Abstract
Anthracnose caused by Colletotrichum gloeosporioides is an economically important disease which affects greater yam (Dioscorea alata L.) worldwide. Apart from airborne conidia, the pathogen propagules surviving in soil and planting material are the major sources of inoculum. A nested PCR assay has been developed for specific detection of C. gloeosporioides in soil and planting material. In conventional (single-round) PCR, the limit of detection was 20 pg, whereas in nested PCR the detection limit increased to 0.2 pg of DNA. The primers designed were found to be highly specific and could be used for accurate identification of the pathogen up to species level. The protocol was standardized for detection of the pathogen in artificially and naturally infected field samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The greater yam (Dioscorea alata L.), is one of the staple food crops for tropics and subtropics. They are nutritionally rich in carbohydrates, certain valuable vitamins such as thiamine, riboflavin, niacin, and ascorbic acid and also contain 1.9–3.9% protein as in rice or maize [1]. Anthracnose or Die back (Fig. 1) is one of the most serious foliar epiphytotic diseases of D. alata worldwide. The causal agent of the disease is an ascomycete fungus, Colletotrichum gloeosporioides (sexual stage Glomerella cingulata) [2]. Colletotrichum species cause anthracnose, which can cause considerable damage in a large number of crops such as cereals, coffee, and legumes [3] and even in human subcutaneous hyalohyphomycosis [4]. The rate of disease through sequence of spatial patterns was affected by the susceptibility of the cultivar, age of leaf, age of vine, stage of the epidemic, rainfall, and agronomic practices such as fungicide application [5]. Our preliminary observations on yam anthracnose revealed that the pathogen surviving on plant debris in soil and infected planting material acts as foci of anthracnose development in the field. The damage due to disease can be reduced to a great extent just by use of healthy planting materials. In case of yams, the tubers are being used for planting. The incipient infection of C. gloeosporioides cannot be visualized by naked eye at the time of planting. It is recommended to use healthy planting material and follow strict field sanitation by removing plant debris and weeds for management of the disease. Though need based fungicide application is also recommended, the poor farmers growing yams in India hardly use any fungicides. It is also reported that yam anthracnose is transmitted through true botanical seeds [6]. In this context, the role of diagnosis is very important in selecting healthy planting material for effective management of anthracnose. Traditional methods of diagnosis of fungi which relies primarily on differences in morphological features such as colony color, size, and shape of conidia and appressoria, optimal temperature for growth, growth rate, and the presence or absence of setae are often time consuming [7]. Though serological techniques have also been developed in different fungal species, but the lack of specificity of some antibodies and the necessity of obtaining monoclonal antibodies complicate the technique [8, 9].
The application of PCR has now made it possible to amplify the low copy number of DNA molecules and their detection. In contrast to more conventional methods, field samples can be tested directly through PCR and isolates do not require culturing. PCR method is rapid and can be used for detecting the presence of the fungus in the soil or planting material, therefore allowing implementation of early control measures. The ability to design PCR primers to target specific regions of DNA has led to rapid, accurate, and sensitive detection which is a greater understanding for managing Colletotrichum diseases. The development and use of molecular markers for the detection and exploitation of DNA polymorphism is one of the most significant developments in the field of molecular diagnosis [10]. The species-specific primers based on conserved regions in the genome are now widely used to detect and differentiate organisms with greater degree of sensitivity and reliability. The ribosomal DNA of higher eukaryotes, is having high copy number in the genome consisting of tandem repeats of 18S (smaller subunit), 5.8S (smallest subunit), and 28S (large subunit). These regions can be exploited for diagnostic purpose and the primers can be designed against species-specific regions from rDNA. Moreover, the high copy number increases the sensitivity of this detection technique. Successful primer design for PCR-based detection of a pathogen requires that the target region be unique to the organism of interest and conserved across populations of the organism. Ribosomal DNA sequence data has been widely used to design pathogen specific molecular primers due to their high gene copy number and the nature of the sequences containing both conserved and variable regions [11, 12]. Apart from the discriminatory potential, the highcopy number of rDNA genes in any genome permits a highly sensitive detection. Earlier research works on the smaller subunit rDNA (18S) region of C. gloeosporioides and found its inappropriateness for designing of species-specific primer [13]. A PCR-based detection assay for C. gloeosporioides using a forward primer (CgInt) from the conserved region of rDNA and a universal primer ITS 4 [14] is already reported. But chances of cross reactivity still exist as the reverse primer is universal for all fungi. The detection of pathogen in environmental samples may produce erroneous results due to nonspecific binding of the primer pairs. Hence, the present study was undertaken to design species-specific primers from highly conserved region of C. gloeosporioides and to develop PCR protocol for early detection of the pathogen from soil and yam tubers. The potential applications for the designed PCR primers are discussed.
