Abstract
Maged1 is a member of the type II melanoma antigen (MAGE) family of proteins, which is highly conserved in the brain between mouse and human. Recently, Maged1 has been reported to be involved in depression and impaired sexual behavior. However, the role of Maged1 in learning and memory remains unknown. The aim of the present study was therefore to investigate whether Maged1 deficiency can impair learning and memory formation. By behavioral tests and electrophysiological recording, we observed that 5–6-month-old Maged1 knockout mice displayed the reduced basal synaptic transmission, pronounced hippocampal dysfunction, impaired spatial learning, and a deficit in long-term potentiation induction. Data from immunohistochemical and Western blot showed the reduced dendritic spine density and the number of synapses in the hippocampus of the Maged1 knockout mice, and Maged1 deficiency prevented the interaction of Maged1 with cAMP response element-binding protein (CREB). Furthermore, by chromatin immunoprecipitation and luciferase assay, we observed the downregulated activity of CREB and the suppressed CREB-dependent transcription after deficiency of Maged1, which lead to the decreased levels of brain-derived neurotrophic factor. Taken together, our results provide the evidence that Maged1 is involved in synaptic transmission and hippocampus-dependent learning and memory formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The melanoma antigen (MAGE) family of proteins is well known for being tumor-specific antigens and comprised of more than 60 genes [1]. These genes (>50) share a highly conserved ~190-amino-acid MAGE homology domain (MHD) [2, 3]. The distinct expression patterns of MAGE genes divide the MAGE family into two types: type I and type II. Type I MAGEs are highly expressed cancer antigens and play an important role in tumorigenesis and cancer cell survival [4, 5]. Type II MAGEs are widely expressed in a variety of tissues during development [6]. The structure of type II MAGE proteins is more complicated: in addition to MHD, many contain other domains that provide these proteins with other potential functions. Maged1 is a member of type II MAGEs, which is first identified as a binding partner to the intracellular domain of p75 neurotrophin receptor facilitating neuronal apoptosis [7, 8]. It has also been reported to act as a transcriptional regulator by interacting with transcription factors and thereby mediating several functions ranging from apoptosis, cell-cycle progression, cell adhesion, and angiogenesis to developmental morphogenesis [7, 9–14].
Maged1 is expressed throughout the neural tube during the early stage of neurogenesis and is restricted within the ventricular zone, subplate, and cortical plate during the later stage of neurogenesis [15]. Recent studies have suggested a functional relevance for Maged1 activity in the brain [16–18]. Maged1 interacts with a specific group of nuclear receptors that regulate the amplitude of the mammalian circadian clock [16] and is involved in depression-like behavior through modulating the ubiquitylation of serotonin transporters [17], whereas loss of Maged1 results in deficits of social interactions and impairs sexual behavior by reducing the production of mature oxytocin in the hypothalamus of mice [18]. Thus, we hypothesized that Maged1 is involved in cognition function, which remains largely to be determined. The purpose of the present study was to investigate whether Maged1 deficiency can impair synaptic plasticity, learning, and memory.
Materials and Methods
Animals
The male Maged1 knockout (Maged1−/−) mice used in the present study have been previously described [15]. Maged1 is X-linked; therefore, Maged1−/− and wild-type (WT) male littermates are generated by crossing Maged1+/− females with WT C57BL/6 J males. All experiments were done with mice backcrossed for >15 generations on a C57BL/6 J genetic background. Animal studies were carried out in an Association for Assessment and Accreditation of Laboratory Animal Care International-credited specific pathogen-free animal facility, and all animal protocols were approved by the Animal Care and Use Committee of the Model Animal Research Center.
Electrophysiological Analysis
Electrophysiological recording was performed as previously described [19]. Briefly, transverse hippocampal slices (400 μm thick) were prepared from 5–6-month-old male mice. After decapitation, the brain was removed and placed in oxygenated (95 % O2/5 % CO2) artificial cerebrospinal fluid (ACSF) at 4 °C. Slices were cut using a Leica VT1000S vibratome (Leica Instruments Ltd., Wetzlar, Germany) and maintained at 32 °C for 1 h in a holding chamber filled with oxygenated ACSF. After an equilibration period of at least 2 h at room temperature, a single slice was transferred to the recording chamber where it was held between two nylon nets and continuously perfused with oxygenated ACSF (23–25 °C) at a flow rate of 2.5–3 ml/min.
Golgi Impregnation
Golgi-Cox-stained brains were cut to 200-μm-thick cross sections using a vibratome and analyzed using the Olympus FV1000 microscope and Olympus software. The number of spines on the apical and basal dendrites of hippocampal CA1 pyramidal neurons was counted by a researcher who was blinded to the genotype. For each experimental group, a minimum of ten cells per slice were analyzed.
Immunohistochemistry
Immunohistochemical analysis was performed as described previously [19]. Briefly, anti-synaptophysin (SVP)-38 antibodies were used at a 1:2,000 dilution (Sigma) and confocal images (1 μm) were scanned from brain sections. Brain sections that had the strongest intensity were scanned first. All other images included in the analysis were scanned using the same settings. Staining was quantified using the Leica software.
