Abstract
Several studies demonstrated the role of the endogenous opioid system in the development of susceptibility to alcohol dependence. Recently, we reported that binge intragastric administration of ethanol induces selective alterations of pronociceptin and prodynorphin gene expression in the rat amygdala complex depending on the days of exposures and on the development of tolerance and dependence. The aim of the present study was to investigate the potential epigenetic mechanisms leading to these alcohol-induced changes in gene expression. Specific histone modifications and DNA methylation at opioid peptide precursor promoters were analyzed by chromatin immunoprecipitation and real-time methylation-specific PCR, respectively. We found a linkage between gene expression alterations and epigenetic modulation at pronociceptin and prodynorphin promoters following alcohol treatment. In animals treated for 1 day, we observed a reversed correlation, with a decrease of histone 3 lysine 27 trimethylation (repressive mark) and an increase of histone 3 lysine 9 acetylation (activating mark), associated with both gene expression up-regulation. In rats treated with alcohol for up to 5 days, we found an increase in histone 3 lysine 9 acetylation in the pronociceptin promoter providing further evidence of the already proposed possible role for histone deacetylases for addiction treatment. No significant alterations in DNA methylation and histone 3 lysine 4 trimethylation following different alcohol exposures were present, suggesting the selectivity of epigenetic effects induced by alcohol. These data demonstrate that ethanol induces selective epigenetic changes, thus better defining the role of opioid peptides in the ethanol-induced effects in the amygdala complex.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
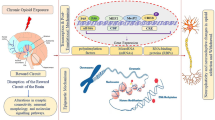
Alcohol exerts its effects interacting with many neurotransmitter systems; much evidence show a strong link between alcohol consumption and the endogenous opioid system. In both humans and animals, ethanol (EtOH) exposure induces changes in opioid peptide systems possibly reflecting their hedonic effects (Oswald and Wand 2004). In particular, EtOH-related alterations of dynorphin (DYN)/κ-opioid receptor (KOP) and nociceptin (NOC)/nociceptin opioid receptor (NOP) systems have been reported, as well as their role within the amygdala complex (AM). This brain area is well known as a key structure playing a crucial role in the alcohol-related behavior and anxiety (Koob 2003, 2006; Pandey et al. 2006).
It has been demonstrated that EtOH induces hyperactivity of the DYN/KOP system and that KOP antagonists reduce alcohol consumption in alcohol-dependent but not in non-dependent animals (Walker and Koob 2008; Walker et al. 2011). Moreover, acute systemic administration of EtOH in rats induces a dose-dependent increase of DYN release in the central nucleus of the AM (Lam et al. 2008).
The NOC/NOP system is involved in the reinforcing or conditioning effects of EtOH and likely plays an important role in the regulation of anxiety related to EtOH seeking (Mogil and Pasternak 2001; Kuzmin et al. 2007). It has been shown that systemic EtOH administration modifies NOC levels in the AM and that NOC injection in the central nucleus of AM inhibits EtOH self-administration in alcohol preferring animals (Lindholm et al. 2002; Economidou et al. 2008). Recently, we observed selective alterations of prodynorphin (PDYN) and pronociceptin (PNOC) opioid gene expression, in the rat AM, following different protocols of alcohol exposure (D'Addario et al. 2011a).
It has also been proposed that epigenetic mechanisms could be relevant in EtOH cellular effects (Shukla and Aroor 2006; Kim and Shukla 2006). Moreover, chronic EtOH exposure, inducing a more open structure of chromatin, can evoke the switch from hetero- to euchromatin (Mahadev and Vemuri 1998). Chromatin remodeling has been also proposed in the AM as an important mechanism involved in the anxiolytic and anxiogenic effects of both EtOH exposure and withdrawal (Pandey et al. 2008).
Two types of functionally interactive epigenetic modifications, i.e., DNA methylation and histone modifications (acetylation, methylation, and phosphorylation), can regulate gene transcription (Grunstein 1997; Turner 2002; Egger et al. 2004; Hsieh and Gage 2005; Verdone et al. 2005). Increasing evidence supports the link between transcriptional activity and changes of histone 3 (H3) lysine methylation (Boggs et al. 2002; Zegerman et al. 2002; Lachner et al. 2001) as well as H3 acetylation (Hebbes et al. 1988). A key role in regulation of gene expression is also played by the methylation of DNA occurring typically at cytosine bases in cytosine–phosphate–guanine (CpG) sites (Gruenbaum et al. 1981). Many findings suggest that hypomethylation of DNA surrounding the promoter region is a prerequisite for gene activation and that a high methylation status may lead to the loss of transcriptional activity (Hsieh 1994; Furuta et al. 2008).
