Abstract
Antimicrobial peptides (AMPs), also known as host defense peptides, are petite molecules with inherent microbicidal properties that are synthesized by the host’s innate immune response. These peptides serve as an initial barrier against pathogenic microorganisms, effectively eliminating them. Human defensin (HD) AMPs represent a prominent group of peptides involved in the innate immune response of humans. These peptides are primarily produced by neutrophils and epithelial cells, serving as a crucial defense mechanism against invading pathogens. The extensive research conducted has focused on the broad spectrum of antimicrobial activities and multifaceted immunomodulatory functions exhibited by human defensin AMPs. During the process of co-evolution between hosts and bacterial pathogens, bacteria have developed the ability to recognize and develop an adaptive response to AMPs to counterattack their bactericidal activity by different antibiotic-resistant mechanisms. However, numerous non-pathogenic commensal bacteria elicit the upregulation of defensins as a means to surmount the resistance mechanisms implemented by pathogens. The precise mechanism underlying the induction of HD by commensal organisms remains to be fully understood. This review summarizes the most recent research on the expression of human defensin by pathogens and discusses the various defense mechanisms used by pathogens to counter host AMP production. We also mention recent developments in the commensal induction of defensin AMPs. A better knowledge of the pathogens’ defensin AMP resistance mechanisms and commensals’ induction of AMP expression may shed light on the creation of fresh antibacterial tactics to get rid of bacterial infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The growth and dissemination of drug-resistant bacteria have been attributed to the fields of human and veterinary medicine and agriculture, particularly in developing nations [63]. According to the World Health Organization (WHO), there has been a significant increase in global resistance to conventional antimicrobials, which poses a substantial and concerning threat to public health [97]. Hence, the matter of antibiotic resistance has necessitated the exploration of alternate options to traditional antibiotics that possess unique mechanisms of action and are less susceptible to bacterial resistance. The pursuit of novel antibiotics has sparked significant interest in antimicrobial peptides (AMPs), which are alternatively referred to as host defense peptides [8, 100].
AMPs are a class of bioactive peptides synthesized by various organisms and serve as crucial constituents of their innate immune system. In Eukaryotes, AMPs function as the primary defense mechanism to evade microbial invasions, while in Prokaryotes, they are produced as a competitive strategy to restrict the proliferation of other microorganisms [99]. According to the antimicrobial peptide database (https://aps.unmc.edu/), a total of 3569 AMPs from six life kingdoms were observed. About 2600 active AMPs were identified from animals in Tables 1 and 2.
Natural AMPs have robust and wide-ranging efficacy against several categories of microorganisms, including bacteria, yeasts, fungi, viruses, and parasites [63, 97]. These AMPs demonstrate bacteriostatic, microbicidal, and cytolytic characteristics. In this particular scenario, the utilization of endogenous AMPs as the most recent iteration of compounds exhibiting noteworthy antimicrobial activity is garnering growing interest. This review attempts to describe how bacteria express human defensin and the variety of defense strategies they employ against host AMP synthesis. We also discuss the most recent developments in the induction of defensin AMPs by commensals. It may be possible to develop new antibacterial strategies to combat bacterial infection with the help of a better understanding of the dynamics of human defensin AMP expression, pathogen resistance, and commensal induction.
AMPs and Its Expression
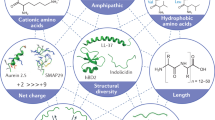
AMPs are important effector molecules that kill bacteria, fungi, and viruses in a wide range. In addition to killing bacteria directly, AMPs have been shown to change many important cell behaviours, such as chemotaxis, cytokine production, epithelial cell proliferation, promoting cell migration, angiogenesis, apoptosis, and wound healing [111]. AMPs are small peptides with anywhere from 12 to 50 amino acids. They are made up of cationic residues like arginine and lysine, which are thought to be responsible for most of their antimicrobial action [105]. Also, these peptides have both water-loving and water-hating parts. Because these peptides are amphipathic, they can work against pathogens where they interact with and become part of the cell walls and membranes of microorganisms [105].
AMPs are frequently categorized based on their origins, encompassing microorganisms, plants, and animals. Additionally, their classification is determined by their secondary structure, biosynthetic pathways, and mode of action [110]. Humans produce various types of AMPs, such as cathelicidins, thrombocidins, and defensins [11, 124, 131]. AMPs have been found to provide protection against microbial infections in humans. They have been detected in various anatomical sites or epithelial surfaces, including the integumentary system, ocular structures, auditory organs, oral cavity, respiratory tract, pulmonary structures, gastrointestinal tract, and the urogenital system. Marchini et al. [101] found human cathelicidin LL-37 in the skin of newborn babies, but human beta-defensin 2 (hBD2) is often found in older people. Gschwandtner et al. [53] found that foetal keratinocytes have a lot more human S100 proteins, hBD2, human beta-defensin 3 (hBD3), and cathelicidin than adult skin cells. Also, human skin that is healthy has different amounts of psoriasin (S100A7), RNase 7, and hBD3 [181]. According to Gläser et al. [50], the upregulation of psoriasin occurs when there is a disruption in the skin barrier. According to McDermott [106], human tears have been found to contain lysozyme and lactoferrin. In a recent study, the role of NLRP3 in antimicrobial peptide expression was studied in infected bladder epithelial cells [90]. It is now recognized that certain defensins also play a critical role in sperm fertilization [163, 164, 189]. AMPs have received more attention because of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) outbreak. This is because scientists are looking for new antiviral molecules to fight COVID-19 disease [81].
According to van der Does et al. [171], numerous AMPs possess immunomodulatory properties and can exhibit both anti-inflammatory and pro-inflammatory effects. The direct interaction between AMPs and the microbiota is thought to occur due to their antimicrobial activity and their expression in epithelial cells. Therefore, it is plausible that AMPs have emerged along the course of evolution as significant modulators and influencers of the microbiota. The accuracy of this concept has been effectively demonstrated through the transgenic overexpression of a single AMP, specifically human defensin 5, within the Paneth cells of the small intestine in mice. The findings of Salzman et al. [133] showed significant alterations in the makeup of microbiota. Furthermore, it appears that the absence of oxygen in the gastrointestinal tract promotes the efficacy of AMP human beta-defensin (hBD)-1 through the process of disulphide bond reduction. The study conducted by Schroeder et al. [140] showed that the effectiveness of hBD1, a human beta-defensin, was significantly enhanced, resulting in its powerful antimicrobial activity against anaerobic gut commensals.
