Abstract
DNA methylation analysis, an epigenetic specification, has been explored for partial determination of cancer cell phenotypes. The development of metastasis in cancerogenesis has led its feasible association with the epigenetic modulations. We generated highly aggressive non-small cell lung cancer cell lines (HTB56 and A549) by using in vivo selection approach. These were, then, subjected to DNA methylation analysis (genome-wide). We also explored the therapeutic effects of azacytidine, an epigenetic agent, on DNA methylation patterns as well as the in vivo phenotypes. During the development of highly aggressive cell lines, we observed widespread modulations in DNA methylation. Reduced representation bisulfite sequencing was used and compared with the less aggressive parental cell lines to identify the differential methylation, which was achieved up to 2.7 % of CpG-rich region. Azacytidine inhibited DNA methyltransferase and reversed the prometastatic phenotype. We found its high association with the preferential loss of DNA methylation from hypermethylated sites. After persisted exposure of azacytidine, we observed that DNA methylation affected the polycomb-binding sites. We found close association of DNA methylome modifications with metastatic capability of non-small cell lung cancer. We also concluded that epigenetic modulation could be used as a potential therapeutic approach to prevent metastasis formation as prometastatic phenotype was reversed due to inhibition of DNA methyltransferase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the second most common cancer and leading cause of cancer-related death in both males and females [1]. On the basis of histology of lung cancer, non-small cell lung cancer is supposed as the most frequent one because it represents around 85 % of all cases. Epigenetic mechanisms are responsible for gene expression regulation. Epigenetic modifications play important roles in many pathophysiological and physiological conditions along with carcinogenesis with no modification in DNA sequence [2]. Modifications in histone and DNA methylation may impact on cancer aggressiveness and its phenotype [3, 4]. Moreover, genetic modifications can be outnumbered by these alterations and usually occur early in carcinogenesis [5]. Epigenetics along with genetics intersect for promoting the carcinogenesis during each and every stage of development of cancer. In many cases, epigenetic modifications are reversible and dynamic and that is why it represents interesting targets for treatment of cancer [6]. Recently, it was explored that DNA methyltransferase inhibitory drugs such as decitabine and azacytidine can modify cellular cancer phenotypes over prolonged times even after the end of drug exposure [7]. Decitabine and azacytidine can induce DNA hypomethylation at specific gene loci, and it can lead to sustained gene reactivation [8]. However, DNA methylation analysis (genome-wide) after drug-induced DNA methyltransferase inhibition has not been explored till now, and the actual mechanisms of action of these epigenetic modulators still defined poorly [9].
The most frequent causes of cancer-related death after complete tumor resection are metastases. However, there is similarity between gene expression profiles of cancer stem cells and gene expression profiles of metastatic cancer cells [10, 11]. A large number of additional genetic mutations are not responsible for these expression profiles, but epigenetic modifications are responsible for predispose for metastasis [12]. Thus, we may say that formation of metastasis in most cases occurs in short time span in comparison with primary tumor initiation and growth until diagnosis, which takes several years of time. As a result, DNA methyltransferase inhibitory drugs can potentially target the metastasis-associated epigenetic state. In the present work, we have explored the profile of DNA methylation modifications, forming with the transition from a cellular state with low metastatic capacity toward a highly aggressive state. We also analyzed whether these changes were reversible or irreversible with respect to DNA methyltransferase inhibition.
Materials and methods
Cell lines and azacytidine treatment
As per previous reports [13, 14], HTB56 and A549 lung adenocarcinoma cells were cultured. STR marker analysis was further used for the identification of cell lines. HTB56 and A549 cells were exposed to 5-azacytidine (Sigma, Beijing, China) at a concentration of 1 µmol/L to 100 nmol/L. Under these conditions, the cells were grown for 8 days. Meanwhile, cells were supplemented with fresh medium in every 2 days. The cells were washed three times with PBS and released for extra 1 week in regular medium.
In vitro functional assays
As per previous reports, 3[H]-thymidine-incorporated proliferation assays [15], cell viability assays [16], human tumor cloning assays [17], and migration assays [13, 15] were performed.
