Abstract
Background
MicroRNAs (miRNAs) are small, non-coding RNAs that are involved in carcinogenesis through posttranscriptional gene regulatory activity. The current study aimed to evaluate serum miR-21 expression levels as potential biomarkers for diagnosis and prognosis of colorectal cancer (CRC) patients.
Methods
Quantitative real-time RT-PCR was applied to determine the relative expression level of miR-21 in serum. At the same time, the sensitivity and specificity of this marker were evaluated by receiver operating characteristic (ROC) curve analysis.
Results
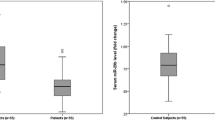
miR-21 expression levels of serum were 3.4 and 1.25 in patient and control, respectively (p < 0.05). The sensitivity and specificity of miR-21 were found to be 95.8% and 91.7%, respectively. The high expression level of serum miR-21 were associated with higher local recurrence, TNM staging, PT staging, venous invasion, liver metastasis, and recurrence (p < 0.05).
Conclusion
The results of this study indicated that miR-21 expression levels in serum can be considered as a novel non-invasive biomarker for early detection and prognosis of CRC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is situated in fourth place in the world regarding neoplastic frequency, it constitutes 9.5% of all cancers worldwide [1]. Whereas in Egypt, it contributes 6.5% of all cancers [2]. In the European Union, colon cancer has the third position among the digestive cancers, with a relative equality regarding the sex ratio, the highest incidence being met in the 6–7 decades of life [3]. In the USA, colorectal cancer (CRC) is the third most common cancer, but the second most deadly one [4].
The 5-year survival rate for patients with localized disease is 89.8%; therefore, colorectal cancer is a potentially curable disease if diagnosed early. However, only 39.6% of patients are diagnosed at this stage [5].
Moreover, as most colorectal cancers develop through a stepwise adenoma–carcinoma sequence or the serrated pathway, most patients would be cured if the disease were detected and resected at a precancerous stage. Therefore, detection of early colorectal neoplasms including precancerous lesions and early colorectal cancers is essential in reducing mortality associated with colorectal cancer. Current guidelines recommend colonoscopy and fecal occult blood tests for colorectal cancer screening [6, 7]. Although colonoscopy is regarded as the gold standard for detecting colorectal neoplasms, this approach has several limitations. It is an invasive and expensive procedure and requires an unpleasant bowel preparation; its efficacy depends on the skill and experience of the endoscopist, and a significant percentage of adults prefer non-invasive options for colorectal cancer screening [8].
Fecal occult blood testing is a commonly used non-invasive test for colorectal cancer screening and has demonstrated a reduction in colorectal cancer mortality from 33 to 15% [8, 9]. However, fecal tests are not recommended for detecting precancerous lesions, due to the limited sensitivity and specificity, which is further compromised by inappropriate handling of specimens or poor adherence to recommended guidelines [8, 10].
The most commonly used blood-based CRC biomarker, carcinoembryonic antigen, which is used for postoperative surveillance and for monitoring response to therapy, lacks sensitivity and specificity for screening or for the detection of recurrent CRC [11]. Thus, alternative minimally invasive or non-invasive tests to detect early colorectal neoplasms are urgently needed.
MicroRNAs (miRNAs) are small (18–25 nucleotides) non-coding RNAs that regulate the translation of specific genes through sequence-specific binding to the 3′ untranslated region of target mRNAs. MiRNAs reportedly have important roles in various cellular processes that are commonly involved in cancer; these processes include cell growth, differentiation, invasion, angiogenesis, and epithelial mesenchymal transition [12, 13]. Due to their oncogenic or tumor-suppressive properties, certain miRNAs participate in carcinogenesis [14, 15]. miRNAs have been identified in body fluids, such as plasma, serum, saliva, urine, and feces [16]. They have been shown to be actively released from cells in microvesicles, exosomes, or bound to proteins. miRNAs are inherently stable accounting for the emerging use as potential biomarkers for human disease and as targets for disease intervention [17, 18].
