Abstract
LINGO-1(LRR and Ig domain–containing NOGO receptor interacting protein 1) is a viable target for spinal cord injury (SCI) repair due to its potent negative regulation in neuron survival and axonal regeneration. Although promising, the intracellular mechanism underlying LINGO-1 regulation is unclear. Here, we identified miR-615 as a potential microRNA (miRNA) that directly targets LINGO-1 by binding its 3′-untranslated region (3′-UTR) and caused the translation inhibition of LINGO-1. MiR-615 negatively regulated LINGO-1 during neural stem cell (NSC) differentiation and facilitated its neuronal differentiation in vitro. Interestingly, compared to the control, neurons differentiated from miR-615-treated NSCs were immature with short processes. Further results showed LINGO-1/epidermal growth factor receptor (EGFR) signaling may be involved in this process, as blockade of EGFR using specific antagonist resulted in mature neurons with long processes. Furthermore, intrathecal administration of miR-615 agomir in SCI rats effectively knocked down LINGO-1, increased neuronal survival, enhanced axonal extension and myelination, and improved recovery of hindlimbs motor functions. This work thus uncovers miR-615 as an effective miRNA that regulates LINGO-1 in NSC and SCI animals, and suggests miR-615 as a potential therapeutic target for traumatic central nervous system (CNS) injury.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Traumatic SCI is an overwhelming CNS disease involved in approximately 180 thousands new cases every year, and more than 2 million individuals suffered with SCI worldwide [1]. SCI usually leads to permanent or temporary loss of motor/sensory capacity, resulting in devastating functional and neurological deficits including paraplegia or tetraplegia. In spite of ongoing efforts over the past decades, the restoration of neurological functions post-SCI still remains a great challenge due to severe neuron loss, limited axon regeneration, and the aggressive pathological microenvironment impeding tissue repair [2].
LINGO-1 is a transmembrane protein selectively expressed on neurons and oligodendrocytes in CNS and the spinal cord, which is originally identified as a critical component of a cell-surface receptor complex mediating axon growth [3]. LINGO-1 has been extensively studied in multiple CNS disorders and SCI due to its prominent inhibitory role in neuronal death, oligodendrocyte differentiation, and axonal regeneration. Various methods of LINGO-1 antagonism resulted in increased surviving neurons, enhanced axonal regeneration and myelination, and improved functional recovery [4,5,6,7,8]. Additionally, emerging evidence suggested LINGO-1 also plays an important role in NSCs [9]. LINGO-1 inhibition by anti-LINGO-1 antibodies or RNAi facilitated NSCs’ proliferation and neuronal differentiation, and LINGO-1-RNAi-treated NSC transplantation significantly improved functional restoration of SCI rats [9, 10]. In spite of the convincing evidence that LINGO-1 antagonism is a promising approach for SCI repair, the underlying endogenous regulatory mechanism of LINGO-1 has however rarely previously been investigated.
MiRNAs are endogenous, small (~ 23 nt) RNAs that play a critical gene regulatory role by binding to the 3′-UTR of target mRNAs, leading to translation suppression or direct destructive cleavage [11]. Approximately 60% of all genes in the human genome are controlled by miRNAs [12]. Microarray analysis of miRNA expression patterns showed extensive miRNAs dysregulated following SCI [13,14,15,16]. Further bioinformatics assay suggested most of these miRNAs targeted genes associated with inflammation, regeneration, apoptosis, and demyelination process that are known to play essential roles in SCI pathogenesis [13, 17]. Thus, we speculated whether LINGO-1 is likewise regulated by certain miRNAs.
To investigate this hypothesis, we attempted to screen possible miRNAs targeting LINGO-1 via bioinformatics analysis. MiR-615 was identified as a potential miRNA that share a consensus sequence with LINGO-1 3′-UTR. The modulation of LINGO-1 by miR-615 was further verified using luciferase reporter assays and Western blot analysis. Many of the researches have demonstrated that miR-615 plays a tumor suppressor role in several cancers. MiR-615 inhibited cell proliferation, migration, and invasion by targeting EGFR in human glioblastoma multiforme [18]; and in human esophageal squamous cell carcinoma (ESCC), miR-615-5p/insulin-like growth factor 2 (IGF2) axis was involved in cell motility and invasion [19]. In addition, miR-615-5p also acts as an inhibitor of VEGF-AKT/eNOS-mediated endothelial cell angiogenic responses in tissue injury [20]. Recent evidence showed miR-615 dynamically expressed in embryonic stem cells (ESCs) [21] and neurolastumor [22], and during the ethanol-induced NSC differentiation, it was detected to be 2-fold of that in undifferentiated NSCs [23]. Moreover, miRNA expression profiling results from various neurological disorders, such as Alzheimer’s disease (AD) [24], neonatal hypoxic–ischemic brain injury [25], and spinocerebellar ataxia type2 (SCA2) [26], also demonstrated the differential expression of miR-615. However, no reports are available on the functions of miR-615 in SCI.
In this study, miR-615 was identified as a potential miRNA negatively regulating LINGO-1 via bioinformatic analysis. The effects of miR-615 on NSCs by targeting LINGO-1 were explored. Furthermore, the neural regeneration and functional recovery of SCI rats were evaluated after intrathecal administration of miR-615. This research provides evidence that miR-615 functions as a negative regulator of LINGO-1, and possibly presents a promising therapeutic target for SCI repair.
Materials and Methods
Bioinformatic Analysis
To discover possible miRNAs targeting the 3′-UTR of LINGO-1, we searched several common online prediction websites, including Target Scan: http://www.targetscan.org/, miRanda: http://www.microrna.org/, MieroCosm Targets: http://www.ebi.ac.uk/enright-srv/microcosm/cgi-bin/targets/v5/genome.pl, and miRDB: http://www.mirdb.org/. The candidate miRNAs that bind with LINGO-1 3′-UTR were further scanned through RNA sequence comparison using BLAST on the NCBI database (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
Preparation of miR-615 Mimics and miR-615 Agomir
The miR-615 mimics/inhibitor, miR-615 agomir, and their negative control (NC) were synthesized by RiboBio (Guangzhou, China). The sequences are as follows: miR-615 mimics—5′GGGGGUCCCCGGUGCUCGGAUC3′, 3′CCCCCAGGGGCCACGAGCCUAG5’; NC-miR-615 mimics—5′UUUGUACUACACAAAAGUACUG3′, 3′AAACAUGAUGUGUUUUCAUGAC5′; miR-615 inhibitor—5′GAUCCGAGCACCGGGGACCCCC3′, 3′CUAGGCUCGUGGCCCCUGGGGG5′; NC-miR-615 inhibitor—5′ CAGUACUUUUGUGUAGUACAAA 3′, 3′GUCAUGAAAACACAUCAUGUUU5′. For the animal experiments, miR-615 mimics and NC-miR-615 mimics were conducted with special process of chemical modification to synthesize miR-615 agomir and NC-agomir, respectively, which present higher biostability and bioactivity to ensure the efficacy in vivo.
NSC Isolation, Culture, and Differentiation
NSCs were obtained from the fetal brain of embryonic 14-day Sprague-Dawley (SD) rats as previous studies described [10]. Briefly, under sterile condition, the brain was dissected and dissociated in pre-cooling Hanks’ Balanced Salt Solution. Then, the cell suspension was collected and centrifuged at 1000 rpm (revolutions per minute) for 5 min. The supernatant was discarded and the cell pellet was resolved in DMEM/F-12 medium (Gibco, Life Technologies, USA) to obtain single-cell suspension. The resuspended cells were subsequently plated on 25-cm2 culture flask (Corning, Acton, MA) and cultured with DMEM/F-12 medium supplemented with 1% N-2 CTS™ supplement (100X; Gibco, Life Technologies, USA), 2% B-27™ supplement (50X; Gibco, Life Technologies, USA), 20 ng/ml epidermal growth factor (EGF; Peprotech, New Jersey, USA), 20 ng/ml fibroblast growth factor (FGF; Peprotech, New Jersey, USA), and 1% penicillin and streptomycin (10,000 U/ml, Gibco, Life Technologies, USA). NSCs were non-adherently cultured into neurospheres at a 5% CO2, 37 °C incubator, and passaged every 3–5 days by digesting with Accutase (Millipore, Bedford, MA) in the medium.
For NSC differentiation, cells were digested and resuspended into single-cell suspension, and subsequently plated on culture dishes coated by poly-l-lysine (Sigma-Aldrich Inc., USA). Cells were cultured in differentiation-inducing medium (DMEM/F-12 medium supplemented with 2% B-27™, 1% fetal bovine serum (FBS; Gibco, Life Technologies, USA)) to initiate cell differentiation. Cell medium was changed every 2 days. All NSCs used in this study were between passages 2 and 4.
