Abstract
Background
This study is focused on the identification of gene mutations in H-ras which are probably associated with tumor recurrence in oral squamous cell carcinoma (OSCC) following conventional therapy.
Methods
Surgically removed biopsies from OSCC patients without recurrence (n = 43) and biopsies from recurrent cases (n = 19) were analyzed. Also, gingival tissues (n = 5) from normal individuals were processed and considered as control. DNA was extracted and amplified using primers for exons 1 and 2 for the H-ras gene, and then DNA products were analyzed using Sanger’s sequencing technique. Besides, H-ras expression was compared in samples by immunostaining (IHC), using anti-ras antibody.
Results
Demographic data show that smoking habit in patients and recurrent tumors was ~ 44.1 and 78%, respectively. The major site of malignancy was tongue tissue (40–60%). The rate of pathological stage III/IV were 41.8 and 100% in primary tumors and recurrence malignancy respectively. The sequencing data showed that a specific mutation in H-ras gene, Gly12Ala (G6266A) in recurrence samples and primary cases was detected in ~ 66.6% and 10% respectively. Accumulation of H-ras protein in tissues was relatively high scores (> 5) in both primary and recurrence tumors. The H-ras mutation detected was associated with increased level of H-ras protein accumulated in the malignant cells (IHC data).
Conclusion
These data may suggest that regardless of the causes and factors involved, Gly12Ala (G6266A) is associated with recurrence in high-grade OSCC tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
According to the World Health Organization (WHO) statistics, the prevalence of oral cavity cancer is relatively high in the developing countries [1]. Accordingly, the incidence of oral cancer is increasing in the Iranian population and ranks as the 10th most prevalent cancer among other cancers [2]. Head and neck carcinoma is a group of heterogeneous disease that involve different regions in the oral cavity with different histological characteristics, including paranasal sinuses and salivary glands [3]. Evidences show that the etiological factors of oral cancers may differ in populations living in different geographical regions. Environmental factors as well as lifestyle are considered as major factors contributing to such differences in the incidence of oral cancer [4]. For instance, cigarette smoking could be a common risk factor in different populations [5], but the contribution of human papilloma virus (HPV) infection as a risk factor may differ in different regions [6]. Studies carried out on Iranian patients diagnosed with OSCC show that this malignancy is associated with smoking but not with at least types 16 and 18 of HPV [7]. However, evidence show that HPV infection is probably a major risk factor for oral cancer in the European population [8, 9].
To our knowledge, there are no particular molecular and genetic markers for diagnosis and monitoring treatments for oral SCC. However, alterations in oncogenes and tumor suppressor genes, including changes in their protein products, have been reported [10]. Genetic mutations in TP53, PIK3CA, NOTCH1 and CDKN2 genes [11] are the most frequently reported from different regions [12]. Accordingly, over-expression of FGFR1/FGFR3/EGFR/H-RAS PTEN/CKND2a and myc genes which are linked to epigenetic changes, particularly methylation in promoter region of p16 and TGM-3 genes have been assigned to progression of oral SCC [13].
H-ras is among the oncogenes that most frequently undergo genetic mutations in OSCC [14]. H-ras gene belongs to the ras gene family (H-, K- and N-ras), which plays important role in development of tumors [15]. A point mutations (missense) in codons 12 and 13 of exon 2 of this gene can lead to decreased GTPase activity of H-ras protein, causing constitutive activation of this protein [16, 17]. This mutation can lead to abnormal cell proliferation and progression [18].
Genetic analysis of H-ras is probably more relevant to the initiation and progression of oral SCC. The missense mutations in codons 12,13, and 61 have been reported in oral cancer in patients in India [16], and in the USA [19]. The type of gene mutations in oral cancer patients has also been identified, mainly as point mutations in FGFR exons [20]. The conventional protocols used for oral cancer treatment in Iran are mainly surgery, radiotherapy, and chemotherapy [21]. However, recurrence in some of the patients is an emerging issue [22, 23]. During the period of sample collection for analysis of genetic and molecular markers in Iranian patients, we noticed that a considerable number of patients suffered from cancer recurrence. Earlier we reported that the H-ras expression at mRNA levels is substantially higher in OSCC samples, particularly in moderately differentiated (MD)/ poorly differentiated (PD) tumors compared to normal gingival [24] Based on this information, the present study was carried out to examine the possible impact of H-ras (exon-1 and exon-2) gene mutations on recurrence of this type of malignancy in Iranian patients.