Circulating miRNAs can withstand unfavorable physiological conditions, such as extreme variations in pH, temperature, and multiple freeze/thaw cycles [19]. Furthermore, the profiles of circulating miRNAs show consistent expression levels across physiologically healthy individuals [20]. To date, several interesting circulating miRNAs have been a candidate as CRC molecular biomarkers; however, it is difficult to draw a definite conclusion [21, 22].
The current study aimed to evaluate serum miR-21 expression levels as potential biomarkers for diagnosis and prognosis of CRC patients.
Subjects and Method
Study Subjects
The study was conducted in Medical Biochemistry Department and the scientific, medical research center (ZSMRC) and department of general surgery. Approval for the study was obtained from the International Review Board (IRB), Faculty of Medicine, Zagazig University. This is a case-control study and 98 adult subjects were included in it. Informed written consents were obtained from all of them to use their samples and clinical data in this study according to the Declaration of Helsinki using a dedicated form. Forty-eight subjects with histopathological confirmation of the diagnosis of CRC, adequate hepatic, renal, cardiac, and respiratory functions were included in the study and individuals with a personal history of malignant tumors in other organs, inflammatory bowel disease, familial adenomatous polyposis, or Lynch syndrome, patients (or their guardians) refusing to participate in study were excluded. In addition, 48 subjects who were enrolled as a healthy control group age and sex-matched with the patients. They randomly selected from various clinics of same ethnic origin and no family history.
Blood Sampling
Whole blood samples were collected from each participant into 5-ml RNase-free tubes. The serum was then separated. The miRNeasy (Cat number: Q217004; Qiagen, Germany) were used to extract miRNAs from serum and detection of serum CEA level were determined by means of an enzyme immunoassay test kit (catalog no. 201–12-1715).
Real-Time RT-PCR
Serum levels of microRNA-21 (miR-21) were examined by real-time RT-PCR using TaqMan MicroRNA Assays (Applied Biosystems, catalog number 4427975). A fixed volume (2 μL) of total RNA was reverse transcribed using TaqMan MicroRNA Reverse Transcription Kits (Applied Biosystems, catalog no. 4366596) in a total volume of 15 μL with the following conditions: 16 °C for 30 min, 42 °C for 30 min, 85 °C for 5 min and maintained at 4 °C. Real-time PCR was conducted using MicroRNA Assay Kits and TaqMan Universal Master Mix II, no UNG (Applied Biosystems, catalog no. 4440040) and performed in duplicate on the StepOne Plus system (Applied Biosystems) with the following cycling conditions: 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Cycle threshold (Ct) values were calculated using Step One Software v2.3 (Applied Biosystems). Expression levels of miRNAs were normalized to those of RNU6 and determined by the 2−ΔΔCt method [23]. ΔCt was calculated as follows: ΔCt = Ct (miRNA of interest)–Ct (RNU6). Then, ΔΔCt was calculated by using a sample from a healthy volunteer as a calibrator: ΔΔCt = ΔCt (tested sample)–ΔCt (calibrator).
Statistical Analysis
Data were analyzed using SPSS version 22, and data were expressed as mean ± SD for quantitative parametric variable, median for non-parametric one, categorical variable expressed as frequency and percentage, Student’s t test, Mann-Whitney, and chi-squared test were used when appropriate. p < 0.05 was considered statistically significant. The analysis was based on the accuracy of the identified miRNAs to diagnose the presence of CRC as determined using receiver operator characteristic (ROC) curves as area under the curve (AUC) value and sensitivity and specificity.
Results
Demographic and Serum miR-21 Expression among the Studied Groups
The studied groups were comparable as regards age and gender distribution.
In comparison to the control group, the miR-21 expression levels in serum in CRC patients were upregulated (p < 0.05). A significant association was found between the miR-21 expression level in serum of CRC patients using the Mann-Whitney statistical test (p < 0.05) (Table 1).