Thus, the aim of present study was to investigate the epigenetic mechanisms leading to the PDYN and PNOC gene expression changes that we previously reported in the rat AM following EtOH exposure and withdrawal (D’Addario et al. 2011a). We analyzed PDYN and PNOC gene promoters with regard to DNA methylation and selected histone modifications: H3 trimethylation of lysine-4 (H3K4me3), H3 acetylation of lysine-9 (H3K9Ac) as activating markers, and H3 trimethylation of lysine-27 (H3K27me3) as repressive marker, to determine the possible link with alcohol-induced gene transcription changes.
Materials and Methods
Animals and Treatments
Briefly, all animal experiments were carried out in accordance with the European Communities Council Directive of 24 November 1986 (86/609/EEC) and National (Ministry of Health) laws and policies (authorization no., 204/2008-B). Care was taken to minimize the number of experimental animals and to take measures to limit their suffering. Male Sprague Dawley rats (Harlan, Italy) (weighing 300–350 g), housed two per cage in standard Macrolon® cages, were maintained on a 12/12-h light/dark cycle with unrestricted access to rat chow and water in a temperature- and humidity-controlled room. Rats were allowed to acclimatize for at least 1 week before the start of experiments.
On the basis of previous reports (D'Addario et al. 2011a; Zhou et al. 2000), a binge alcohol paradigm was chosen to more closely resemble human drinking behavior with several hours of heavy alcohol exposure by the oral route. EtOH or water was administered intragastrically by oral gavage as previously described (D'Addario et al. 2011a). Briefly, animals received daily intragastric administrations of alcohol (20 % in water) at a dose of 1.5 g/kg or equal volume of water in their home cages according to this binge pattern regimen: three times daily at 1.5-h intervals (10:00, 11:30, and 13:00 hours), for a total daily dose of 4.5 g kg−1 day−1. Two groups of rats received EtOH (1D group, n = 7) or water (control group, n = 7) for 1 day, they were killed by decapitation 30 min after the treatment.
EtOH or water (control group) was also administered to other groups of animals (n = 7 for each group) for 5 days and, depending on the end-point and based on the behavioral studies described below, they were named 5D (rats killed 30 min after the last EtOH or water administration) or W-1D (rats killed after 1 day of withdrawal) (D'Addario et al. 2011a).
Brains were placed into an ice-cold matrix and then sliced with a razor blade into coronal sections. Slices containing the amygdala complex (2 mm starting at 1.5 posterior the bregma) were obtained. Sample punches of the amygdala complex (including basolateral complex and central nucleus) were dissected under stereomicroscope in accordance with rat brain atlas (Paxinos et al. 1980), frozen immediately on dry ice and stored at −80 °C until analysis.
The epigenetic analyses were performed on the 1D, 5D, and W-1D animal groups, since at 3 and 7 days after EtOH withdrawal no changes in opioid gene expression were observed (D'Addario et al. 2011a). In a limited number of rats (n = 4), we found that the repeated (3 × 1.5 g/kg) EtOH administration induced blood alcohol levels in a range of 330–425 mg/dl, similar to our previous observation (D'Addario et al. 2011a). In this study, we did not want to introduce the confounding factor of the stress of blood sampling as this may affect epigenetic studies. Moreover, behavioral observations of intoxication and withdrawal resulted to be consistent with previously reported data (D'Addario et al. 2011a) and here not shown to avoid redundancy. We should also specify that part of the AM tissues (n = 4 for each group) belonging to the same set of animals used to perform the already published gene expression analysis (D'Addario et al. 2011a) have been also used in the present study to better clarify the possible role of epigenetic regulation in the gene expression modifications.
Analysis of Histone Modifications by Chromatin Immunoprecipitation and Real-time PCR
Chromatin was prepared from frozen tissues as previously described with minor modifications (Dahl and Collas 2007): proteins were cross-linked to DNA by addition of formaldehyde at a final concentration of 1 % in phosphate-buffered saline containing a broad-range protease inhibitor cocktail (PIC; Roche Diagnostic, Mannheim, Germany) and sodium butyrate (Sigma, St. Louis, MO, USA), for 8 min at room temperature. The cross-linking reaction was quenched by adding glycine to a final concentration of 0.125 M and incubating for 5 min at room temperature, the sample was washed and lysed by resuspending with a pipette in 120 μl of lysis buffer (50 mM Tris–HCL, pH 8, 10 mM EDTA, and 1 % SDS) containing PIC and sodium butyrate (20 mM). The sample was incubated on ice and sonicated for 30 s to shear the DNA to fragments ranging in size from 150 to 700 bp, as analyzed by agarose gel electrophoresis. The lysate was centrifuged at 12,000×g for 10 min at 4 °C and the supernatant transferred into a chilled tube, leaving around 30 μl of buffer with the pellet. Lysis buffer (30 μl) was added, the tube vortexed, centrifuged as before and the new supernatant pooled with the previous one and sonicated again two times for 30 s on ice.