Furthermore, a growing body of evidence suggests that other AMPs, such as RegIIIγ and its human equivalent HIP/PAP [18], as well as the cathelicidin LL-37/CRAMP [187], play a role in preserving the equilibrium of the gut microbiota. Staphylococcus epidermidis, a prevalent commensal bacterium found on the skin, has the ability to stimulate the production of innate defense mediators, including IL-1 beta and the antimicrobial peptide hBD2, in keratinocytes. The induction of AMPs in keratinocytes can also be accomplished by the production of IL-17A by IL-17A + CD8 + T cells, which had been previously activated by CD103 + dendritic cells upon detection of commensal bacteria, such as S. epidermidis [114].
Moreover, certain studies have revealed that psychological stress significantly reduces the expression of AMPs, particularly cathelicidins and defensins, in the dermal layer of mice. This, in turn, heightens their vulnerability to skin infections. Furthermore, it has been observed that the signalling pathway involving transforming growth factor-beta, originating from neurons, regulates the expression of AMPs (specifically cnc-2) in the skin of Caenorhabditis elegans, a nematode species (the skin of Caenorhabditis elegans [1, 190]. Several factors induced the AMPs; however, the bacteria have established the resistant mechanism to escape from the AMPs. One of the major resistant mechanisms that bacteria frequently establish is the downregulation of AMP expression by inhibiting the NF-kB activation in epithelial cells [19, 174]. Building upon this point, recent research has explored the activation of overexpression of AMPs using non-pathogenic stimuli, including biotic and abiotic factors. Non-pathogenic commensal and probiotic bacteria have been employed in this context to induce AMP expression, overcoming the inhibition mechanisms established by bacterial pathogens.
Mechanism of AMP Expression
Pathogen recognition receptors (PRRs) like toll-like receptors (TLRs), which recognize common structural pathogen motifs called pathogen-associated molecular patterns (PAMPs), can turn on AMP expression [111]. Some TLRs reside on the outside of the cell (TLR-1, TLR-2, TLR-4, TLR-5, TLR-6, TLR-10) and recognize unique bacterial compounds like LPS or flagellin. Other TLRs live in the endosomes (TLR-3, TLR-7, TLR-8, TLR-9) and recognize nucleic acids from outside the cell [109]. TLRs activate an intracellular signalling pathway when they bind to an endogenous or external ligand. This cascade can be 88-dependent or MyD88-independent. MyD88 is an adapter molecule that is used by TLRs as a co-stimulatory protein. It helps control how cells respond to PAMPs (pathogen-associated molecular patterns). Depending on which TLRs are turned on, the signalling pathway that follows may include the mitogen-activated protein kinase (MAPK) or other protein kinases. In the big picture, this chain of events turns on transcription factors, such as nuclear factor kappa B (NF-κB), activator protein 1 (AP-1), and interferon regulatory factors 3 and/or 7 (IRF3/7). Because of this, both the innate and adaptive immune responses of the host are triggered, which leads to the production of cytokines that cause inflammation, type 1 interferons, and AMPs (Fig. 1).
Illustration of pathogen-activated antimicrobial peptide expression in human epithelial cells. Initially, the specific toll-like receptor TLR-4 are expressed when pathogenic stimuli are attached to it, eventually using MyD88 and MAL adaptor proteins to activate the intracellular protein kinases (IRAK1 and IRAK4). These protein kinases initiate a signalling cascade of TRAF6, MAPK, and AKT signalling, which initiates nuclear factor kappa B (NF-κB) and activator protein 1 (AP-1). These central regulatory markers activate AMP expression and inflammatory cytokine expression. Abbreviations: MyD88, myeloid differentiation primary response 88; MAL, MyD88 adaptor-like protein; IRAL1, interleukin-1 receptor-associated kinase 1; IRAL4, interleukin-1 receptor-associated kinase 4; TRAF6, tumour necrosis factor receptor (TNFR)-associated factor 6 (TRAF6); MAPK, mitogen-activated protein kinase; AKT, protein kinase B (PKB), also known as Akt
The activation of the NF-κB cascade plays a crucial part in modulating the host’s responses to microbial infection. The innate immune response serves as an initial barrier of defense against various pathogens and is regulated by the NF-κB signalling system. Activation of this pathway subsequently facilitates the upregulation of specific target genes [87]. NF-κB has a crucial role in regulating the immune response and inflammation by increasing the expression of several chemokines (such as CXCL1, CXCL2, CXCL3) and cytokines (including TNF-α, IL-1β, IL-6, IL-8) [60]. The activation of NF-κB has been observed to exert an influence on cellular proliferation and apoptosis through its targeting of Bcl2, IAPs, and cyclins. Furthermore, the activation of NF-κB is crucial for initiating antimicrobial effectors, including AMPs, which play a significant role in eradicating harmful microorganisms [43].
The molecule NF-kB plays a crucial role in activating the immune system; yet, it needs meticulous regulation to prevent the occurrence of excessive inflammation [83]. However, some bacterial pathogens have established resistant mechanisms to hinder AMP expression by the host immune system. Tumour necrosis factor alpha-induced protein 3 (TNFAIP3), commonly referred to as A20, is a zinc finger protein that has a role in negatively regulating NF-kB signalling pathways [19]. The induction of A20 is facilitated by stimuli that activate NF-kB through the presence of NF-kB sites in the A20 promoter [78]. Consequently, this leads to a decrease in NF-kB activity. The termination of NF-kB signalling can be achieved by directly deactivating the upstream signalling molecules, such as TNF, interleukin 17 (IL-17), toll-like receptor 4 (TLR-4), and nucleotide-binding oligomerization domain-containing protein 2 (NOD2) [19, 48].
Analogous to the mechanisms employed in the stimulation of AMPs activation through TLR activation, it has been observed that inflammatory cytokines and mediators can also elicit the expression of AMPs, thereby regulating the immune defense of the host. Upon encountering pathogens, innate immune cells initiate a series of signalling events that trigger the release of inflammatory cytokines. These cytokines serve as molecular messengers, communicating with cells of the adaptive immune system. These cytokines function not only in the modulation of the host’s immune response but also in the upregulation of AMP expression [73]. The cytokines belonging to the interleukin-1 (IL-1) family are widely recognized as potent stimulators of AMPs. There are many small molecules, like vitamin D and fatty acids, that are known to increase the production of AMP. Proteins in the vitamin D receptor (VDR) pathway are found in many epithelial and immune cells, and the VDR pathway is a key part of how AMP production is turned on. There is proof that LL-37 and hBD2 AMP expression goes up directly when the VDR pathway is turned on [51]. Among the different types of AMPs, human defensins are promising AMPs. In humans, defensins showed a broad range of antibacterial activity and were expressed in most of the host cells under different infectious conditions. Nowadays, human defensin-based antimicrobial strategies have gained significant interest due to their wide antimicrobial and anti-inflammatory activity. In this review, we discussed the activation of human defensin antimicrobial peptides in humans by bacterial and viral pathogens.