MiR-21 as a Non-invasive Biomarker for CRC Diagnosis
ROC curve analysis indicated that the serum miR-21 expression level could be considered as a promising marker for the diagnosis of CRC patients with a sensitivity and specificity of 95.8% and 91.7%, respectively (an area under the ROC curve, AUC 0.94) (Fig. 1).
Relationship between Serum miR-21 Expression Levels and Clinicopathological Parameters
The clinicopathological variable associated with high miR-21 expression levels were TNM staging, PT classification, size > 5, venous invasion, liver metastasis, and also in recurrence while location, LN metastasis, peritoneal invasion, and chemotherapy were not statistically significant (Table 2, Figs. 2, 3, and 4).
Discussion
Nowadays, non-invasive biomarkers are being widely explored as a reliable tool for cancer diagnosis. Colorectal cancer (CRC) is the second leading cause of cancer death in the world [24].
Early detection of tumor improves the overall survival rate of CRC patients, which highlights an urgent need to find specific, sensitive, and non-aggressive molecular biomarkers suitable for the early diagnosis of CRC [25]. In recent years, there has been an increased interest in finding prognostic biomarkers for CRC to evaluate the expression profiles of single or multiple miRNA in tumor tissues [26, 27].
The findings of the present study demonstrated that miR-21 is up-regulated in serum samples from CRC patients compared to controls. Also, Liu et al. [21] indicated that miR-21 serum level in patient with CRC and advanced adenoma were significantly higher than those in healthy controls.
The sensitivity and specificity of miR-21 expression levels, as a CRC molecular biomarker, were assessed in the serum of CRC patients using ROC curve analysis. MiR-21 expression levels in serum showed high sensitivity and specificity, so had a significant diagnostic value for CRC patients. In a study carried out by Toiyama et al. [26], it turned out that the sensitivity and specificity of serum miR-21 expression levels were 82.8% and 90.6% for CRC detection, respectively. While Huang et al. [28] founded that combined ROC analysis using to target microRNAs yield an increased AUC of 0.88 from 0.84 with 83% sensitivity and 84% specificity in discriminating CRC from controls.
In accordance with our results Liu et al. [21], Toiyama et al. [26] indicated that the miR-21 expression level is increased in different TNM stages of CRC, from the early to later stages.
While Bastaminejad et al. [29] found no obvious differences were detected in miR-21 expression levels between stages III/IV, while a significant increase was found between stages I/II. This difference could be attributed to the number of cases studied in each stage. It is important to note that if the cancer is detected at early stages I/II, the survival rate increases. Considering this fact, using ROC analysis, the sensitivity and specificity of miR-21 expression were assessed in serum of CRC patients to distinguish stages I/II from stages III/IV. Comparison between ROC curves showed statistically significant difference between TNM stage I/II and those TNM stage III/IV. The results of this study suggested that miR-21 expression level in serum could be considered as a valuable marker for CRC prognosis.
It is well known that miR-21 is one of the prominent miRNAs implicated in the genesis, progression, and promotion of tumor growth and the proliferation of human cancer [30]. Until now, the source of circulating serum miRNAs was unclear.
MiR-21 is one of the most representative oncogenic miRNAs, being overexpressed in various types of tumor. Recently, several papers have reported the usefulness of serum miR-21 levels as a biomarker for diagnosis in CRC patients [17]. However, the clinical significance of serum miRNA as a biomarker for diagnosis is still controversial, and some papers have indicated that serological miR-21 does not have any impact on diagnosis in CRC patients [31, 32]. One reason for this controversy may be the instability of miRNA in plasma/serum.
Contrary to the results of this study, the patients with local recurrence and mortality in Menendez et al. [33] found that it exhibited decreased expression of serum miR-21 and when the miR-21 expression levels and survival were analyzed with the Kaplan–Meier method. Although, a reduction in the expression of miR-21 exhibited a trend toward a correlation with poorer survival.