Immunoprecipitation and Immunoblotting
The hippocampus was collected and lysed. For immunoprecipitation, tissue homogenates (400 μg of protein) were diluted fourfold with immunoprecipitation buffer containing as described before [20]. Samples were preincubated for 1 h with 25 μl protein A/G-Sepharose CL-4B (Amersham Pharmacia, Buckinghamshire, UK) and then centrifuged to remove any nonspecific adhesion to the A/G. The supernatant was incubated with 2–4 μg of anti-cAMP response element-binding protein (CREB) (Cell Signaling, 9104) antibody for 4 h or overnight at 4 °C. Protein A/G-Sepharose (25 μl) was then added, and the incubation continued for 2 h. Samples were centrifuged at 4,000 × g, and the pellets were washed three times with immunoprecipitation buffer. Bound proteins were eluted by adding 2× sample buffer and boiled at 100 °C for 5 min. Samples were then centrifuged, and supernatants were used for immunoblotting.
For immunoblotting, tissue homogenates were subjected to 10 % SDS-PAGE and electrotransferred onto a PVDF membrane (Amersham Pharmacia). The membrane was probed with anti-SVP (1:5,000, Sigma), anti-brain-derived neurotrophic factor (BDNF) (1:500, Santa Cruz), anti-CREB (1:1,000, Cell Signaling), β-actin (1:500, Santa Cruz), or GAPDH (1:1,000, Santa Cruz) antibodies at 4 °C overnight.
Real-Time PCR
Total RNA was extracted using TRIzol (Invitrogen), and random hexamers were used to prime reverse-transcription reactions with SuperScript III (Invitrogen). Real-time quantitative PCR was performed using an ABI 7300 detection system (Applied Biosystems) with SYBR Green I reagents (Takara). The primers were the same as reported earlier [19], and all other primers, including primers for real-time PCR, are listed below. Transcript levels for each gene were normalized to GAPDH cDNA levels according to standard procedures. Data were derived from three independent amplifications.
BDNF: 5′-atgggactctggagagcctgaa-3′; 3′-gccagccaattctctttttgc-5′
Mecp2: 5′-aagtctggccgatctgctgg-3′; 3′-atttgggcttcttaggtggtttc-5′
Bmal1: 5′-tggagggactccagacattc-3′; 3′-tgggactactggatccttgg-5′
CREB: 5′-ctacaccagcttccccggt-3′; 3′-acggaaacagccgagctc-5′
Maged1: 5′-agaacttacgaccctcaccta-3′; 3′-aagataatgcccagaatcact-5′
GAPDH: 5′-ctcccaggaagaccctgctt-3′; 3′-ggaacagggaggagcagaga-5′
Chromatin Immunoprecipitation
Chromatin immunoprecipitation (ChIP) was performed with mouse brain fixed with 4 % paraformaldehyde (PFA) solution and stored at −80 °C before use. Brains were cut into small (~1 mm3) pieces and chemically cross-linked by 1 % PFA in PBS solution for 15 min at room temperature and then homogenized, resuspended, and lysed in TNN lysis buffer (100 mM Tris/HCl pH 8.0, 250 mM NaCl, 0.5 % NP-40) with RNAse, followed by digestion of chromatin DNA with micrococcal nuclease (Neb). TNN lysates were then incubated with nuclease for 10–15 min at 37 °C (1 μl, 2,000 U) plus 1 ml lysate plus 10 μl of 0.5 M CaCl2. The reaction was stopped with 100 μl of 0.5 M EDTA on ice and then sonicated to shear cross-linked DNA. We used a Branson S450 digital sonifier at 20 % power and delivered ten 30-s pulses (60-s pause between pulses) during which all samples were immersed in an ice bath. The resulting whole-cell extract was incubated overnight at 4 °C with 30 μl of Dynal Protein G magnetic beads that had been preincubated with 5 μg of the appropriate antibody. Beads were washed five times with TNN lysis buffer and once with 1× ChIP buffer. Bound complexes were eluted from the beads by heating to 65 °C. Whole-cell extract DNA (reserved from the sonication step) was treated to reverse cross-link. Immunoprecipitated DNA and whole-cell extract DNA were then purified by treatment with 50 mM NaCl and proteinase K for 2 h at 65 °C with rotation. Purified DNA samples were normalized and subjected to PCR analysis. Antibodies used for pulldowns were CREB which was from Cell Signaling and Maged1 which was from Oncogene. After immunoprecipitation, recovered chromatin fragments were subjected to semiquantitative PCR or real-time PCR for 40 cycles, using primer pairs specific for 150–500-bp segments as described in Table 1.