Human Defensins
Defensins encompass a group of compact, cationic peptides with a molecular weight ranging from 2 to 5 kDa. These peptides serve as host defense molecules and possess a structural framework characterized by a β-sheet core. The term “defensin” was coined by Lehrer in 1985 after the discovery of homologous peptides in human neutrophils by the same laboratory. This core structure is reinforced by the presence of three conserved intramolecular disulphide bonds. The initial discovery of the first mammalian defensin, also known as the microbicidal cationic protein, occurred in 1980 when Lehrer and his colleagues successfully isolated it from rabbit lung macrophages [122, 123]. These peptides, which are of mammalian origins and possess broad antimicrobial activity against bacteria, viruses, and fungi, are characterized by their cationic nature and disulphide stabilization [46, 142]. Mammalian defensins are categorized into three subfamilies, namely α-, β-, and θ-defensins, based on their disulphide structure [44, 86] as shown in Table 3. In the human population, two types of defensins have been found, namely α- and β-defensins (Fig. 2). The θ-defensins, characterized by their distinctive circular shape that is reinforced by three parallel disulphide bonds arranged in a ladder-like pattern, are exclusively present in the leukocytes of Rhesus macaques [159]. Currently, a total of six human α-defensins have been discovered. These defensins can be categorized into two main classes based on their expression patterns and gene structures. The first class is known as myeloid defensins or human neutrophil peptides (HNPs) 1 to 4. The second class is referred to as human (enteric) defensins (HDs) 5 and 6 [6, 85]. Human neutrophil peptides (HNPs) are stored within the azurophilic granules of neutrophils in humans. These HNPs, namely HNPs 1–3, together with their less prevalent counterpart HNP4, collectively constitute approximately 5–7% of the total protein content in neutrophils [165]. Granules containing human neutrophil peptides (HNPs) often undergo limited secretion and are frequently targeted to fuse with phagolysosomes, which are organelles with high levels of HNPs. This process enables the direct elimination of phagocytosed microorganisms by the potent antimicrobial activity of HNPs [40, 165].
Types and structure of human a alpha-defensin and b beta-defensin AMPs. The structure of the defensins was retrieved from Protein Data Bank (https://www.rcsb.org/)
Upon holocrine secretion and neutrophil infiltration during inflammation, HNPs are released into the extracellular milieu through the degranulation of activated neutrophils [42, 45]. HD5 and HD6 are constitutively expressed in and secreted by Paneth cells at the bottom of the small intestinal crypt [6, 29, 49]. While the concentration of HD5 at the luminal surface of the small intestine is estimated to be as high as 50–250 µg/ml, it is significantly lower at the colonic mucosal surface due to the distance from secretion [49]. HD5 ranging from 1 to 50 µg/ml is also found in the vaginal fluid from healthy women [130] and induced in the male and female reproductive tract in response to sexually transmitted infections (STIs) [72, 129, 150]. Although more than 30 β-defensin genes exist in the human genome, only a few have been extensively characterized at the genomic and functional levels [143].
In contrast to the regulation of α-defensin expression, which mostly occurs at the secretory level, the expression of β-defensins is primarily regulated transcriptionally and is limited to the keratinocytes of the skin and epithelial cells. For example, the expression of human β-defensin 1 (HBD1) is consistently present, whereas HBD2 and HBD3 are stimulated by microbial infections and pro-inflammatory cytokines in different epithelial and mucosal tissues [116, 125]. Since their first identification in the early 1980s, defensins have been the subject of extensive research due to their wide-ranging antibacterial properties and diverse immunomodulatory actions in both normal and disease states.
Numerous comprehensive analyses have elucidated a diverse array of intricate molecular and cellular mechanisms through which defensins exert their antibacterial, antiviral, and antifungal activities while also serving as multifaceted immune effectors in the contexts of inflammation, development, and cancer [5, 61, 65, 85, 143]. Defensins are widely recognized as versatile components of the innate immune system, effectively combating microbial infections and earning them the nickname of the “Swiss army knife”. However, the collection of new research has revealed that defensins may have a significant function in the interactions between hosts and pathogens and in the development of tumours. This suggests that the processes via which defensins operate are more intricate than previously believed. The increasing acknowledgment of the dual nature of defensins, with their potential benefits and drawbacks contingent upon their spatial and temporal contexts, motivates us to conduct a comprehensive analysis of the most current scholarly works pertaining to their roles in safeguarding and contributing to the development of illnesses in the context of overall well-being.
Defensins in Viral Infection
Defensins have been found to possess the ability to inactivate and impede the replication of many viruses directly. Extensive research has been conducted to clarify the numerous processes via which defensins exert their antiviral effects [61, 71]. The intricate nature of the involvement of defensins in host-virus interactions becomes apparent when considering the case of human immunodeficiency virus (HIV-1). Initial research findings have indicated that defensins can effectively intervene at various stages of host-virus interactions. The inhibitory effects of HNP1–3, HD5, retrocyclins 1, and retrocyclins 3 have been demonstrated to efficiently impede the adherence of enveloped herpes simplex virus 2 (HSV-2) to host cells. This is achieved by interfering with the interactions between HSV-2 gB and its receptor HSPGs [173]. Defensins can impede the fusion process between virions of various enveloped viruses and their respective host cells.
HNP1 is widely acknowledged for its direct antiviral activity against human immunodeficiency virus (HIV) [21, 98]. In addition, it hinders the uptake of HIV-1 by impeding Env-mediated viral fusion and reducing the expression of CD4 and coreceptor CXCR4 on the surface of host cells [36]. This mechanism, which involves the inhibition of HIV-1 infection by HBD2 and HBD3, has sparked controversy [82, 156]. The inhibitory effect of defensins on viral infection after entry has been documented in various non-enveloped virus families, most notably in the case of human papillomavirus (HPV) [14]. The α-defensins have been shown to inhibit the intracellular uncoating of human papillomavirus (HPV) and its release from cytoplasmic vesicles. This is achieved by stabilizing the viral capsid structure, thereby preventing interactions between viral proteins and the genome with host factors that are necessary for productive infection [14, 56, 160, 177, 178].
