activated by binding to GTP, HRAS transmits downstream signals to various cellular pathways, including the Raf-MEK-ERK pathway. Mutations in the RAS gene result in the mutant RAS protein losing its capacity to exchange GTP with GDP, leading to sustained activation of the protein. HRAS protein becomes constitutively active therefore it is continuously signaling for cell growth and proliferation, even when it should not be. This uncontrolled signaling can lead to the formation of tumors and promote cancer progression [23].
In this systematic review, our objective is to offer insights into the potential role of H-Ras in the development of oral cancer and to analyze the status of H-Ras gene mutations in OSCC to gauge the possibility of targeting H-Ras for therapeutic benefits in OSCC patients.
Methods
Protocol and Registration
This systematic review adhered to the Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) guidelines. It has also been registered with PROSPERO (CRD42023485202).
Criteria for Studies to be Considered (PICOS)
-
Participants Any age and gender with clinically and histologically diagnosed cases of oral squamous cell carcinoma.
-
Interventions H-Ras gene mutation.
-
Control Normal Individuals without OSCC.
-
Outcomes Expression and Mutation of H-Ras gene in OSCC patients.
-
Studies Original studies.
Data Sources
A thorough electronic scientific literature search was carried out in PUBMED, SCOPUS, and GOOGLE SCHOLAR databases from 2007 to 2021.
Search Strategy
The search was created by combining the keywords with AND/OR Boolean without any language or time restrictions. Keywords used for the electronic literature search were (HRAS mutation) OR (mutation of HRAS) OR (mutation of HRAS in OSCC) (HRAS in oral cancer) OR (mutation of HRAS in oral cancer) (oral cancer) AND (HRAS).
Inclusion Criteria
-
(a)
Original studies containing primary data
-
(b)
Studies including humans with oral squamous cell carcinoma as subjects (i.e., no cell line models or animal models) were permitted.
Exclusion Criteria
-
(a)
Experimental studies, reviews, letters, abstracts, opinion articles, case reports, case series, and book chapters were excluded from the analysis.
-
(b)
Studies using non-human subjects.
-
(c)
Studies in languages other than English were excluded.
Screening and Study Selection
Two blinded reviewers independently chose the studies to be included [PD, RD]. The search was conducted in January 2022. After the title and abstract were reviewed, 120 papers were chosen for full-text review. The first reviewer (PD) identified irrelevant papers based on their title and abstracts and excluded them. Two reviewers (PD, RD) independently screened the full texts of all possible qualified studies and examined them for duplicates.
Literature Search
The search strategy yielded 120 articles. Fifty duplicate articles were excluded, and 40 articles were excluded following title and abstract analysis. Assessment of the full text was done for 30 articles. After aggregating and filtering all the results according to our predefined criteria, 15 articles were excluded as some studies were done on animals and few studies were on cell lines. After the screening, a total of 9 [24,25,26,27,28,29,30,31,32] articles were included in our literature review. The total number of participants included was 697 across different countries. Additional articles were also included in the study to increase the comprehension. (Fig. 1).
Data Extraction and Qualitative Synthesis
The following parameters were obtained from the studies that were included: Author, publication year, country of study, the total number of cases, control sample, method used, mutation of the gene, mutation about tumor location, and outcome of the study. Using a Microsoft Excel spreadsheet, the data were recorded and summarized. No statistical analysis was performed.
Quality Assessment
Refer to Table 1.
Results
Refer to Table 2.
Discussion
High mortality and morbidity associated with oral cancer particularly OSCC have intrigued researchers to direct studies towards identifying genetic/molecular changes that are responsible for carcinogenesis. Since genetic changes form the basis of carcinogenetic changes it becomes crucial to identify and gauge them. Ras genes are crucial players in several key pathways of cell growth. It is well understood that their mutation can significantly affect the transformation of normal tissues to malignancy [33, 34]. Therefore, it becomes crucial to understand and explore this gene and its mechanism of downstream signaling.
Ras Gene: The Basic Knowhow!
The Ras-Raf-MEK-ERK-MAP kinase signaling pathway couples cellular response to growth signals and Ras genes are an active component of this system. Therefore, mutations causing activation of this pathway can lead to the development of cancer [35]. Mutations in the Ras gene commonly occur at specific regions known as “hotspots,” which results in constant activation of downstream signaling pathways driven by Ras [36].
Mutations in Ras genes have been reported with significant variations across different types of human cancer and ethnicities [37]. The Ras gene family has 3 functional Ras genes—H-Ras (Harvey-Ras), K-Ras (Kristen-Ras- isoform A and isoform B), and N-Ras (Neuroblastoma-Ras) which encodes four nuclear receptors. Although compared to the K-Ras and N-Ras genes, the H-Ras gene undergoes fewer mutations. But K-Ras and N-Ras mutations are not frequently seen in head and neck malignancies, there the H-Ras mutations predominate [38].