Dual-Luciferase Reporter Assay
Rat 3′-UTR of LINGO-1 (LINGO-1 3′-UTR-F: 5′AGAGAATTC CACCCCGCAAGTTCAACA3′; LINGO-1 3′-UTR-R: 5′CGTTCTAGA CAAGTTACAGAGGAAGTTTGTAACT3′) that contains miR-615 target sequence (GGACCCC) was cloned into p-miR-Repoter vector to construct a wild-type Dual-Luciferase Reporter vector (WT-LINGO-1-3′UTR). The miR-615 binding sequence on LINGO-1 3′-UTR was mutated and also cloned into p-miR-Repoter vector to generate a mutated Dual-Luciferase Reporter vector (Mu-LINGO-1-3’UTR). HEK 293T cells were seed in 48-well plates at 4–5 × 104/well and then transiently co-transfected with either WT or mutant LINGO-1-3′-UTR vector (200 ng) and miR-615 mimics using Lipofectamine 2000 (Thermo Fisher Scientific, USA) for 48 h. Cells were collected and the luciferase activity in lysates was measured by a Dual-Luciferase Reporter Assay System (Promega, WI, USA) according to the manufacture’s protocols. Each value from the firefly luciferase activity was normalized to the Renilla luciferase activity.
Cell Transfection
NSCs were cultured with serum free to passage 3 and resuspended into single-cell suspension prior to transfection, subsequently seed on poly-l-lysine-coated 24 wells at density of 1 × 105/well and cultured with 1% FBS supplemented medium for 2 days. When cells were at a confluence of 50~70%, NSC transfection was performed using Lipofectamine 2000 according to the manufacturer’s instructions. NSCs were randomly transfected with miR-615 mimics, miR-615 inhibitor, and their negative control (RIBO Biotech Company, Guangzhou, China) by Lipofectamine 2000, respectively. At 48 h post-transfection, further analysis of the samples such as Western blotting and RNA isolation was performed.
Proliferation Assay
NSCs were digested into single-cell suspension and seeded on recoated 96-well plates at a density of 5 × 103/well. At 48 h post-transfection, 20 μM 5-ethynyl-2′-deoxyuridine (EdU) was added into the cell culture medium. After incubation for another 24 h, cells were fixed in pre-cooling 4% paraformaldehyde (PFA; (Biosharp, Hefei, China)) in PBS for 30 min at 1 day and 3 days post-transfection, followed by incubated with Apollo® reaction cocktails at room temperature (RT) for 30 min, and then counter-stained with 4′-6-diamidino-2-phenylindole (DAPI; 1:5000; Sigma, USA) according to protocols of Cell-Light EdU DNA Cell Proliferation Kit (RIBO Biotech Company, Guangzhou, China). The pictures of EdU-positive cells were captured randomly under a fluorescence microscope and the ratio of EdU-positive cells to total cells was calculated by two experimenters blind to groups.
Establishment of Spinal Cord Injury Model
All procedures using laboratory animals were conducted in compliance with the Guide for the Care and Use of Laboratory Animals (National Research Council, 1996) and approved by the Animal Care and Use Committee of Guangdong Medical University.
Adult SD rats (female, weighing 180–200 g, aged 8–10 weeks) used in this study were ordered from Laboratory Animal Center of Southern Medical University (Guangdong, China). Rats were housed in standard cages under a regulated environment (12-h light/dark cycle) with free access to food and water for about 1 week. To produce T10 complete transection model, animals were anesthetized and the hair of the back was shaved. For the surgery, the anesthetized rats were placed on prone position, and an about-2-cm midline incision was made at the level of T10 under the aseptic condition. After the muscles were separated away from the vertebral arch, the spinous process of thoracic vertebra level 10 (T10) was dissected out and a laminectomy was performed at the same level. Then an about-0.5-cm midline longitudinal incision was carefully made to expose the spine cord. The T10 spinal cord was completely transected using a microscissor. Finally, the paraspinal muscles and skin were sutured separately after hemostasis. The body temperature of rats was maintained at 37.0 ± 0.5 °C with a heating pad during surgery.
The inclusion/exclusion criteria of SCI animals
Before the surgery, BBB score was performed and all animals scored 21. At 24 h after SCI, BBB score was evaluated again and rats that scored 0 were included. Animals with severe wound infection and erosion, bleeding, and death were excluded.
Transplantation of miR-615 Agomir and NC-Agomir into SCI Rats
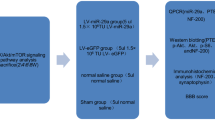
The intrathecal catheter and minipump implantation surgery was performed at T12/T13 immediately after SCI. Rats were randomly divided into three groups: control group (n = 24), miR-615 agomir group (n = 24), and NC-agomir group (n = 24). All operation was performed by two experimenters who were blinded to the animal groups.
In brief, miR-615 agomir and NC-agomir were dissolved in 0.9% saline to achieve a concentration of 20 nmol/mL and filled into the osmotic mini-pumps (Alzet 1030D, CA, USA), which then were continuously delivered (1 μL/h) into the spinal cord of SCI rats for 3 days through a subdural-implanted catheter connected to the subcutaneously implanted osmotic mini-pumps, as previous studies described [16]. The control animals were treated with PBS (1 μL/h) for 3 days. To ensure immediate delivery after implantation, each pump was primed overnight at 37 °C. After the implantation, muscles and skin were sutured in layers sequentially, and rats were kept in appropriate environment for recovery. The entire surgical procedures were conducted at a warm environment. All animals were intraperitoneally administrated with penicillin G and streptomycin (10,000 U/ml; Gibco, Life Technologies, USA) once a day in the first week post-operation. Distended bladders were expressed by manual massage on the lower abdomen twice daily until voluntary or autonomic emptying fully recovered.
Behavior Test
The Basso-Beattie-Bresnahan (BBB) locomotor test (n = 24 per group) was conducted to evaluate the hindlimb functions as described previously [27]. BBB test is an evaluative criteria relevant to the lumbar injury model, which grades ranging from 0 (no observable movement of the hind limbs) to 21 (normal hind limb locomotion). Animals were placed individually in a quiet, open field to ensure voluntary movement. The locomotor function of the ankle, knee, and hip joints was evaluated and the BBB scores were recorded weekly until to the 8-week endpoint. The BBB scores were evaluated and recorded by two experimenters who know the criteria well but were blinded to the animal groups. The final score of each animal for statistical evaluation was obtained by averaging the values from both investigators.
Axonal Retrograde Tracing
At 2 days before perfusion, rats (n = 8) from each group were randomly selected to perform axonal retrograde tracing study using Fluorogold (FG; Fluorochrome.LLC, USA). Animals were re-anesthetized and a dorsal laminectomy was performed at T11–12. About 2 μl Fluorogold was slowly injected into a second site that is approximately 5-mm caudal to the lesion site. After 2 days, the labeled animals were perfused and a 15-mm-long spinal segment encompassing the T10 transection site was extracted and sectioned longitudinally at a freezing microtome (Leica CM1950, Germany). FG-labeled cells rostral to the T10 transection site in sections were observed and randomly counted under the fluorescence microscope.
Immunofluorescence Assay
For immunocytochemistry, cells were fixed with 4% pre-cooling PFA (Biosharp, Hefei, China) for 30 min and permeabilized in 0.3% Trixion-100 (Biosharp, Hefei, China) at RT for 10 min, subsequently blocked in 5% bovine serum albumin (BSA; Gen-view Scientific INC., USA) for 30 min. For immunohistochemistry, frozen sections collected from the harvested spinal segments were firstly balanced in RT for 10–20 min, and soaked in PBS for 10 min. After incubated in proteinase K (1:2000; Sigma-Aldrich, USA) for 10 min at 37 °C for antigen retrieval, tissues were washed in PBS for three times and incubated in blocking reagent (5% BSA and 0.2% TritionX-100) for 30 min. After blocked, cells and tissue sections were incubated with primary antibodies overnight at 4 °C. Post-washing in PBS for three times, the primary antibodies were identified with Alexa Fluor 488/568 goat-anti rabbit or mouse antibodies (1:500; Life Technologies, USA), and the nucleus were counter-stained with DAPI. Finally, the slides were mounted and observed under a fluorescence microscope.
Primary antibodies used in this study were as follows: βIII-tubulin (1:200; Millipore, Bedford, USA), glial fibrillary acidic protein (GFAP; 1:200; Sigma-Aldrich, USA), 2′,3′-cyclic nucleotide-3′-phosphodiesterase (CNPase; 1:200; Abcam, England), Neuronal Nuclei (NeuN; 1:200; Millipore, USA), LINGO-1 (1:400; Abcam, England), neurofilament 200 (NF-200; 1:400; Sigma-Aldrich, USA).