Materials and Methods
Patients and Collection of Biopsies
This cross-sectional study was conducted at the Cancer Institute, Imam Khomeini Hospital, Tehran, Iran from July 2019 to August 2020. Sixty-two patients diagnosed as cases of OSCC were enrolled in the study, among these patients, nineteen recurrent cases were identified. All the recurrent patients were receiving both chemotherapy and radiotherapy according to the standard cancer therapy guidelines [25, 26]. The treatment protocol used for management of almost all these patients is surgery combined with radiotherapy and chemotherapy (cisplatin and cetuximab). The patients were selected based on a questioner.
Surgically removed tumor biopsies were collected from OSCC patients without recurrence (n = 43) and tumor recurrent cases (n = 19). Also, gingival tissue biopsies from a group of age and sex-matched apparently healthy candidates (n = 5), were collected and processed and considered as control. Representative images from the site of the tumors as well as histological examinations (H&E staining) are presented in Fig. 1.
A written informed consent was obtained from each patient before undergoing surgery for the use of their clinical data for research purposes. This study was conducted in accordance with the Helsinki declaration and was approved by the Medical Ethics Committee of the Cancer Institute of Imam Khomeini Hospital, Tehran University of Medical Sciences (ID99-10-21-400). Surgically removed tumor biopsies were collected from each patient and cut into small pieces. A small piece of the tissue was fixed in formalin solution (10%) and sent to the pathology department for routine sectioning, staining, IHC and histopatholgical examination for determination of tumor grade and stage. Another portion was transferred to microtubes and immediately transferred to a deep freezer (-80 °C) until homogenization and DNA extraction.
DNA Extraction
DNA was extracted from tumor biopsies using the QIAamp DNA FFPE Tissue Kit (Cat. No.56,404, Qiagen, Germany). The quality of each DNA sample was determined using a NanoDrop (2000 C Spectrophotometer, Thermo Scientific, Wilmington, DE, USA. DNA samples with an absorption ratio (260 nm/280 nm) greater than 1.92 was considered acceptable and the sample was processed for further analysis. The quality of DNA samples was further checked by electrophoresis analysis on a 1% agarose gel.
PCR Analysis
For detection of any possible mutation in exon − 1 and exon-2 of the H-ras gene, the gene was amplified using specific pair of primers as shown below:
Exon − 1.
Forward: 5´GCCCTGCTCGGAGATGC3´.
Reverse: 5´ GGACCGTGCCCAGCG3´.
Exon − 2.
Forward: 5´TGTGGGTTTGCCCTTCAG 3´.
Reverse: 5´ ATGAGGAAGCAGGAGACAG3´.
The reaction mixture consisted of 10 µl master mix (2X PCR Master Mix Red, MgCl2, 1.5mM- 1.25 ml, Ampliqon, Cat.No.180301-50), 0.5 µl of each of the forward primer (µΜ), 0.5 µl reverse primer (0.5 µl), and 1 µl DNA template. DNA was amplified for 40 cycles at 95 °C for 30 s, 61 °C for 30 s, and 72 °C for 35 s, followed by 10 minutes’ final extension at 72 °C.
The DNA product was analyzed on agarose gel electrophoresis (1%). A DNA ladder of 100 bp (GeneAll, Cat. No. GA-010, Korea) stained with Gel Red® (Biotium, Cat. No. 41,003, USA) was also analyzed as a standard.
Direct DNA Sequencing
The PCR products initially prepared from DNA samples (n = 62) together with the primer samples used in the initial PCR were processed for direct sequencing. Sequencing was carried out on a Chromas Pro sequencing machine (Technelysium, South Brisbane, Australia, version 2.6.6), and the nucleotide blast program of NCBI. The sequencing data was aligned with the reference sequence (NG_007666.1) available in the GenBank database (www.ncbi.nlm.nih.gov/genbank). Furthermore, the clinical significance of mutations detected was characterized by using the ClinVar database (www.ncbi.nlm.nih.gov/clinvar/).
Immunohistochemistry (IHC) for H-ras
For this assay, paraffin-embedded tissue blocks were cut into 2 μm sections. Sections were deparaffinized and dewaxed in xylene, rehydrated in descending alcohol concentrations. From each sample, at least 3 sections were prepared, and then the sections were placed in citrate buffer (pH 6.0) for antigen retrieval. Then endogenous peroxidase activity was blocked by incubation with hydrogen peroxide for 10 min. The slides were incubated with primary antibody (Mouse monoclonal antibody against the recombinant H-Ras protein, Cat.No.:sc-29, from the Santa Cruz Biotechnology Inc) for 60 min at room temperature.