Materials and Methods
Biological Materials
Colletotrichum gloeosporioides isolates were obtained from lesions on leaf, petiole, and vine from diseased plants sampled from different locations of India and maintained at Central Tuber Crops Research Institute (CTCRI), Thiruvananthapuram, India. The pure cultures were obtained by single spore isolation and maintained at 20 °C on potato dextrose agar medium (PDA; 250 g/L potato, 20 g/L dextrose and 20 g/L agar). The isolates of C. gloeosporioides collected from distant and different states of India, were given preference to the study. They represented the major yam growing areas Kerala, West Bengal, Bihar, Orissa, Gujarat, and Maharashtra. The isolates were selected after screening with RAPD primers (data not shown). The isolates with greater levels of variability were selected for further characterization.
Extraction of Genomic DNA
For DNA isolation, separate Erlenmeyer flasks (250 mL) containing 100 mL of autoclaved potato dextrose broth were inoculated with 1 cm disk of actively growing cultures of C. gloeosporioides. The cultures were placed on a rotary shaker (200 rpm) and incubated at 27 °C for 2–3 days. Mycelia were harvested by filtration through cheese cloth blotted dry with sterile paper towels and used immediately for DNA extraction using previously standardized method [15]. The DNA dissolved in TE were treated with RNAase (10 mg/mL), incubated at 37 °C for 30 min and stored at −20 °C until further use. DNA was quantified using Quant-iT™ Assay kits (Invitrogen, USA) according to manufacturer instruction and visualized by 0.8% agarose gel electrophoresis.
PCR Amplification
The genomic DNA was amplified using the universal ITS1 and ITS4 primer pairs used to amplify the nuclear rDNA region [16]. Each 25 μL of PCR reaction consisted of 100 ng of template DNA, 100 μM each deoxynucleotide triphosphate, 20 ng of each primer, 1.5 mM MgCl2, 1 × Taq buffer (10 mM Tris–HCl pH 9.0, 50 mM KCl, 0.01% gelatin), 1 U of Taq DNA polymerase (Bangalore GeNei, India). Amplifications were performed in Eppendorf thermal cycler (Eppendorf AG, Hamburg, Germany). The maximum and optimum annealing temperature per primer pair was empirically investigated by gradient PCR starting with a range of annealing temperatures from 50 to 60 °C and followed by PCR at the standardized annealing temperature. Amplified products were resolved on a 1.5% agarose gel containing 0.5 mg/mL ethidium bromide and photograph was scanned through Gel Doc System (Alpha imager, Alpha Innotech, USA). The PCR products were eluted using QIAquick Gel extraction kit (QIAGEN, Tokyo, Japan) and cloned into the pGEM-T® vector (Promega, WI, USA). They were transformed into DH5α cells and sequenced by using T7 and SP6 promoter primers. The representative sequences have been submitted to GenBank under the accession numbers: FJ940734, FJ940735, and HM161642.
Development of Colletotrichum gloeosporioides Specific Primers and PCR Conditions
The rDNA sequences of 96 isolates of C. gloeosporioides were collected after similarity search with BLAST and aligned with available ITS sequences of other fungal species using the Clustal W program in BioEdit software [17]. This also included the rDNA sequences of C. gloeosporioides infecting other crops. The rDNA sequences of other species of Colletotrichum, other fungi were selected for alignment. Regions of dissimilarity between the consensus of C. gloeosporioides and other fungal species were identified and primers specific for C. gloeosporioides were developed using the Primer Premier 6.0 (build 60006). C. gloeosporioides specific primer pairs CgsF1 (GGCGGGTAGGGTCTCCGTGAC)/CgsR1 (TTTGAGGGCCTACATCAGCT) were designed and assessed against a range of PCR conditions to determine the optimum cycling regime for detection of C. gloeosporioides.
Standardization of PCR Conditions
-
(1)
Annealing temperature: the maximum and optimum annealing temperature per primer pair was empirically investigated starting with a range of annealing temperatures, from 55 to 65 °C.