Quantitative Real-Time PCR
The expression level of miR-615 in NSCs during differentiation and in the spinal tissues of SCI rats (n = 4) at 7 days post-transplantation surgery was evaluated by quantitative real-time polymerase chain reaction (qRT-PCR). Total RNA from NSCs or 10-mm-long spinal cord segments containing the injury epicenter was extracted using Trizol (Invitrogen, Life Technologies, USA) according to the manufacturer’s instructions. Total RNA from each sample was reversely transcripted to cDNA using the PrimeScript RT reagent Kit (TaKaRa, Tokyo, Japan). QRT-PCR was conducted using the SYBR Premix Ex Taq (TaKaRa, Tokyo, Japan) and specific primers for miR-615 (RiboBio, Guangzhou, China) on the Applied Biosystems 9700 Thermocycler (Thermo Fisher Scientific, USA). U6 (RiboBio, Guangzhou, China) was used as the internal control. The relative expression of miR-615 was calculated using the comparative 2−ΔΔCt method and normalized to U6. Primer sequences for genes were as follows: miR-615, F: 5′AAGGGGGTCCCCGGT3′, R: 5′GTGCGTGTCGTGGAGTCG3′; U6, F: 5′GCTTCGGCAGCACATATACTAAAAT3′, R: 5′CGCTTCACGAATTTGCGTGTCAT3′.
Western Blotting Analysis
The expression of proteins in cells and spinal tissues was detected using Western blot analysis. Total protein from each sample was extracted using RIPA (Solarbio, Beijing, China) supplemented with PMSF (RIPA: PMSF = 100:1; Solarbio, Beijing, China). The proteins were separated through sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS/PAGE; KeyGEN BioTECH, Nanjing, China) and transferred to polyvinylidene difluoride (PVDF; Millipore, Mississauga, Canada) membranes. Next, after blocked in 5% nonfat milk for 2 h at RT, the membranes were then incubated with primary antibodies at 4 °C overnight, followed by incubated in horseradish peroxidase (HRP)–conjugated secondary antibodies (1:1000) at RT for 2 h. Proteins were visualized via chemiluminescence (Pierce, USA) and the relative intensities of bands were determined using Image J. Each value of protein level was normalized to GAPDH or β-actin, respectively. The value of phosphorylated protein was normalized to the corresponding total protein. Data was provided in terms of the mean ± SD of the percentage ratio of the control.
The primary antibodies used in this experiment were obtained from the following sources: rabbit anti-LINGO-1; rabbit anti-CNPase (1:1000; abcam, England); rabbit anti-EGFR (1:1000; Bioworld technology, USA); rabbit anti-p-EGFR (1:1000; Bioworld technology, USA); rabbit anti-RhoA (1:1000; Bioworld technology, USA); rabbit anti-p-RhoA (1:1000; Bioworld technology, USA); GFAP (1:1000; Sigma-Aldrich, USA); NF-200 (1:1000; Sigma-Aldrich, USA); myelin basic protein (MBP) (1:1000); βIII-tubulin (1:1000; Millipore, Bedford, USA); GAPDH (1:5000; Thermo Fisher, Wilmington, USA); β-actin (1:5000; EarthOx, USA).
Myelination Observation
For myelination assessment, rats (n = 4 per group) were anesthetized at 8 weeks post-surgery and perfused with 2.5% glutaraldehyde (Kemiou, Tianjing, China; 25% glutaraldehyde: 4% PFA = 1:9). The harvested tissues were post-fixed in 2.5% glutaraldehyde, subsequently in 1% osmium tetroxide, and then cleared in propylene oxide post-dehydration in a series of graded alcohols. After embedded in pure Epon, transverse semithin sections (1 μm) were obtained and stained with 1% toluidine blue. For electron microscopy (EM), 50-nm ultrathin sections were required and double-stained with 2% uranyl acetate and led citrate for EM analysis. The number of myelinated axons was counted and the morphology of myelin was observed under EM in a blinded manner.
Clarity
Spinal cord tissues were optically cleared according to the CLARITY protocols as described previously [28]. At 8 weeks after injury, rats (n = 4 per group) were anesthetized and transcardially perfused with 0.9% saline followed by 4% PFA. After the abdominal aorta was identified and clipped with vessel forceps, rats were sequentially perfused with 50-ml ice-cold reagent A (0.05% bisacrylamide, 4% acrflamide, 0.25% VA-044, 8% PFA in 0.1 M PBS). Then 10-mm-long spinal segment centered at the injury site was rapidly extracted and post-fixed in reagent A at 4 °C for 6 h under the condition of protection from light. For hydrogel–tissue hybridization, the tissues were then transferred into modified reagent B (0.05% bisacrylamide, 4% acrflamide, 0.25% VA-044 in 0.1 M PBS) and incubated at 37 °C for 3 h. After the hydrogel hybridization was triggered, excessive gel was softly removed from the surface of spinal cord. Transverse sections of 200–500 μm were collected and soaked into clearing solution (8% SDS in 0.1 M PBS, pH 7.5) to allow passive clear, on a rotating incubator at 37 °C. The clearing solution was changed every 5 days until the tissues were adequately cleared. After cleared about 2 months, the cleared sections were carefully taken out and washed in 0.1 M PBS for 3 days. The sections were then incubated with the primary antibody for 2 days at 37 °C in the water bath, followed by incubated with the secondary antibody for additional 2 days. The primary antibodies were the following: NF-200 (1:200; Sigma-Aldrich, USA) and GFAP (1:200; Sigma-Aldrich, USA). The secondary antibodies were goat anti-mouse Alexa Fluor 488 (1:200; Life Technologies, USA) and goat anti-rabbit Alexa Fluor 568 (1:200; Life Technologies, USA). Finally, post-rinsed in PBS for 2 days, sections were mounted with isosorbide and imaged under a confocal microscope.
Statistical Analysis
All statistical data were analyzed using SPSS (Version 20.0; Abbott Laboratories, Chicago, IL). In vitro tests, cell differentiation, Western blot, and PCR analysis were evaluated by one-way ANOVA followed by Tukey’s multiple comparison test, and EdU assay was performed via two-way ANOVA followed by Tukey post-test. In vivo experiments, the BBB scores and Western blot analysis of CNPase were analyzed by two-way ANOVA between groups over time followed by Tukey’s multiple comparison test, and other analysis were all obtained using one-way ANOVA followed by Tukey post-test. Homogeneity of variance test was used to measure the data. Each data was presented as mean ± standard deviation (SD). All tests were two-tailed and the value of P < 0.05 was considered statistically significant.
Results
MiR-615 Negatively Regulated LINGO-1 During NSC Differentiation In Vitro
Most miRNAs are thought to control gene expression by base-pairing with the miRNA-recognizing elements found in the 3′-UTR of their messenger target. To identify the potential miRNAs targeting LINGO-1, several miRNA target site prediction databases were searched to screen miRNAs that share a specific consensus sequence with the 3′-UTR of LINGO-1. Results indicated five potential RNA fragments, among which miR-615 was preliminarily designated as the candidate miRNA via further literature review and RNA sequences comparison analysis using BLAST on the NCBI database (http://blast.ncbi.nlm.nih.gov/Blast.cgi). MiRNA prediction analysis suggested miR-615 (miR-615-5p) could bind with the target sites (GGACCCC) in the 3′-UTR of LINGO-1, which are located in 202–223 bp (Fig. 1a).