Mechanism of Action of Ras Gene in Oral Carcinogenesis
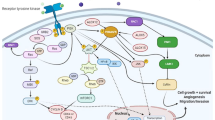
The Ras gene serves as an upstream controller of the Raf-MEK-ERK pathway. Alterations in the Ras gene and subsequently in the RAS protein have been identified as significant contributors to the initiation and advancement of oral cancer [39, 40]. Ras is activated by various factors like EGF, tumor necrosis factor (TNF) and Protein Kinase C (PKC) activators [41]. Extracellular signals bind to the receptors. Then the activated receptor binds to Grb2, which interacts with the proline-rich sequence of Son of Sevenless (SOS) to form the Receptor-Grb2-SOS Complex. When SOS binds to the tyrosine phosphorylation site on the receptor or receptor substrate protein, it triggers the movement of cytoplasmic SOS to the membrane. This relocation leads to a significant concentration of SOS near Ras. SOS and Ras-GDP interaction take place therefore SOS and Ras-GDP promote the replacement of GDP with GTP. This activates Ras and initiates the Ras pathway (Fig. 2).
Evidence of H-Ras Mutation in Oral Carcinogenesis
Literature has several reports where H-Ras mutations have been identified as a significant contributor to the pathogenesis of oral cancer. Research conducted in three distinct regions of India revealed diversity in Ras mutations, encompassing variations in both the percentage of mutations and the specific types of Ras genes affected. Das et al. studied fifty oral cancer specimens using selective oligodeoxynucleotide hybridization and restriction fragment length polymorphism analysis of polymerase chain reaction amplified products and observed mutation of the H- Ras gene in 28% of its total cases [42]. A previous study by Munirajan et al. analyzed the mutation of the H-Ras gene by polymerized chain reaction-single-strand conformation polymorphism (PCR-SSCP) and direct sequencing of 46 oral SCCs. During this study, the author discovered that the H-Ras gene had mutations at codons 12 (in 6 cases), codon 13 (in 1 case), and codon 59 (in 1 case) in 8 out of 46 cases, i.e. 17% of all its cases [43]. Similarly, Saranath et al. examined 57 primary oral cancer samples from Indian patients and observed 37% mutation in H- Ras gene. Among these, eight samples exhibited mutations at codon 12, one at codon 13 and thirteen at codon 61 [44].
In four out of nine, i.e. (44.4%) studies included in this systematic review; it was observed that mutation at Codon 12 was reported in a significant number of cases. Additionally, mutations were also reported at other codons like codon 13, codon 62 and codon 61 [24,25,26, 31]. Ten cases harbored a mutation in H- Ras gene in a study done by Murugan et al. Out of these ten cases, two novel mutations were identified. The first mutation involved the insertion of three nucleotides (GGC) between codons 10 and 11 (10Gly11), whereas codon 62 (E62G) harbored the second mutation, a missense mutation [25].
Mutation Analysis Through Polymerase Chain Reaction and DNA Sequencing
Employing next-generation sequencing (NGS) in the context of head and neck squamous cell carcinoma (HNSCC) has resulted in the discovery of previously unidentified mutated oncogenes. This advancement has also contributed to the creation of predictive biomarkers. In clinical samples, various coding exons were examined. Primer pairs for PCR and sequencing were designed after which PCR products were purified and sequenced. Sequences were analyzed using Sequencing Analysis software. In six of the included studies, 8.4% of the cases were found to have a mutation in the H-Ras gene [24,25,26,27, 31, 32]. The H-Ras gene has six exons and codes for a polypeptide of 189 amino acids and a molecular weight of 21 kDa. as reported by Sanchez-Montenegro et al. [45]. As indicated by the data from the catalogue of somatic mutations in cancer (COSMIC), among the 114 documented mutations of H-Ras in OSCC, the majority are characterized as point mutations occurring in codons 12, 13, or 61 and some other locations. The cases reported were 59 cases (52%), 22 cases (19%), 21 cases (18%) and 12 cases (11%), respectively [46].