After removing a few microliters to serve as “input” DNA, for each immunoprecipitation, 8 μg of chromatin was diluted 10-fold in RIPA buffer (10 mM Tris–HCl, pH 7.5, 1 mM EDTA, 0.5 mM EGTA, 1 % Triton X-100, 0,1 % SDS, 0.1 % Na-deoxycholate, 140 mM NaCl) containing PIC and incubated overnight by rotation for 2 h at 4 °C with agitation either with no antibody as control or with 1–4 μg of antibody, previously coated with Protein A beads (Invitrogen, Carlsbad, CA, USA), against either H3K4me3 (ab8580, Abcam, Cambridge, MA, USA), H3K27me3 (17-622, Millipore, Bedford, MA, USA), H3K9Ac (17-658, Millipore, Bedford, MA, USA), or normal rabbit IgG (17-658; Millipore, Bedford, MA, USA) as a negative control.
The beads and associated immune complexes were washed three times with RIPA buffer and once with Tris–EDTA buffer. The immune complexes were eluted with elution buffer (20 mM Tris–HCl, 5 mM EDTA, 50 mM NaCl, 1 % SDS) containing proteinase K (50 μg/ml; Quiagen, Valencia, CA, USA) at 68 °C for 2 h, and DNA was recovered by phenol extraction, ethanol precipitated, and resuspended in 50 μl of sterile water. This procedure has been described in more detail elsewhere (Dahl and Collas 2007). Thereafter, real-time qPCR (RT-qPCR) quantification of the genomic sequences from regions in the rat PDYN and PNOC proximal promoter associated with the immunoprecipitated proteins were carried out. The primers used for PCR amplification were designed using Primer 3 software (Rozen and Skaletsky 2000) (see Fig. 1 for details):
PDYN (from −348 to −175 bp)
Forward: (5′-CTGTCTCCTCCCATCTCTGC) and the antisense primer: (5′-TAGCTGCTCCAGGTGATGTG).
PNOC (from −645 to −460 bp)
Forward: (5′-CAGACAGGGAGGACATGGAT) and the antisense primer: (5′-GGACTGCAAAGTGCAGACAA).
The relative abundance was assessed by RT-qPCR using iQ SYBR Green Supermix (Hercules, CA, USA) on a DNA Engine Opticon 2 Continuous Fluorescence Detection System (MJ Research, Waltham, MA, USA). To provide precise quantification of the initial target in each PCR reaction, the amplification plot was examined and the point of early log phase of product accumulation defined by assigning a fluorescence threshold above background, defined as the threshold cycle number or Ct. Differences in threshold cycle number were used to quantify the relative amount of the PCR targets contained within each tube. After PCR, a dissociation curve (melting curve) was constructed in the range of 60 to 95 °C (Lyon et al. 2001) to evaluate the specificity of the amplification products. The relative expression of different amplicons was calculated by the delta-delta Ct (DDCt) method and converted to relative expression ratio (2−DDCt) for statistical analysis (Livak and Schmittgen 2001). All chromatin immunoprecipitation (ChIP) data were normalized to the input DNA amounts (Ct values of immunoprecipitated samples were normalized to Ct values obtained from “input”). In addition, results on DNA of EtOH-treated animals were compared with DNA from vehicle-treated animals (control group). Each ChIP experiment was repeated at least three times.
Analysis of DNA Methylation by Methylation Specific Primer Real-time PCR
Genomic DNA was extracted using TRI Reagent solution (Ambion, Austin, TX, USA) and subjected to bisulfite modification using a commercially available modification Kit (Zymo Research, Irvine, CA, USA). Methylation specific PCR analysis was performed by fluorescence-based real-time PCR using a DNA Engine Opticon 2 Continuous Florescence Detection System (MJ Research). Amplified sequences were located within the promoter region of PDYN and intron 1 of PNOC genes, containing respectively 7 and 11 CpG sites, and in a non-CpG-containig region of beta-actin gene (BACT), that served as control gene (Fig. 1). The percentage of methylation was calculated by the 2−DDCt method (Livak and Schmittgen 2001; Pfaffl 2001), where DDCt = (Ct, Target − Ct, BACT) sample − (Ct, Target − Ct, BACT) fully methylated DNA, multiplied by 100. To generate positive methylated DNA controls, we incubated rat genomic DNA with 4 U M.SssI (CpG methylase) in the presence of S-adenosylmethionine (Zymo Research, Irvine, CA, USA). After methylase treatment, the positive control would have 100 % methylated cytosines in all cytosine sites (Wu et al. 2005). Again, following PCR, to check for the specificity of the amplification products, a melting curve was constructed in the range of 60 to 95 °C (Lyon et al. 2001).