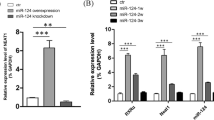
MiR-615 negatively regulated LINGO-1 during NSC differentiation by directly targeted LINGO-1. a Schematic diagram of predicted target sites (202–223 bp) of miR-615 at the 3′-UTR of LINGO-1, and sequence alignment of the putative miR-615 binding sites. The 7-nt seed sequence was underlined. b The relative luciferase activity was analyzed after WT/mutant-pGL3-LINGO-1 3′-UTR vectors were co-transfected into HEK293 cells with miR-615 mimics, miR-615 inhibitor, and their negative control. Firefly luciferase values were normalized for Renilla luciferase. c Western blotting analysis of LINGO-1 in NSCs transfected with miR-615 mimics, miR-615 inhibitor, and their negative control. β-Actin was used as the internal control. d Histogram showing the relative amount of LINGO-1 protein level. The expression of miR-615 and LINGO-1 during NSC differentiation was detected using qRT-PCR and Western blot, respectively. e MiR-615 significantly increased in cultures differentiated for 1 day, 3 days, and 5 days compared to proliferating cells, but it appeared a downward trend during the differentiation. U6 was used as an internal control. f Western blot analysis and g relative quantification of LINGO-1 protein during NSC differentiation. There was a sustained increase in LINGO-1. GAPDH was used as an internal control. Date was shown as mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001
To determine whether LINGO-1 was regulated by miR-615 through direct binding to its 3′-UTR, the WT and mutant LINGO-1 3′-UTR segment containing the potential target sites for miR-615 was constructed and cloned into the downstream region of dual-luciferase reporter gene. MiR-615 mimics, miR-615 inhibitor, or their negative controls were co-transfected into HEK293T cells with LINGO-1-3′-UTR-luciferase reporter vector to detect the luciferase enzyme activity. The relative luciferase activity was significantly repressed when miR-615 mimics was co-transfected with WT-LINGO-1-3′UTR-luciferase vector (Fig. 1b); on the contrary, miR-615 inhibitor abolished this repression and resulted in a prominent increase of the luciferase activity (Fig. 1b), confirming the specificity of this action. This result suggested miR-615 directly contributes to mRNA repression via specific binding with LINGO-1 3′-UTR.To further elucidate the effect of miR-615 on LINGO-1, NSCs were transfected with miR-615 mimics, miR-615 inhibitor, and their negative controls, respectively. Western blotting analysis revealed a significant reduction of LINGO-1 in NSCs treated with miR-615 mimics compared with the control (Fig. 1c, d); conversely, miR-615 inhibitor obviously enhanced LINGO-1 expression (Fig. 1c, d). Taken together, these results suggested miR-615 negatively regulated LINGO-1 via direct binding with its 3′-UTR.
Although little is known about the functions of miR-615 currently, recent miRNs profiling analysis reported miR-615 dynamically expressed during the differentiation of ESCs and ethanol-induced NSCs [21, 23]. In light of the supporting evidences that LINGO-1 antagonism promoted neuronal differentiation of NSCs [9, 10], it left open the possibility that the process of NSC differentiation may also be mediated by miR-615. To investigate the expression of miR-615 and LINGO-1 during NSC differentiation, cell lysates from proliferating NSCs (0 day) and differentiated NSCs (differentiated for 1, 3, 5, and 7 days) were harvested and analyzed via RT-PCR and Western blotting, respectively. Results showed that though presenting a down-trend after differentiation, the endogenous level of miR-615 was significantly increased during the 5-day differentiation compared with proliferating NSCs (Fig. 1e), suggesting that miR-615 was upregulated in the early differentiation of NSCs. In spite of the low protein level of LINGO-1 at 0 day, Western blot results revealed LINGO-1 expressed in both the proliferating and differentiated NSCs (Fig. 1f). A continuous increase of LINGO-1 was observed during NSC differentiation, and the maximum protein level was detected at cells differentiated for 7 days (Fig. 1f). Quantification of LINGO-1 protein revealed a 7-fold increase at 7 days relative to that of undifferentiated cells (Fig. 1g). The opposite expression tendency between miR-615 and LINGO-1 observed above indicated miR-615 may negatively regulated LINGO-1 during NSC differentiation. Critically, other certain factors may also control LINGO-1 in addition to miR-615 during the differentiation.
MiR-615 Facilitates NSC Proliferation and Neuronal Differentiation In Vitro
To identify whether miR-615 influences NSC proliferation, EdU proliferation assay was performed at 1 day and 3 days after transfection. The number of cells incorporated EdU and total cell numbers illustrated by DAPI-positive cells were counted and the rate of EdU-positive cells was calculated (Fig. 2a, b). At 1 day, massive cell proliferation was observed in each group, but no significant difference was found with respect to the percentage of EdU-positive cells (Fig. 2a, b). At 3 days, however, though cell proliferation was largely declined generally, the rate of EdU-positive cells in miR-615 mimics group was prominently increased compared to other groups (Fig. 2a, b). These data indicated miR-615 could promote NSC proliferation.
MiR-615 promoted NSC proliferation and neuronal differentiation in vitro. a Edu assay was conducted to investigate the effect of miR-615 on NSC proliferation at 1 day and 3 days post-transfection. b Edu-positive cells were counted and plotted as the ratio of the total cell number. To investigate the effect of miR-615 on NSC differentiation, cells were cultured with miR-615 mimics or miR-615 inhibitor for 5 days. c Specific antibodies against βIII tubulin, GFAP, and CNPase were used to identified neurons, astrocytes, and oligodendrocytes, respectively. d Cells positive for specific markers were calculated and plotted as the ratio of the total cell number. e Western blot analysis and f relative amount of βIII tubulin, GFAP, and CNPase expression in differentiated NSCs. GAPDH was used as an internal control. Data was shown as mean ± SD. * represents miR-615 mimics VS NC-mimics, *p < 0.05, **p < 0.01, ***p < 0.001; # represents miR-615 mimics VS miR-615 inhibitor, #p < 0.05, ##p < 0.01, ###p < 0.001. Scale bars (a) = 20 μm, scale bars (c) = 50 μm
Most miRNAs play essential roles in stem cell self-renewal and differentiation by negative regulation of certain genes. To investigate the effect of miR-615 on the fate determination of NSCs in vitro, NSCs were transfected with miR-615 mimics or miR-615 inhibitor and cultured in differential medium. At 5 days post-transfection, cells were fixed and immunostained against neural markers specific for neurons (βIII tubulin), astrocytes (GFAP), and oligodendrocytes (CNPase). Compared with NC-mimics group, in which βIII tubulin-positive cells have a rather mature neuronal morphology with multiple long extending processes, the βIII tubulin-positive cells in miR-615 mimics group represented relatively immature phenotype with short processes (Fig. 2c). But neurons in miR-615 inhibitors developed mature and had an identified morphology with βIII tubulin-positive cells in the control (Fig. 2c). Inversely, astrocyte differentiation was significantly inhibited and promoted by miR-615 mimics or miR-615 inhibitor, respectively, but no difference was observed with respect to the morphology of GFAP-positive cells among different groups (Fig. 2c). Interestingly, we also found CNPase-positive oligodendrocytes in miR-615 mimics group were slightly more differentiated than cells cultured with miR-615 inhibitor or the controls (Fig. 2c). In addition, LINGO-1 inhibition by miR-615 dynamically affected the proportion of different type of cells differentiated from NSCs. MiR-615 mimics resulted in a 2-fold increased number of βIII tubulin-positive cells compared to other groups (Fig. 2d), and decreased percentage of GFAP-positive astrocytes, whereas no significant difference was observed in the percentage of oligodendrocytes among groups though increased number of CNPase-positive cells was detected in miR-615 mimics group (Fig. 2d). Western blotting analysis revealed an obviously increased expression of βIII tubulin and CNPase, and decreased enrichment of GFAP in cells treated with miR-615 mimics (Fig. 2e, f), while miR-615 inhibitor treatment showed the opposite results. Taken together, the above results indicated miR-615 facilitated neuronal differentiation and impeded astrocyte formation from NSCs in vitro.
Identification of Potential Signaling Pathways Involved in LINGO-1 Regulation by miR-615
Our results have demonstrated miR-615 negatively mediated LNGO-1 during NSC differentiation. Then, certain signal proteins associated with LINGO-1 in NSCs were measured post-transfection. As illustrated in Fig. 3, the protein level of LINGO-1 and its downstream signal protein p-RhoA were both significantly decreased after transfected with miR-615 mimics (Fig. 3a, b), suggesting miR-615 decreased the activation of LINGO-1/RhoA pathway. In contrast, when miR-615 was inhibited, LINGO-1 increased notably, therefore enhancing p-RhoA expression (Fig. 3a, b). Moreover, the upregulation of EGFR was also observed in miR-615 mimics group (Fig. 3a, b). Studies have reported EGFR suppression significantly promoted axonal regeneration and neurite outgrowth [29], which stimulated us whether EGFR was implicated in the generation of neurons with short process. Then NSCs were transfected with miR-615 mimics, NC-mimics, EGFR inhibitor PD168393, and miR-615 mimics plus PD168393. MiR-615 mimics treatment resulted in significantly decreased LINGO-1 and elevated EGFR level (Fig. 3c, d). As expected, LINGO-1 and EGFR were both suppressed by the co-transfection of miR-615 mimics and PD168393 (Fig. 3c, d). Compared with the neurons with short process in miR-615 mimics group, miR-615 mimics with PD168393 led to βIII tubulin-positive cells with long extending neurites (Fig. 3e); data revealed the average neurite length of neurons in this group was largely greater than that of miR-615 mimics and NC-mimics ones as well (Fig. 3f). The maximum proportion of neurite length ranges in miR-615 mimics mainly located in smaller range (0~60 μm) (Fig. 3g), while most processes were longer than 100 μm when EGFR was blocked by PD168393 (Fig. 3g). Together, these results suggested LINGO-1 inhibition by miR-615 may contribute to neuronal differentiation with short process via LINGO-1/RhoA or EGFR signal pathways.