Several studies have reported the expression of epithelial-associated defensins (hBD1, hBD2, hBD3, HD5, and HD6) in HPV infection. Several defensins, including hBD1, hBD2, and HD5, are known to be expressed in epithelial cells of the urogenital tract. Therefore, it is likely that these epithelial defensins may also play a role in genital HPV infection. Botez et al. [10] showed an intense expression of hBD1 in the membrane and cytoplasm of cells from the basal layer of patients with HPV-16 and HPV-18 types, which suggest an inhibitory action of this molecule on transduction of these HPV types. hBDs appeared to be ineffective at limiting viral disease progression since papillomas persisted despite the high-level expression of hBDs [27]. A novel mechanism by which high-risk HPV-16, through its well-known virulence factor E6, utilizes the suppression of p53, in the presence of ΔNp63α to induce selective overexpression of hBD3 [35]. In a recent study, HPV infection upregulates HBD2 and HBD3 production in HAEC, and this phenomenon is significantly dependent on the oncogenic potential. Smaller increases in the HBD2 level observed after HR-HPV infection may facilitate processes of cancer genesis of the anogenital region [157].
The study conducted by Chang et al. [21] showed that the presence of HD5 and HD6, which are produced by Neisseria gonorrhoeae infection in a cervicovaginal tissue culture system, leads to an elevation in HIV infectivity. Importantly, this effect was observed to occur in a manner that is independent of CD4 and HIV coreceptor interactions [72]. According to Rapista et al. [132], the proteins HD5 and HD6 facilitate the process of HIV infection by interacting with the viral particle, augmenting its ability to attach to specific host cells. According to Ding et al. [37], the defensins have the ability to counteract the anti-HIV effects of polyanion microbicide candidates, which are substances that inhibit HIV entry. HNP1, a well-researched α-defensin with diverse anti-HIV properties, has been found to possess the ability to disrupt the integrity of epithelial cells. This disruption facilitates the traversal of HIV over epithelial barriers, hence promoting viral infection and dissemination [169].
According to a recent study conducted by Van Cleemput et al. [170], it has been observed that horse herpesvirus type 1, an alpha herpesvirus, exhibits resistance towards equine β-defensins 2–3. Similar to HIV-1, the augmentation of infection by HAdV-D and -F through HNP1 and HD5 is associated with heightened viral attachment to target cells, irrespective of receptor binding [149]. According to Wilson et al. [180], the infection of MAdV-2 is observed to be higher in enteroids that express mouse α-defensins, as opposed to those that do not produce them. This ex vivo investigation provides evidence that α-defensin-mediated viral infection takes place not just in conventional cell cultures but also in situations that mimic physiological settings.
Defensins in Bacterial Infection
Defensins possess the ability to eliminate bacteria or impede their growth by employing various antimicrobial mechanisms, including direct disruption of the bacterial membrane [85] and inhibition of bacterial cell wall synthesis [113, 139]. Defensins have been found to possess the ability to mitigate bacterial infection by the neutralization of produced toxins, as demonstrated in studies conducted by Kim et al. [70], Jin et al. [66], Kudryashova et al. [80], and Lehrer et al. [84]. According to Harder et al. [58], HBD1 and HBD2 demonstrate activity mostly against Gram-negative bacteria. However, HBD3, which possesses a substantially higher cationic charge, exhibits robust bactericidal effects against both Gram-positive and Gram-negative strains [59]. The broad bactericidal activity of HBD3 can be attributed to its highly cationic properties, allowing it to kill bacteria in a structure-independent way [176, 182]. It is worth mentioning that the process of disulphide reduction can transform the bactericidal activity of HBD1, which is initially modest, into a highly effective antimicrobial peptide that can effectively combat opportunistic pathogenic fungi and Gram-positive commensal bacteria [140]. There are also highly favourable evaluations of the antifungal efficacy of defensins, as documented by Ordonez et al. [118] and Parisi et al. [121].
This review proposes to examine the involvement of human defensins in the interactions between hosts and bacteria, with a specific emphasis on contrasting their protective and pathogenic roles. The study conducted by Wilson et al. [179] demonstrates the protective function of mouse intestine α-defensins or cryptdins against Salmonella infection in vivo. It is noteworthy that enteric HD6, despite showing limited bactericidal and membranolytic activity in laboratory settings, effectively shields mice from Salmonella infection by capturing bacteria using a distinctive self-assembled structure known as “nanonets”. This structure prevents direct interaction between the pathogen and the intestinal epithelium of the host [28].
The secretion of HNP1–3 by neutrophils that infiltrate during Staphylococcus aureus infection elicits the release of TNF-α and IFN-γ from macrophages. Consequently, this leads to an augmentation in the phagocytic activity against pathogens. The absence of fimbriae in Shigella provides the pathogen with a distinct bacterial surface. This surface allows HD5 to form structures composed of multiple units, which facilitate Shigella’s adhesion to the host’s epithelial cells. This increased bacterial adhesion, in turn, greatly enhances Shigella’s invasion of host cells. Consequently, this leads to a significantly heightened infection both in vivo and ex vivo. HNP1 has been observed to play a role in facilitating Shigella infection of epithelial cells, as demonstrated by Liao et al. [89].
According to Liao et al. [89], while HNP1 exhibits a lower capacity compared to HD5 in facilitating Shigella adherence, its potent ability to disrupt the epithelial barrier further contributes to the progression of Shigella infection. It is noteworthy to mention that HD5 amplifies the pathogenicity of Shigella in macrophages. Although HD5 enhances the process of phagocytosis of Shigella by macrophages, which is typically detrimental to invading pathogens, it is ineffective in preventing the subsequent escape of Shigella from the phagosome and its multiplication within the host cell. Consequently, the infected macrophages undergo necrosis due to the proliferation of Shigella, leading to the massive release of intracellular bacteria [184].
The antibacterial action of human α-defensins depends on their hydrophobicity and selective cationic, which are segregated within a dimeric structure stabilized by intramolecular disulphide bonds [85]. Numerous mutational investigations have successfully revealed the key functional elements of α-defensins that contribute to the facilitation of viral and bacterial infections [89, 132, 168, 169, 184, 185].
AMP Resistance Mechanism of Bacterial Pathogens
Extensive research has been carried out to explore the molecular mechanism involved in the downregulation of host AMP expression by bacterial pathogens over the past couple of decades. In this chapter, we summarized the recent breakthrough in the direct and indirect mechanisms adopted by human bacterial pathogens to suppress AMP activation by humans, which includes surface charge alteration, external sequestration by secreted or surface-associated molecules, energy-dependent membrane efflux pumps, peptidase degradation, and the immunomodulatory molecule-mediated downregulation of AMP (Fig. 3). By inhibiting or escaping from the AMP attack, the bacterial pathogens colonize and infect the human epithelial cells. However, understanding and inhibiting the pathogen’s resistance mechanism could result in finding a new antimicrobial strategy to treat the bacterial infection [30].