The sections were placed in secondary antibody (Cat. # M1U539 G, L10BIO C ARE Medical, USA) solution for 15 min, in MACH1 polymer for 15 min, and in DAB staining (3,3’-Diaminobenzidine) for 15 min. The sections were then counter stained with hematoxylin, dehydrated, cleared, and mounted.
The primary antibody for detection of H-ras accumulation in the tissues was optimized at a concentration of 1–2 µg/100–500 µg of total protein (1:50 dilution). IHC scoring was carried out by two independent pathologists, and H-Ras protein expression was assessed by giving scoring numbers of 0 to 8 according to proportion score and intensity.
Scoring System Used for IHC
Semi-quantitative analysis of H-ras-stained tissue sections was performed based on the Allred scoring system guidelines (44). To achieve the final scores, individual scores of the percentage of H-ras-positive cancer cells (0–5) and the staining intensity of the cytoplasmic H-ras (0–3) (Fig. 1) were summed up. The percentage of H-ras-positive cancer cells was set as follows: 1—less than 1% of positive cancer cell; 2—from 1 to 10% of positive cancer cell; 3—from 11 to 33% of positive cancer cell; 4—from 34 to 66% of positive cancer cell; and score 5—more than 67% of positive cancer cell. Whereas, the staining intensity in the cytoplasm was scored as: 1—weak; 2—medium; and 3—strong [27]. The addend combinations (Proportion score + Intensity score) for the Allred scoring system. The scoring system used for comparing the accumulation of H-ras protein in the cells is based on the intensity and classified as follows; Score 0 for negative, scores 2–3 for weak, scores 4–6 for moderate, and scores 7-8, for strong.
Results
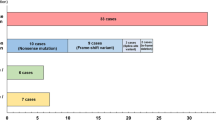
The results presented here are confined to the oral SCC cancer samples (n = 62) referred to the hospital. As shown in Table 1, the majority of the cases (63.1%) had tumors in tongue tissue, four cases (21.0%) had tumors in buccal mucosa, and three cases (15.7%) had tumors in the hard palate. The pathological stage of tumors in the recurrence cases was diagnosed as stage III (n = 6, 31.5%) and stage IV (n = 13, 68.4%) on the basis of the pTNM staging system. Studies on the pathological grade of tumors revealed that in 12 cases (63%), tumor cells were moderately differentiated and in 7 cases (36.8%) were poorly differentiated.
Identification of Mutations (Direct Sequencing)
The data of the sequencing of exon 2 of the H-Ras gene (NG_007666.1) in the recurrence cases (n = 19) as well as normal controls (n = 5) is presented in Fig. 2.
Mutations detected in exons 2 H-ras gene of samples from recurrent OSCC patients
(A) H-ras exon 2 Gly12Val. The arrow indicates the substitution of a guanine by a thymine. (B) H-ras exon 2 Gly13Asp and Gly12Ala. The arrow indicates the two substitutions 1: A guanine by a cytosine 2: guanine by an adenine. (C) H-ras exon 2 Gly12Ala. The arrow indicates the substitution of a guanine by an adenine. (D) H-ras exon 2 “Silent mutation” the arrow indicates the substitution of a cytosine by a thymine. The sequence chromatogram fragment of the H-ras gene as determined by using the classic Sanger’s method
The prevalence of the gene mutation found in most of the recurrence cases was a typical point mutation, considered as a missense change. The G12A and G12V mutations were identified in 17 samples (89.4%) obtained from recurrence patients (Table 2). Five of the samples (OSCC-A-2, OSCC-A-37, OSCC-A-15, OSCC-A-10 and OSCC-A-12) had mutations in amino acid codon 12 of exon 2 of the H-Ras gene that were G→T transversion (Glycine to Valine) (Fig. 2 A and Table 3). As an exceptional case, two samples registered under the code; OSCC-A-3 and OSCC-A-45, had two mutations that refer to codon 6263 where Guanine was substituted by a Cytosine (transversion) and codon 6266 Guanine was substituted by an Adenine (transition). In these samples, there were two substitutions, Glycine to Alanine and Glycine to Aspartate, in codons 12 and 13 respectively, were observed (Fig. 2B; Table 3). Twelve samples had missense mutations in codon 12 of exon 2 of H-Ras. The translational outcome of the G →A transition was a substitution of a Glycine with an Alanine (Fig. 2 C and Table 3).