MiR-615 may regulate NSCs via LINGO-1/RhoA or EGFR pathway. a Western blot analysis and b relative quantification of LINGO-1 and relevant signal proteins in NSCs at 6 days post-transfection. β-Actin was used as an internal control. c Western blot analysis and d relative amount of LINGO-1 and EGFR in NSCs transfected with miR-615 mimics or miR-615 mimics plus PD18393 (EGFR inhibitor). GAPDH was used as an internal control. e Representative immunofluorescent staining showing βIII-tubulin neurons in the control, miR-615 mimics, and miR-615 mimics plus PD18393 group. Scar bar = 10 μm. f The mean neurite length of βIII-tubulin neurons was measured and plotted. g The proportion of neurite length ranges was counted and plotted. Data was shown as mean ± SD. * represents miR-615 mimics VS NC-mimics, *p < 0.05, **p < 0.01, ***p < 0.001; # represents miR-615 mimics VS miR-615 inhibitor, #p < 0.05, ##p < 0.01, ###p < 0.001
Transplantation of miR-615 Agomir Contributed to Axonal Regeneration and Functional Recovery of SCI Rats by Targeting LINGO-1
To further investigate the effect of miR-615 in SCI animals in vivo, the chemically modified miR-615 mimics named miR-615 agomir and NC-agomir were constructed, and grafted into SCI rats through intrathecal injection immediately post-surgery. QRT-PCR results showed miR-615 remarkably reduced post-SCI over time, and it significantly upregulated in miR-615 agomir-treated rats at 7 days after transplantation (Fig. 4a). On the contrary, the protein level of LINGO-1 was gradually increased post-injury, and this was remarkably reversed by miR-615 overexpression as expected (Fig. 4b, c). The BBB scores were evaluated and recorded weekly post-surgery by two blind observers. A spontaneous functional recovery was observed in all groups, which was demonstrated by the gradually increased BBB scores. Rats treated with miR-615 agomir exhibited significantly improved hindlimb activity scores and this significant difference was maintained until to the 8-week experiment endpoint (Fig. 4d). No obvious difference was detected between the control and NC-mimics groups (Fig. 4d).
MiR-615 effectively contributed axonal regeneration and functional recovery following SCI by targeting LINGO-1. a qRT-PCR analysis revealed that miR-615 obviously decreased post-SCI, and it significantly upregulated in rats transplanted with miR-615 agomir at 7 days after surgery. U6 was used as an internal control. b Western blot analysis and c relative amount of LINGO-1 in SCI rats. β-Actin was used as an internal control. d BBB-locomotor scores were evaluated weekly post-surgery. The BBB scores of rats treated with miR-615 agomir was significantly increased since the 2 weeks post-injury compared to the controls. e Longitudinal sections encompassing the injury center with immunofluorescent staining exhibited that regenerative nerve fibers (NF-200-labeled, green) partly bridged the lesion at 2 weeks after injury in the miR-615 agomir group. Scale bars = 1000 μm. Enlarged image of the box area (white box) illustrating a large number of parallel nerve fibers extended through the glial scar. Scale bars = 50 μm. f Representative immunofluorescent staining showing surviving neurons and regenerative nerve fibers in the lesion labeled by NF-200 at 8 weeks post-SCI. Scale bars = 20 μm. g Comparison of the relative number of surviving neurons in each group. h Representative images of Fluorogold-labeled motoneurons in the T8-T9 spinal segment. The significantly increased FG-positive cells were observed in the miR-615 agomir group at 8 weeks post-SCI. Scale bars = 50 μm. i The relative number of FG-positive cells in each group. Data was shown as mean ± SD, * represents VS the control, *p < 0.05, **p < 0.01, ***p < 0.001; # represents VS NC-agomir, #p < 0.05, ##p < 0.01, ###p < 0.001
At 2 weeks post-transplantation, the tissue continuity of the lesion center was then evaluated by NF-200 (green). In the control and NC-agomir treatment, very few regenerative nerve fibers and varying amount of big cavities were observed (Fig. 4e). However, in spite of several small cavities, there was abundance of newborn nerve fibers out-extending into the damage crack in miR-615 agomir-treated rats, which partly bridged the damage gap and repaired the tissue continuity (Fig. 4e). Next, the surviving neurons in the lesion sites were analyzed at 8 weeks post-surgery. In addition to significantly increased numbers, neurons in miR-615 agomir animals were multipolar with large cell bodies, but those in the controls displayed sunken stomata (Fig. 4f, g). The plasticity of regenerative axons from neurons was further investigated via FG retrograde tracing. Only a small percentage of neurons were labeled in both the control and NC-agomir rats at 8 weeks post-surgery, while in the miR-615 agomir group a large number of FG-positive neurons were detected (Fig. 4h), which was approximately triple of that in the control (Fig. 4h, i).
Transplantation of miR-615 Agomir Promoted the Out-Extension of Axons Through the Glial Scar
Glial scar formation is recognized as a major physical impediment for axonal regeneration after SCI. Then we investigate whether miR-615 has any effects on axonal regrowth and scar formation. In the control and NC-agomir group, obvious glial scar and few nerve fibers were observed, whereas plentiful regenerative axons extended in parallel in the scar tissues in the miR-615 agomir group (Fig. 5a). The protein levels of NF-200 and MBP were both increased in rats treated with miR-615 agomir, and accompanied with downregulation of GFAP compared to the control (Fig. 5b, c). To better observe the out-spouting of regenerative nerve fibers into the scar, the lesion center tissues were hyalinized using CLARITY and scanned under the confocal microscope after staining. As the three-dimensional images of cleared spinal tissues reveal in Fig. 5d, there were few regenerative axons and obvious hyperplasia of glial scar at the lesion in the controls, and the long-distance regeneration of nerve fibers was notably prevented by the dense scar. On the contrary, miR-615 agomir alleviated glial scar formation and facilitated the regenerative nerve fibers extending through the inhibitory scar tissues for long-distance outgoing growth (Fig. 5d).
MiR-615 promoted the outgrowth of regenerative fibers cross the glial scar. a Representative immunofluorescent staining showing the regenerative nerve fibers (NF-200-labeled, green) and glial scar (GFAP-labeled, red) in the lesion at 8 weeks post-injury. Scale bars = 50 μm. b Western blot analysis and c relative amount of NF-200, MBP, and GFAP in SCI rats at 8 weeks post-surgery. GAPDH was used as an internal control. d A 5-mm-long spinal segment encompassing the transection site was optically cleared and spatially scanned using laser-scanning confocal microscope after conducted with immunofluorescent labeling procedure. Representative CLARITY images showed many of regenerative nerve fibers crossed through the reduced glial scar in the miR-615 agomir group, while few axons and dense scar tissues were observed in the controls. Scale bars = 50 μm. Higher magnification view from box area (white box) demonstrating multiple regenerative nerve fibers passed through the glial scar for out-extension in rats treated with miR-615 agomir. Scale bars = 20 μm. Data was shown as mean ± SD, * represents VS the control, *p < 0.05, **p < 0.01, ***p < 0.001; # represents VS NC-agomir, #p < 0.05, ##p < 0.01, ###p < 0.001
Transplantation of miR-615 Agomir Promoted Axonal Remyelination
Axonal myelination is critical for neurological functions. To explore the possible effects of miR-615 on remyelination, CNPase, a major myelin protein, was measured at the early phase after SCI. The protein level of CNPase was significantly enhanced in the miR-615 agomir group relative to the control and NC-agomir treatment (Fig. 6a, b); notably, it increased continuously at the first week post-injury, suggesting miR-615 may contribute to axonal myelination. We further evaluated remyelination via toluidine blue staining and EM. Apparent loss of myelination was observed in the control and NC-agomir groups (Fig. 6c–e), while miR-615 agomir led to slight demyelination and much more small regenerative myelinated axons (Fig. 6c). The number and percentage of myelinated axons in miR-615 agomir treatment were both obviously increased (Fig. 6d, e). No significant difference was observed between the control and the NC-agomir groups (Fig. 6d, e). EM photographs showed multi-layer compacted myelination in miR-615 agomir rats, while the lamellae of myelin sheath were loose and fragmentary in the controls (Fig. 6f). All these results suggested miR-615 contributed to remyelination after SCI.
