Abstract
Background and aims
Researches have shown that miRNAs have been proposed as novel diagnostic biomarkers for classification and prognostic stratification of HCC. However, whether or not miR-431 contributes to the progression of HCC remains unknown. Therefore, we aimed to investigate the clinicopathological significance of miR-431 in HCC.
Methods
MiR-431 expression in 95 HCC cases and corresponding adjacent non-cancerous tissues was evaluated by quantitative reverse transcription polymerase chain reaction (qRT-PCR). Furthermore, statistical analysis was performed to identify the correlations between expression of miR-431 and a variety of clinicopathological parameters and patient recurrence. The area under the receiver operating characteristic curve (AUC) was used to evaluate the accuracy of miR-431 as a biomarker for HCC diagnosis and prediction of disease deterioration.
Results
MiR-431 was markedly down-regulated in the HCC samples (1.1885 ± 0.75867) compared with corresponding adjacent tumor tissues (1.7957 ± 0.89333, P < 0.001). The AUC of low miR-431 expression to diagnose HCC was 0.668 (95 % CI 0.592–0.744, P < 0.001). MiR-431 down-expression was correlated with multiple malignant characteristics, including lymph node metastasis (r = −0.455, P < 0.001), clinical TNM stage (r = −0.223, P = 0.030), MTDH (r = −0.292, P = 0.006), vaso-invasion (r = −0.204, P = 0.047), MVD (r = −0.281, P = 0.006) and HCV (r = 0.215, P = 0.037). Additionally, the recurrent time of lower miR-431 expression group was 56.602 ± 3.914 months, much longer than that in the high expression group (50.009 ± 2.731 months), however, no significant difference was noted (χ 2 = 0.005, P = 0.943).
Conclusions
The down-expression of miR-431 is partially responsible for a series of clinicopathological features which may be tightly correlated with the progression of HCC. Thus, expression of miR-431 may be proposed as a new factor in association with the progression of HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the fifth most common type of malignant tumor and ranks as the third leading cause of cancer-related mortality rate in the world, with estimated 600,000 new cases per annum [1]. HCC is a complicated disease involving numerous factors, and patients with cirrhosis, hepatitis B or C virus may be at high risk of HCC [2, 3]. Although surgical resection is recognized as the first priority for HCC treatment, the frequent postsurgical recurrence is still an obstacle needed to be overcome urgently. Besides, the general response of HCC patients to chemotherapy remains far from satisfaction [4]. Early diagnosis of HCC and effective treatment are likely to prolong the survival time of liver cancer patients [5]. Therefore, discovering new biomarkers that identify patients at high risk for recurrence and combining other postsurgical adjuvant therapies to obtain better outcomes would be applied to improve their survival.
MicroRNAs (miRNAs) comprise a family of small, endogenous, noncoding functional RNA molecules of 18–25 nucleotides in length. They are responsible for post-transcriptional regulation and participate in nearly all biological processes [6]. MiRNAs regulate gene expression by binding to the 3′-untranslated regions (3′-UTRs) of target messenger RNAs and decrease or stop their translation to proteins [7]. Numerous studies have revealed differences in the expression of various miRNAs between tumors and normal tissues [8], suggesting that miRNAs can function as either tumor suppressors or oncogenes in human cancers. In recent years, the therapeutic potential of miRNAs in HCC has been reported in various studies [9–11].
Previous studies have proposed that in RSa cells (a human embryo-derived fibroblasts line) which were highly sensitive to the ability of HuIFN-β (human interferon-β) to inhibit cell viability, HuIFN-β-induced miR-431 expression may down-regulate insulin-like growth factor 1 receptor and insulin receptor substrate 2 (putative miR-431 target genes) expression, and consequently inhibit cell proliferation by suppressing the MAPK pathway [12]. Another research also demonstrated that miR-431 could bind to versican 3′-UTR [13]. It also reported that transgenic mice expressing the versican 3′-UTR developed HCC and increased expression of versican isoforms V0 and V1, which play important roles in HCC development. As little is known accurately about the functions of miR-431 in HCC patients, herein we constructed a series of clinicopathological data of 95 clinical samples to quantify the expression of miR-431 in HCC tissues and adjacent tissues to identify the relationship between miR-431 and clinicopathological features.