IHC Data for H-ras Protein Detection
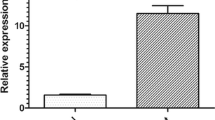
IHC data presented in Fig. 1, shows that H-ras oncoprotein is mainly localized in the cytoplasmic region in tumors obtained from OSCC cases. According to the scoring system used in this study, the majority of samples (15/19) showed relatively high scores (Score > 5) (Table 3), which is significantly higher compared to normal biopsies (p < 0.05). Of note, the control samples are not included in Table 3. However, all the control samples have the sum of score less than 5 (Fig. 3).
Discussion
In this paper, we report a missense mutation in exon 2 of the H-ras gene which could be linked to the recurrence of oral carcinoma in Iranian patients after undergoing conventional treatments. During our study, we encountered individuals 19/62 (30.6%) with cancer recurrence within 6 months of treatment. It was interesting to see if there were differences in gene mutations. Hence, the gene mutations in H-ras (exons 2) were compared in cancer patients with those who experienced a recurrence of malignancy (Table 2).
Such differences could be due to several factors which are responsible for oral cancer causation and/or due to the response to treatments by some individuals. Oral carcinoma is a multi-factorial malignancy and the incidence and progression highly depends on environmental and lifestyle factors. Therefore, the incidence of oral cancer can differ in different ethnic groups living in different geographic regions. Iran is a unique society with special dietary habits and lifestyle. In recent years the incidence of oral cancer has been increasing in younger individuals (< 45 years), with unknown etiology [28]. Earlier, we reported that this type of cancer is not associated with HPV infection, but most likely tobacco smoking, which was found to be associated with the incidence and development of this type of cancer [7]. Our study on Iranian patients with oral carcinoma is focused on the expression of molecular markers often considered for diagnosis such as H-ras, p16 and Cox-2 genes [7, 24]. The H-ras mutation detected was corroborated by the increased level of H-ras protein accumulated in the malignant cells as shown in IHC data. The accumulation of H-ras protein in tumor tissues was high (Score > 5) in both primary tumors and recurrence tumors (Fig. 1). This finding may suggest that gene mutations in exons 2 can lead to overexpression of H-ras protein as a consequence of which abnormal proliferation and progression of tumor can occur. Ras protein activate signaling pathways and networks controlling cell proliferation, differentiation and survival [18]. Although H-ras mutations which render the protein constitutively active are common in different types of cancers, there is a distinctive pattern of mutation frequencies of H-ras mutations in each cancer type [29]. The missense mutation in codons 12,13 and 61 has been reported in oral cancer in patients in India [16, 17], USA[19], Japan [30] and China [31] with distinct patterns and etiological factors.
The importance of the present study is the detection of a considerably high frequency (> 60%) of H-ras G6266A mutation (Gly12 Ala) in oral SCC recurrence cases. Moreover, the evidence shows that patients with recurrent oral SCC show higher pathological stages (III and IV) of tumors compared to patients with primary tumors. Substitution of Glycine at the 12th amino acid position by Alanine or Valine was the most common mutation observed. Since Glycine plays an important role in secondary structure of proteins, it is assumed that this causes a disruption in protein structure and function.
There are different factors contributing to the recurrence of OSCC, among which the treatment protocols in different countries and regions can be considered important [21]. The limited information provided in this paper may suggest that probably smoking habit is another important issue that may contribute to the recurrence of oral SCC, as it was shown that the number of smoker cases suffering with tumor recurrence is greater as compared to those suffering from primary malignancy. This data is implicated in the diagnosis of oral SCC as well as the prediction of patients vulnerable to recurrence of oral SCC. H-ras gene mutations linked to oral SCC can help to improve treatment protocols. This information, although important, needs to be verified with a larger sample size. However, no sufficient evidence is available to prove the relationship of H-ras mutations with the treatment protocols applied to these patients.
Data Availability
All primary data are available on request.
Code Availability
Software application used for the study are available on request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J Clin. 2021;71(3):209–49.
Maleki D, Ghojazadeh M, Mahmoudi S-S, Mahmoudi S-M, Pournaghi-Azar F, Torab A, et al. Epidemiology of oral cancer in Iran: a systematic review. Asian Pac J Cancer Prev. 2015;16(13):5427–32.