MiR-615 facilitated axonal myelination following SCI. a Western blot analysis and b relative amount of CNPase in spinal tissues at 3 days, 5 days, and 7 days after injury. GAPDH was used as an internal control. c Toluidine blue staining of myelinated axons showed that miR-615 agomir transplantation led to much more small, regenerative myelinated axons compared to the control at 8 weeks post-injury. Scare bars = 50 μm. d The number and e percentage of myelinated axon in the spinal cord was counted and plotted. f Scanning electron microscope images of viable myelinated axons in spinal cord. Scare bars = 1 μm. Each date was shown as mean ± SD. * represents VS control, *p < 0.05, **p < 0.01 ***p < 0.001; # represents VS NC-agomir, #p < 0.05, ##p < 0.01, ###p < 0.001
Discussion
Taken together, our data indicate that miR-615 is a potential miRNA-modulating LINGO-1 and regulates the neuronal differentiation of NSCs. Moreover, miR-615 promotes axonal regeneration and functional recovery of SCI rats via LINGO-1 inhibition, suggesting a promising role for miR-615 in LINGO-1 regulation and repair of traumatic CNS diseases.
Extensive miRNAs are dysregulated following SCI, most of which accelerate or attenuate the aggressive pathological progress by targeting certain genes [14]. Here, via vast bioinformatics analysis and dual-luciferase reporter assays, miR-615 was identified as a possible miRNA targeting LINGO-1 by directly binding with its 3′-UTR. MiR-615 is part of the Hox cluster of genes responsible for regulating spatial patterning during development [30]. It has been demonstrated that miR-615 plays a critical role in multiple cancers, neurodegeneration, and angiogenesis post-injury [18,19,20]. Interestingly, consistent with our study, miR-615 also exerts suppressive effects on tumor cell and endothelial cell by inhibition of certain genes via targeted the 3′-UTR. MiR-615 significantly upregulated at the earlier differentiation of ESCs, NSCs, and neuroblastoma cell lines [21,22,23], implying it may be associated with neural fate determination. LINGO-1 expressed in proliferating NSCs and dynamically increased as cell differentiation [9], which was also observed in the present study. Consistently, there was an increase in miR-615 during the early differentiation (0~1 day) of NSCs, and it exhibited a downward trend along with cell differentiation. The opposite expression tendency between miR-615 and LINGO-1 during NSC differentiation further verified the negative modulation of LINGO-1 by miR-615. However, it is notable that the increase of miR-615 was contradictory with LINGO-1 upregulation at 0~1 day during differentiation. Recent studies [31, 32] have described that miR-615 could be occupied by the ESC transcription factors, and it was transcriptionally silenced in ESCs but expressed in a tissue-specific fashion in differentiated cells, which was a possible explanation for the transient enhancement of miR-615. Additionally, other certain factors may also be implicated in the early NSC differentiation. Investigations on the temporal expression of miR-615 in NSCs is worth further exploration and is expected to provide new sights for miR-615 mechanism.
LINGO-1 is a potent negative regulator of neuronal survival and neural commitment. Target loss of LINGO-1 in NSCs enhanced cell proliferation and skew NSC differentiation to generate neurons [10, 33, 34]. Similarly, blockade of LINGO-1 by miR-615 significantly facilitated NSC proliferation and neuronal differentiation, but inhibited the production of astrocytes. In progressive retinal neurodegeneration, circular RNAs (circRNAs) acted as miR-615 sponge to sequester and inhibit miR-615 activity, leading to increased Meteorin (METRN), a secreted protein expressed in undifferentiated neural progenitors and selectively promoted astrocyte formation from mouse cerebrocortical neurospheres [35], which further confirmed our present results. However, in addition to inhibit LINGO-1, whether increased miR-615 exerted neuronal differentiation effects via regulation of Meteorin required investigation. Interestingly, in keeping with the phenomenon observed previously that neutralization of LINGO-1 results in proliferation of immature neurons [9], generated neurons from miR-615-treated NSCs retained similar phenotype with short processes compared to the control. But the long-term effect remains to be further evaluated and elucidated.
LINGO-1/RhoA signaling pathway plays a vital role in the inhibition of neuronal sprouting and axonal regeneration. Silence of LINGO-1 promoted neuronal differentiation and neurite outgrowth, and the change of RhoA was the same as that in LINGO-1, implying this process possibly realized via RhoA inactivation [10, 33, 36]. Consistently, enhanced NSC proliferation and neuronal differentiation was observed after LINGO-1 blockade by miR-615 and consequent RhoA decrease, suggesting miR-615 may regulate NSC differentiation through LINGO-1/RhoA pathway. Furthermore, EGFR, a critical receptor involved in multiple cellular events of CNS development and injuries, was upregulated. Studies suggested that endogenous LINGO-1 could directly reduce EGFR level or inhibit its phosphorylation, and hence suppress the EGFR/PI3-K/AKT pathway, resulting in decreased neuronal survival and growth [37, 38]. Activated EGFR signaling sustained the self-renewal capability of NSCs, and heightened its proliferation, survival, and migration [39,40,41]. Thus, the increased EGFR may contribute to NSC proliferation here. Myelin inhibitors trigger phosphorylation of EGFR in a calcium-dependent manner and lead to signaling inhibition of axon outgrowth [29], thus stimulating us to hypothesize whether the generation of neurons with short process attributes to EGFR upregulation. Antagonism of EGFR using AG1478 or PD168393 (EGFR inhibitor) could neutralize inhibitory influences, and promote neurite outgrowth and axonal regeneration [29, 42, 43]. In our experiments, overexpression of miR-615 resulted in heightened EGFR level and increase of βIII tubulin-positive cells with short processes. Notably, EGFR blocker PD168393 significantly facilitated neurite extension of those neurons, which is consistent with studies that PD168393 enhances elongation of HNPC processes [29]. These results indicated miR-615 may promote generation of neurons with short process from NSCs via EGFR upregulation resulting from LINGO-1 inhibition. But others recently pointed the disinhibition of neurite outgrowth was as a consequence of the off-target effects of EGFR antagonist, rather than its direct inhibitory roles [44]. Therefore, whether the influence of LINGO-1 inhibition by miR-615 on NSCs actually correlates with RhoA or EGFR signaling pathway has not been studied in full and requires further investigation.
Above all, we have provided supporting evidence that miR-615 is a potential miRNA regulating NSCs via targeting LINGO-1. To further explore the modulation of LINGO-1 by miR-615 in vivo, miR-615 agomir was constructed and transplanted into SCI rats. As expected, miR-615 effectively inhibited LINGO-1 expression in spinal tissues. Trauma to the spinal cord causes damage to axonal tracts responsible for motor and sensory functions. Although certain axonal regeneration occurs following SCI, it is insufficient to compensate for the destructive neural network and neurological dysfunction. LINGO-1 expressed in spinal cord axonal tracts and a 5-fold increase of LINGO-1 mRNA was observed at 2 weeks post-injury [45]. Attenuation of LINGO-1 function using LINGO-1 RNAi, anti-LINGO-1 antibodies, or soluble LINGO-1 all resulted in enhanced axonal regrowth, improved plasticity by promoting sprouting from intact collateral axons, and consequently promoted functional restoration of SCI animals [5, 6, 10]. In the present study, an obviously increased BBB scores were detected in rats treated with miR-615 agomir, and accompanied with substantial regenerative nerve fibers that partly recovered the tissue continuity. LINGO-1 binds with NgR1 complex in neurons and axons to inhibit neuronal survival and axonal regeneration by activation of RhoA pathway [46]. Blockade of LINGO-1 could enhance neural survival and improve axonal plasticity through the promotion of neuron sprouting post-SCI. Here, miRNA-615 enhanced neuronal survival surrounding the lesion and improved axonal regeneration, which was demonstrated by the significantly increased FG-positive neurons with capability of retrograde transportation through functional axons. However, the long-distance outgrowth of axons is inconsistent with the generation of neurons with short process during early differentiation in vitro. The difference of microenvironment in vivo and in vitro, as well as the action time, may be a possible explanation.
In addition to the intrinsic regenerative incompetence of mature neurons, the failure of axonal regeneration is also attributed to the non-permissive environment such as myelin-associated inhibitors and glial scar formation [47, 48]. Despite the neuroprotective and reparative roles in acute stages of SCI, glial scar is well known for its detrimental effects, which poses a long-lasting physical and chemical barrier on axonal regrowth at latter time after injury [48]. Attenuation of glial scar formation has shown an overt promotion of axon regeneration [49,50,51]. In this study, miR-615 reduced the glial scar formation and significantly enhanced the regeneration of neurofilament-positive fibers across the glial scar. To more deeply investigate the regeneration of nerve fibers into glial scar, CLARITY, an emerging technique that achieves transparency of intact tissue, was conducted. Via the three-dimensional structural scanning using confocal microscope, we observed plentiful regenerative nerve fibers went through the glial scar for out-extension within the damage center in the miR-615 agomir group, whereas axonal regeneration was largely blocked by the dense glial scar in the control rats. These data suggested miR-615 regulates glial scar formation and facilitates axonal regeneration following SCI. But the underlying mechanism still needs to be further deeply explored.