Materials and methods
Patients and samples
In this prospective study, a total of 95 HCC samples (with a mean age of 52 years old, ranged from 29 to 82 years old and 70 included in the follow-up) and corresponding adjacent tissues were recruited from March, 2010 to December, 2011 at the First Affiliated Hospital of the Guangxi Medical University (Nanning, Guangxi, China). The non-tumorous tissues were obtained from a section of the resected specimen at the farthest distance from tumor (>2 cm from tumor). Of the 70 HCC patients included in the follow-up, 59 had recurrent tumors and 11 were dead or censored at the end of the follow-up. The median time of follow-up was 32.78 months (range 2.68–68.00 months). The clinicopathological characteristics (including age, tumor size, serum level of AFP and other information) were presented in Table 1. The study was approved by the Research Ethics Committee of First Affiliated Hospital, Guangxi Medical University, China in accordance with institutional protocol and informed consents were obtained from all patients. All samples were reviewed and diagnosed by two independent pathologists.
RNA isolation and qRT-PCR assay
Total RNA including miRNA was extracted from tumor sections using the miRNeasy FFPE Kit (QIAGEN, KJ Venlo, the Netherlands) according to our previous reports [14–16]. RNA concentrations were determined by NanoDrop 2000 (Wilmington, DE, USA). A combination of RUN6B and RUN48 was the housekeeping genes for detection of miR-431 expression [15, 16]. The primers for miR-431, RNU6B and RNU48, were included in TaqMan MicroRNA Assays (4427975, Applied Biosystems, Life Technologies, Grand Island, NY, USA). Sequence of miRNA and references used in the paper are as follows: miR-431 (Applied Biosystems Cat. No. 4427975-001979): UGUCUUGCAGGCCGUCAUGCA; RNU6B, (Applied Biosystems Cat. No. 4427975-001093): CGCAAGGAUGACACGCAAAUUCGUGAAGCGUUCCAUAUUUUU; RNU48 (Applied Biosystems Cat. No. 4427975-001006): GAUGACCCCAGGUAACUCUGAGUGUGUCGCUGAUGCCAUCACCGCAGCGCUCUGACC. The reverse primers were also used for reverse transcription with TaqMan MicroRNA Reverse Transcription Kit (4366596, Applied Biosystems, Life Technologies, Grand Island, NY, USA) in a total volume of 10 µL. Real-time qPCR for miRNA was performed with Applied Biosystems PCR7900. The miR-431 abundance in each sample was normalized to its references. The expression of miR-431 in the FFPE experiments was calculated with the formula 2−Δcq [14–17].
Statistical analysis
Statistical analysis was performed by using SPSS20.0 software and the GraphPad Prism Data are presented as mean ± SD. The t test (Student’s t test) or ANOVA (analysis of variance) was used to determine the differences between two or more independent groups. Receiver operating characteristic (ROC) curve was constructed to evaluate the specificity and sensitivity of predicting HCC and non-tumorous tissues by miR-431 expression levels. And the various cut-off values were identified by the Youden’s Index. Kaplan–Meier method was used to estimate recurrence, and the log-rank test was used to compare the recurrence between groups. Values of P < 0.05 were considered to indicate a statistically significant difference.
Results
Down-regulation of miR-431 in HCC tissues
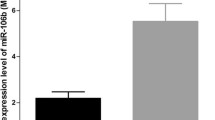
To investigate the expression of miR-431, we performed quantitative real-time PCR in 95 pairs of HCC tissues and matched adjacent tissues. MiR-431 was markedly down-regulated in the HCC samples (1.1885 ± 0.75867), and expressed at a much higher level in corresponding non-cancerous tissues (1.7957 ± 0.89333, P < 0.001, Fig. 1a). To evaluate the diagnostic value of miR-431, the ROC curve was used to analyze the sensitivity and specificity. As shown in Fig. 1b, the ROC curve in HCC and non-cancerous tissues indicated the AUC of the miR-431 was 0.668. At a cut-off value of 2.25, and the sensitivity and specificity were 0.389 and 0.895, respectively (Fig. 1b).