Mutation Analysis Through Quantitative Real-Time PCR (qPCR)
Roodi et al. employed quantitative real-time PCR (qPCR) on OSCC and reported that the mRNA level expression of H-Ras was significantly higher, showing a threefold elevation (p = 0.044) when compared to the normal mucosal tissue of the oral cavity [29]. The levels of H-Ras mRNA fold change in patients (n = 56) were investigated later by Krishna et al.; they varied from 0.46 to 6.38, with a median of 1.28 and a mean standard deviation [SD] of 1.62 ± 1.02. The mRNA fold change in the control group, in contrast, ranged from 0.34 to 2.47, had a median value of 0.97, and had a mean SD of 1.10 ± 0.50. In particular, the H-Ras mRNA level showed a substantial increase in oral cancer compared to the control group (p 0.001) [30]. The elevated presence of H-Ras in tissue biopsies from OSCC, as documented by these investigations, suggests the involvement of this oncogene in cancer development among these individuals.
Identification of Mutation Through Multiplex PCR and Primer Extension Analysis
Chang et al. examined tissues obtained from a group of 79 patients diagnosed with OSCC and analyzed the samples using multiplex PCR and primer extension techniques to investigate the frequency of RAS gene mutations in exons 2 and 3. Among these patients, H-Ras mutations were detected in 10/79 individuals (12.66%). Among the 10 identified H-Ras mutations, all were linked to residue 12. These included 9 cases of GGC → AGC mutation (G12S) and 1 instance of GGC → TGC mutation (G12C) [28].
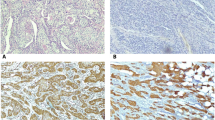
Immunohistochemical Analysis of H-RAS Protein
It is understood that to judge whether the genetic changes translate into altered protein expression too, the immunohistochemical (IHC) analysis of the H-RAS protein is considered. Krishna et al. used IHC to analyze the H-RAS protein in tissue samples from OSCC. They found that 39/65 cases (60%) showed positive H-RAS expression, but no statistically significant differences were observed between the case and control groups in the subcategories of H-RAS expression. Also, a majority of tissue samples exhibited moderate levels of positive immunostaining for H-RAS [30].
A few earlier studies have revealed that the H-RAS protein has a role in the metabolism of normal cells and is not just present in cancerous tumors [37]. The above results by Krishna et al. [30] align with the research conducted by Cutilli et al., where they investigated the tumor suppressor p53 and H-Ras oncogene through immunohistochemical and genetic analysis in Oral and maxillofacial neoplasms. They found that the H-RAS protein was significantly overexpressed immunohistochemically in the majority of patients (12 out of 15 cases; 80%) [47]. Similar results were found by McDonald et al., 68% of cases of head and neck squamous cell carcinoma stained positive for H-RAS protein [48].
Correlations of H-Ras Mutation with Other Key Regulators of Oral Carcinogenesis
H-Ras and G1 Cell Cycle Regulators
Additionally, a significant relationship was found between the H-Ras mutation and the G1 phase of the cell cycle regulatory proteins cyclin D1 and CDK4. The signal transduction pathway Ras–Raf–MEK–ERK plays a crucial role in governing cell-cycle advancement across various cell categories through the modulation of transcription factors. Ras exerts its influence during various stages of the cell cycle, encompassing the early G1 phase, the transition from G1 to S, and the G2/M phase [49]. In a previous study, it was documented that elevated p16 expression and reduced cyclin D1 levels were linked to a positive outcome in cases of oral carcinoma [50]. There was also an increase in the expression of p16 and Rb proteins. Williams and colleagues [51] documented that the absence of the Rb tumor suppressor results in reduced proliferation in tumor cells having mutated Ras genes. The noticeable increase in Rb expression within Ras-mutated cases might create a conducive setting for Ras-driven oncogenesis.
H-Ras and Cox-2 Expression
According to Roodi et al., there was a notable and statistically significant increase in Cox-2 expression which is an enzyme within the prostaglandin pathway, in tumor tissue (an increase of 11.5-fold, p < 0.0001). They demonstrated the relationship between the mRNA levels of Cox-2 and H-Ras. As compared to normal gingival biopsy tissues of healthy individuals minimal Cox-2 expression at the mRNA level was detected. However, they also proposed that activation and overexpression of H-Ras during carcinogenesis might upregulate Cox-2 and elucidate the role of inflammatory responses in oncogene mutations and the advancement of cancer [29]. The Ras family presents itself as a susceptible target for diverse environmental mutagens and lifestyle-related elements like smoking and alcohol consumption. It was observed that Cox-2 is easily stimulated in reaction to such factors. These findings align with the observations made by Lee et al., who studied invasive rat liver epithelial cells and proposed that H-Ras might exert specific control over MMP-9 and Cox-2 by triggering the ERKs and IKK-IkBa-NFkB signaling pathway [52].
Clinicopathological Correlation
Evaluation of clinical and pathological data seems crucial along with mutational status to explore any existence of an association between H-Ras mutation and tumor grade (Table 2).