The primers used were:
M_PDYN
Forward: 5′-TTTTATAGTTTAATTACGTAAAGTCGT-3′; Reverse: 5′-AACTCATTACATACCTCATACAACCG-3′;
M_PNOC
Forward 5′-TTTTTGAAATTTAAGGGTTTATTTC-3′; Reverse 5′-AATTTACTACACTCCAATAACYGAT-3′
BACT
Forward 5′-TTTTGAATTTGGGGGTGTGT-3′; Reverse 5′-AAAACCAAACCAACTTCCTTATTCT-3′
Statistical Analysis
EtOH effects on histone modification and DNA methylation alterations were statistically analyzed using the two-tailed Student’s t test, using Prism version 5 (Graph-Pad Software, San Diego, CA) and p values of <0.05 were considered to be statistically significant.
Results
Rats have been subjected to repeated binge EtOH intoxications able to induce tolerance to alcohol effects measured by the progressive decrease of intoxication signs as well as dependence revealed by the occurrence of clear withdrawal signs. Results observed in 1D, 5D, and W-1D groups were consistent with previously reported data and are not shown here (D'Addario et al. 2011a).
Using this experimental procedure, we investigated whether alcohol could evoke epigenetic changes (histone modifications and/or DNA methylation) in the PDYN and PNOC promoter regions, associated with the previously observed gene expression changes caused by different EtOH exposures (D'Addario et al. 2011a).
In the AM of the animals treated for 1 day with EtOH (1D group), we found a significant decrease of H3K27me3 in both PDYN (0.46 ± 0.09 versus control group = 1.00 ± 0.14, p < 0.05) and PNOC (0.45 ± 0.11 versus control group = 1.00, p < 0.05) promoter regions, as well as a significant increase in H3K9Ac in PDYN promoter (1.70 ± 0.30 versus control group = 1.00 ± 0.22, p < 0.05) and in PNOC promoter (1.80 ± 0.26 versus control group = 1.00 ± 0.25, p < 0.05) (Fig. 2). We also observed an increase of H3K9Ac at PNOC gene promoter in the animals treated for 5 days (5D group) (2.40 ± 0.48 versus control group = 1.00 ± 0.14, p < 0.05) (Fig. 3). In contrast, we did not observe any histone modifications changes in the W-1D group in either gene promoters (Fig. 4). Moreover, EtOH exposure did not induce at PDYN and PNOC gene promoters any statistically significant alteration of H3K4me3 (see Figs. 2, 3, and 4) as well as of DNA methylation levels (Table 1) in all groups under study.
RT-qPCR analysis of H3K27me3-, H3K9Ac-, and H3K4me3-immunoprecipitated DNA fragments at a PDYN and b PNOC promoters. Histogram shows specific histone modification levels, normalized to total input DNA, in rats treated with EtOH intragastrically (total daily dose, 4.5 g kg−1 day−1; EtOH, n = 7) or vehicle (cont, n = 7) for 1 day (group 1D). Data are expressed as means ± SE of triplicate independent samples. *P < 0.05 vs. control group
RT-qPCR analysis of H3K27me3-, H3K9Ac-, and H3K4me3-immunoprecipitated DNA fragments at a PDYN and b PNOC promoters. Histogram shows specific histone modification levels, normalized to total input DNA, in rats treated with EtOH intragastrically (total daily dose, 4.5 g kg day; EtOH, n = 7) or vehicle (cont, n = 7) for 5 days (group 5D). Data are expressed as means ± SE of triplicate independent samples. *P < 0.05 vs. control group
RT-qPCR analysis of H3K27me3-, H3K9Ac-, and H3K4me3-immunoprecipitated DNA fragments at a PDYN and b PNOC promoters. Histogram shows the level of specific histone modifications, normalized to total input DNA, in rats treated with EtOH intragastrically (total daily dose, 4.5 g kg−1 day−1; EtOH, n = 7) or vehicle (cont, n = 7) for 5 days, assessed after 1 day of withdrawal (group W-1D). Data are expressed as means ± SE of triplicate independent samples
Discussion
The present study shows that EtOH administration triggers selective epigenetic changes on PDYN and PNOC gene promoter regions in the rat AM.