Cell Cultures and Transient Transfection
HEK293 cells were grown in DMEM medium supplemented with 10 % fetal bovine serum at 37 °C and 5 % CO2 as described previously. The cells were transiently transfected with equal molar amounts of various luciferase-fusion constructs or pGL3-basic in serum-free growth media using Lipofectamine-Plus reagent according to the manufacturer’s protocol (Invitrogen). pRL-tK (Promega) containing the Renilla luciferase gene was cotransfected in order to normalize transfection efficiencies. For experiments that examined the effects of exogenously expressed Maged1, mutant Maged1 fragment, luciferase-fusion reporter constructs, and pRL-tK were cotransfected. The transfected cells were then lysed 48 h after transfection, and the luciferase activities of the cells’ lysates were determined using the dual-luciferase reporter assay system (Promega).
Fear-Conditioning Tests
Context-Dependent Fear Conditioning
Training consists of a 3-min exposure of mice to the conditioning box (context) followed by a foot shock (2 s, 0.7 mA). The memory test was performed 24 h later by reexposing the mice for 3 min into the conditioning context. Freezing, defined as a lack of movement except for breathing associated with a crouching posture, was recorded every 10 s by two trained observers (who were blinded to the experimental conditions) over a period of 3 min (a total of 18 sampling intervals). The number of observations indicating freezing obtained as a mean from both observers was expressed as a percentage of the total number of observations.
Tone-Dependent Fear Conditioning
Training consisted of a 3-min exposure of mice to the conditioning box, followed by a tone (30 s, 20 kHz, 75 dB) and a foot shock (2 s, 0.7 mA). The memory test was performed 24 h later by exposing the mice for 3 min to a novel context followed by an additional 3-min exposure to a tone (10 kHz, 75 dB). Freezing was recorded every 10 s by the two nonbiased observers as described above.
Morris Water Maze Test
The water maze paradigm was performed in a circular tank (diameter 1.2 m) filled with opaque water. A platform (11 cm × 11 cm) was submerged below the water’s surface in the center of the target quadrant. The swimming path of the mice was recorded by a video camera and analyzed by software. For each training session, the mice were placed into the maze consecutively from four random points of the tank. Mice were allowed to search for the platform for 60 s. If the mice did not find the platform within 60 s, they were gently guided to it. Mice were allowed to remain on the platform for 15 s. Three training trials were performed every day, and the latency for each trial was recorded for analysis. During the memory test (probe test), the platform was removed from the tank, and the mice were allowed to swim in the maze for 60 s.
Statistical Analysis
Data are shown as means ± SEM. Comparisons were performed by Student’s t test or two-way analysis of variance followed by the Duncan new multiple range method or Newman-Keuls test, as appropriate. A p value of <0.05 was considered as statistically significant.
Results
Deficiency in Maged1 Reduced Learning and Memory Formation
To directly evaluate the physiological role of Maged1 in the brain, the Maged1−/− mice were trained using Pavlovian fear-conditioning paradigms prior to a memory test 24 h later. The Maged1−/− mice showed the decreased freezing behavior in both context- and tone-dependent fear learning compared with the WT mice (Fig. 1a, b, p = 0.036 and p = 0.015, respectively). The reduction in freezing behavior of the Maged1−/− mice was not attributed to motor defects or impaired pain sensation because the WT mice and the Maged1−/− mice had similar responses to electric foot shock at baseline and after training (Fig. 1c).
Maged1 deficiency reduces learning and memory formation. a, b Maged1−/− mice showed significantly decreased freezing behavior during the context- and tone-dependent memory tests compared with the WT mice (Maged1−/−, n = 14; WT, n = 19). c The responses to electric shock (ES) did not differ between the WT mice and Maged1−/− mice at baseline and after training. d The Maged1−/− mice showed increased escape latency throughout the training process in the Morris water maze at days 5 and 6 compared with the WT mice. e The Maged1−/− mice spent less time in the target quadrant at day 7 than the WT mice (O, opposite quadrant; R, right quadrant; L, left quadrant; T, target quadrant). f Visual function and swimming ability were not affected in the Maged1−/− mice. Data are shown as means ± SEM; *p < 0.05 compared with the WT mice
To further evaluate the integrity of hippocampus-dependent memory formation in the Maged1−/− mice, the Morris water maze paradigm was utilized. The Maged1−/− mice showed an increased escape latency throughout the training process at days 5 and 6 (Fig. 1d, p = 0.041 and p = 0.031, respectively). A probe trial was further performed to quantify the time spent in each quadrant of the swimming pool when the hidden platform was removed from the pool. The Maged1−/− mice spent less time in the target quadrant than the WT mice at day 7 (Fig. 1e, p = 0.042). Comparable visual and motor functions were observed between the Maged1−/− and WT mice in the visible platform test (Fig. 1f).
Deficiency in Maged1 Decreased Dendritic Spine Density
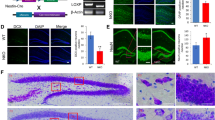
The synapse is widely assumed to be the cellular site for learning and memory. We therefore assessed whether Maged1 regulates the density of dendritic spines and synapse numbers. The density of dendritic spines along individual dendrites of hippocampal CA1 pyramidal neurons and dentate gyrus granule cells was reduced in the Maged1−/− mice compared with the WT mice (p = 0.005, Fig. 2a). The spine shape was not affected by the expression of Maged1. The presynaptic terminals of functional synapses were reduced in the CA1 stratum radiatum of the Maged1−/− mice compared with the WT mice (Fig. 2b, c, p = 0.007 and p = 0.006, respectively).