Remyelination is an indispensable repair process for SCI. Spontaneous remyelination following injury usually fails in most case, and the persistent demyelination ultimately results in progressive and irreversible neurological deficits and function loss [52]. LINGO-1 selectively expressed on oligodendrocyte progenitor cells (OPCs) and oligodendrocytes, where it plays an important inhibitory role in oligodendrocyte differentiation and axonal myelination. Blockade of LINGO-1 in experimental autoimmune encephalomyelitis (EAE) or SCI mice significantly increased myelinated axons within spinal cord, and led to notable functional improvement correlated with the enhanced axonal remyelination [4, 6, 8]. In line with these studies, our present data showed LINGO-1 silencing by miRNA-615 was a promising approach to improve axonal remyelination and functional rehabilitation, as evidenced by the better myelination via EM observation and upregulation of CNPase, an essential myelin-associated protein.
Conclusion
In summary, we have identified miRNA-615 as a potential miRNA regulating LINGO-1, and indicated the negative modulation of LINGO-1 by miRNA-615 in NSCs and traumatic SCI rats. Overexpression of miRNA-615 in NSCs enhanced its proliferation and neuron differentiation via LINGO-1 inhibition. Furthermore, transplantation of miRNA-615 into SCI rats effectively blocked LINGO-1, and contributed to axonal regeneration and functional rehabilitation. These results suggest miRNA-615 is a promising therapeutic target for LINGO-1 modulation and behavior recovery following SCI. However, the potential mechanisms underlying the generation of neurons with short process deserve further investigation. More importantly, it is certain that other factors regulating LINGO-1 exist and more efforts are needed to clarify the interactive effects.
References
Yang H, Liu CC, Wang CY, Zhang Q, An J, Zhang L, Hao DJ (2016) Therapeutical strategies for spinal cord injury and a promising autologous astrocyte-based therapy using efficient reprogramming techniques. Mol Neurobiol 53(5):2826–2842. https://doi.org/10.1007/s12035-015-9157-7
Salewski RP, Mitchell RA, Shen C, Fehlings MG (2015) Transplantation of neural stem cells clonally derived from embryonic stem cells promotes recovery after murine spinal cord injury. Stem Cells Dev 24(1):36–50
Zhang Z, Xu X, Zhang Y, Zhou J, Yu Z, He C (2009) LINGO-1 interacts with WNK1 to regulate Nogo-induced inhibition of neurite extension. J Biol Chem 284(23):15717–15728. https://doi.org/10.1074/jbc.M808751200
Wu H-F, Cen J-S, Zhong Q, Chen L, Wang J, Deng DY, Wan Y (2013) The promotion of functional recovery and nerve regeneration after spinal cord injury by lentiviral vectors encoding Lingo-1 shRNA delivered by Pluronic F-127. Biomaterials 34(6):1686–1700
Ji B, Li M, Wu WT, Yick LW, Lee X, Shao Z, Wang J, So KF et al (2006) LINGO-1 antagonist promotes functional recovery and axonal sprouting after spinal cord injury. Mol Cell Neurosci 33(3):311–320. https://doi.org/10.1016/j.mcn.2006.08.003
Mi S, Hu B, Hahm K, Luo Y, Kam HE, Yuan Q, Wong WM, Wang L et al (2007) LINGO-1 antagonist promotes spinal cord remyelination and axonal integrity in MOG-induced experimental autoimmune encephalomyelitis. Nat Med 13(10):1228–1233. https://doi.org/10.1038/nm1664
Andrews JL, Fernandez-Enright F (2015) A decade from discovery to therapy: Lingo-1, the dark horse in neurological and psychiatric disorders. Neurosci Biobehav Rev 56:97–114. https://doi.org/10.1016/j.neubiorev.2015.06.009
Zhang Y, Zhang YP, Pepinsky B, Huang G, Shields LB, Shields CB, Mi S (2015) Inhibition of LINGO-1 promotes functional recovery after experimental spinal cord demyelination. Exp Neurol 266:68–73. https://doi.org/10.1016/j.expneurol.2015.02.006
Lööv C, Fernqvist M, Walmsley A, Marklund N, Erlandsson A (2012) Neutralization of LINGO-1 during in vitro differentiation of neural stem cells results in proliferation of immature neurons. PLoS One 7(1):e29771. https://doi.org/10.1371/journal.pone.0029771.g001
Chen N, Cen JS, Wang J, Qin G, Long L, Wang L, Wei F, Xiang Q et al (2016) Targeted inhibition of leucine-rich repeat and immunoglobulin domain-containing protein 1 in transplanted neural stem cells promotes neuronal differentiation and functional recovery in rats subjected to spinal cord injury. Crit Care Med 44(3):e146–e157. https://doi.org/10.1097/CCM.0000000000001351
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136(2):215–233. https://doi.org/10.1016/j.cell.2009.01.002
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105. https://doi.org/10.1101/gr.082701.108
Liu NK, Wang XF, Lu QB, Xu XM (2009) Altered MicroRNA expression following traumatic spinal cord injury. Exp Neurol 2(219):424–429. https://doi.org/10.1016/j.expneurol.2009.06.015
Strickland ER, Hook MA, Balaraman S, Huie JR, Grau JW, Miranda RC (2011) MicroRNA dysregulation following spinal cord contusion: implications for neural plasticity and repair. Neuroscience 186:146–160. https://doi.org/10.1016/j.neuroscience.2011.03.063
Hutchison ER, Okun E, Mattson MP (2009) The therapeutic potential of microRNAs in nervous system damage, degeneration, and repair. NeuroMolecular Med 11(3):153–161. https://doi.org/10.1007/s12017-009-8086-x
Hu JZ, Huang JH, Zeng L, Wang G, Cao M (2013) Anti-apoptotic effect of MicroRNA-21 after contusion spinal cord injury in rats. J Neurotrauma 30:1349–1360. https://doi.org/10.1089/neu.2012.2748
Liu G, Keeler BE, Zhukareva V, Houle JD (2010) Cycling exercise affects the expression of apoptosis-associated microRNAs after spinal cord injury in rats. Exp Neurol 226(1):200–206. https://doi.org/10.1016/j.expneurol.2010.08.032 Copyright (c) 2010 Elsevier Inc. All rights reserved
Ji Y, Sun Q, Zhang J, Hu H (2018) MiR-615 inhibits cell proliferation, migration and invasion by targeting EGFR in human glioblastoma. Biochem Biophys Res Commun 499(3):719–726. https://doi.org/10.1016/j.bbrc.2018.03.217
Yang B, Xie R, Wu SN, Gao CC, Yang XZ, Zhou JF (2018) MicroRNA-615-5p targets insulin-like growth factor 2 and exerts tumor-suppressing functions in human esophageal squamous cell carcinoma. Oncol Rep 39(1):255–263. https://doi.org/10.3892/or.2017.6079
Icli B, Wu W, Ozdemir D, Li H, Cheng HS, Haemmig S, Liu X, Giatsidis G et al (2019) MicroRNA-615-5p regulates angiogenesis and tissue repair by targeting AKT/eNOS (protein kinase B/endothelial nitric oxide synthase) signaling in endothelial cells. Arterioscler Thromb Vasc Biol 39(7):1458–1474. https://doi.org/10.1161/atvbaha.119.312726
Tripathi R, Saini HK, Rad R, Abreugoodger C, Van DS (2011) Messenger RNA and microRNA profiling during early mouse EB formation. Gene Expr Patterns 11:334–344. https://doi.org/10.1016/j.gep.2011.03.004
Stallings RL, Foley NH, Bray IM, Das S, Buckley PG (2011) MicroRNA and DNA methylation alterations mediating retinoic acid induced neuroblastoma cell differentiation. Semin Cancer Biol 21:283–290. https://doi.org/10.1016/j.semcancer.2011.07.001
Ji HP, Halder D, Mi RC, Jin CC, Lee YS (2012) Expression profiles of miRNAs during ethanol-induced differentiation of neural stem cells. BioChip J 1(6):73–83. https://doi.org/10.1007/s13206-012-6110-y
Nuneziglesias J, Liu CC, Morgan TE, Finch CE, Zhou XJ (2010) Joint genome-wide profiling of miRNA and mRNA expression in Alzheimer's disease cortex reveals altered miRNA regulation. PLoS One 2(5):e8898
Kojima T, Ueda Y, Sato A, Sameshima H, Ikenoue T (2013) Comprehensive gene expression analysis of cerebral cortices from mature rats after neonatal hypoxic-ischemic brain injury. J Mol Neurosci 49:320–327. https://doi.org/10.1007/s12031-012-9830-5
Roshan R, Ghosh T, Scaria V, Pillai B (2009) MicroRNAs: novel therapeutic targets in neurodegenerative diseases. Drug Discov Today 14(23–24):1123–1129. https://doi.org/10.1016/j.drudis.2009.09.009