Expression levels of miR-431 in HCC tissues (n = 95) and corresponding non-cancerous tissues (n = 95) by using quantitative real-time PCR. a MiR-431 expression levels in HCC tissues and non-cancerous tissues (***P < 0.001). b ROC curve for miR-431 for HCC patients (95) vs. adjacent tumor group (n = 95), AUC = 0.668
Effects of miR-431 on the clinical parameters in HCC patients. Results are presented as means ± SD. *P < 0.05 **P < 0.01, ***P < 0.001. The relationship between the expression of miR-431 and lymph node metastasis, clinical TNM stage, and the expression level of HCV, MTDH and MVD. HCV: 1. Negative 2. Positive; MTDH: 1. Negative 2. Positive; MVD: 1. Low 2. High; Clinical TNM Stage: 1. I–II 2. III–IV; Metastasis: 1. Without metastasis 2. With metastasis
Correlation between miR-431 expression levels and clinicopathological characteristics in patients with HCC
The relationship between levels of miR-431 and the clinicopathological features
The parameters of 95 HCC patients were summarized in Table 1. Compared with those with lymph node metastasis (0.8361 ± 0.68831), the expression of miR-431 was significantly higher in those HCC patients without lymph node metastasis (1.5639 ± 0.64628, P < 0.001). When compared with HCC patients of advanced stages (III and IV, 1.0996 ± 0.75715), the relative expression of miR-431 in early stages patients (I and II, 1.4836 ± 0.70153, P = 0.037) notably increased. In association with HCV, HCC patients with positive expression of HCV (1.4141 ± 0.76540) had a higher level of miR-431 than those with negative expression (1.0740 ± 0.73494, P = 0.038). Moreover, miR-431 level in HCC patients with MTDH negative (1.4208 ± 0.73775) was higher, compared with those with MTDH positive (0.8600 ± 0.70250, P < 0.001). Besides, the relative expression of miR-431 in HCC patients with low level of MVD (1.3951 ± 0.62353) was remarkably higher in comparison with those with high level of MVD (0.9863 ± 0.82863, P = 0.008, Fig. 2). By contrast, no statistically significant relations were detected between the expression of miR-431 and the following factors: gender, age, differentiation, HCC cirrhosis, portal vein invasion, tumor capsular infiltration and other clinicopathological information (Table 1). Simultaneously, we conducted a further analysis by using Spearman correlation test. Negative correlation was found between miR-431 and metastasis (r = −0.455, P = 0.000), MTDH (r = −0.292, P = 0.006), vaso-invasion (r = −0.284, P = 0.047), MVD (r = −0.281, P = 0.06), as well as clinical TNM stage (r = −0.223, P = 0.030). However, no significant correlation was found between miR-431 expression and the rest clinicopathological parameters.
Capability of miR-431 to function as a HCC tumor marker for diagnosis and progression prediction
The ROC curve analysis was conducted to identify the diagnostic value of miR-431 in HCC patients. As was mentioned in Fig. 1b, the AUC of the miR-431 in HCC and non-cancerous tissues was 0.668, at a cut-off value of 2.25, and the sensitivity and specificity were 0.389 and 0.895, respectively. As was shown in Fig. 3, the AUC of miR-431 to predict metastasis was 0.763, at a cut-off of 0.865 (95 % CI 0.669–0.586, P < 0.001). The AUC for clinical TNM stage was 0.652, at a cut-off of 1.555 (95 % CI 0.527–0.777, P = 0.031). As for vaso-invasion, an AUC of 0.621 was detected, with its cut-off as 1.265 (95 % CI 0.506–0.737, P = 0.048).
Evaluation of the relationship between miR-431 expression and recurrence of HCC patients
Among the 70 patients, 41 had low miR-431 expression (lower than the median level of 1.26), while 29 had high miR-431 expression.As for time to recurrence, low miR-431 expression group was 50.009 ± 2.731 months, while the high expression group was 56.602 ± 3.914 months. Compared with those with low level of miR-431, HCC patients with high level of miR-431 had nearly 6 months longer survival time. However, no significant difference of recurrent time was found between the low and high miR-431 groups (χ 2 = 0.005, P = 0.943).
Discussion
MiR-431 was initially cloned from brain tissue of mouse embryos and identified as central nervous system specific miRNA for the first time by Wheeler et al. [18]. Whole mount in situ hybridization revealed that miR-431 is particularly strongly expressed in the pons. The pons is particularly rich in synapses as ninety percent of the descending axons passing through the midbrain synapse on neurons in the pons [18]. However, up till now, limited information is available on physiological function of miR-431. A previous study in correlation with atuism revealed that alteration of miR-431 expression was discovered in autism patients [19]. Meanwhile, another report verified that neither mutations nor SNPs in miR-431 and miR-21 were found among atuism patients by analyzing the sequence virations of miR-431 [20]. Wu et al. demonstrated [21] that nerve injury-induced miR-431 could stimulate regenerative axon growth by silencing Kremen1 (an antagonist of Wnt/beta-catenin signaling pathway) through interacting with the 3′-UTR of Kremen1 mRNA [21]. Articles mentioned above all emphasized the correlations of miR-431 with nerve system.