Maged1 deficiency decreases dendritic spine density. a Representative images of Golgi staining in the CA1 region of the hippocampus. The density of dendritic spines was less in the Maged1−/− mice (Maged1−/−, n = 21 slices; WT, n = 23 slices; scale bar = 10 μm). b Representative confocal images of the SVP-immunoreactive signal in the CA1 region. The presynaptic terminals of functional synapses were reduced in the Maged1−/− mice (Maged1−/−, n = 17 slices; WT, n = 22 slices; scale bar = 50 μm). c Western blots from hippocampus lysate showed downregulation of SVP in the Maged1−/− mice (n = 3 per group). Data are shown as means ± SEM; ***p < 0.01 compared with the WT mice
Deficiency in Maged1 Impaired Hippocampal Synaptic Plasticity
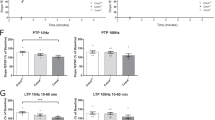
The reduction of dendrite spines in the Maged1−/− mice prompted us to investigate whether the induction of synaptic plasticity was affected in the Maged1−/− mice. We recorded field excitatory postsynaptic potentials (fEPSPs) from CA1 pyramidal cells in acute hippocampal slices from the mice. Input-output curves were determined using measurements of fEPSP slope in response to a series of stimulation intensities from 0.2 to 1.0 mA (0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, and 1 mA), demonstrating that the Maged1−/− mice exhibited a significant decrease in the input-output relationship compared with the WT mice (Fig. 3a). Secondly, we determined whether the Schaffer CA3-CA1 hippocampal collateral pathway had any deficits in activity-dependent synaptic plasticity by measuring long-term potentiation (LTP). LTP was induced by two trains of high-frequency stimulation (HFS). After recording a stable baseline, HFS was applied to fibers of the CA3 presynaptic neurons, and fEPSP was recorded extracellularly in the stratum radiatum of the CA1 region for 90 min poststimulation. Short-term synaptic plasticity, measured as posttetanic potentiation after HFS, was unchanged in the Maged1−/− mice. However, comparison of the Maged1−/− mice to the WT mice revealed a significant decrease in LTP 90 min after LTP induction (WT: 136.4 % versus Maged1−/−: 101.9 % at 90 min after HFS application, p = 0.021; Fig. 3c). No significant difference was observed in paired-pulse facilitation between the WT mice and Maged1−/− mice (Fig. 3b).
Maged1 deficiency impaired hippocampal synaptic plasticity. a The amplitude of the evoked fEPSP was significantly decreased in the Maged1−/− mice compared with the WT mice. b No difference was observed in paired-pulse facilitation (PPF) between the Maged1−/− mice and WT mice. c LTP induced by two HFS in the CA1 region was impaired in the Maged1−/− mice. By 40 min, the fEPSP from the Maged1−/− mice decayed to the baseline (n = 9 slices, 102.2 ± 5.4 % compared with baseline) whereas fEPSP from the WT mice remained potentiated (n = 9 slices, 130.6 ± 4.9 % compared with baseline). Insert in c shows superimposed sample sweeps from the first 5 min (gray) and last 5 min (black) of the recording. Data are shown as means ± SEM; *p < 0.05 compared with the WT mice
Deficiency in Maged1 Regulated CREB Activity
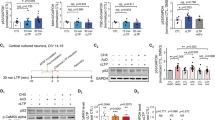
To further gain insight into the molecular mechanism underlying the plasticity and memory impairments of the Maged1−/− mice, we examined the expression of genes involved in learning and memory. In particular, we focused on BDNF and CREB, as these two genes have been shown to play critical roles in synaptic plasticity and, at the cellular level, to positively modulate synapse formation [21]. We found that both mRNA and protein levels of BDNF were significantly decreased in the Maged1−/− mice hippocampus compared with the WT mice hippocampus (Fig. 4a, b), while the mRNA and protein levels of CREB were not changed in the Maged1−/− mice (Fig. 4a, c). We next assayed the interaction between CREB proteins and Maged1. We observed the co-immunoprecipitation of Maged1 with CREB in the brain of the WT mice, but not in the Maged1−/− mice (Fig. 4d). Because the expression of CREB was not altered in the Maged1−/− mice, one possible explanation for the decreased levels of BDNF is that CREB activity is downregulated. To determine if Maged1 regulated the CREB activity, we carried out a cell line-based CREB activity reporter assay using a luciferase reporter driven by a minimal promoter with CREB-binding upstream elements. Compared with co-transfection of vector alone, co-transfection of Maged1 with CRE-Luc resulted in a significant increase in CREB activity that was blocked if shRNAMaged1 was delivered together with Maged1 (Fig. 4e, p = 0.001).