Reduced representation bisulfite sequencing for DNA methylation analysis (genome-wide)
As per published protocols [18, 19], reduced representation bisulfite sequencing library was prepared using total 0.2–1 µg of DNA. During the experiment, Illumina TruSeq adapters (Illumina, San Diego, CA, USA) were used for ligation purpose, EZ DNA Methylation™ Kit (Zymo Research Europe, Freiburg, Germany) was used for bisulfate conversion, and PfuTurbo Cx DNA polymerase (Agilent Technologies, Santa Clara, CA, USA) was used for amplification purpose. Human genomic sequences (hg19) and other tracks (e.g., RefSeq genes) were downloaded from the University of California, Santa Cruz Genome Browser database [20]. Adapter sequences were removed using Cutadapt version 0.9.3 [21], and sequences were mapped to hg19 genome using Bismark version 0.5 [22]. Methylation calls from Bismark were extracted with a modified script that removed 3′-MspI sites. The conversion rate of all non-CpG cytosine positions was calculated from the Bismark methylation_extractor output. Besides, samples with good conversion rate (at least 99 % of all non-CpG cytosine positions) were converted to uracil (Table 1). BiSeq package [23] was used for analysis of methylation data in R/Bioconductor. However, differentially methylated region detection was restricted to regions with a high CpG site density covered across all samples. Smoothed methylation levels (between 0 and 1) were calculated every 10 bps within each CpG cluster. Regions with methylation differences of at least 30 % were called differentially methylated regions (minimum one base pair). We supposed that the number m of region centers within differentially methylated regions is binomially distributed with o = number of region centers in CpG clusters and s = sum of differentially methylated region widths divided by the sum of CpG cluster widths to determine whether regions of interest were overrepresented or underrepresented in differentially methylated regions. We then carried out a two-sided binomial test that m ≠ s × o.
Illumina methylation bead arrays
Infinium Human Methylation 27 BeadChip (Illumina) was used as per manufacturer’s instructions. Genome Studio version 2011.1, along with Methylation Module version 1.9.0 and Illumina Genome Viewer Module version 1.9.0 softwares, was used for data preprocessing and methylation level extraction.
Gene expression analysis
Human Gene 1.0 ST Array (Affymetrix) based on highly metastatic and parental A549 cell lines was used as per manufacturer’s instructions. Affymetrix GeneChip Scanner 3000 was used for scanning of arrays at 1.56-µm resolution. After importing the raw gene expression data to the Affymetrix expression console, it was subjected to robust multi-array average (RMA). RankProd software [24] was used for differential gene expression calculation.
Single nucleotide polymorphism analysis
As per published reports [25, 26], high-resolution single nucleotide polymorphism array (Affymetrix, 6.0) method was used for the screening of copy-number alterations in the genome of highly metastatic and parental non-small cell lung cancer cell lines using HTB56 and A549. Single nucleotide polymorphism array raw data are available at NCBI (GSE44549).
Exome capture and high-throughput sequencing
As per manufacturer’s standard protocol, the SureSelect XT Human All Exon V5 + UTR kit (Agilent) for Illumina sequencing was used. However, HiScanSQ instrument (Illumina) was used for exome-enriched libraries, which were subjected to 2 × 100 cycles paired-end sequencing.
Global DNA methylation analysis
DNAzol (Invitrogen, Beijing, China) was used for the extraction of genomic DNA from highly metastatic and parental A549 cell lines. As per previously published protocol [27], global methylation levels were determined by capillary electrophoresis.
Exome-sequence processing—read mapping, variant calling, and effect determination
For this purpose, Burrows–Wheeler alignment algorithm [PMID:19451168] was applied to map raw reads from the Illumina HiSeq 2000 to the hg19 genome followed by removal of duplicate reads. To remove low-quality mutations, standard hard filtering parameters (Best Practice Variant Detection using GTAK v4 documentation) were applied. Ingenuity© Variant Analysis (CA, USA) online software was used to transfer the detected variations.
Analysis of double-strand break by H2AX staining
According to the manufacturer’s instructions (cell signaling), the determination of H2AX-positive cells was done. H2AX-positive cells were considered as apoptotic cell population.