Tumor Grade
A potential link between H-Ras mutations and the differentiation status of OSCC, with an emphasis on well-differentiated tumors in some studies, has been found. Murugan et al. in their study observed that there was a significant portion of H-Ras mutations that were identified in well-differentiated squamous cell carcinomas (p = 0.0356). One mutation was discovered in a tumor that was only moderately differentiated, out of the total of ten. Nine of the mutations were found in well-differentiated tumors [25]. Consistent with this, Krishna et al. illustrated that among 45 well-differentiated cases, 26 (57.7%) displayed elevated levels of H-RAS protein. While among 17 moderately differentiated cases, 10 exhibited similar higher expression [30]. Nonetheless, these variations did not yield statistically significant differences. Batta and Pandey also demonstrated that out of a total of 8 mutated cases of the H-Ras gene, 5 cases of well-differentiated and 3 cases of moderately differentiated oral squamous cell carcinoma were seen [31]. Whereas Dimitra Koumaki et al. in their study revealed that 2 case of H-Ras mutated case were moderately differentiated oral squamous cell carcinoma [26]. Roodi et al. when considering tumor grade, analysis revealed a statistically significant increase in H-Ras expression among tumors diagnosed as moderately to poorly differentiated compared to well-differentiated tumors (p = 0.033) [29]. However, the variations in findings and the lack of statistical significance in some instances indicate the complexity of this relationship and the need for further research to elucidate the precise role of H-Ras in differentiating oral squamous cell carcinomas.
Habit Association
Individuals who practiced numerous risk behaviors, such as chewing tobacco, were shown to have a substantial number of H-Ras positive cases. Six of the included studies exhibited habit association with the H-Ras gene [25, 28,29,30,31,32]. Earlier research has indicated a noteworthy correlation between chewing tobacco and heightened rates of H-Ras mutations in individuals with OSCC [43, 44]. Significantly, nitroso-containing compounds have gained acknowledgment due to their propensity to provoke H-Ras mutations, especially in experimental animal models simulating skin and breast cancer [53, 54]. This lends credibility to the idea that nitroso compounds originating from tobacco might contribute to the emergence of OSCC. In a study by Murugan, it was observed that patients with tobacco-related habits such as chewing and smoking exhibited a greater occurrence of H-Ras positive expression [25]. The genetic modifications and heightened immunoexpression levels of Ras genes within tumors could potentially mirror the underlying causes and ethnic backgrounds of the patients. Chang et al. study noted that a majority of oral cancer patients were engaged in tobacco use or multiple risk behaviors [55]. Another investigation by Xu J et al. revealed a link between tobacco use and oral cancer cases with H-Ras mutations in particular codons 12, 13 and 61 [56]. Therefore, it is reasonable to suggest that the presence of the mutant form of the protein may have contributed to the participants in Krishna et al. study having an enhanced expression of the H-Ras protein. Smokers were 1.46 times more likely than non-smokers to test positive for H-Ras [30]. Similarly, Uchibori et al. also observed that H-Ras mutations were associated with chewing tobacco (p < 0.05) [32].
Gender Correlation
Upon gender-based analysis, a noteworthy contrast emerged in the distribution of H-Ras mutations. Specifically, females exhibited a heightened frequency of H-Ras mutations. Two of the included studies showed a slightly higher mutation in females than males. The predominant mutations identified in the study done by Sathyan et al. (65% being G4A transitions and 20% G4T transversions) have been associated with exposure to carcinogens found in tobacco. This suggests that the elevated occurrence of H-Ras mutations among women could potentially be attributed to the widespread habit of tobacco chewing among them [24]. But in contrast to this Krishna et al. in his study observed a higher H-RAS protein expression in males rather than females (73.1%) [30]. Whereas, Roodi et al. reported no sex-dependent differences in their study [29].
The Road Ahead-Targeting H-Ras
Even a decade ago, RAS inhibitors were extremely difficult to find, to the extent that RAS was labeled as ‘undruggable’. But unlike K-Ras and N-Ras, H-Ras is prenylated exclusively by Farnesyltransferase and therefore its inhibitor (Farnesyltransferase inhibitor—FTIs) could be useful for treating H-Ras-mutant cancers. Also, as evident from the above data significant mutation in the H-Ras gene is observed in individuals diagnosed with oral squamous cell carcinoma, it can be deduced that targeting H-Ras seems promising in oral cancer therapeutics.
Tipifarnib is an FTI that prevents H-Ras binding to the cell membrane. Therefore, recently it has emerged as a promising treatment for H-Ras mutant cancers. Currently, two phase 2 clinical trials are ongoing where Tipifarnib is being used on HNSCC patients [57].