Illustration represents the different resistance mechanisms established by the bacterial pathogens to avoid the human AMP attack. A Electrostatic repulsion of AMPs by increasing the surface charges through modification with cationic molecules. Defensins are positively charged, while the bacterial outer surface is negatively charged; by enhancing the overall positive surface charge with cationic molecules, bacteria avoid the AMP affinity with the outer cell wall. B Trapping of AMPs by surface molecules: bacteria produce proteins and polysaccharides on the outer surface, which directly bind with the AMPs and inactivate them. C Efflux pump systems for AMP resistance: human bacterial pathogens use driven efflux pumps to resist AMPs by pushing AMPs out of the cell membrane by efflux pump transporter proteins. D Proteolytic degradation of AMPs: bacterial pathogens produce some protein-degrading enzymes that actively lyse and inactivate the AMPs. E Regulatory networks of AMP resistance: bacterial pathogens activate the AMP-resistant gene expression according to the extracellular metal ion concentration, which enhances the bacterial membrane rigidity and slows the bacterial growth to avoid the host AMP expression. F Downregulation of host AMP expression by bacterial pathogens: some bacterial pathogens produce some transcriptional factor MxiE and the type III secretion system to modulate the host AMP expression. These factors downregulate the AMP expression by activating the TLR-2 and associated MyD88 and A20 expression, which eventually inhibits the NF-kB expression, resulting in the inhibition of AMP expression. Abbreviations: TLR-2, toll-like receptor 2; MyD88, myeloid differentiation primary response 88; A20, tumour necrosis factor alpha-induced protein 3 (TNFAIP3), commonly referred to as A20
Bacterial Surface Modification
Commonly, human AMPs such as defensins and cathelicidin are cationic in nature, which provides the electrostatic affinity. In contrast, the bacterial outer surface was negatively charged with hydroxylated phospholipids, including phosphatidylglycerol (PG), cardiolipin, and phosphatidylserine [93]. The amphipathic nature of AMPs enables integration with the lipid bilayer of both Gram-positive and Gram-negative bacterial cell walls, virus, and fungi; forms transmembrane pores; and eventually induces cell death [57, 119].
Gram-positive bacteria, in contrast to their Gram-negative counterparts, do not possess an outer membrane. However, the ability of AMPs to bind with the cytoplasmic membrane of Gram-positive bacteria is hindered due to the presence of a thick peptidoglycan layer, which is responsible for its cross-linking. In Gram-negative bacteria, AMPs are required to navigate through the outer membrane envelope, which is primarily made up of negatively charged lipopolysaccharide (LPS) [138]. Additionally, AMPs must cross the periplasmic space located beneath the outer membrane. Both Gram-negative and Gram-positive bacteria develop a prevalent strategy to enhance their overall positive surface charge by modifying themselves with cationic molecules. This modification leads to the electrostatic repulsion of cationic AMPs, thereby preventing AMPs from disrupting the cytoplasmic membrane. Various Gram-negative bacteria employ different mechanisms to decrease the overall negative charge of the lipid A component of lipopolysaccharides (LPS). This is achieved through the incorporation of 4-amino-4-deoxy-L-arabinose (L-Ara4N), phosphoethanolamine (pEtN), or palmitoyl groups [54]. Certain Gram-positive pathogens possess the ability to modify their surface charge through the process of altering teichoic acids (TAs). These TAs are comprised of linear anionic glycopolymers consisting of polyglycerol phosphate and polyribitol phosphate, which are connected by phosphodiester bonds [13, 115].
The process of integrating D-alanyl esters into the cell matrix, facilitated by the activity of four proteins encoded by the dltABCD operon, results in the exposure of a positively charged amino group. This leads to a decrease in the overall negative charge of teichoic acids (TAs) and subsequently reduces the electrostatic attraction between cationic AMPs and the bacterial cell envelope [76, 79]. The MprF, also referred to as LysS, is a peptide resistance factor that exists in multiple forms. The mprF gene encodes it and is a remarkably conserved integral membrane protein with a molecular weight of approximately 97 kDa. Notably, this protein is present in both Gram-positive and Gram-negative bacteria. MprF shows a well-preserved hydrophilic cytoplasmic domain at its C-terminus and a substantial flippase domain at its N-terminus [38]. This protein is accountable for reducing the overall negative charge on the surface of Gram-positive bacteria by integrating L-lysine or L-alanine into the peptidoglycan layer of the cell wall [38, 152].
Adding amine substituents L-Ara4N or pEtN to the LPS complex lowers its net negative surface charge and AMP affinity. This makes Gram-negative bacteria like Salmonella spp. less susceptible to AMP. Adding pEtN to dephosphorylated lipid A of LPS makes H. pylori, a Gram-negative pathogen that causes peptic ulcers and a higher risk of gastrointestinal cancer, more resistant to AMP and less likely to activate the innate immune system [33, 166].
Trapping of AMPs by Surface Molecules
Bacteria may have proteins and polysaccharides on their surface or released into the extracellular space that can bind directly to AMPs. This stops AMPs from reaching their target in the cytoplasmic membrane and prevents the formation of lytic pores. Bacterial pathogens also use the release of bound AMP from the surface of the bacteria as a secondary way to neutralize AMP. Plasminogen is the inactive form of plasmin, a host serine protease that breaks down blood clots and changes the shape of tissues. A plasminogen-activating protein made by S. aureus is called staphylokinase (SK). SK binds and consequently turns off mCRAMP and α-defensins that are synthesized by human neutrophils, such as HNP1–3 [12, 66]. AMP action against S. aureus was cut by more than 80%. Also, S. aureus strains that produce SK are less likely to be killed by α-defensins in a mouse model of arthritis. Adding purified SK to strains that lack SK increased their survival in the presence of α-defensin in a lab setting [66]. Streptococcal inhibitor of complement (SIC), a released hydrophilic GAS protein, binds to and disables human LL-37, α-defensin, and lysozyme to help bacteria stay alive [41, 126].