The association between prognosis and serum miR-21 expression might be due to the anti-angiogenic function of miR-21. Studies of miR-21 have shown a correlation between tissue and serum overexpression with a poorer prognosis [34, 35]. To date, there are no studies where the under expression of miR-21 is related to the CRC outcome; therefore, one possible interpretation of the data is that miR-21 demonstrates different behaviors depending on the environment. MiR-21 is overexpressed in most malignancies and favors tumor growth and invasion. However, the results obtained in some studies resemble those published by Sabatel et al. [36]. Who reported that the expression of miR-21 was associated with the inhibition of angiogenesis; the mRNA of which contains a binding site for the 3′UTR of miR-21. Although the measurement of serum miR-21 has demonstrated a relationship with CRC outcome, there are several possible biases.
Our study is a preliminary study in the Egyptian population; we concluded that circulating miR-21 has potential diagnostic value with adequate sensitivity and specificity for CRC and could be considered as a valuable marker for CRC diagnosis. Also, miR-21 may be a promising prognostic tumor marker, and its determination could select candidate patients to a more aggressive adjuvant treatment or use the novel therapeutics options for the correction of the abnormal miRNAs expressions.
References
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55(74–108):3.
El-Bolkainy TN, ASakr M, Nouh AA, El-Din NH. A comparative study of rectal and colonic carcinoma: demographic, pathologic and TNM staging analysis journal of the Egyptian. Natl Cancer Inst. 2006;18:258–63.
Mastalier B, Botezatu C. Colentina surgical clinic experience in treatment of rectal cancer. Romanian statistical review - supliment trimestrul II. 2012:126–34.
Minna J, Schiller JH (2008): Harrison’s principles of internal medicine. New York: McGraw-Hill, 285(16): 12011–12027.
Howlader NNA, Krapcho M, Garshell J, Miller D, Altekruse SF, Kosary CL, et al., editors. SEER Cancer Statistics Review, 1975–2011. Bethesda: National Cancer Institute; 2014. p. 2014.
Rex DK, Johnson DA, Anderson JC, Schoenfeld PS, Burke CA, Inadomi JM. American College of Gastroenterology guidelines for colorectal cancer screening 2009. Am J Gastroenterol. 2009;104:739–50.
Qaseem A, Denberg TD, Hopkins RH Jr, Humphrey LL, Levine J, Sweet DE, et al. Screening for colorectal cancer: a guidance statement from the American College of Physicians. Ann Intern Med. 2012;156:378–86.
Levin B, Lieberman DA, McFarland B, Andrews KS, Brooks D, Bond J, et al. Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: a joint guideline from the American Cancer Society, the US multi-society task force on colorectal Cancer, and the American College of Radiology. Gastroenterology. 2008;134:1570–95.
Mandel JS, Bond JH, Church TR, Snover DC, Bradley GM, Schuman LM, et al. Reducing mortality from colorectal cancer by screening for fecal occult blood. Minnesota Colon Cancer Control Study. N Engl J M. 1993;328:1365–71.
Quintero E, Castells A, Bujanda L, Cubiella J, Salas D, Lanas A, et al. Colonoscopy versus fecal immunochemical testing in colorectal-cancer screening. N Engl J Med. 2012;366:697–706.
Fakih MG, Padmanabhan A. CEA monitoring in colorectal cancer.What you should know. Oncology (Williston Park). 2006;20:579–87.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 2006;6(4):259–69.
Wiemer EA. The role of microRNAs in cancer: no small matter. Eur J Cancer. 2007;43:1529–44.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Chevillet JR, Lee I, Briggs HA, He Y, Wang K. Issues and prospects of microRNA-based biomarkers in blood and other body fluids. Molecules. 2014;19:6080–105.
Kanaan Z, Rai SN, Eichenberger MR, Roberts H, Keskey B, Pan J, et al. Plasma miR-21: a potential diagnostic marker of colorectal cancer. Ann Surg. 2012;256:544–51.
Mishra PJ. Non-coding RNAs as clinical biomarkers for cancer diagnosis and prognosis. Expert Rev Mol Diagn. 2014;14:917–9.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105:10513–8.