In a single-arm, open-label phase II trial (NCT02383927) by Alan et al. [58] the effectiveness of tipifarnib in mHRAS-related malignancies in 30 patients with recurrent and/or metastatic (R/M) HNSCC was evaluated. The trial considered an ad hoc analysis of the initial 16 HNSCC patients with mHRAS variant allele frequency (VAF) data and also enrolled participants with mHRAS VAF equal to or exceeding 20% (high VAF). The primary endpoint focused on the objective response rate, while secondary endpoints included the evaluation of safety and tolerability. The patients were administered oral doses of tipifarnib at either 600 or 900 mg twice daily on days 1–7 and 15–21 within 28 day cycles. During the administration of tipifarnib, the median progression-free survival was recorded as 5.6 months and the median overall survival (OS) reached 15.4 months [58].
Concurrently, in a recent study, Coleman et al. also observed an increased trend toward improvement in overall survival in cohorts exhibiting oncogenic mutations in H-Ras in HNSCC (3–4%) receiving treatments such as tipifarnib which otherwise had poor clinic outcomes without targeted therapy [59].
The FDA has granted breakthrough therapy approval for Tipifarnib in H-Ras mutant cancer. Future research could explore targeting the α4–α5 interface using a nanobody such as NS1 to disrupt HRAS self-association by binding directly to the α4–α5 interface. This approach aims to decrease the activation of downstream pathways and inhibit cell proliferation while preserving RAS localization and GTPase activity unaffected [60].
Recently, a novel mechanism for RAS auto-inhibition called “membrane occlusion” has emerged as a potential approach to target RAS protein–protein interactions. In this process, RAS forms direct interactions with the lipid membrane, effectively isolating the effector binding interface of RAS from the cytosol. In this process, a tiny molecule called Cmpd2 is used to help occlude the membrane and reduce RAS binding to the RAF’s RAS-binding domain. Cmpd2 can bind to the region where RAS and the lipid membrane interact. Because the RAS-effector interface is substantially conserved, this approach has the potential to successfully block all RAS-initiated downstream signaling pathways [61].
Limitations
This systematic review acknowledges limitations associated with the type and quality of the relevant literature under examination. The qualitative analysis of the literature emphasizes the necessity for standardization in both study design and methods. This standardization is crucial for achieving comparable results, thereby enhancing the growing body of evidence and providing clarity to the existing knowledge. Certainly, the literature exhibits significant heterogeneity, encompassing not only methodological variations but also differences in population characteristics, lifestyle habits, and culture. These diverse factors have the potential to exert influence on the outcomes. Another limitation of this systematic review is that the inclusion criteria restricted the search to studies in the English language, those with accessible full texts, and published studies. Consequently, the potential for selection and publication bias is likely to be present in the findings.
Conclusion
The associated mortality and morbidity of oral cancer have directed for exploration of key genetic changes that can be targeted for therapeutic benefits. Ras genes are vital participants in major cell growth pathways. Therefore, any mutation in the Ras gene can significantly affect the transformation of normal tissues to malignancy. Mutations in the Ras gene commonly reported in hotspots at codons 12, 13, and 61 resulting in the activation of downstream signaling pathways cause abnormal and uncontrolled cell growth. The current targeted therapies though appear promising but newer treatments focussing on specific mutant types can facilitate anticancer therapy targeting H-Ras in OSCC.
Data Availability
No datasets were generated or analysed during the current study.