-
(2)
PCR cycles: the specificity and efficiency of selected primer pairs in the PCR reaction was tested from 20 to 40 cycles.
-
(3)
Specificity and sensitivity: after successful amplification of the target DNA, the primer pairs were checked for non specific binding against genomic DNA of a wide range of other species of Colletotrichum, other ascomycete, basidiomycete, oomycete, bacteria, plant and virus. In order to assess the primer sensitivity, a serial tenfold dilution of C. gloeosporioides gDNA from 200 ng/mL stock was used.
-
(4)
Primer pairs were checked against with the gDNA from many accessions of C. gloeosporioides from different parts of India isolated from D. alata and other hosts of the pathogen.
-
(5)
The primer pairs were used for in silico PCR analysis against sequence obtained from GenBank of different C. gloeosporioides accessions across the world.
Standardization of Conditions for PCR and Nested PCR Assays
The primer pairs CgsF1 and CgsR1 were used for PCR and nested PCR with the following reaction parameters, 25 μL of PCR reaction consisted of 100 ng of template DNA, 100 μM each deoxynucleotide triphosphate, 20 ng of each primer, 1.5 mM MgCl2, 1 × Taq buffer (10 mM Tris–HCl pH 9.0, 50 mM KCl, 0.01% gelatin), 1 U of Taq DNA polymerase (Bangalore GeNei, India). The PCR profile was denaturation at 94 °C for 2 min, followed by 30 cycles of 94 °C for 30 s, 62 °C for 40 s, and 72 °C for 40 s, then a final extension at 72 °C for 5 min. For nested PCR, Primer pairs ITS1 and ITS4 were used in the first round PCR and then 1 μL of the first round was used as template in the second round of amplification with C. gloeosporioides specific primers, performed according to the standardized PCR protocol described above. The PCR reaction was conducted in triplicates in three different thermal cyclers (Bio-Rad C-1000, Corbett Research Palm Cycler and Eppendorf Thermal Cycler) to insure the reproducibility of the detection.
PCR Amplification of Pathogen from Artificially and Naturally Infected Soil and Tuber Samples
Plant parts of D. alata such as leaf, tuber, and vine were inoculated with spore suspension of C. gloeosporioides, kept for infection to develop and the total genomic DNA was isolated from them according to the protocol previously described [18]. In addition, field samples of yam tubers and leaves that either was asymptomatic, contained visible anthracnose spots, or were dead from infections caused by C. gloeosporioides were collected from field plots in CTCRI and were used for DNA isolation and PCR assay. Soil DNA was also isolated from samples collected from the farms, where infected plants had been harvested and analyzed by the specific primers. DNA isolated from pure culture of C. gloeosporioides served as positive control and specific pathogen free plants maintained in glass house served as negative control.
Results
Primer Design, Specificity, and Sensitivity
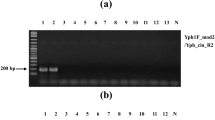
The DNA isolated using the designed protocol was excellent in quality (A260/280:1.5–1.9) and quantity (0.75 ± 0.10 μg DNA/mg of fungal mycelium). Amplification of gDNA with ITS1 and ITS4 primers yielded a single product of ~580 bp in all the three isolates. The rDNA sequence analysis of C. gloeosporioides isolates revealed 97–100% nucleotide sequence homology with each other and 96–100% with different gene sequences of C. gloeosporioides and less similarity to other fungal species present in the nucleotide database. The primers were designed from 100% sequence homology region of C. gloeosporioides isolates and from regions of the greatest sequence dissimilarity among other species. The designed C. gloeosporioides specific CgsF1 (GGCGGGTAGGGTCTCCGTGAC)/CgsR1 (TTTGAGGGCCTACATCAGCT) gave an approximately 310 bp product at standardized 59 °C annealing temperature in all isolates of C. gloeosporioides. The DNA from C. gloeosporioides only recorded amplification with the designed primer in the standardized protocol which indicated that no corresponding sites of the designed primer existed in the genomic DNA of the other organisms (Fig. 2). The designed primer pairs amplified C. gloeosporioides from different places of India in the standardized PCR protocol (Fig. 3). The specific primer pair produced an amplicon of 310 bp on a 1.5% agarose gel after 25 cycles when starting with 200 ng of genomic DNA of C. gloeosporioides. In conventional PCR, the lowest amount of DNA for detection was 20 pg (Fig. 4a) and for nested PCR after the first round amplification with the ITS1 and ITS4 primers, the detection limit was 0.2 pg for each primer pair (Fig. 4b). The results were reproducible in different thermal cyclers in the standardized protocol.