Date to the recent 2 or 3 years, scholars have successively observed the potential contribution that miR-431 may play important roles in regulating the progression of carcinoma. Recent observation has identified that expression of miR-431 may be correlated with regulation of cell viability by using microarray-based comparative transcriptome analysis [12]. MiR-431 was upregulated by the addition of human fibroblast interferon (HuIFN-β) in a non-cancerous HuIFN-β sensitive cell lines RSa, with concomitant suppression of IGF1R and IRS2 (putative miR-431 target genes) signaling and inhibition of cell proliferation by suppressing the MAPK pathway [12]. Compared with normal subjects, miR-431 was suppressed in PBMC from initial GD patients, and then recovered in remission. Furthermore, miR-431 expression could be directly inhibited by acute and chronic T3 exposure in PBMC in vitro. Considering these results, miR-431 could serve as a novel biomarker of GD and potential targets for GD treatment [22]. Only one study has indirectly identified the correlation between miR-431 and HCC.As mentioned in the study by Fang et al. [13], HepG2 cells transfected with versican 3′-UTR, which was found to promote versican expression by regulating microRNAs, displayed increased proliferation, survival, migration, invasion, colony formation, and enhanced endothelial cell growth, but decreased apoptosis. Furthermore, 3′-UTR of versican could bind and arrest the functions of miR-431, resulting in upregulation of versican, fibronectin, and CD34 mRNA and protein levels.
HCC still remains as a major disease that causes substantial mortality and plays a negative role in human health, though there has been much advancement in early prognosis and medical or surgical treatment. As better understanding of molecular of pathogenesis of HCC has been proposed in HCC research progresses and novel treatment options. Therefore, the relationship between expression of miR-431 and the progression of HCC probably would provide a new target in the treatment of HCC patients.
Up to now, our study was the first one to reveal directly the relationship between expression of miR-431 and the progression of HCC. In our study, we observed that down-expression of miR-431 may promote the development of HCC through the negative correlation with metastasis, clinical TNM stage, MTDH, MCV and vaso-invasion.
First of all, low level of miR-431 was closely associated with factors of HCC development, such as metastasis, lymph node metastasis and advanced clinical TNM stages. This verified that miR-431 may inhibit the deterioration of HCC. Furthermore, MVD, MTDH and vaso-invasion are factors related to angiogenesis, which play a vital role in tumor growth and progression. In this study, these three factors are all negatively relevant to expression of miR-431. Thus, the expression of miR-431 may inhibit the progression of HCC by affecting the levels of MVD and MTDH, as well as the status of vaso-invasion. Besides, HCV is positively associated with the expression of miR-431. However, a mount of researches have shown that HCV-associated HCC is induced by viral protein-mediated oxidative stress and DNA damage [23]. Another new research has also proposed that HCV nonstructural protein 3(NS3), which plays a role in regulating cell proliferation in carcinogenesis, could activate MMP-9 and COX-2 expression to enhance the invasion and migration of hepatoma cells [24]. Considering these, the role of miR-431 in HCV-related HCC was deserved further research to elucidate the influence of HCV. Although patients with lower level of miR-431 showed a longer time of recurrence than the higher level patients, no statistical significance was observed. A larger sample size is required to figure out the correlation between miR-431 and recurrence in HCC.
Conclusion
In accordance with previous researches, our current study significantly proposes that miR-431 could function as a new tumor-suppressive molecular biomarker for HCC, contributing to the carcinogenesis and deterioration of HCC. Considering the new value of miR-431 in HCC, further observation to investigate and identify the potential functions of miR-431 in HCC is urgent.
References
Hu Q, Lou GG, Liu YC, Qian L, Lv BD. The tumor necrosis factor-alpha-308 and -238 polymorphisms and risk of hepatocellular carcinoma for Asian populations: a meta-analysis. Curr Ther Res Clin Exp. 2014;76:70–5.
McGivern DR, Lemon SM. Virus-specific mechanisms of carcinogenesis in hepatitis C virus associated liver cancer. Oncogene. 2011;30:1969–83.
Ye SL, Takayama T, Geschwind J, Marrero JA, Bronowicki JP. Current approaches to the treatment of early hepatocellular carcinoma. Oncologist. 2010;15(Suppl 4):34–41.
Tanaka S, Arii S. Molecular targeted therapies in hepatocellular carcinoma. Semin Oncol. 2012;39:486–92.