Maged1 enhances the transcriptional activity of CREB. a Real-time PCR showed that the level of BDNF and Bmall1 mRNA was reduced in the hippocampus from the Maged1−/− mice (n = 3 per group). b Representative western blots are shown in the upper panel. The level of BDNF was decreased in the hippocampus of the Maged1−/− mice (n = 3 per group). c The CREB level was not changed in the hippocampus of the Maged1−/− mice. d Maged1 deficiency impaired the interaction of Maged1 and CREB. No co-immunoprecipitation of Maged1 with CREB was observed in the Maged1−/− mice. e Maged1 positively regulated CREB-mediated transcriptional activity. CREB activity reporter construct was co-transfected with an empty vector or Maged1 or shRNAMaged1. Data are shown as means ± SEM; **p < 0.01 compared with the WT mice (control) in Fig. 4a, b or compared with the vector group in Fig. 4e
Deficiency in Maged1 Inhibited CREB-Dependent Transcription
We next performed ChIP and assayed, by q-PCR, the extent to which the following promoter regions were occupied by CREB: BDNF, Atf, Nrxn, Grin, Prkcz, and Erg-1. In each case, we found the decreased levels of promoter occupancy in the Maged1−/− samples (Fig. 5a). Consistent with this, in the Maged1−/− mice, the amount of Maged1 binding to BDNF1 and BDNF4 was also reduced (Fig. 5b).
Maged1 deficiency inhibits CREB-dependent transcription. a The association of CREB with its target gene in the brain was reduced in the Maged1−/− mice. Following ChIP with CREB antibody from the WT or Maged1−/− mice, a quantitative real-time PCR analysis was performed for the presence of specific CREB target genes. b After ChIP with antibodies to Maged1 from the hippocampus, a quantitative real-time PCR analysis was performed for the presence of specific BDNF1 and 4. Data are shown as means ± SEM; *p < 0.05 compared with the WT mice
Maged1 Interacted with CREB Through a Hexapeptide Repeat Region
We then investigated which domain was responsible for the activation of CREB and binding to CREB. The primary structure of Maged1 comprises a MAGE/necdin homology domain and a unique 25-hexapeptide repeat region. Mutant proteins were constructed that had deletions in the characterized domains (Fig. 6a). Deletion of the unique repeat region completely abolished the ability of Maged1 to co-activate CREB activity (Fig. 6b). Truncated Maged1 was further assayed for interaction with CREB by co-immunoprecipitation. Deletion of the hexapeptide repeats was found to block the interaction of Maged1 and CREB (Fig. 6c).
Maged1 interacts with CREB through a hexapeptide repeat region. Construct strategies for truncated Flag-tagged Maged1. b Effects of truncated Maged1 on CREB activity in HEK 293 T cells. Maged1 lacking the 25 hexapeptide repeats (Maged1 ΔB) impaired the ability to activate CRE promoter. c Co-immunoprecipitation assays of HEK 293 T cells using Flag-tagged truncated Maged1 as indicated. Deletion of the unique hexapeptide repeat domain abolished the interaction between Maged1 and CREB. Data are shown as means ± SEM; ***p < 0.001 compared with the vector group; ##p < 0.01 compared with the Maged1 group
Discussion
The MAGE genes encode many proteins that control cell cycle, differentiation, and survival. In the present study, we provided the following evidence that Maged1 was required for memory formation in mice: (1) Maged1 regulates both synapse formation and transmission. Loss of Maged1 expression reduces the number of dendritic spines. This results in reduced spatial learning capacities in the Maged1−/− mice. (2) Maged1 is further required for the regulation of CREB activity. In the absence of Maged1, the interaction of Maged1 with CREB is prevented, leading to the downregulation of CREB-dependent transcription. (3) The 25-hexapeptide repeat region in Maged1 is essential for the interaction of Maged1 with CREB. In HEK293T cells transfected with truncated Maged1, deletion of the unique repeat region completely abolished the ability of Maged1 to co-activate the CRE promoter, while the constructs in which the MAGE domain was deleted retained co-activation ability. Taken together, our data showed that Maged1 binds with CREB and regulates CREB-dependent transcription. Interfering with this binding leads to impaired learning and memory in mice.
Recent data showed that Maged1 was involved in depression-like behavior and the disruption of community interactions through its ability to regulate the serotonin transporter ubiquitylation and the mature oxytocin production, respectively [16, 17]. Consistently, similar depression-like phenotypes were observed in the present study (data not shown). Although Mouri et al. reported unchanged cognitive functions in their fear-conditioning test, this may be attributable to the differences in the age of the animals tested (5–6 months in the present study and 7 weeks in Mouri’s study) and the stimulation protocol (one foot shock of 0.7 mA in the present study and four foot shocks of 0.6 mA in Mouri’s study). Indeed, the impaired learning and memory was also observed in patients who had depression or a related disorder and impaired learning and memory is one of the clinical symptoms used for the diagnosis of these diseases. In the present study, we also observed the decreased levels of BDNF, which is one important factor in learning and memory and is reported to be reduced in the brain of patients suffering from depression. Furthermore, it was expected that there should be more neurons in the Maged1−/− mice because Maged1 is a binding partner for the p75 neurotrophin receptors [7, 8]. However, there were decreased BDNF levels, and thus a reduction of ligand, as found in the brains of Maged1−/− mice. This suggested the possibility that, in addition to the direct effects of Maged1 on serotonin transporter, oxytocin and a BDNF decrease could also contribute to the phenotypes observed in the Maged1−/− mice.