References
Johnson NW, Jayasekara P, Amarasinghe AA (2000) Squamous cell carcinoma and precursor lesions of the oral cavity: epidemiology and etiology. Periodontol 2011(57):19–37
Chi AC, Day TA, Neville BW (2015) Oral cavity and oropharyngeal squamous cell carcinoma-an update. CA Canc J Clin 65:401–421. https://doi.org/10.3322/caac.21293
Albuquerque R, Lopez-Lopez J, Mari-Roig A, Jane-Salas E, Rosel Lo-Llabres X, Santos JR (2011) Oral tongue squamous cell carcinoma (OT SCC): alcohol and tobacco consumption versus non-consumption. A study in a Portuguese population. Braz Dent J 22:517–521
Moyses RA, López RV, Cury PM, Siqueira SA, Curioni OA, Gois Filho JF et al (2013) Significant differences in demographic, clinical, and pathological features in relation to smoking and alcohol consumption among 1,633 head and neck cancer patients. Clinics (Sao Paulo) 68:738–744
Ribeiro AC, Silva AR, Simonato LE, Salzedas LM, Sundefeld ML, Soubhia AM (2009) Clinical and histopathological analysis of oral squamous cell carcinoma in young people: a descriptive study in Bra zillions. Br J Oral Maxillofac Surg 47:95–98
Argiris A, Eng C (2003) Epidemiology, staging, and screening of head and neck cancer. Cancer Treat Res 114:15–60
Majchrzak E, Szybiak B, Wegner A, Pienkowski P, Pazdrowski J, Luczewski L et al (2014) Oral cavity and oropharyngeal squamous cell carcinoma in young adults: a review of the literature. Radiol Oncol 48:1–10
Toner M, O’Regan EM (2009) Head and neck squamous cell carcinoma in the young: a spectrum or a distinct group? Part 1. Head Neck Pathol 3:246–248
Falaki F, Dalirsani Z, Pakfetrat A, Falaki A, Saghravanian N, Nosratzehi T et al (2011) Clinical and histopathological analysis of oral squamous cell carcinoma of young patients in Mashhad, Iran: a retrospective study and review of literature. Med Oral Patol Oral Cir Bucal 16:e473–e477
Troeltzch M, Knösel T, Eichinger C, Probst F, Troeltzsch M, Woodlock T et al (2014) Clinicopathologic features of oral squamous cell carcinoma: do they vary in different age groups? J Oral Maxillofac Surg 72:1291–1300
Patel SC, Carpenter WR, Tyree S, Couch ME, Weissler M, Hack man T et al (2011) Increasing incidence of oral tongue squamous cell carcinoma in young white women, age 18 to 44 years. J Clin Oncol 29:1488–1494
Kaminagakura E, Villa LL, Andreoli MA, Sobrinho JS, Vartanian JG, Soares FA et al (2012) High-risk human papillomavirus in oral squamous cell carcinoma of young patients. Int J Cancer 130(1726):32
Colevas AD, Yom SS, Pfister DG et al (2018) NCCN guidelines insights: head and neck cancers, version 1.2018. J Natl Compr Canc Netw 16:479–490. https://doi.org/10.6004/jnccn.2018.0026
Krishna A, Singh S, Kumar V et al (2015) Molecular concept in human oral cancer. Natl J Maxillofac Surg 6(1):9–15. https://doi.org/10.4103/0975-5950.168235
Neville BW, Damm DD, Allen CM et al (2009) Oral and maxillofacial pathology. Elsevier, St. Louis
Stransky N, Egloff AM, Tward AD et al (2011) The mutational landscape of head and neck squamous cell carcinoma. Science 333:1157–1160. https://doi.org/10.1126/science.1208130
Parada LF, Tabin CJ, Shih C et al (1982) Bladder carcinoma oncogene is homologue of Harvey sarcoma virus ras gene. Nature 297:474–478. https://doi.org/10.1038/297474a0
Young A, Lou D, McCormick F (2013) Oncogenic and wild-type ras play divergent roles in the regulation of mitogen-activated protein kinase signaling. Cancer Discov 3:112–123
Nakadate Y, Kodera Y, Kitamura Y, Shirasawa S, Tachibana T, Tamura T et al (2014) KRAS mutation confers resistance to antibody-dependent cellular cytotoxicity of cetuximab against human colorectal cancer cells. Int J Cancer 134:2146–2155
Grabocka E, Bar-Sagi D (2016) Mutant KRAS enhances tumor cell fitness by upregulating stress granules. Cell 167:1803–1813
Shields JM, Pruitt K, McFall A, Shaub A, Der CJ (2000) Understanding ras: ‘It ain’t over ‘til it’s over.’ Trends Cell Biol 10:147–154
Krishna A, Singh S, Kumar V, Pal US (2015) Molecular concept in human oral cancer. Natl J Maxillofac Surg 6:9–15
Repasky GA, Chenette EJ, Der CJ (2004) Renewing the conspiracy theory debate: does Raf function alone to mediate ras oncogenesis? Trends Cell Biol 14:639–647. https://doi.org/10.1016/j.tcb.2004.09.014