Several bacterial pathogens have surface shells made up of high molecular mass polysaccharides that help them stay alive inside living things and trap cationic AMPs to stop them from interacting with the surface of the microbe. When AMPs are present, they upregulate the capsular polysaccride synthesis. This makes K. pneumoniae more resistant to polymyxin B, prototamine sulphate, defensin 1, β-defensin 1, and lactoferrin [15]. Delivering K. pneumoniae that is not encased capsular polysaccharide extracts from S. pneumoniae serotype 3 and P. aeruginosa made it more resistant to α-defensin HNP1 [92].
Efflux Pump Systems for AMP Resistance
Efflux pumps are transport proteins found in bacteria that help substrates move from inside cells to the outside world. Adenosine triphosphate–binding cassette (ABC)–human bacterial pathogens use driven efflux pumps to resist AMPs by pushing AMPs out of the cell membrane, where they work, and into the extracellular environment [128]. These pumps have been studied a lot because they play a big part in bacteria being resistant to antibiotics. Several three-component ABC transporters that play a role in AMP resistance have been identified in Gram-positive bacteria, such as SpaFEG in B. subtilis [154] and CprABC in Clostridium difficile [155]. The BceAB transporter system found in B. subtilis [102] and S. aureus [107] is a common two-component system that plays a role in AMP resistance.
The MtrCDE efflux pump is part of the resistance-nodulation-division efflux family and depends on energy. Pathogens N. gonorrhoeae and N. meningitidis use MtrCDE to move AMPs out of the bacterial cytoplasm and periplasmic space to make them less sensitive to LL-37, mCRAMP, PC-8, tachyplesin-1, protegrin-1, and defensins [54], 167, 144]. The sapABCDF operon in S. typhimurium encodes the ABC importer Sap, which is sensitive to AMPs. This makes the bacteria more resistant to protamine, AMP melittin from bees, and crude extracts from human neutrophil granules [103]. Aside from AMP resistance in H. influenzae [103] and H. ducreyi [112], the Sap transporter also helps other Gram-negative species.
Proteolytic Degradation of AMPs
AMPs are not easily broken down by surface-attached or released proteases made by bacteria that cause disease [145]. However, some bacterial proteases that can work on a wide range of substrates help diseases spread by effectively cutting and inactivating AMPs. Proteases from several human pathogens, such as E. faecalis (metallopeptidase gelatinase) [137], S. aureus (aureolysin) [147] and P. mirabilis (50-kDa metalloprotease) [3], cut the human AMP LL-37 and defensins into pieces that do not work. The metalloprotease ZapA is a key part of P. mirabilis’s ability to infect and cause damage. It breaks down antibodies, extracellular matrix molecules, and complement components C1q and C3. It also helps the virus avoid AMP by cutting human β-defensin 1, LL-37, and protegrin-1 [3]. AMPs LL-37 and β-defensin 1 are cut and turned off by ZmpA and ZmpB zinc-dependent metalloproteases in Burkholderia cenocepacia [74].
Having a lot of the outer membrane protease OmpT in enterohemorrhagic E. coli makes it more resistant to AMP by breaking down LL-37 efficiently at dibasic sites [161]. Some other pathogens, like B. anthracis (LL-37), Porphyromonas gingivalis (α- and β-defensins, cecropin B), and Prevotella spp. (brevinin), have been shown to efficiently cleave and inactivate AMPs [2, 17, 162].
Regulatory Networks of AMP Resistance
Bacterial pathogens use two-part regulatory systems to change gene expression in response to metal ion levels outside of cells, metabolic needs, and growth phase, or to trick the host’s innate immune system, which is made up of neutrophils and macrophages in the host tissue. This changes the expression of genes needed for survival and disease progression [30]. AMPs are very hard for some bacteria to kill because they coordinate the upregulation of AMP resistance factors. A two-part system in S. typhimurium called PhoPQ has been studied a lot. It reacts to changes in pH, the amount of magnesium ions (Mg2+), and the presence of cationic AMPs [75]. Besides that, PhoPQ changes the surface charge of LPS and boosts macrophage resistance by increasing the activity of the AMP-degrading outer membrane enzyme PgtE [39, 55].
Human β-defensin 3 (HBD3) causes the cell wall stress response pathway to be turned up in S. aureus to offset the changes that HBD3 makes to peptidoglycan synthesis [188]. When S. aureus is exposed to low levels of magainin 2 and gramicidin D, it makes the membranes more solid, which makes them less susceptible to these AMPs [146]. A rise in the levels of membrane-saturated fatty acids and phosphatidylethanolamine and a fall in the levels of phosphatidylglycerol make the cell membrane less flexible, which makes it more resistant to nisin [32, 172].
Downregulation of Host AMP Expression by Bacterial Pathogens
Low amounts of AMPs are made by epithelial cells and host immune cells at rest, but when bacteria infect them, AMP expression is usually greatly increased. Some bacterial pathogens avoid being killed by the innate immune system because they mess with or lower the amounts of AMP expression in the host [30]. Shigella dysenteriae and S. flexneri lower the levels of LL-37 and β-defensin 1 in intestinal epithelial cells during early infection. This is done through a process that depends on the transcriptional factor MxiE and the type III secretion system. This helps the bacteria survive, colonize, and invade the gastrointestinal tract [64, 151]. P. aeruginosa is a human pathogen that is often found in the lungs of people with cystic fibrosis. It causes the host cysteine protease cathepsins B, L, and S to work to break down and inactivate β-defensins 2 and 3, which stops AMP from getting rid of the bacteria in the airway fluid [158]. According to Chakraborty et al. [20], enterotoxigenic E. coli and V. cholerae exotoxins lower the expression of host cell HBD1 and LL-37. N. gonorrhoeae, on the other hand, lowers the expression of AMP genes [52]. Human pathogens Burkholderia spp. are linked to opportunistic infections in people with cystic fibrosis and chronic granulomatous disease [95]. It is thought that the presence of L-Ara4N in the LPS molecule is what gives this genus its high level of AMP resistance [31, 95]. The alternative sigma factor RpoE controls the expression of Burkholderia genes during stress and, depending on the temperature, helps the cells fight AMP [95, 96].
A recent investigation by Simanski et al. discussed how commensal S. epidermidis can cause A20 to be produced. A20 lowered IL-1b (an inflammatory cytokine) and hBD2 mRNA. This may help S. epidermidis stay on the skin [148]. By changing A20 expression, N. meningitidis slows down the activation of hBD2. There was a rise in A20 expression when N. meningitidis was present but not when L. reuteri was present. NF-kB activity was linked to A20 expression in the opposite way, as expected [174]. When Lactobacillus was given to mice that were sick with N. meningitidis, the disease got better, and the bacteria spread less [4].