Maged1 which is discovered in bone marrow stromal cells, also known as Dlxin-1 or NRAGE, is a member of the type II MAGE-D family and contains a unique domain of 25 consecutive hexapeptide repeats with a consensus sequence of tryptophan-glutamine-x-proline-x-x (WQxPxx, where x is any amino acid) that is hypothesized to be instrumental in its function [7, 15, 22, 23]. Maged1 functions as an adaptor that mediates multiple apoptotic pathways through distinct input signals, such as bone morphogenetic protein, Ror receptor kinases, the anti-apoptotic factor CHE-1, and the axon guidance receptor UNC5H [10, 24–26]. Through binding with RING domain protein XIAP and IAP proteins, Maged1 mediates p38 activation and thus facilitates the apoptosis of neural progenitors [14, 24]. Furthermore, Maged1 was reported to regulate Dlx/Msx-mediated transcription by cooperating with necdin, another MAGE family protein [27, 28]. The high intensity of Maged1 signals is detected in progenitor cells in the neuroepithelia and subventricular regions in the developing embryo and is also detected in most adult tissues, predominantly in the cortex, hippocampus, olfactory bulb, hypothalamic and amygdaloidal nuclei in adult mice [11, 18, 29]. Previous studies reported that Maged1 knockout mice showed depressive-like behavior, disruption of community interactions, and impairments of circadian rhythm, further demonstrating the functional importance of Maged1 in neurons.
Here, we observed that Maged1 interacted with CREB and regulated learning and memory, which found to be the additional functions of Maged1 in the central nervous system. Memory formation begins with the activation of signalling pathways at the membrane of dendritic spines. The signalling cascades reach the nucleus, where transcription factor functions are modulated that results in changes in gene expression. Newly synthesized proteins cause long-term changes in cell function, such as the growth and maturation of new dendritic spines. During memory formation, the CRE-mediated regulation of gene transcription is one of the important ways the organisms adapt to environments. Transcription factors from the CREB/CRE modulator (CREM)/activating transcription factor 1 (ATF-1) gene family bind to CREs in the promoter regions of different genes and mediate the response of the cell to extracellular stimuli [30–32]. CREB was the first characterized CRE-binding factor [33, 34]. Numerous studies have established a strong connection between CREB and neuronal plasticity [35–38]. Loss-of-function manipulations of the most abundant CREB isoforms expressed in the mouse brain (CREBα/Δ) resulted in impaired LTP and deficits in several types of memories, including spatial, contextual, and cued memory [39–42]. These results were then followed by many studies which confirmed the essential role of CREB in memory in numerous additional species, brain regions, and learning systems. Several reports also described that the overexpression of activated forms of CREB resulted in the enhancement of long-term memory [43–47]. However, Balschun et al. [48] generated a mouse in which the expression of all CREB isoforms was either absent or limited and found no effect on late-phase LTP. The observation that different genetic manipulations of CREB produce diverse effects on learning and memory indicated that CREB functioned as a rate-limiting “molecular switch” in its biochemical pathway.
Taken together, as a multifunctional protein, after having withdrawn from the cell cycle, Maged1 could perform alternative “non-canonical functions” that regulate neuronal plasticity by modulating CREB-related transcription, including the expression of BDNF. This would then impair synapses and thus impair learning and memory. A more profound understanding of the Maged1 mechanism by which Maged1 enhances transcriptional activity can provide an excellent model system to gain better insight into the mechanisms of learning and memory and the pathophysiology of associated diseases.