Specificity of PCR with the primer pair CgsF1/CgsR1 for detection of C. gloeosporiodes. Lane M 100 bp ladder, Lane 1 and Lane 17 Colletotrichum gloeosporioides, Lane 2 Dioscore alata, Lane 3 Dioscorea esculenta, Lane 4 Colletotrichum dematium, Lane 5 Colletotrichum lindemuthianum, Lane 6 Colletotrichum falcatum, Lane 7 Colletotrichum truncatum, Lane 8 Colletotrichum capsici, Lane 9–Lane 14 Sclerotium rolfsii isolates, Lane 15 Sclerotium hydrophilum, Lane 16 Sclerotium oryzae, Lane 18 Phytophthora colocasiae, Lane 19 Phytophthora capsici, Lane 20 Phytophthora meadii, Lane 21 and Lane 22 Phytophthora palmivora, Lane 23 Phytophthora parasitica, Lane 24 Phytophthora boehmeriae, Lane 25 Trichoderma harzianum, Lane 26 Mucor sp, Lane 27 Pythium ultimum, Lane 28 Rhizoctonia solani, Lane 29 Bacillus subtilus, Lane 30 Badna virus, Lane 31 Erwinia carotovora, Lane 32 no DNA template
Detection of Pathogen from Infected Soil and Tuber Samples
Detection of pathogen was observed in artificially infected and naturally infected tubers and leaves of yam by the nested and conventional PCR. The DNA samples isolated from soil samples collected in the same farms, where infected plants had been showed the specific detection of C. gloeosporioides in them (Fig. 5). The naturally infected tuber and leaves also recorded amplification with the primer set. The nested PCR has shown to be more sensitive and could be used for detection of pathogen even before the onset of visible symptoms. The detection time for the nested and conventional PCR was 16–24 h post inoculation, respectively. The healthy and control samples did not record any amplification.
Discussion
The objectives of this study were to develop a sensitive and accurate method based on conventional PCR and nested PCR for detection of C. gloeosporioides. These results indicated that the specific primer pairs are useful in detecting C. gloeosporioides directly from infected host tissues, even before the onset of visible symptoms on the host, pathogen contaminated soil samples, seed tubers etc. Microscopic identification or cultural characterization of C. gloeosporioides is an acceptable approach and fungus can be cultured from different samples such as infected leaf, stem, tubers, or from soil but an experienced operator is needed to obtain reliable results since many species of Colletotrichum have similar growth characteristics, morphology and is having cosmopolitan distribution with a wide host range. Immunological test methods are quick and detect mainly living material, but the laborious development of assays and possible cross-reactions with related species or the host plant are serious drawbacks [19]. The use of rDNA gene for diagnosis of Colletotrichum species have been already been proven, this is due to the presence of species-specific region present between two coding sequence. The earlier approach for molecular diagnosis and detection of C. gloeosporioides was based on a single primer CgInt designed from the rDNA gene as the forward primer and the universal primer ITS4 as the reverse primer [14]. In the present study, both the forward and the reverse primers are from highly conserved regions for C. gloeosporioides. This eliminated the possible chances of cross reactivity with any other organism particularly while handling environmental samples such as soil, plant parts etc. PCR has emerged as a powerful tool for the diagnosis of the diseases because it is more sensitive, robust, rapid, and less labor-intensive than traditional diagnostic methods. Unlike identification based on culture techniques, PCR is more rapid and can work even when there is a limited amount of sample [20]. The PCR assay can be performed in 3–4 h, including sample processing, PCRs, gel electrophoresis, and staining. Visual identification may require placement of infected tissue on culture medium or in moist chambers to observe growth and/or sporulation characteristic of C. gloeosporioides, which may take 24 h or more. The use of RAPD primers to screen the isolates helped in the identification of the isolates with variability after sequencing the rDNA region. For designing the species-specific primers to C. gloeosporioides, rDNA sequences were selected for several reasons. First, they have evolved quickly, showing variation among related taxa and even among species of the same genus [21]. Second, rDNA is found in many copies in the genome [22]; in fact, genes coding for rDNA in fungi are found on a single chromosome in repeated units arranged in tandem with 60–220 copies represented in the haploid genome [23]. The rDNA based species-specific diagnostic primers have been used for detection of other species of Colletotrichum [24–28], and other plant pathogens [29–31]. In this attempt, we have developed a PCR assay based on sequence analysis of the ITS region of rDNA of C. gloeosporioides, which allows sensitive and accurate detection than that is possible based on cultural characterization and serological kits. The designed primers was successful in amplifying a range of C. gloeosporioides isolates from almost all major yam growing areas in India, representing that they can amplify C. gloeosporioides across its populations without having any cross-reactivity with DNA from different oomycetes species, several fungal species and the D. alata plant. The primers could also be employed for detection of this pathogen from its other hosts. The nested PCR assay was proved to have a greater sensitive detection and could be employed for indexing seed tubers and identification of disease free planting sites where the pathogen inoculum is found to be less during the post harvest period. Even though, reliability of the PCR assay in detecting C. gloeosporioides isolates from other countries was not tested due to the lack of isolates from those region the sequence obtained from NCBI database was used for in silico PCR analysis for C. gloeosporioides isolates from across the globe to check the reproducibility in different geographical regions of the world. The primers and the assay methods we have described will be useful for yam planting material testing programs, particularly since laboratory procedures for detection are needed as an adjunct to visual identification. Although, the use of a conventional PCR assay is the first step in pathogen detection and epidemiology, more accurate information can only be determined by quantitative measurements via the development of real-time quantitative PCR assays, which could be the next step of this research.
References
Hahn, S. K., Osiru, D. S. O., Akoroda, M. O., & Otoo, J. A. (1987). Yam production its future prospects. Outlook on Agriculture, 16(3), 105–110.
Ayodele, M.A., Hughes, J. d’A. & Asiedu, R. (2000). Yam anthracnose disease: Field symptoms and laboratory diagnostics. International Institute of Tropical Agriculture.
Bailey, J. A., & Jeger, M. J. (1992). Colletotrichum: Biology, pathology and control (p. 388). Wallingford: CAB International.
Guarro, J., Svidzinski, T. E., Zaror, L., Forjaz, M. H., Gene, J., & Fischman, O. (1998). Subcutaneous hyalohyphomycosis caused by Colletotrichum gloeosporioides. Journal of Clinical Microbiology, 36, 3060–3065.
Sweetmore, A., Simons, S. A., & Kenward, M. (1994). Comparison of disease progress curves for yam anthracnose (C. gloeosporioides). Plant Pathology, 43, 206–215.
Jackson, G.V.H., Newhook, F.J. & Winch, J. (2002). Pest advisory leaflet. New Caledonia: Plant Protection Service No. 12, Secretariat of the Pacific Community.
Smith, B. J., & Black, L. L. (1990). Morphological, cultural and pathogenic variation among Colletotrichum species isolated from strawberry. Plant Disease, 74, 69–76.
Choe, J. P., Hwang, U. W., & Kim, W. (1999). Putative secondary structures of unusually long 226 strepsipteran SSU rRNAs and its phylogenetics implications. Molecular Cells, 9, 191–199.
McCartney, H. A., Foster, S. J., Fraaije, B. A., & Ward, E. (2003). Molecular diagnostics for fungal plant pathogens. Pest Management Science, 59, 129–142.
Semagn, K. (2006). An overview of molecular methods for plants. African Journal of Biotechnology, 5(25), 2540–2568.
Wallenhammar, A., & Arwidsson, O. (2001). Detection of Plasmodiophora brassicae by PCR in naturally infested soils. European Journal of Plant Pathology, 107, 313–321.
Grote, D., Olmos, A., Kofoet, A., Tuser, J. J., Bertolini, E., & Cambra, M. (2002). Specific and sensitive detection of Phytophthora nicotianae by simple and nested-PCR. European Journal of Plant Pathology, 108, 197–207.
Azim, T., Jeeva, M. L., Pravi, V., Archana, P. V., & Mishra, A. K. (2010). Exploration of smaller subunit ribosomal DNA for detection of Colletotrichum gloeosporioides causing anthracnose in Dioscorea alata L. Journal of Root Crops, 36(1), 83–87.
Mills, P. R., Sreenivasaprasad, S., & Brown, A. E. (1992). Detection and differentiation of Colletotrichum gloeosporioides isolates using PCR. FEMS Microbiology Letters, 98(1–3), 137–143.
Jeeva, M. L., Sharma, K., Mishra, A. K., & Misra, R. S. (2008). Rapid extraction of genomic DNA from Sclerotium rolfsii causing collar rot of Amorphophallus. Genes, Genomes and Genomics, 2(1), 60–62.