The expression of AMP can be restored by substances from outside the body, like butyrate, a short-chain fatty acid made by the microbiota in the gut [64]. Putting lactobacilli into the body as a possible cure for AMPs that are not working right is an interesting idea that needs more research. In this context, the use of non-pathogenic commensal bacteria could increase AMP expression by overcoming the pathogens persuaded AMP inhibition in host cells. Therefore, in this chapter, we discussed the effects of commensal bacteria on human defensin expression over the pathogens’ dampening activity on AMP expression.
Effects of Commensal Bacteria on Defensins
The micro-ecosystem discussed in this context arises from the mutualistic relationship between the host organism and its microbiota, and plays a crucial role in preserving the homeostasis of a healthy individual. Commensal bacteria play a crucial role in supplying the host organism with vital nutrients. According to Mazmanian et al. [104], gut microorganisms have a crucial role in metabolizing indigestible substances, protecting against the colonization of opportunistic infections, contributing to the development of intestinal architecture, and stimulating the immune system, among other functions. In contrast, the host organism supplies the bacteria with essential nutrients and a consistent ecological setting. In a specific instance of microevolution, both host and indigenous microbes have undergone adaptations to establish a mutually beneficial relationship, as documented by Sekirov et al. [141].
It is evident that commensal bacteria, which inhabit different regions of the human body, such as the gastrointestinal tract and respiratory system, are roughly equivalent in number to human cells (approximately a 1:1 ratio). These bacteria have a significant influence on the regulation of immunophysiological processes, encompassing metabolism, development, and protection against pathogens [22, 117]. Multiple recent studies have demonstrated that commensal organisms have a role in enhancing the ability of the gut to resist infections, resulting in a mutually advantageous relationship between the host and the commensal microbiota [69, 153]. Nevertheless, it is possible for disparities to arise within microbial communities, which have been associated with several disorders, including inflammatory bowel disease, allergies, asthma, diabetes, and obesity [26].
Emerging research indicates that probiotic bacteria have the potential to enhance the stability of the gut barrier function by stimulating the production of AMPs, specifically defensins. According to Schlee et al. [136], it has been observed that lactobacilli and other probiotics have the ability to stimulate the intestinal barrier defense system in patients without triggering any inflammatory reactions. A recent study provided evidence that in vitro, E. coli Nissle 1917, along with L. acidophilus, L. fermentum, and P. pentosaceus, as well as VSL no. 3 and other probiotic strains, exhibit a high degree of specificity in activating colonic cells to synthesize defensins [136, 175]. A previous study conducted by Schlee et al. [135] showed that the bacterial culture supernatant of E. coli Nissle 1917 exhibited a 13-fold higher capacity to induce hBD2 compared to the pelleted bacteria. This observation suggests that a soluble component is the primary inducer responsible for this effect.
Limited studies were conducted in order to explore the mechanism of commensal-mediated AMP expression in humans to date. A recent study showed that the non-pathogenic commensal bacteria Escherichia coli Nissle 1917 significantly increased the HBD2 by inducing the NF-kB and AP-1 signalling pathway [175]. Similarly, Liu et al. [91] reported that the probiotic bacteria B. siamensis LF4 can effectively induce the expression of AMPs by TLRs/NLRs-MyD88-dependent signalling and both the downstream MAPKs and NF-κB pathways (Fig. 4). The commensal lactobacilli induce hBD2 expression, while N. meningitidis dampens it in human epithelial cells through NF-kB signaling [174]. However, more studies need to be conducted to find out the more efficient commensal bacteria to induce AMP expression against the dampening effect of pathogens.
Illustration represents the commensal-mediated AMP expression in humans. Non-pathogenic commensal bacteria may enhance the expression of AMPs by counteracting the inhibition of AMPs imposed by pathogens in host cells. Commensal bacteria activate the TLR-4-associated MyD88 signalling, which activates the expression of MAPK, followed by AP-1 followed by NF-kB activation. Abbreviations: TLR-2, toll-like receptor 2; MyD88, myeloid differentiation primary response 88; MAPK, mitogen-activated protein kinase; AP-1, activator protein 1; NF-kB, nuclear factor kappa B (NF-κB)
Limitations
There are a lot of specific ways that defensins could be used to make future medicines, but there are also some problems that are holding them back from becoming real medicines. Some of these problems are the high cost of generation, sensitivity to osmotic pressure, and the fact that it breaks down blood cells in general. This problem has a direct effect on clinical trials, which need a lot of preclinical assessments. Because of these physicochemical features, defensins and other AMPs have not been used as drugs yet. Before these peptides can be used in clinical settings, they need to be better at getting to where they need to go, staying stable inside the body, being delivered precisely, being released under control, and not making the immune system react negatively. To get around these problems, a lot of work has gone into improving deep plasma proteomics and peptides’ half-lives in a number of ways, such as by creating peptidomimetics, counting cyclization, lipidation, a mix of hybrid peptides, or the use of nanocarriers [68, 108]. Moreover, most of the pathogens develop a resistant mechanism against the AMPs.
Using commensal probiotics as effective strategies for the eradication of pathogen colonization has some limitations. The finding of a suitable human-associated commensal probiotic bacterial strain is very important. The bacterial load that is to be administrated to the humans should be studied before the application. Moreover, the expression of AMPs may kill the probiotic bacteria along with the pathogens; in this, the pathogens’ colonization may occur when the commensal population is reduced. However, the use of commensals may cause dysbiosis in the human gut microbiome. Certine primary and clinical study is needed for the safe and effective use of commensals to induce AMPs to eradicate the pathogenic infection.
Conclusion
In reaction to infectious diseases, defensins work as short, cationic, broad-range immunostimulatory to turn on both innate and adaptive immunity. Antimicrobial action against bacteria and viruses was found in a number of human defensins. Because of these traits, they show great promise for being used to make medicines against bacterial and viral infections. However, the bacterial pathogens developed different resistance mechanisms to escape, inactivate, and inhibit the host AMPs. Several bacterial pathogens showed host immunomodulatory activity to inhibit the defensin AMPs in humans. When commensals colonized along with the pathogens on the human cells, they induced AMP production, especially defensins in humans. This immunomodulatory strategy developed by commensal bacterial strain could eradicate the pathogen’s colonization by overcoming their resistance mechanisms. This commensal-mediated AMP expression could be a safer and effective method for the treatment of infectious diseases.
Data Availability
All data were presented in the manuscript.