Ehtesham H, Safdari R, Mansourian A, Tahmasebian S, Mohammadzadeh N, Pourshahidi S. Developing a new intelligent system for the diagnosis of oral medicine with case-based reasoning approach. Oral Dis. 2019;25(6):1555–63.
Kang H, Kiess A, Chung CH. Emerging biomarkers in head and neck cancer in the era of genomics. Nat reviews Clin Oncol. 2015;12(1):11.
Lee SU, Moon SH, Choi SW, Cho KH, Park JY, Jung YS, et al. Prognostic significance of smoking and alcohol history in young age oral cavity cancer. Oral Dis. 2020;26(7):1440–8.
Wittekindt C, Wagner S, Sharma SJ, Würdemann N, Knuth J, Reder H, et al. Referateband: HPV–A different view on Head and Neck Cancer. Laryngo-rhino-otologie. 2018;97(Suppl 1):S48.
Allameh A, Moazeni-Roodi A, Harirchi I, Ravanshad M, Motiee-Langroudi M, Garajei A, et al. Promoter DNA methylation and mRNA expression level of p16 gene in oral squamous cell carcinoma: correlation with clinicopathological characteristics. Pathol Oncol Res. 2019;25(4):1535–43.
Foy J-P, Bertolus C, Boutolleau D, Agut H, Gessain A, Herceg Z, et al. Arguments to support a viral origin of oral squamous cell carcinoma in non-smoker and non-drinker patients. Front Oncol. 2020;10:822.
Dalla Torre D, Burtscher D, Soelder E, Offermanns V, Rasse M, Puelacher W. Human papillomavirus prevalence in a Mid-European oral squamous cell cancer population: A cohort study. Oral Dis. 2018;24(6):948–56.
Dubot C, Bernard V, Sablin M, Vacher S, Chemlali W, Schnitzler A, et al. Comprehensive genomic profiling of head and neck squamous cell carcinoma reveals FGFR1 amplifications and tumour genomic alterations burden as prognostic biomarkers of survival. Eur J Cancer. 2018;91:47–55.
Zhang H, Song Y, Du Z, Li X, Zhang J, Chen S, et al. Exome sequencing identifies new somatic alterations and mutation patterns of tongue squamous cell carcinoma in a Chinese population. J Pathol. 2020;251(4):353–64.
Nakagaki T, Tamura M, Kobashi K, Omori A, Koyama R, Idogawa M, et al. Targeted next-generation sequencing of 50 cancer-related genes in Japanese patients with oral squamous cell carcinoma. Tumor Biology. 2018;40(9):1010428318800180.
Shojaeian S, Moazeni-Roodi A, Allameh A, Garajei A, Kazemnejad A, Kabir K, et al. Methylation of TGM-3 promoter and its association with oral squamous cell carcinoma (OSCC). Avicenna J Med Biotechnol. 2021;13(2):65.
Koumaki D, Kostakis G, Koumaki V, Papadogeorgakis N, Makris M, Katoulis A, et al. Novel mutations of the HRAS gene and absence of hotspot mutations of the BRAF genes in oral squamous cell carcinoma in a Greek population. Oncol Rep. 2012;27(5):1555–60.
Murugan AK, Munirajan AK, Tsuchida N. Ras oncogenes in oral cancer: the past 20 years. Oral Oncol. 2012;48(5):383–92.
Jayaprakash C, Varghese VK, Jayaram P, Chakrabarty S, Kudva A, Ray S, et al. Relevance and actionable mutational spectrum in oral squamous cell carcinoma. J Oral Pathol Med. 2020;49(5):427–34.
Patel K, Bhat FA, Patil S, Routray S, Mohanty N, Nair B, et al. Whole-exome sequencing analysis of oral squamous cell carcinoma delineated by tobacco usage habits. Frontiers in oncology. 2021;11.
Prior IA, Hood FE, Hartley JL. The frequency of Ras mutations in cancer. Cancer Res. 2020;80(14):2969–74.
Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011;333(6046):1157–60.
von Mässenhausen A, Deng M, Billig H, Queisser A, Vogel W, Kristiansen G, et al. Evaluation of FGFR3 as a therapeutic target in head and neck squamous cell carcinoma. Target Oncol. 2016;11(5):631–42.