References
Feng Y, Gao J, Yang M (2011) When MAGE meets RING: insights into biological functions of MAGE proteins. Protein Cell 2:7–12
Barker PA, Salehi A (2002) The MAGE proteins: emerging roles in cell cycle progression, apoptosis, and neurogenetic disease. J Neurosci Res 67:705–712
Chomez P, De Backer O, Bertrand M, De Plaen E, Boon T, Lucas S (2001) An overview of the MAGE gene family with the identification of all human members of the family. Cancer Res 61:5544–5551
Doyle JM, Gao J, Wang J, Yang M, Potts PR (2010) MAGE-RING protein complexes comprise a family of E3 ubiquitin ligases. Mol Cell 39:963–974
Goldman B, DeFrancesco L (2009) The cancer vaccine roller coaster. Nat Biotechnol 27:129–139
Ohman Forslund K, Nordqvist K (2001) The melanoma antigen genes—any clues to their functions in normal tissues? Exp Cell Res 265:185–194
Salehi AH, Roux PP, Kubu CJ, Zeindler C, Bhakar A, Tannis LL, Verdi JM, Barker PA (2000) NRAGE, a novel MAGE protein, interacts with the p75 neurotrophin receptor and facilitates nerve growth factor-dependent apoptosis. Neuron 27:279–288
Salehi AH, Xanthoudakis S, Barker PA (2002) NRAGE, a p75 neurotrophin receptor-interacting protein, induces caspase activation and cell death through a JNK-dependent mitochondrial pathway. J Biol Chem 277:48043–48050
Bragason BT, Palsdottir A (2005) Interaction of PrP with NRAGE, a protein involved in neuronal apoptosis. Mol Cell Neurosci 29:232–244
Di Certo MG, Corbi N, Bruno T, Iezzi S, De Nicola F, Desantis A, Ciotti MT, Mattei E, Floridi A, Fanciulli M, Passananti C (2007) NRAGE associates with the anti-apoptotic factor Che-1 and regulates its degradation to induce cell death. J Cell Sci 120:1852–1858
Kendall SE, Goldhawk DE, Kubu C, Barker PA, Verdi JM (2002) Expression analysis of a novel p75(NTR) signaling protein, which regulates cell cycle progression and apoptosis. Mech Dev 117:187–200
Nikopoulos GN, Martins JF, Adams TL, Karaczyn A, Adams D, Vary C, Oxburgh L, Verdi JM (2009) NRAGE: a potential rheostat during branching morphogenesis. Mech Dev 126:337–349
Calabrese G, Bennett BJ, Orozco L, Kang HM, Eskin E, Dombret C, De Backer O, Lusis AJ, Farber CR (2012) Systems genetic analysis of osteoblast-lineage cells. PLoS Genet 8:e1003150
Rochira JA, Matluk NN, Adams TL, Karaczyn AA, Oxburgh L, Hess ST, Verdi JM (2011) A small peptide modeled after the NRAGE repeat domain inhibits XIAP-TAB1-TAK1 signaling for NF-kappaB activation and apoptosis in P19 cells. PLoS One 6:e20659
Bertrand M, Huijbers I, Chomez P, De Backer O (2004) Comparative expression analysis of the MAGED genes during embryogenesis and brain development. Dev Dyn 230:325–334
Wang X, Tang J, Xing L, Shi G, Ruan H, Gu X, Liu Z, Wu X, Gao X, Xu Y (2010) Interaction of MAGED1 with nuclear receptors affects circadian clock function. EMBO J 29:1389–1400
Mouri A, Sasaki A, Watanabe K, Sogawa C, Kitayama S, Mamiya T, Miyamoto Y, Yamada K, Noda Y, Nabeshima T (2012) MAGE-D1 regulates expression of depression-like behavior through serotonin transporter ubiquitylation. J Neurosci 32:4562–4580
Dombret C, Nguyen T, Schakman O, Michaud JL, Hardin-Pouzet H, Bertrand MJ, De Backer O (2012) Loss of Maged1 results in obesity, deficits of social interactions, impaired sexual behavior and severe alteration of mature oxytocin production in the hypothalamus. Hum Mol Genet 21:4703–4717
Gao J, Wang WY, Mao YW, Graff J, Guan JS, Pan L, Mak G, Kim D, Su SC, Tsai LH (2010) A novel pathway regulates memory and plasticity via SIRT1 and miR-134. Nature 466:1105–1109
Gao J, Duan B, Wang DG, Deng XH, Zhang GY, Xu L, Xu TL (2005) Coupling between NMDA receptor and acid-sensing ion channel contributes to ischemic neuronal death. Neuron 48:635–646
Ji M, Dong L, Jia M, Liu W, Zhang M, Ju L, Yang J, Xie Z, Yang J (2014) Epigenetic enhancement of brain-derived neurotrophic factor signaling pathway improves cognitive impairments induced by isoflurane exposure in aged rats. Mol Neurobiol. doi:10.1007/s12035-014-8659-z
Rochira JA, Cowling RA, Himmelfarb JS, Adams TL, Verdi JM (2010) Mapping of NRAGE domains reveals clues to cell viability in BMP signaling. Apoptosis 15:63–70
Tian XX, Rai D, Li J, Zou C, Bai Y, Wazer D, Band V, Gao Q (2005) BRCA2 suppresses cell proliferation via stabilizing MAGE-D1. Cancer Res 65:4747–4753
Kendall SE, Battelli C, Irwin S, Mitchell JG, Glackin CA, Verdi JM (2005) NRAGE mediates p38 activation and neural progenitor apoptosis via the bone morphogenetic protein signaling cascade. Mol Cell Biol 25:7711–7724
Williams ME, Strickland P, Watanabe K, Hinck L (2003) UNC5H1 induces apoptosis via its juxtamembrane region through an interaction with NRAGE. J Biol Chem 278:17483–17490