A nonspecific EtOH-induced chromatin remodeling has already been described in the whole rat brain several years ago, when alterations of the chromatin have been reported in different cerebral cells. In particular, the nuclei of neurons, astrocytes, and oligodendrocytes showed a chromatin relaxation, whereas the microglia presented a condensed chromatin following EtOH exposure (Mahadev and Vemuri 1998).
More recently, Pandey and co-workers highlighted the relationship between chromatin remodeling in the AM and the EtOH-induced anxiolytic or anxiogenic effects during EtOH dependence or withdrawal, respectively. In particular, EtOH administration caused an increased H3 and H4 acetylation and a decreased histone deacetylase activity in the AM whereas an inverse epigenetic pattern occurred during EtOH withdrawal (Pandey et al. 2008).
Selective histone modifications have been demonstrated in vitro: in rat hepatocytes, EtOH induces a post-translational dose-dependent increase of total H3K9Ac (Park et al. 2003). Moreover, a specific histone methylation pattern (H3K4me2 and H3K9me2) has been linked to the up- and down-regulation of selected genes (Pal-Bhadra et al. 2007).
The present data from in vivo experiments revealed an inverse relationship between the H3K27me3, repressive mark, and H3K9Ac, activating mark, in both PDYN and PNOC promoters in the AM of animals treated for 1 day with EtOH (Fig. 2). These epigenetic changes are consistent with our previous data of gene expression showing an up-regulation of PDYN and PNOC (D'Addario et al. 2011a) and also confirm, for the PDYN gene, the presence of a covalently modified chromatin domain, already observed in our laboratory in SH-SY5Y cells exposed to alcohol and acetaldehyde (D'Addario et al. 2011b)
In the 5D EtOH-treated group, no changes of histone modifications have been observed in the PDYN promoter region (Fig. 3a), consistent with the lack of gene expression alterations in these animals at the same time point. In contrast, in the PNOC promoter, only the activating histone modification H3K9Ac was found to be significantly increased (Fig. 3b), in agreement with the previously reported up-regulation of PNOC gene expression. This different trend of epigenetic modifications observed in the PDYN and PNOC gene promoters following 1 and 5 days of EtOH administration could be explained taking into account the different role of DYN and NOC in EtOH-related effects occurring at the AM level and may suggest a different role of the two opioid systems in EtOH-related anxiety.
It has been demonstrated that DYN plays a role in the anxiety related to addictive behaviors (Bruchas et al. 2010; Wee and Koob 2010) and that systemic EtOH administration produces a significant increase of extracellular concentrations of DYN in the AM (Lam et al. 2008). In contrast, the anxiolytic-like effect of NOC is well known (Jenck et al. 2000) and its role in the AM-involving circuitry is also ascertained by evidence that this peptide inhibits norepinephrine release in AM basolateral nucleus (Kawahara et al. 2004). Thus, it is plausible that the significant increase of H3K9Ac, we found in the PNOC promoter at 5D time-point is due to the persistence of the anxiolytic-like effect of NOC during EtOH dependence; in contrast, the DYN-related signal, bringing anxiogenic properties, progressively turns off as the EtOH dependence arises. Moreover, these data are consistent with the above-mentioned study showing that the anxiolytic effects produced by EtOH are associated with an increase in histone acetylation (Pandey et al. 2008).
In the W-1D group, no histone modifications were observed. This result may be explained by different hypothesis. First, it should be taken into account the complexity of gene regulation by histone modifications (Barski et al. 2007), as well as the fact that the multiple modifications may occur cooperatively to prepare the chromatin for transcriptional activation (Wang et al. 2008). Thus, we cannot rule out that other histone modifications, that we did not survey here, might occur and it has to be taken into account that our data focused to the effects of a few types of chromatin modification on the global gene expression. In addition, other studies reported only a partial matching between addictive drug-induced gene expression and histone modification changes in mice (Renthal, et al. 2009) and in human samples (Zhou et al. 2011).
Moreover, in addition to epigenetic mechanisms, the gene expression is also regulated by several components of the complex transcriptional machinery.