Sathyan KM, Nalinakumari KR, Kannan S (2007) H-Ras mutation modulates the expression of major cell cycle regulatory proteins and disease prognosis in oral carcinoma. Modern Pathol. https://doi.org/10.1038/modpathol.3800948
Murugan AK, Nguyen TH et al (2009) Detection of two novel mutations and relatively high incidence of H-RAS mutations in Vietnamese oral cancer. Oral Oncol. https://doi.org/10.1016/j.oraloncology.2009.05.638
Koumaki D, Kostakis G, Koumaki V et al (2012) Novel mutations of the HRAS gene and absence of hotspot mutations of the BRAF genes in oral squamous cell carcinoma in a Greek population. Oncol Rep 27:1555–1560. https://doi.org/10.3892/or.2012.1653
Chiosea SI, Grandis JR, Lui VWY et al (2013) PIK3CA, HRAS, and PTEN in human papillomavirus positive oropharyngeal squamous cell carcinoma. BMC Cancer 13:602. https://doi.org/10.1186/1471-2407-13-602
Chang YS, Hsu HT, Ko YC et al (2014) Combined mutational analysis of RAS, BRAF, PIK3CA, and TP53 genes in Taiwanese patients with oral squamous cell carcinoma. Oral Surg Oral Med Oral Pathol Oral Radiol. https://doi.org/10.1016/j.oooo.2014.03.016
Roodi AM, Allameh A, Harirchi I et al (2016) Studies on the contribution of Cox-2 expression in the progression of oral squamous cell carcinoma and H-Ras activation. Pathol Oncol Res. https://doi.org/10.1007/s12253-016-0114-1
Krishna A, Singh S, Singh V et al (2018) Does Harvey-Ras gene expression lead to oral squamous cell carcinoma? A clinicopathological aspect. J Oral Maxillofacial Pathol. 22:1–65. https://doi.org/10.4103/jomfp.JOMFP_246_17
Batta N, Pandey M (2019) Mutational spectrum of tobacco associated oral squamous carcinoma and its therapeutic significance. World J Surg Oncol 17:198. https://doi.org/10.1186/s12957-019-1741-2
Uchibori M, Osawa Y, Ishii Y et al (2021) Analysis of HRAS mutations in Japanese patients with oral squamous cell carcinoma. Adv Oral Maxillofacial Surg. 1:100021. https://doi.org/10.1016/j.adoms.2021.100021
Molinolo AA, Amornphimoltham P, Squarize CH, Castilho RM, Patel V, Gutkind SJ (2009) Dysregulated molecular networks in head and neck carcinogenesis. Oral Oncol 45(4–5):324–334
Shaw RJ, Cantley LC (2006) Ras, PI(3)K and mTOR signaling controls tumor cell growth. Nature 441(7092):424–430
Chang F, Steelman LS, Shelton JG et al (2003) Regulation of cell cycle progression and apoptosis by the Ras/Raf/MEK/ERK pathway (Review). Int J Oncol 22:469–480. https://doi.org/10.3892/ijo.22.3.469
Barbacid M (1987) RAS genes. Annu Rev Biochem 56:779–827. https://doi.org/10.1146/annurev.bi.56.070187.004023
Paterson IC, Eveson JW, Prime SS (1996) Molecular changes in oral cancer may reflect aetiology and ethnic origin. Eur J Cancer B Oral oncol 32B(3):150–153
Lyu H, Li M, Jiang Z et al (2019) Correlate the TP53 mutation and the HRAS mutation with immune signatures in head and neck squamous cell cancer. Comput Struct Biotechnol J 17:1020–1030. https://doi.org/10.1016/j.csbj.2019.07.009
Shaw RJ, Cantley LC (2006) Ras, PI(3)K and mTOR signalling controls tumor cell growth. Nature 441:424–430. https://doi.org/10.1038/nature04869
Muñoz-Maldonado C, Zimmer Y, Medová M (2019) A comparative analysis of individual RAS mutations in cancer biology. Front Oncol 9:1088. https://doi.org/10.3389/fonc.2019.01088
Terrell EM, Morrison DK (2019) Ras-mediated activation of the raf family kinases. Cold Spring Harb Perspect Med 9:033746. https://doi.org/10.1101/cshperspect.a033746
Das N, Majumder J, DasGupta UB (2000) Ras gene mutations in oral cancer in eastern India. Oral Oncol 36:76–80. https://doi.org/10.1016/S1368-8375(99)00058-5
Munirajan AK, Mohanprasad BK, Shanmugam G et al (1998) Detection of a rare point mutation at codon 59 and relatively high incidence of H-ras mutation in Indian oral cancer. Int J Oncol 13:971–974. https://doi.org/10.3892/ijo.13.5.971
Saranath D, Chang SE, Bhoite LT et al (1991) High frequency mutation in codons 12 and 61 of H-ras oncogene in chewing tobacco-related human oral carcinoma in India. Br J Cancer 63:573–578. https://doi.org/10.1038/bjc.1991.133