Matsuda T, Suzuki H, Oishi I, Kani S, Kuroda Y, Komori T, Sasaki A, Watanabe K, Minami Y (2003) The receptor tyrosine kinase Ror2 associates with the melanoma-associated antigen (MAGE) family protein Dlxin-1 and regulates its intracellular distribution. J Biol Chem 278:29057–29064
Sasaki A, Masuda Y, Iwai K, Ikeda K, Watanabe K (2002) A RING finger protein Praja1 regulates Dlx5-dependent transcription through its ubiquitin ligase activity for the Dlx/Msx-interacting MAGE/Necdin family protein, Dlxin-1. J Biol Chem 277:22541–22546
Kuwajima T, Nishimura I, Yoshikawa K (2006) Necdin promotes GABAergic neuron differentiation in cooperation with Dlx homeodomain proteins. J Neurosci 26:5383–5392
Barrett GL, Greferath U, Barker PA, Trieu J, Bennie A (2005) Co-expression of the P75 neurotrophin receptor and neurotrophin receptor-interacting melanoma antigen homolog in the mature rat brain. Neuroscience 133:381–392
Mayr B, Montminy M (2001) Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat Rev Mol Cell Biol 2:599–609
Montminy M (1997) Transcriptional regulation by cyclic AMP. Annu Rev Biochem 66:807–822
Lonze BE, Ginty DD (2002) Function and regulation of CREB family transcription factors in the nervous system. Neuron 35:605–623
Montminy MR, Bilezikjian LM (1987) Binding of a nuclear protein to the cyclic-AMP response element of the somatostatin gene. Nature 328:175–178
Hoeffler JP, Meyer TE, Yun Y, Jameson JL, Habener JF (1988) Cyclic AMP-responsive DNA-binding protein: structure based on a cloned placental cDNA. Science 242:1430–1433
Stevens CF (1994) CREB and memory consolidation. Neuron 13:769–770
Silva AJ, Kogan JH, Frankland PW, Kida S (1998) CREB and memory. Annu Rev Neurosci 21:127–148
Tully T, Bourtchouladze R, Scott R, Tallman J (2003) Targeting the CREB pathway for memory enhancers. Nat Rev Drug Discov 2:267–277
Frank DA, Greenberg ME (1994) CREB: a mediator of long-term memory from mollusks to mammals. Cell 79:5–8
Yin JC, Wallach JS, Del Vecchio M, Wilder EL, Zhou H, Quinn WG, Tully T (1994) Induction of a dominant negative CREB transgene specifically blocks long-term memory in Drosophila. Cell 79:49–58
Bourtchuladze R, Frenguelli B, Blendy J, Cioffi D, Schutz G, Silva AJ (1994) Deficient long-term memory in mice with a targeted mutation of the cAMP-responsive element-binding protein. Cell 79:59–68
Lamprecht R, Hazvi S, Dudai Y (1997) cAMP response element-binding protein in the amygdala is required for long- but not short-term conditioned taste aversion memory. J Neurosci 17:8443–8450
Guzowski JF, McGaugh JL (1997) Antisense oligodeoxynucleotide-mediated disruption of hippocampal cAMP response element binding protein levels impairs consolidation of memory for water maze training. Proc Natl Acad Sci U S A 94:2693–2698
Josselyn SA, Shi C, Carlezon WA Jr, Neve RL, Nestler EJ, Davis M (2001) Long-term memory is facilitated by cAMP response element-binding protein overexpression in the amygdala. J Neurosci 21:2404–2412
Jasnow AM, Shi C, Israel JE, Davis M, Huhman KL (2005) Memory of social defeat is facilitated by cAMP response element-binding protein overexpression in the amygdala. Behav Neurosci 119:1125–1130
Perazzona B, Isabel G, Preat T, Davis RL (2004) The role of cAMP response element-binding protein in Drosophila long-term memory. J Neurosci 24:8823–8828
Yin JC, Del Vecchio M, Zhou H, Tully T (1995) CREB as a memory modulator: induced expression of a dCREB2 activator isoform enhances long-term memory in Drosophila. Cell 81:107–115
Barco A, Marie H (2011) Genetic approaches to investigate the role of CREB in neuronal plasticity and memory. Mol Neurobiol 44:330–349
Balschun D, Wolfer DP, Gass P, Mantamadiotis T, Welzl H, Schutz G, Frey JU, Lipp HP (2003) Does cAMP response element-binding protein have a pivotal role in hippocampal synaptic plasticity and hippocampus-dependent memory? J Neurosci 23:6304–6314
Acknowledgments
We thank Dr. Ying Xu for kindly providing all kinds of plasmids of Maged1. This work is partially supported by the National Basic Research Program of China (2013CB835100, 2011CBA01104), National Natural Science Foundation of China (nos. 81222013 and 30871264), and Specialized Research Funds for the Doctoral Program of Higher Education (20070284042, 20110091120029) to J Gao. J-J Yang is supported by the National Natural Science Foundation of China (81271216) and A-L Xu by the National Natural Science Foundation of China (nos. 81200186 and 81171233) and Natural Science Foundation of Shandong (ZR2011BL026).
Author information
Authors and Affiliations
Corresponding authors
Additional information
J. Yang, B. Lai, and A. Xu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yang, J., Lai, B., Xu, A. et al. Maged1 Co-interacting with CREB Through a Hexapeptide Repeat Domain Regulates Learning and Memory in Mice. Mol Neurobiol 51, 8–18 (2015). https://doi.org/10.1007/s12035-014-8677-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-014-8677-x