The possibility also exists, for PNOC gene, that a long-term maintenance of epigenetic chromatin state (Boggs et al. 2002; Peters et al. 2002), which determines accessibility for transcription factors, might be still present even when the histone modifications are decreasing. In fact, PNOC gene expression is still increased after 1 day of EtOH withdrawal, but not at the same high level as observed during dependence, and it drops to control levels after 3 days of alcohol abstinence (D'Addario et al. 2011b). In this frame, H3K9Ac seems to fade earlier and thus precede gene expression normalization, which occurs after 3 days of withdrawal, since no changes in histone modification were observed in the W-1D group (Fig. 4).
It is currently unknown if and how alterations in chromatin remodeling drive transient or long-lasting effects on gene regulation (Madhani et al. 2008); from the present data, we may deduce that during EtOH withdrawal, at least for the PNOC gene, there is a transient genomic memory triggered by a histone modification (H3K9Ac) occurring immediately at exposure. It is known that an epigenetic change involves feedback and interacting loops: transcriptional activators bind DNA and start the recruitment of transcribing or repressing complexes including nucleosome modifiers (Ptashne and Gann 2001), thus turning genes on even in the absence of the original activator.
In contrast, the increase of PDYN gene expression observed at W-1D is not supported by the increase of H3K9Ac at 5D, probably reflecting a different time-dependent role of DYN in the neurochemical EtOH-related effects.
Moreover, the present data showed that not all of the activating histone modifications are altered since H3K4me3 was unchanged in all experimental groups, thus suggesting a selectivity of epigenetic effects of EtOH.
No significant alterations in DNA methylation following different schedule of EtOH exposures were observed. These findings are in agreement with our previous in vitro data showing no changes in DNA methylation of PDYN promoter in SH-SY5Y cells exposed to EtOH (D'Addario et al. 2011b), but are not matching with changes in DNA methylation in the promoter region of different genes reported in alcohol dependent patients (Bönsch et al. 2005; Hillemacher et al. 2009; Taqi et al. 2011). The discrepancy probably reflects differences of alcohol exposure between the experimental protocol performed in our animal study and the wide variability of EtOH consumption in alcohol dependent patients.
Altogether, our present and previous data further define the role of opioid peptides in the EtOH-induced effects at the AM level; AM is known to represent a major neural substrate for alcohol-related behaviors and anxiety and an important brain area for the acute positive reinforcement of EtOH addiction and the negative reinforcement associated with EtOH abstinence (Koob 2003, 2006).
The DYN/KOP and NOC/NOP systems have been already proposed as targets for new therapeutic strategies. The administration of KOP antagonists is able to inhibit alcohol intake in dependent animals, but not in non-dependent ones (Walker et al. 2011); furthermore, the administration of NOC significantly inhibits EtOH self-administration in alcohol preferring animals (Economidou et al. 2008). The present data provide more specific target at molecular level, which may possibly allow the development of new drugs to counteract EtOH-induced biochemical changes.
Finally, these data concerning the effects of alcohol in the rat AM, underline the relevance of histone acetylation, which was already shown to modulate the effect of cocaine (Kumar et al. 2005). Thus, as previously suggested (Renthal and Nestler 2009), future directions for the treatment of drug addiction will be to deeply investigate the role of histone deacetylases, already proposed as possible target for the treatment of several diseases (e.g., cancer and bipolar disorder) (Spiegel et al. 2011; Machado-Vieira et al. 2011). In our case, drugs acting directly on histone deacetylases at PNOC gene level could be of potential interest for the treatment of alcohol dependence.