In vivo mouse experiments
By strictly following the relevant regulations for animal experiments, we used 9- to 11-week-old non-obese diabetic or severe combined immunodeficient mice. Two non-obese diabetic/severe combined immunodeficient mice per cell line were irradiated with a single dose of 3.6 Gy from a cobalt 60 unit 1 day before injection. A total number of 2.5 × 106 parental cells (HTB56_R0 or A549_R0) were injected intravenously into the tail vein of two different mice. Mice were killed after 9 weeks for the first round of selection, after 7 weeks for the second round of selection, and after 5 weeks for the third round of selection. After digestion for 35 min with an adequate amount of trypsin (around 5.5 × 104 cells) in cell culture, nodules of both lungs were obtained and pooled. As per reported works [13, 15], total of 1.5 × 106 (HTB56) or 2.5 × 105 (A549) or cells either highly or low metastatic and either exposed or non-exposed were injected intravenously into the tail vein and followed up for 5 weeks. Then, mice were killed for counting the lung nodules. To block the cage effects, treatment groups were randomized in all the experiments.
Statistical analysis
The SPSS (version 18 (IBM) software) was used for statistical analyses. Kruskal–Wallis test was used for the analysis of overall differences between multiple groups of statistical significances. However, Student’s t test was used to analyze the differences between two groups. A P value of <0.05 was considered significant.
Results
In vivo selection of a highly aggressive phenotype
The epigenetic therapy can be used for the alteration of specific phenotypes and properties, which would be associated with epigenetic modifications. Non-small cell lung cancer cell lines were generated with increased propensity for the formation of tumor nodules after intravenous injections (Fig. 1a). We know that the important parts of the metastatic process are extravasation and growth at a distant site. On account of this, we explored these parts as surrogate markers for potential metastatic capability and in vivo aggressiveness. For this purpose, two cell lines were used: First cell line was HTB56 cells which is an anaplastic carcinoma cell line and formed fewer but larger nodules in vivo and the second one was A549 lung adenocarcinoma cells which formed multiple small nodules in non-obese diabetic or severe combined immunodeficient mice after intravenous injection. We observed that removal of tumor nodules from the lungs followed by reinjection was responsible for a drastic increase in metastatic capacity (Fig. 1a). After three rounds of in vivo selection for both cell lines, we achieved a highly aggressive phenotype, which was very stable over time (Fig. 1b, P < 0.001 for each cell line). These highly aggressive cells showed enhanced clonogenic growth in vitro and additional in vitro, which led to accelerated proliferation, a metastatic potential. A high number of potential driver mutations in the parental cells were verified by whole exome sequencing (HTB56_R0: n = 182; A549_R0: n = 251) as well as additional mutations were observed in the highly aggressive cell lines [only 8 (HTB56_R3 or 5 (A549_R3); whole data not shown]. Hence, the strong differences about metastatic capacity in vivo phenotypes could play a big role, and only 3.7 % (HTB56) or 1.4 % (A549) of the mutations was newly acquired during in vivo selection process.
DNA methylation changes and aggressiveness of lung cancer cell lines. a Experimental outline for the generation of highly aggressive lung cancer cell lines. HTB56 or A549 cells were intravenously injected into the tail vein of non-obese diabetic or severe combined immunodeficient mice. After 9 weeks, the mice were killed and lung nodules reminiscent of metastases were removed. Tumor cells were expanded in vitro and subsequently intravenously injected. The high metastatic potential cell lines (after three rounds of selection) were compared with the parental cell lines. b Verification of the aggressiveness of the low and highly metastatic potential cell lines in vivo. Representative photographs of the resected lungs are shown. Black arrows indicate metastases. c Genome-wide DNA methylation analyses were performed by reduced representation bisulfate sequencing. The scatter plots indicate changes in DNA methylation between high metastatic (y-axis) and low metastatic cells (x-axis). Colors represent the density of CpG sites ranging from red (high density) to blue (low density). Notably, the majority of CpG sites were either unmethylated or fully methylated. Also, more changes in DNA methylation were observed in HTB56 cells compared with A549 cells