White, T. J., Bruns, T., Lee, S., & Taylor, J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. Innis, D. Gelfand, J. Sninsky, & T. White (Eds.), PCR Protocols: a Guide to Methods and Applications (pp. 315–322). Orlando, Florida: Academic Press.
Hall, T. A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Sharma, K., Mishra, A. K., & Misra, R. S. (2008). A simple and efficient method for extraction of genomic DNA from tropical tuber crops. African Journal of Biotechnology, 7(8), 1018–1022.
Ward, E., Foster, S. J., Fraaije, B. A., & McCartney, H. A. (2004). Plant pathogen diagnostics: Immunological and nucleic acid-based approaches. Annuals of Applied Biology, 145, 1–16.
Schubert, R., Bahnweg, G., & Nechwatal, J. (1999). Detection and quantification of Phytophthora species which are associated with root to rot diseases in European deciduous forests by species to specific polymerase chain reaction. European Journal for Pathology, 29, 169–188.
Cooke, D. E. L., Drenth, A., Duncan, J. M., Wagels, G., & Brasier, C. M. (2000). A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genetics and Biology, 30, 17–32.
Lee, S. B., & Taylor, J. W. (1992). Phylogeny of five fungus-like protoctista Phytophthora species, inferred from the internal transcribed spacers of ribosomal DNA. Journal of Molecular Biology and Evolution, 9, 636–653.
Martin, F. N. (1990). Variation in the ribosomal DNA repeat unit within single oospore isolates of the genus Pythium. Genome, 33, 585–591.
Torres-Calzada, C., Tapia-Tussell, R., Quijano-Ramayo, A., Martin-Mex, R., Rojas-Herrera, R., Higuera-Ciapara, I. (2011). A species-specific polymerase chain reaction assay for rapid and sensitive detection of Colletotrichum capsici. Molecular Biotechnology, 49(1), 48–55.
Chen, L. S., Chu, C., Liu, C. D., Chen, R. S., & Tsay, J. G. (2006). PCR-based detection and differentiation of anthracnose pathogens, Colletotrichum gloeosporioides and C. truncatum, from vegetable soybean in Taiwan. Journal of Phytopathology, 154, 654–662.
Wang, W., Tang, J. H., & Wang, Y. C. (2008). Molecular detection of Colletotrichum lindemuthianum by duplex PCR. Journal of Phytopathology, 156, 431–437.
Sreenivasaprasad, S., Sharada, K., Brown, A. E., & Mills, P. R. (1996). PCR-based detection of C. acutatum on strawberry. Plant Pathology, 45, 650–655.
Cullen, D. W., Lees, A. K., Toth, I. K., & Duncan, J. M. (2002). Detection of Colletotrichum coccodes from soil and potato tubers by conventional and quantitative real-time PCR. Plant Pathology, 51, 281–292.
Ippolito, A., Schena, L., Nigro, F., Ligorio, V. S., & Yaseen, T. (2004). Real-time detection of Phytophthora nicotianae and P. citrophthora in citrus roots and soil. European Journal of Plant Pathology, 110, 833–843.
Jeeva, M. L., Mishra, A. K., Pravi, V., Misra, R. S., & Hegde, V. (2010). A species-specific polymerase chain reaction assay for rapid and sensitive detection of Sclerotium rolfsii. Australasian Plant Pathology, 39, 1–7.
Mishra, A. K., Jeeva, M. L., Pravi, V., Misra, R. S., & Hegde, V. (2010). Rapid and sensitive detection of Phytophthora colocasiae associated with leaf blight of taro by species-specific polymerase chain reaction assay. Annals of Microbiology, 60, 209–215.
Acknowledgments
The funding provided for research study by the National Fund for Basic, Strategic, and Frontier Application Research in Agriculture (NFBSFARA), ICAR, New Delhi, was gratefully acknowledged. The authors thanked the Director, Central Tuber Crops Research Institute, Thiruvananthapuram for providing the infrastructure facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mithun Raj, Jeeva, M.L., Hegde, V. et al. Polymerase Chain Reaction Assay for Rapid, Sensitive Detection, and Identification of Colletotrichum gloeosporioides causing Greater Yam Anthracnose. Mol Biotechnol 52, 277–284 (2012). https://doi.org/10.1007/s12033-012-9496-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-012-9496-9