References
Barski A, Cuddapah S, Cui K et al (2007) High-resolution profiling of histone methylations in the human genome. Cell 129:823–883
Boggs BA, Cheung P, Heard E et al (2002) Differentially methylated forms of histone H3 show unique association patterns with inactive human X chromosoes. Nat Genet 30:73–76
Bönsch D, Lenz B, Kornhuber J, Bleich S (2005) DNA hypermethylation of the alpha synuclein promoter in patients with alcoholism. Neuroreport 16:167–170
Bruchas MR, Land BB, Chavkin C (2010) The dynorphin/kappa opioid system as a modulator of stress-induced and pro-addictive behaviors. Brain Res 1314:44–55
D'Addario C, Caputi FF, Rimondini R et al (2011a) Different alcohol exposures induce selective alterations on the expression of dynorphin and nociceptin systems related genes in rat brain. Addict Biol. doi:10.1111/j.1369-1600.2011.00326.x
D'Addario C, Johansson S, Candeletti S et al (2011b) Ethanol and acetaldehyde exposure induces specific epigenetic modifications in the prodynorphin gene promoter in a human neuroblastoma cell line. FASEB J 25:1069–1075
Dahl JA, Collas P (2007) Q2ChIP, a quick and quantitative chromatin immunoprecipitation assay, unravels epigenetic dynamics of developmentally regulated genes in human carcinoma cells. Stem Cells 25:1037–1046
Economidou D, Hansson AC, Weiss F et al (2008) Dysregulation of nociceptin/orphanin FQ activity in the amygdala is linked to excessive alcohol drinking in the rat. Biol Psychiatry 64:211–218
Egger G, Liang G, Aparicio A, Jones PA (2004) Epigenetics in human disease and prospects for epigenetic therapy. Nature 429:457–463
Furuta T, Shuto T, Shimasaki S et al (2008) DNA demethylation-dependent enhancement of Toll-like receptor-2 gene expression in cystic fibrosis epithelial cells involves SP1-activated transcription. BMC Mol Biol 9:39
Gruenbaum Y, Stein R, Cedar H, Razin A (1981) Methylation of CpG sequences in eukaryotic DNA. FEBS Lett 124:67–71
Grunstein M (1997) Histone acetylation in chromatin structure and transcription. Nature 389:349–352
Hebbes TR, Thorne AW, Crane-Robinson C (1988) A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J 7:1395–1402
Hillemacher T, Frieling H, Hartl T et al (2009) Promoter specific methylation of the dopamine transporter gene is altered in alcohol dependence and associated with craving. J Psychiatr Res 43:388–392
Hsieh CL (1994) Dependence of transcriptional repression on CpG methylation density. Mol Cell Biol 14:5487–5494
Hsieh J, Gage FH (2005) Chromatin remodeling in neural development and plasticity. Curr Opin Cell Biol 17:664–671
Jenck F, Ouagazzal AM, Pauly-Evers M, Moreau JL (2000) OrphaninFQ: role in behavioral fear responses and vulnerability to stress? Mol Psychiatry 5:572–574
Kawahara Y, Hesselink MB, van Scharrenburg G, Westerink BH (2004) Tonic inhibition by orphanin FQ/nociceptin of noradrenaline neurotransmission in the amygdala. Eur J Pharmacol 485:197–200
Kim JS, Shukla SD (2006) Acute in vivo effect of ethanol (binge drinking) on histone H3 modifications in rat tissues. Alcohol Alcohol 41:126–132
Koob GF (2003) Alcoholism: allostasis and beyond. Alcohol Clin Exp Res 27:232–243
Koob GF (2006) The neurobiology of addiction: a neuroadaptational view relevant for diagnosis. Addiction 101:23–30
Kumar A, Choi KH, Renthal W et al (2005) Chromatin remodeling is a key mechanism underlying cocaine induced plasticity in striatum. Neuron 48:303–314
Kuzmin A, Kreek MJ, Bakalkin G et al (2007) The nociceptin/orphanin FQ receptor agonist Ro 64-6198 reduces alcohol self-administration and prevents relapse-like alcohol drinking. Neuropsychopharmacology 32:902–910
Lachner M, O' CD, Rea S et al (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410:116–120
Lam MP, Marinelli PW, Bai L, Gianoulakis C (2008) Effects of acute ethanol on opioid peptide release in the central amygdala: an in vivo microdialysis study. Psychopharmacology 201:261–271
Lindholm S, Ploj K, Franck J, Nylander I (2002) Nociceptin/orphanin FQ tissue concentration in the rat brain. Effects of repeated ethanol administration at various post-treatment intervals. Prog Neuropsychopharmacol Biol Psychiatry 26:303–306
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
Lyon E, Millson A, Lowery MC et al (2001) Quantificationof HER2/neu gene amplification by competitive PCR using fluorescent melting curve analysis. Clin Chem 47:844–851
Machado-Vieira R, Ibrahim L, Zarate CA Jr (2011) histone deacetylase and mood disorders: epigenetic programming in gene-environment interactions. CNS Neurosci Ther 17:699–704
Madhani HD, Francis NJ, Kingston RE et al (2008) Epegenomics: a roadmap, but to where? Science 322:43–44