Genome-wide modifications in DNA methylation in the highly metastatic state
Since we know that in human cancer, functionally relevant DNA methylation modifications occur at CpG-rich regions; hence, we used a well-established modified reduced representation bisulfite sequencing procedure [18] to focus on CpG islands and the associated shores. Thus, we identified modifications in DNA methylation in both the non-small cell lung cancer cell lines with increased aggressiveness in vivo. We found 1.8 × 107 reads which were, on an average, uniquely mapped to the human genome in each cell line (detailed data not shown). We defined, as per the spatial density of covered CpG sites, 20143 CpG clusters that spanned 7,771,217 bp of the human genome. These clusters range from 37 to 2137 bp (median 339 bp) length. The CpGs in CpG clusters were unaffected with DNA methylation (Fig. 1c). The median methylation level was 1.08 % for CpGs in the metastatic HTB56_R3 cells and 0.86 % for CpGs in the parental HTB56_R0 cells. Similar numbers were obtained in A549 cells with a small decrease toward higher CpG methylation in metastatic cells (1.04 %) compared with the parental cells (0.75 %). A total of 2,519 regions (2.6 % of the tested genomic region) were found to be differentially methylated in highly metastatic HTB56_R3 cells and 2009 regions (0.9 % of the tested genomic region) in A549_R3 cells. However, DNA hypomethylated regions were more frequent than DNA hypermethylated differentially methylated ones as in highly metastatic HTB56_R3 (21.6 %) and A549_R3 (38.7 %) cells. A total of 181 (14.9 %) of HTB56_R3 differentially methylated regions overlapped and were similarly altered in metastatic A549_R3 cells. Hypomethylation in both cell lines was observed for 139 regions, whereas concordant DNA hypermethylation was observed in 41 regions.
On the other hand, association between the DNA methylation modifications in the cell lines and the transcription factor binding sites determined by ChIP-Seq (28) was also analyzed. An increased likelihood to be DNA hypermethylated and hypomethylated, respectively, in both cell lines was shown as the binding sites of Suz12 indicate for the polycomb complex formation in embryonic stem cells, similar to previous report [28]. It can be said that the stability of DNA methylation might primarily depend on transcription factor binding such that regions without active transcription factor binding are more prone to undergo modifications in DNA methylation.
Azacytidine-treated DNA methyltransferase inhibition in vivo reversal of metastatic-prone phenotype
We found a good association between the modified DNA methylation patterns with the selection of a more aggressive phenotype. DNA hypermethylation covered around 33 % of the modifications in methylation. Azacytidine (5-azacytidine, precisely) was used to target DNA methylation by trapping and inhibiting DNA methyltransferase. It has one drawback that at high concentrations it is toxic and creates difficulty to differentiate the epigenetic effect from toxic effect. As a result, epigenetic effects targeting the phenotypic memory of tumor cells should persist longer than toxic effects. Thus, we exposed the highly metastatic cells to azacytidine for 8 days followed by removal of drug by washing and releasing of cells into regular media for an extra week (Fig. 2a). Release experiments (Fig. 2a) were performed at concentrations as low as 250 nmol/L. We found the presence of breaks of DNA double strand in all non-small cell lung cancer cell lines regardless of previous azacytidine exposure (Fig. 2a). Next, the in vivo growth at a distant site for metastatic capacity of azacytidine-exposed tumor cells was analyzed. A549 cells were injected into non-obese diabetic or severe combined immunodeficient mice at the end of a weeklong exposure after the end of azacytidine exposure.