Zhao YJ, Ju Q, Li GC. Tumor markers for hepatocellular carcinoma. Mol Clin Oncol. 2013;1:593–8.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Elmen J, Lindow M, Schutz S, Lawrence M, Petri A, Obad S, et al. LNA-mediated microRNA silencing in non-human primates. Nature. 2008;452:896–9.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M, et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature. 2005;438:685–9.
Park JK, Kogure T, Nuovo GJ, Jiang J, He L, Kim JH, et al. miR-221 silencing blocks hepatocellular carcinoma and promotes survival. Cancer Res. 2011;71:7608–16.
Tanaka T, Sugaya S, Kita K, Arai M, Kanda T, Fujii K, et al. Inhibition of cell viability by human IFN-beta is mediated by microRNA-431. Int J Oncol. 2012;40:1470–6.
Fang L, Du WW, Yang X, Chen K, Ghanekar A, Levy G, et al. Versican 3′-untranslated region (3′-UTR) functions as a ceRNA in inducing the development of hepatocellular carcinoma by regulating miRNA activity. FASEB J. 2013;27:907–19.
Chen G, Kronenberger P, Teugels E, Umelo IA, De Greve J. Targeting the epidermal growth factor receptor in non-small cell lung cancer cells: the effect of combining RNA interference with tyrosine kinase inhibitors or cetuximab. BMC Med. 2012;10:28.
Dang Y, Luo D, Rong M, Chen G. Underexpression of miR-34a in hepatocellular carcinoma and its contribution towards enhancement of proliferating inhibitory effects of agents targeting c-MET. PLoS One. 2013;8:e61054.
Rong M, Chen G, Dang Y. Increased miR-221 expression in hepatocellular carcinoma tissues and its role in enhancing cell growth and inhibiting apoptosis in vitro. BMC Cancer. 2013;13:21.
Chen G, Umelo IA, Lv S, Teugels E, Fostier K, Kronenberger P, et al. miR-146a inhibits cell growth, cell migration and induces apoptosis in non-small cell lung cancer cells. PLoS One. 2013;8:e60317.
Wheeler G, Ntounia-Fousara S, Granda B, Rathjen T, Dalmay T. Identification of new central nervous system specific mouse microRNAs. FEBS Lett. 2006;580:2195–200.
Abu-Elneel K, Liu T, Gazzaniga FS, Nishimura Y, Wall DP, Geschwind DH, et al. Heterogeneous dysregulation of microRNAs across the autism spectrum. Neurogenetics. 2008;9:153–61.
Salem AM, Ismail S, Zarouk WA, Abdul Baky O, Sayed AA, Abd El-Hamid S, et al. Genetic variants of neurotransmitter-related genes and miRNAs in Egyptian autistic patients. Sci World J. 2013;2013:670621.
Wu D, Murashov AK. MicroRNA-431 regulates axon regeneration in mature sensory neurons by targeting the Wnt antagonist Kremen1. Front Mol Neurosci. 2013;6:35.
Liu R, Ma X, Xu L, Wang D, Jiang X, Zhu W, et al. Differential microRNA expression in peripheral blood mononuclear cells from Graves’ disease patients. J Clin Endocrinol Metab. 2012;97:E968–72.
Wu MJ, Ke PY, Horng JT. RacGTPase-activating protein 1 interacts with hepatitis C virus polymerase NS5B to regulate viral replication. Biochem Biophys Res Commun. 2014;454:19–24.
Sakamuro D, Furukawa T, Takegami T. Hepatitis C virus nonstructural protein NS3 transforms NIH 3T3 cells. J Virol. 1995;69:3893–6.
Acknowledgments
The study was supported partly by the Fund of Guangxi Natural Scientific Research (No. 2013GXNSFBA019191), Guangxi Provincial Health Bureau Scientific Research Project (Z2014054), Youth Science Foundation of Guangxi Medical University (GXMUYSF201311), Guangxi University Science and Technology Research Projects (LX2014075), and the Fund of National Natural Science Foundation of China (NSFC 81360327). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of interest
We declare no conflicts of interest (both financial and personal).
Author information
Authors and Affiliations
Corresponding author
Additional information
L. Pan and F. Ren were contributed equally.
Rights and permissions
About this article
Cite this article
Pan, L., Ren, F., Rong, M. et al. Correlation between down-expression of miR-431 and clinicopathological significance in HCC tissues. Clin Transl Oncol 17, 557–563 (2015). https://doi.org/10.1007/s12094-015-1278-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-015-1278-y