Mahadev K, Vemuri MC (1998) Effect of ethanol on chromatin and nonhistone nuclear proteins in rat brain. Neurochem Res 23:1179–1184
Mogil JS, Pasternak GW (2001) The molecular and behavioural pharmacology of the orphanin FQ/nociceptin peptide and receptor family. Pharmacol Rev 53:381–415
Oswald LM, Wand GS (2004) Opioids and alcoholism. Physiol Behav 81:339–358
Pal-Bhadra M, Bhadra U, Jackson DE et al (2007) Distinct methylation patterns in histone H3 at Lys-4 and Lys-9 correlate with up- & down-regulation of genes by ethanol in hepatocytes. Life Sci 81:979–987
Pandey SC, Zhang H, Roy A et al (2006) Central and medial amygdaloid brain-derived neurotrophic factor signaling plays a critical role in alcohol-drinking and anxiety-like behaviors. J Neurosci 26:8320–8331
Pandey SC, Ugale R, Zhang H et al (2008) Brain chromatin remodeling: a novel mechanism of alcoholism. J Neurosci 28:3729–3737
Park PH, Miller R, Shukla SD (2003) Acetylation of histone H3 at lysine 9 by ethanol in rat hepatocytes. Biochem Biophys Res Commun 306:501–504
Paxinos G, Watson CR, Emson PC (1980) AChE-stained horizontal sections of the rat brain in stereotaxic coordinates. J Neurosci Methods 3:129–149
Peters AH, Mermoud JE, O’Carroll D et al (2002) Histone H3 lisyne 9 methylation is an epigenetic imprint of facultative heterochromatin. Nat Genet 30:77–80
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Ptashne M, Gann A (2001) Transcription initiation: imposing specificity by localization. Essays Biochem 37:1–15
Renthal W, Nestler EJ (2009) Histone acetylation in drug addiction. Semin Cell Dev Biol 20:387–394
Renthal W, Kumar A, Xiao G et al (2009) Genome-wide analysis of chromatin regulation by cocaine reveals a role for sirtuins. Neuron 62:335–348
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, New York, pp 365–386
Shukla SD, Aroor AR (2006) Epigenetic effects of ethanol on liver and gastrointestinal injury. World J Gastroenterol 12:5265–5271
Spiegel S, Milstien S, Grant S (2011) Endogenous modulators and pharmacological inhibitors of histone deacetylases in cancer therapy. Oncogene 31:537–551
Taqi MM, Bazov I, Watanabe H et al (2011) Prodynorphin CpG-SNPs associated with alcohol dependence: elevated methylation in the brain of human alcoholics. Addict Biol 16:499–509
Turner BM (2002) Cellular memory and the histone code. Cell 111:285–291
Verdone L, Caserta M, Di Mauro E (2005) Role of histone acetylation in the control of gene expression. Biochem Cell Biol 83:344–353
Walker BM, Koob GF (2008) Pharmacological evidence for a motivational role of kappa-opioid systems in ethanol dependence. Neuropsychopharmacology 33:643–652
Walker BM, Zorrilla EP, Koob GF (2011) Systemic kappa-opioid receptor antagonism by nor-binaltorphimine reduces dependence-induced excessive alcohol self-administration in rats. Addict Biol 16:116–119
Wang Z, Zang C, Rosenfeld JA et al (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40:897–903
Wee S, Koob GF (2010) The role of the dynorphin-kappa opioid system in the reinforcing effects of drugs of abuse. Psychopharmacology 210:121–135
Wu Y, Halverson G, Basir Z et al (2005) Aberrant methylation at HOXA10 may be responsible for its aberrant expression in the endometrium of patients with endometriosis. Am J Obstet Gynecol 193:371–380
Zegerman P, Canas B, Pappin D, Kouzarides T (2002) Histone H3 lysine 4 methylation disrupts binding of nucleosome remodeling and deacetylase (NuRD) repressor complex. J Biol Chem 277:11621–11624
Zhou Y, Franck J, Spangler R et al (2000) Reduced hypothalamic POMC and anterior pituitary CRF1 receptor mRNA levels after acute, but not chronic, daily "binge" intragastric alcohol administration. Alcohol Clin Exp Res 24:1575–1582
Zhou Z, Yuan Q, Mash DC, Goldman D (2011) Substance-specific and shared transcription and epigenetic changes in the human hippocampus chronically exposed to cocaine and alcohol. Proc Natl Acad Sci USA 108:6626–6631
Acknowledgments
This work was supported by grants from PRIN08 to SC, RFO09 to PR, and TERCAS (2009-2012) to MM. We thank D. Carretta, MD for providing critical revision.
Author information
Authors and Affiliations
Corresponding author
Additional information
Claudio D’Addario and Francesca F. Caputi have equal contribution.
Rights and permissions
About this article
Cite this article
D’Addario, C., Caputi, F.F., Ekström, T.J. et al. Ethanol Induces Epigenetic Modulation of Prodynorphin and Pronociceptin Gene Expression in the Rat Amygdala Complex. J Mol Neurosci 49, 312–319 (2013). https://doi.org/10.1007/s12031-012-9829-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-012-9829-y