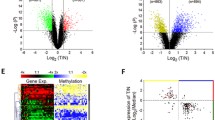
a Experimental outline indicates the timeline of azacytidine exposure and release. Cells were exposed to 5-azacytidine for 8 days at 250 nM and 1 µM (A549) or at 250 nM only (HTB56). After 8 days, all remaining drugs were washed out and cells were released into azacytidine-free media plus fetal calf serum for an additional 1 week to overcome any direct toxic effects and RNA methylation effects. Functional analyses and intravenous injection were performed at day 15. Reduced representation bisulfate sequencing analyses were performed on day 0, 8, and 15 for both cell lines. b Azacytidine induced changes in DNA methylation. a Scatter plot of DNA methylation levels for more aggressive A549 cells versus more aggressive A549 cells after 8 days of azacytidine exposure at 250 nmol/L. b Scatter plot of DNA methylation levels for more aggressive A549 cells treated with azacytidine at 250 nmol/L versus 1 µmol/L doses. c Scatter plot of DNA methylation differences between day 8 and day 0 in A549 versus HTB56 cells. For this plot, only CpGs with at least 30 % methylation level were analyzed. Many CpGs were similarly hypomethylated in both cell lines (high density of points along the diagonal line in the third quadrant). In both cell lines, multiple CpGs were identified that were hypomethylated in one cell line but not the other (indicated by arrows). Colors represent the density of points ranging from red (high density) to blue (low density). c Scatter plot of azacytidine-treated (250 nM) versus untreated high metastatic HTB56 cells. Methylation levels are shown for CpG sites analyzed by reduced representation bisulfate sequencing. Axes are as follows: Methylation levels in HTB56 high metastatic cells, X-axis: Day 0 (d0 = untreated); Y-axis: Day 8 (d8_250 = after 8 days of azacytidine (250 nM)-treatment). Azacytidine reverts the metastasis-prone phenotype in lung cancer cells
To analyze the effects of DNA methyltransferase inhibition by azacytidine on the non-small cell lung cancer methylome, reduced representation bisulfite sequencing method was used. We identified around 7000 differentially methylated regions in each azacytidine-exposed cell line and found an absolute change in methylation of at least 30 % (Figs. 2b, 3a, b). DNA methylation was decreased in all (97 %) of the modified regions (Figs. 2b(a), 2c). Increase in the dose of azacytidine (from 250 nmol/L to 1 µmol/L) in the A549 cells did not further increase DNA hypomethylation, which is an indication of saturation (Figs. 2b(b), 2c). We also found that multiple sites were hypomethylated only in one cell line (A549 but not in other cell line) (Fig. 2b(c)).
Furthermore, we analyzed the influence of specific transcription factor binding sites (as defined in ChIP-Seq data report) on the loss of DNA methylation with the use of azacytidine. Table 2 shows the consistency of this finding across both the cell lines along with for different drug concentrations. There was no dependence of the effect of azacytidine-associated polycomb/Suz12 targets on absolute methylation levels. We also observed that exonic sequences had a lower number of demethylated sequences than expected, which means that other genomic regions were underrepresented in loss of DNA methylation.
We next analyzed the modifications in DNA methylation patterns just after treating with azacytidine, as well as the recovery phase. After 15 days, methylation levels predominantly remained reduced with few increases, as exception (Fig. 4a, b). Interestingly, A549 cells regained DNA methylation slower than HTB56 cells in spite of the fact that A549 cells were pre-exposed to higher doses of azacytidine (Fig. 3c).
a Scatter plot of azacytidine-treated (250 nM) versus untreated high metastatic HTB56 cells to show the methylation levels for CpG sites analyzed by reduced representation bisulfate sequencing. Axes are as follows: Methylation levels in HTB56 high metastatic cells, X-axis: Day 0 (d0 = untreated); Y-axis: Day 15 (d15_250 = after 15 days of azacytidine (250 nM)-treatment). b, c Scatter plot of DNA methylation levels; b for initial metastasis-prone A549 cells versus A549 cells on day 15 (1 week after release from azacytidine treatment). c for azacytidine-treated; a HTB56 and b A549 on day 8 versus cells after release from azacytidine on day 15. HTB56 cells regained DNA methylation faster than A549 cells. Polycomb-binding sites are preferentially demethylated by azacytidine, and methylation remains decreased after the end of azacytidine exposure
Discussion
We found a good association between the rapid developments of a more aggressive cellular behavior with genome-wide DNA methylation modifications in the present work. We achieved relatively few potentially relevant genetic modifications by exome sequencing or single nucleotide polymorphism in comparison with the hundreds of sites modified in DNA methylation. DNA methylation analyses of biologic replicates have concluded high reproducibility of DNA methylation modifications. The continuous selection of more aggressive subclones and further increase in each round has led to stepwise modifications in each round of in vivo selection. These results supported that the observed modifications are not arbitrary. Furthermore, we demonstrated the occurrence of azacytidine-induced preferential loss of DNA methylation at polycomb target genes, which are known as DNA hypermethylation in cancer. However, DNA methyltransferase inhibition reversed a highly aggressive phenotype in vivo and persisted over time.