Hasegawa T, Kobayashi E, Amano R, Saito I, Takeda D, Kakei Y, et al. Time to recurrence associated with poor prognosis in Japanese oral squamous cell carcinoma patients. Journal of Maxillofacial and Oral Surgery. 2021:1–9.
Mohtasham N, Ghazi N, Anvari K, Mohajertehran F, Organji T, Shahabinejad M. Evaluation of the relationship between the invasive front of oral squamous cell carcinoma and clinicopathological parameters. Iran J Pathol. 2021;16(3):316.
Sadri D, Zendedel K, Harirchi I, Movahedifar Z. Survival from oral cancer in Tehran (Iran). Oral Surgery, Oral Medicine. Oral Pathol Oral Radiol. 2015;119(3):e115.
Moazeni-Roodi A, Allameh A, Harirchi I, Motiee-Langroudi M, Garajei A. Studies on the contribution of cox-2 expression in the progression of oral squamous cell carcinoma and H-ras activation. Pathol Oncol Res. 2017;23(2):355–60.
Brunicardi F, Andersen D, Billiar T, Dunn D, Hunter J, Matthews J, et al. Schwartz’s principles of surgery, 10e. McGraw-hill; 2014.
Chen F, Lin L, Liu F, Yan L, Qiu Y, Wang J, et al. Three prognostic indexes as predictors of response to adjuvant chemoradiotherapy in patients with oral squamous cell carcinoma after radical surgery: A large-scale prospective study. Head Neck. 2019;41(2):301–8.
Ilić IR, Stojanović NM, Radulović NS, Živković VV, Randjelović PJ, Petrović AS, et al. The Quantitative ER immunohistochemical analysis in breast cancer: detecting the 3 + 0, 4 + 0, and 5 + 0 Allred score cases. Medicina. 2019;55(8):461.
Tadbir AA, Ebrahimi H, Pourshahidi S, Zeraatkar M. Evaluation of levels of knowledge about etiology and symptoms of oral cancer in southern Iran. Asian Pac J Cancer Prev. 2013;14(4):2217–20.
Kompier LC, Lurkin I, van der Aa MN, van Rhijn BW, van der Kwast TH, Zwarthoff EC. FGFR3, HRAS, KRAS, NRAS and PIK3CA mutations in bladder cancer and their potential as biomarkers for surveillance and therapy. PLoS ONE. 2010;5(11):e13821.
Uchibori M, Osawa Y, Ishii Y, Aoki T, Ota Y, Kimura M. Analysis of HRAS mutations in Japanese patients with oral squamous cell carcinoma. Adv Oral Maxillofacial Surg. 2021;1:100021.
Chang Y-S, Yeh K-T, Hsu NC, Lin S-H, Chang T-J, Chang J-G. Detection of N-, H-, and KRAS codons 12, 13, and 61 mutations with universal RAS primer multiplex PCR and N-, H-, and KRAS-specific primer extension. Clin Biochem. 2010;43(3):296–301.
Acknowledgements
The authors are grateful to all the participants and the staff members of the referral, especially Dr. Ahad Mohammadnejad, the Cancer Research Center, Cancer Institute of Iran, Tehran University of Medical Sciences, Tehran, Iran. The technical assistance in DNA sequencing provided by Dr. Dariush Hamedi, the Behsotun Laboratory is also acknowledged.
Funding
This study was financially supported by Cancer Research Center, Cancer Institute of Iran, Tehran University of Medical Sciences, Tehran, Iran. (Grant#98-3-115-45155). This study was also supported by a grant (Research project #4003715) provided by the Iran National Science Foundation (INSF).
Author information
Authors and Affiliations
Contributions
Azin Hamidavi Asl designed and performed the experiments, Abdolamir Allameh wrote the manuscript, supervised and directed the project. Mohammad Shirkhoda contributed to sample preparation and supported the project. Hana Saffar assisted with IHC technique.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics Approval
This study was approved by the Medical Ethics Committee of the Cancer Institute of Imam Khomeini hospital, Tehran University of Medical Sciences (ID99-10-21-400).
Consent to Participate
Informed consent was obtained from all individuals participated in this study.
Consent for Publication
Written consent from the patient.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hamidavi Asl, A., Shirkhoda, M., Saffar, H. et al. Analysis of H-ras Mutations and Immunohistochemistry in Recurrence Cases of High-Grade Oral Squamous Cell Carcinoma. Head and Neck Pathol 17, 347–354 (2023). https://doi.org/10.1007/s12105-022-01491-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12105-022-01491-0