Jasmin BJ, Campbell RJ, Michel RN (1995) Nerve-dependent regulation of succinate dehydrogenase in junctional and extrajunctional compartments of rat muscle fibres. J Physiol 484(Pt 1):155–164
Tomer R, Ye L, Hsueh B, Deisseroth K (2014) Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat Protoc 9(7):1682–1697. https://doi.org/10.1038/nprot.2014.123
Koprivica V, Cho K-S, Park JB, Yiu G, Atwal J, Gore B, Kim JA, Lin E et al (2005) EGFR activation mediates inhibition of axon regeneration by myelin and chondroitin sulfate proteoglycans. Science 310(5745):106–110
Woltering JM, Durston AJ (2008) MiR-10 represses HoxB1a and HoxB3a in zebrafish. PLoS One 3(1):e1396. https://doi.org/10.1371/journal.pone.0001396
Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, Johnstone S, Guenther MG, Johnston WK et al (2008) Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134(3):521–533. https://doi.org/10.1016/j.cell.2008.07.020
Aranda P, Agirre X, Ballestar E, Andreu EJ, Roman-Gomez J, Prieto I, Martin-Subero JI, Cigudosa JC et al (2009) Epigenetic signatures associated with different levels of differentiation potential in human stem cells. PLoS One 4(11):e7809. https://doi.org/10.1371/journal.pone.0007809
Wang J, Ye Z, Zheng S, Chen L, Wan Y, Deng Y, Yang R (2016) Lingo-1 shRNA and notch signaling inhibitor DAPT promote differentiation of neural stem/progenitor cells into neurons. Brain Res 1634:34–44. https://doi.org/10.1016/j.brainres.2015.11.029
Li X, Zhang Y, Yan Y, Ciric B, Ma CG, Chin J, Curtis M, Rostami A et al (2017) LINGO-1-fc-transduced neural stem cells are effective therapy for chronic stage experimental autoimmune encephalomyelitis. Mol Neurobiol 54(6):4365–4378. https://doi.org/10.1007/s12035-016-9994-z
Wang JJ, Liu C, Shan K, Liu BH, Li XM, Zhang SJ, Zhou RM, Dong R et al (2018) Circular RNA-ZNF609 regulates retinal neurodegeneration by acting as miR-615 sponge. Theranostics 8(12):3408–3415. https://doi.org/10.7150/thno.25156
Gu H, Yu SP, Gutekunst CA, Gross RE, Wei L (2013) Inhibition of the rho signaling pathway improves neurite outgrowth and neuronal differentiation of mouse neural stem cells. Int J Physiol Pathophysiol Pharmacol 5(1):11–20
Inoue H, Lin L, Lee X, Shao Z, Mendes S, Snodgrass-Belt P, Sweigard H, Engber T et al (2007) Inhibition of the leucine-rich repeat protein LINGO-1 enhances survival, structure, and function of dopaminergic neurons in Parkinson’s disease models. Proc Natl Acad Sci U S A 104(36):14430–14435. https://doi.org/10.1073/pnas.0700901104
Oda K, Matsuoka Y, Funahashi A, Kitano H (2005) A comprehensive pathway map of epidermal growth factor receptor signaling. Mol Syst Biol 1:2005 0010. https://doi.org/10.1038/msb4100014
Ayuso-Sacido A, Moliterno JA, Kratovac S, Kapoor GS, O'Rourke DM, Holland EC, Garcia-Verdugo JM, Roy NS et al (2010) Activated EGFR signaling increases proliferation, survival, and migration and blocks neuronal differentiation in post-natal neural stem cells. J Neuro-Oncol 97(3):323–337. https://doi.org/10.1007/s11060-009-0035-x
Li X, Xiao Z, Han J, Chen L, Xiao H, Ma F, Hou X, Li X et al (2013) Promotion of neuronal differentiation of neural progenitor cells by using EGFR antibody functionalized collagen scaffolds for spinal cord injury repair. Biomaterials 34(21):5107–5116. https://doi.org/10.1016/j.biomaterials.2013.03.062
Wang J, Yu RK (2013) Interaction of ganglioside GD3 with an EGF receptor sustains the self-renewal ability of mouse neural stem cells in vitro. Proc Natl Acad Sci U S A 110(47):19137–19142. https://doi.org/10.1073/pnas.1307224110
Novozhilova E, Englund-Johansson U, Kale A, Jiao Y, Olivius P (2015) Effects of ROCK inhibitor Y27632 and EGFR inhibitor PD168393 on human neural precursors co-cultured with rat auditory brainstem explant. Neuroscience 287:43–54. https://doi.org/10.1016/j.neuroscience.2014.12.009
Han Q, Jin W, Xiao Z, Ni H, Wang J, Kong J, Wu J, Liang W et al (2010) The promotion of neural regeneration in an extreme rat spinal cord injury model using a collagen scaffold containing a collagen binding neuroprotective protein and an EGFR neutralizing antibody. Biomaterials 31(35):9212–9220. https://doi.org/10.1016/j.biomaterials.2010.08.040
Douglas MR, Morrison KC, Jacques SJ, Leadbeater WE, Gonzalez AM, Berry M, Logan A, Ahmed Z (2009) Off-target effects of epidermal growth factor receptor antagonists mediate retinal ganglion cell disinhibited axon growth. Brain 132(Pt 11):3102–3121. https://doi.org/10.1093/brain/awp240
Mi S, Lee X, Shao Z, Thill G, Ji B, Relton J, Levesque M, Allaire N et al (2004) LINGO-1 is a component of the Nogo-66 receptor/p75 signaling complex. Nat Neurosci 7(3):221–228. https://doi.org/10.1038/nn1188
Lu Y, Liu X, Zhou J, Huang A, Zhou J, He C (2013) TROY interacts with Rho guanine nucleotide dissociation inhibitor alpha (RhoGDIalpha) to mediate Nogo-induced inhibition of neurite outgrowth. J Biol Chem 288(47):34276–34286. https://doi.org/10.1074/jbc.M113.519744
Sharma K, Selzer ME, Li S (2012) Scar-mediated inhibition and CSPG receptors in the CNS. Exp Neurol 237(2):370–378. https://doi.org/10.1016/j.expneurol.2012.07.009
Yuan YM, He C (2013) The glial scar in spinal cord injury and repair. Neurosci Bull 29(4):421–435. https://doi.org/10.1007/s12264-013-1358-3
Dyck SM, Karimi-Abdolrezaee S (2015) Chondroitin sulfate proteoglycans_ Key modulators in the developing and 3 pathologic central nervous system. Exp Neurol 269:169–187. https://doi.org/10.1016/j.expneurol.2015.04.006
Mckillop WM, Dragan M, Schedl A, Brown AA (2013) Conditional Sox9 ablation reduces chondroitin sulfate proteoglycan levels and improves motor function following spinal cord injury. Glia 61:164–177. https://doi.org/10.1002/glia.22424
Lang BT, Cregg JM, DePaul MA, Tran AP, Xu K, Dyck SM, Madalena KM, Brown BP et al (2015) Modulation of the proteoglycan receptor PTPs promotes recovery after spinal cord injury. Nature 518(7539):404–408. https://doi.org/10.1038/nature13974
Mi S, Pepinsky RB, Cadavid D (2013) Blocking LINGO-1 as a therapy to promote CNS repair: from concept to the clinic. CNS Drugs 27(7):493–503. https://doi.org/10.1007/s40263-013-0068-8
Funding
This work was financially supported by the National Natural Science Foundation of China, No. 81371366 (to HFW); the Characteristic Innovation Project of Colleges and Universities in Guangdong Province, No. 2018KTSCX075 (to HFW); the Key Project of Social Development of Dongguan of China, No. 20185071521640 (to HFW); College Students’ Science and Technology Innovation Training Project, No. 201810571058, No. GDMU2018024, No. GDMU2018056, No. GDMU2018061 (to HFW); College Students’ Innovative Experimental Project in Guangdong Medical University, No. ZZDS001 (to HFW); and College Students’ Science and Technology Innovation Cultivation Project in Guangdong, No. pdjh2019b0217 (to HFW).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Research Involving Human Participants and/or Animals
This article does not contain any studies with human participants performed by any of the authors.
Ethical Approval
All procedures using laboratory animals were conducted in compliance with the Guide for the Care and Use of Laboratory Animals (National Research Council, 1996) and approved by the Animal Care and Use Committee of Guangdong Medical University at which the studies were conducted.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wu, H., Ding, L., Wang, Y. et al. MiR-615 Regulates NSC Differentiation In Vitro and Contributes to Spinal Cord Injury Repair by Targeting LINGO-1. Mol Neurobiol 57, 3057–3074 (2020). https://doi.org/10.1007/s12035-020-01936-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-020-01936-z