We know that there is a close resemblance between the metastatic cells and the parental tumor cells in terms of gene expression profiles, the patterns of somatic mutations, and histology. In comparison with primary tumor which may take multiple years for development, metastatic lesions develop in a much shorter time frame and a core set of driver mutations persists in primary tumors and metastases [30] while there is close resemblance between the pattern of somatic mutations in the primary tumor and the pattern found in metastases. We found analogous results with a very low percentage of additional mutations that were acquired during the selection process to the highly aggressive phenotype. We found that the vast majority of the few acquired mutations with respect to affected genes are mostly arbitrary and that there is no relation with the prometastatic character.
We concluded here stepwise and widespread modifications in DNA methylation patterns in more aggressive lung cancer cells. A likely explanation of the phenotype is based upon epigenetic modifications, which have been concluded on the basis of following observations:
-
(a)
the low number of additional genomic alterations and/or somatic mutations in the more aggressive non-small cell lung cancer cell lines containing almost no gene with a high preponderance for metastases in non-small cell lung cancer;
-
(b)
the stepwise generation of the highly aggressive cells by in vivo selection; and
-
(c)
the use of bulk cell cultures in all experiments with the prominent modifications in DNA methylation.
It was obvious that the observed DNA methylation modifications do not represent a comprehensive feature of metastasis-associated non-small cell lung cancer methylome features. Our aim was to provide evidence for the reversibility, general plasticity, and existence of DNA methylation patterns in the development of aggressive phenotypes. And, to achieve this aim, a relatively simple model with the possibility to recapture a high number of cells was explored, and thus, this xenograft model captured features of highly aggressive tumors and allowed to investigate epigenetic therapy effects.
We found that azacytidine can reverse a metastatic memory in non-small cell lung cancer cells. It might not act by reactivating only a few specific metastasis suppressor genes but by altering the epigenetic landscape. We found that hypermethylation of antimetastatic genes was less frequent in comparison with hypomethylation of prometastatic genes in non-small cell lung cancer cells, and hence, azacytidine found suitable for the model of overall epigenetic disturbance. Our data showed that there is a preference for the hypomethylation of polycomb targets and increased activity near chromosome ends, and hence, we conclude that azacytidine effects were not entirely random throughout the genome and found consistent for both the cell lines. Notably, different concentrations of azacytidine (250 nmol/L or 1 µmol/L) have no effect on this. By using 100 nmol/L concentration of azacytidine in the indicated model in vivo, we confirmed the existence of dose–response, which led to the formation of lung nodules with the same frequency as untreated controls. In vivo selection HTB56 cells showed more modifications in DNA methylation (mostly DNA hypomethylation). The possible reason could be the association of azacytidine sensitivity with the degree of methylation and (or) the rate of methylation modifications in cancer cells.
Next, this is the first ever report of the stability of DNA hypermethylation and its accessibility for DNA hypomethylating. Besides, we showed here that the polycomb target genes show an increased propensity for DNA methylation modifications in pre-established cancer cell lines. Collectively, we concluded here the presence of a strong association between an aggressive phenotype in non-small cell lung cancer cells and profound DNA methylation modifications.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics. CA Cancer J Clin. 2013;63:11–30.
Feinberg AP, Tycko B. The history of cancer epigenetics. Nat Rev Cancer. 2004;4:143–53.
Konishi K, Watanabe Y, Shen L, et al. DNA methylation profiles of primary colorectal carcinoma and matched liver metastasis. PLoS One. 2011;6:e27889.
Baylin SB, Jones PA. A decade of exploring the cancer epigenome-biological and translational implications. Nat Rev Cancer. 2011;11:726–34.
Geutjes EJ, Bajpe PK, Bernards R. Targeting the epigenome for treatment of cancer. Oncogene. 2012;31:3827–44.
Bjornsson HT, Sigurdsson MI, Fallin MD, et al. Intra-individual change over time in DNA methylation with familial clustering. JAMA. 2008;299:2877–83.
Tsai HC, Li H, Van Neste L, et al. Transient low doses of DNA-demethylating agents exert durable antitumor effects on hematological and epithelial tumor cells. Cancer Cell. 2012;21:430–6.