Duttagupta R, Jiang R, Gollub J, Getts RC, Jones KW. Impact of cellular miRNAs on circulating miRNA biomarker signatures. PLoS One. 2011;6:e20769.
Liu GH, Zhou ZG, Chen R, Wang MJ, Zhou B, Li Y, et al. Serum miR-21 and miR-92a as biomarkers in the diagnosis and prognosis of colorectal cancer. Tumor Biol. 2013;34(4):2175–81.
Ma Y, Zhang P, Yang J, Liu Z, Yang Z, Qin H. Candidate microRNA biomarkers in human colorectal cancer: systematic review profiling studies and experimental validation. Int J Cancer. 2012;130(9):2077–87.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods. 2001;25(4):402–8.
Siegel R, Desantis C, Jemal A. Colorectal cancer statistics, 2014. CA Cancer J Clin. 2014;64:104–17.
Bresalier RS, Kopetz S, Brenner DE. Blood-based tests for colorectal cancer screening: do they threaten the survival of the FIT test? Dig Dis Sci. 2015;60(3):664–71.
Toiyama Y, Takahashi M, Hur K, Nagasaka T, Tanaka K, Inoue Y, et al. Serum miR-21 as a diagnostic and prognostic biomarker in colorectal cancer. J Natl Cancer Inst. 2013;105(12):849–59.
Shen J, Stass SA, Jiang F. MicroRNAs as potential biomarkers in human solid tumors. Cancer Lett. 2013;329(2):125–36.
Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127:118–26.
Bastaminejad S, Taherikalani M, Ghanbari R, Akbari A, Shabab N, Saidijam M. Investigation of microRNA-21 expression levels in serum and stool as a potential non-invasive biomarker for diagnosis of colorectal cancer. Iran Biomed J. 2016;21:106–13.
Nedaeinia R, Sharifi M, Avan A, Kazemi M, Rafiee L, Ghayour-Mobarhan M, et al. Locked nucleic acid anti-miR-21 inhibits cell growth and invasive behaviors of a colorectal adenocarcinoma cell line: LNA-anti-miR as a novel approach. Cancer Gene Ther. 2016;23:246–53.
Almeida AL, Bernardes MV, Feitosa MR, Peria FM, Tirapelli DP, Rocha JJ, et al. Serological under expression of microRNA-21, microRNA-34a and microRNA-126 in colorectal cancer. Acta Cir Bras. 2016;31:13–8.
Montagnana M, Benati M, Danese E, Minicozzi AM, Paviati E, Gusella M, et al. (2016): plasma expression levels of circulating miR-21 are not useful for diagnosing and monitoring colorectal cancer. Clin Lab. 2016;62:967–70.
Menendez P, Padilla D, Villarejo P, Palomino T, Nieto P, Menendez JM, et al. Prognostic implications of serum microRNA-21 in colorectal cancer. J Surg Oncol. 2013;108:369–73.
Xi Y, Shalgi R, Fodstad O, et al. Differentially regulated micro-RNAs and actively translated messenger RNA transcripts by tumor suppressor p53 in colon cancer. Clin Cancer Res. 2006;12:2014–24.
Slaby O, Svoboda M, Fabian P, Smerdova T, Knoflickova D, Bednarikova M, et al. Altered expression of miR-21, miR-31, miR-143m and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology. 2007;72:397–402.
Sabatel C, Malvaux L, Bovy N, Deroanne C, Lambert V, Gonzalez MLA, et al. MicroRNA-21 exhibits antiangiogenic function by targeting RhoB expression in endothelial cells. PLoS One. 2011;6:e16979.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ghareib, A.F., Mohamed, R.H., Abd el-Fatah, A.R. et al. Assessment of Serum MicroRNA-21 Gene Expression for Diagnosis and Prognosis of Colorectal Cancer. J Gastrointest Canc 51, 818–823 (2020). https://doi.org/10.1007/s12029-019-00306-w
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-019-00306-w