Sanchez-Montenegro C, Vilanova-Sanchez A, Barrena-Delfa S et al (2017) Costello syndrome and umbilical ligament rhabdomyosarcoma in two pediatric patients: case reports and review of the literature. Case Rep Genet 2017:1587610. https://doi.org/10.1155/2017/1587610
The Catalogue of Somatic Mutations in Cancer. https://scholar.google.com/scholar?q=intitle:The%20Catalogue%20of%20Somatic%20Mutations%20in%20Cancer
Cutilli T, Papola F, Di Emidio P, Corbacelli A (1998) P53 tumor suppressor protein and H-RAS oncogene in maxillofacial tumors: Immunohistochemical and genetic investigation, induction chemotherapy response and prognosis evaluation. J Chemother 10:411–417. https://doi.org/10.1179/joc.1998.10.5.411
McDonald JS, Jones H, Pavelic ZP et al (1994) Immunohistochemical detection of the H-ras, K-ras, and N-ras oncogenes in squamous cell carcinoma of the head and neck. J Oral Pathol Med 23:342–346. https://doi.org/10.1111/j.1600-0714.1994.tb00073.x
Winston JT, Coats SR, Wang YZ et al (1996) Regulation of the cell cycle machinery by oncogenic ras. Oncogene 12:127–134
Jayasurya R, Sathyan KM, Lakshminarayanan K et al (2005) Phenotypic alterations in Rb pathway have more prognostic influence than p53 pathway proteins in oral carcinoma. Mod Pathol 18:1056–1066. https://doi.org/10.1038/modpathol.3800387
Williams JP, Stewart T, Li B et al (2006) The retinoblastoma protein is required for Ras-induced oncogenic transformation. Mol Cell Biol 26:1170–1182. https://doi.org/10.1128/MCB.26.4.1170-1182.2006
Lee KW, Kim MS, Kang NJ et al (2006) H-ras selectively up-regulates MMP-9 and COX-2 through activation of ERK1/2 and NF-kappaB: an implication for invasive phenotype in rat liver epithelial cells. Int J Cancer 119:1775. https://doi.org/10.1002/ijc.22056
Zarbl H, Sukumar S, Arthur AV, Martin-Zanca D, Barbacid M (1985) Direct mutagenesis of of Ha-ras-1 oncogenes by N-nitroso-N-methylurea during initiation of mammary carcinogenesis in rats. Nature 315(6018):382–385
Quintanilla M, Brown K, Ramsden M, Balmain A (1986) Carcinogen specific mutation and amplification of Ha-ras during mouse skin carcinogenesis. Nature 322(6074):78–80
Chang KW, Sarraj S, Lin SC et al (2000) P53 expression, p53 and Ha-ras mutation and telomerase activation during nitrosamine-mediated hamster pouch carcinogenesis. Carcinogenesis 21:1441–1451. https://doi.org/10.1093/carcin/21.5.441
Xu J, Gimenez-Conti IB, Cunningham JE et al (1998) Alterations of p53, cyclin D1, Rb, and H-ras in human oral carcinomas related to tobacco use. Cancer 1998:204–212
Squamous cell carcinoma of head and neck. https://clinicaltrials.gov/
Ho AL, Brana I, Haddad R et al (2021) Tipifarnib in head and neck squamous cell carcinoma with HRAS mutations. J Clin Oncol 10:1856–1864. https://doi.org/10.1200/JCO.20.02903
Coleman N, Marcelo KL, Hopkins JF et al (2023) HRAS mutations define a distinct subgroup in head and neck squamous cell carcinoma. JCO Precision Oncol. https://doi.org/10.1200/PO.22.00211
Khan I, Spencer-Smith R, O’Bryan JP et al (2018) Targeting the α4-α5 dimerization interface of K-RAS inhibits tumor formation in vivo. Oncogene 38:2984–2993. https://doi.org/10.1038/s41388-018-0636-y
Fang Z, Marshall CB, Nishikawa T et al (2018) Inhibition of K-RAS4B by a unique mechanism of action: stabilizing membrane-dependent occlusion of the effector-binding site. Cell Chem Biol 25:1327–1336. https://doi.org/10.1016/j.chembiol.2018.07.009
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Contributions
PD Conceptualization, Methodology, Writing—Original Draft, Formal analysis. RD Writing—Review and Editing, Visualization. RS Review and Editing. AJ Review and Editing, Visualization. SG Supervision, Visualization Shalini Gupta- Supervision, Visualization.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical Approval
This type of study does not require ethical approval.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Devi, P., Dwivedi, R., Sankar, R. et al. Unraveling the Genetic Web: H-Ras Expression and Mutation in Oral Squamous Cell Carcinoma—A Systematic Review. Head and Neck Pathol 18, 21 (2024). https://doi.org/10.1007/s12105-024-01623-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12105-024-01623-8