Raynal NJ, Si J, Taby RF, et al. DNA methylation does not stably lock gene expression but instead serves as a molecular mark for gene silencing memory. Cancer Res. 2012;72:1170–81.
Griffiths EA, Gore SD. Epigenetic therapies in MDS and AML. Adv Exp Med Biol. 2013;754:253–83.
Shats I, Gatza ML, Chang JT, et al. Using a stem cell-based signature to guide therapeutic selection in cancer. Cancer Res. 2011;71:1772–80.
Glinsky GV, Berezovska O, Glinskii AB. Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest. 2005;115:1503–21.
Carmona FJ, Villanueva A, Vidal A, et al. Epigenetic disruption of cadherin-11 in human cancer metastasis. J Pathol. 2012;228:230–40.
Bulk E, Hascher A, Liersch R, et al. Adjuvant therapy with small hairpin RNA interference prevents non-small cell lung cancer metastasis development in mice. Cancer Res. 2008;68:1896–904.
Yu J, Bulk E, Ji P, et al. The EPHB6 receptor tyrosine kinase is a metastasis suppressor that is frequently silenced by promoter DNA hypermethylation in non-small cell lung cancer. Clin Cancer Res. 2010;16:2275–83.
Bulk E, Sargin B, Krug U, et al. S100A2 induces metastasis in non-small cell lung cancer. Clin Cancer Res. 2009;15:22–9.
Bisping G, Wenning D, Kropff M, et al. Bortezomib, dexamethasone, and fibroblast growth factor receptor 3-specific tyrosine kinase inhibitor in t(4;14) myeloma. Clin Cancer Res. 2009;15:520–31.
Kulimova E, Oelmann E, Bisping G, et al. Growth inhibition and induction of apoptosis in acute myeloid leukemia cells by new indolinone derivatives targeting fibroblast growth factor, platelet-derived growth factor, and vascular endothelial growth factor receptors. Mol Cancer Ther. 2006;5:3105–12.
Meissner A, Mikkelsen TS, Gu H, et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature. 2008;454:766–70.
Schoofs T, Rohde C, Hebestreit K, et al. DNA methylation changes are a late event in acute promyelocytic leukemia and coincide with loss of transcription factor binding. Blood. 2013;121:178–87.
Fujita PA, Rhead B, Zweig AS, et al. The UCSC Genome Browser database: update 2011. Nucleic Acids Res. 2011;39:D876–82.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–2.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Hebestreit K, Dugas M, Klein HU. Detection of significantly differentially methylated regions in targeted bisulfite sequencing data. Bioinformatics. 2013;29:1647–53.
Hong F, Breitling R, McEntee CW, et al. RankProd: a bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics. 2006;22:2825–57.
Yamamoto G, Nannya Y, Kato M, et al. Highly sensitive method for genomewide detection of allelic composition in nonpaired, primary tumor specimens by use of Affymetrix single-nucleotide-polymorphism genotyping microarrays. Am J Hum Genet. 2007;81:114–26.
Thoennissen NH, Krug UO, Lee DH, et al. Prevalence and prognostic impact of allelic imbalances associated with leukemic transformation of Philadelphia chromosome-negative myeloproliferative neoplasms. Blood. 2010;115:2882–90.
Rius M, Stresemann C, Keller D, et al. Human concentrative nucleoside transporter 1-mediated uptake of 5-azacytidine enhances DNA demethylation. Mol Cancer Ther. 2009;8:225–31.
Rosenbloom KR, Dreszer TR, Pheasant M, et al. ENCODE whole-genome data in the UCSC Genome Browser. Nucleic Acids Res. 2010;38:D620–5.
Thurman RE, Rynes E, Humbert R, et al. The accessible chromatin landscape of the human genome. Nature. 2012;489:75–82.
Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366:883–92.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Shou-Ping Dai and Chao Xie have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Dai, SP., Xie, C., Ding, N. et al. Targeted inhibition of genome-wide DNA methylation analysis in epigenetically modulated phenotypes in lung cancer. Med Oncol 32, 176 (2015). https://doi.org/10.1007/s12032-015-0615-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-015-0615-x