Abbreviations
- AMPs:
-
Antimicrobial peptides
- HD:
-
Human defense
- WHO:
-
World Health Organization
- RegIIIγ:
-
Regenerating family member 3 gamma
- HIP/PAP:
-
Hepatocarcinoma-intestine-pancreas/pancreatitis associated protein
- CRAMP:
-
Cathelicidin-related AMP
- NF-kB:
-
Nuclear factor kappa B
- PRRs:
-
Pathogen recognition receptors
- TLRs:
-
Toll-like receptors
- PAMPs:
-
Pathogen-associated molecular patterns
- MYD88:
-
Myeloid differentiation primary response 88
- MAPK:
-
Mitogen-activated protein kinase
- AP-1:
-
Activator protein 1
- IRF3/7:
-
Interferon regulatory factors 3 and/or 7
- Bcl2:
-
B-cell lymphoma 2 protein
- IAPs:
-
Inhibitors of apoptosis proteins
- TNFAIP3/A20:
-
Tumour necrosis factor alpha-induced protein 3
- TNF:
-
Tumour necrosis factor
- IL-17:
-
Interleukin 17
- NOD2:
-
Nucleotide-binding oligomerization domain-containing protein 2
- VDR:
-
Vitamin D receptor
- hBD2:
-
Human beta-defensin 2
- HNPs:
-
Human neutrophil peptides
- HD5:
-
Human defensin 5
- HD6:
-
Human defensin 6
- STIs:
-
Sexually transmitted infections
- HSV-2:
-
Herpes simplex virus 2
- CXCR4:
-
Chemokine receptor 4
- HPV:
-
Human papillomavirus
- HAEC:
-
Hirschsprung-associated enterocolitis
- HR-HPV:
-
High-risk human papillomavirus
- HAdV-D and -F:
-
Human adenovirus D and F
- HIV-1:
-
Human immunodeficiency virus
- MAdV-2:
-
Mouse adenovirus
- IFN-γ:
-
Interferon gamma
- MprF:
-
Multiple peptide resistance factor
- SIC:
-
Streptococcal inhibitor of complement
- ABC:
-
Adenosine triphosphate-binding cassette
References
Aberg, K. M., Radek, K. A., Choi, E. H., Kim, D. K., Demerjian, M., Hupe, M., Kerbleski, J., Gallo, R. L., Ganz, T., Mauro, T., Feingold, K. R., & Elias, P. M. (2007). Psychological stress downregulates epidermal antimicrobial peptide expression and increases severity of cutaneous infections in mice. The Journal of Clinical Investigation, 117(11), 3339–3349. https://doi.org/10.1172/JCI31726
Bachrach, G., Altman, H., Kolenbrander, P. E., Chalmers, N. I., Gabai-Gutner, M., Mor, A., Friedman, M., & Steinberg, D. (2008). Resistance of Porphyromonas gingivalis ATCC 33277 to direct killing by antimicrobial peptides is protease independent. Antimicrobial Agents and Chemotherapy, 52(2), 638–642. https://doi.org/10.1128/AAC.01271-07
Belas, R., Manos, J., & Suvanasuthi, R. (2004). Proteus mirabilis ZapA metalloprotease degrades a broad spectrum of substrates, including antimicrobial peptides. Infection and Immunity, 72(9), 5159–5167. https://doi.org/10.1128/IAI.72.9.5159-5167.2004
Belkacem, N., Bourdet-Sicard, R., & Taha, M. K. (2018). Lactobacillus paracasei feeding improves the control of secondary experimental meningococcal infection in flu-infected mice. BMC Infectious Diseases, 18(1), 167. https://doi.org/10.1186/s12879-018-3086-9
Bevins, C. L. (2013). Innate immune functions of alpha-defensins in the small intestine. Digestive Diseases, 31, 299–304.
Bevins, C. L., & Salzman, N. H. (2011). Paneth cells, antimicrobial peptides and maintenance of intestinal homeostasis. Nature Reviews Microbiology, 9, 356–368.
Bharucha, J. P., Sun, L., Lu, W., Gartner, S., & Garzino-Demo, A. (2021). Human Beta-Defensin 2 and 3 Inhibit HIV-1 Replication in Macrophages. Frontiers in Cellular and Infection Microbiology, 11, 535352. https://doi.org/10.3389/fcimb.2021.535352
Bhattacharjya, S., & Straus, S. K. (2020). Design, engineering and discovery of novel a-helical and b-boomerang antimicrobial peptides against drug resistant bacteria. International Journal of Molecular Sciences, 21, 5773. https://doi.org/10.3390/ijms21165773
Bonamy, M., Perrett, T., & Postle, L. (2018). Colouring graphs with sparse neighbourhoods: Bounds and applications. arXiv preprint arXiv, 1810.06704.
Botez, E., Onofrei, P., Laura, S., Bogdan, G., Olinici, D., Oana-Mihaela, C., Alina-Alexandra, P., Dimitriu, D., Solcan, C., Ileana, S., & Carmen, C. E. (2019). Expression of β-defensin-1 and 2 in HPV-induced epithelial lesions. ARS Medica Tomitana, 25, 166–171. https://doi.org/10.2478/arsm-2019-0033
Boyton, R. J., & Openshaw, P. J. (2002). Pulmonary defences to acute respiratory infection. British Medical Bulletin, 61, 1–129.
Braff, M. H., Jones, A. L., Skerrett, S. J., & Rubens, C. E. (2007). Staphylococcus aureus exploits cathelicidin antimicrobial peptides produced during early pneumonia to promote staphylokinase-dependent fibrinolysis. The Journal of Infectious Diseases, 195(9), 1365–1372. https://doi.org/10.1086/513277
Brown, S., Santa Maria, J. P., Jr, & Walker, S. (2013). Wall teichoic acids of gram-positive bacteria. Annual Review of Microbiology, 67, 313–336. https://doi.org/10.1146/annurev-micro-092412-155620
Buck, C. B., Day, P. M., Thompson, C. D., Lubkowski, J., Lu, W., Lowy, D. R., & Schiller, J. T. (2006). Human α-defensins block papillomavirus infection. Proceedings of the National Academy of Sciences of the United States of America, 103(5), 1516–1521.
Campos, M. A., Vargas, M. A., Regueiro, V., Llompart, C. M., Albertí, S., & Bengoechea, J. A. (2004). Capsule polysaccharide mediates bacterial resistance to antimicrobial peptides. Infection and Immunity, 72(12), 7107–7114. https://doi.org/10.1128/IAI.72.12.7107-7114.2004
Cardot-Martin, E., Casalegno, J. S., Badiou, C., Dauwalder, O., Keller, D., Prevost, G., & Dumitrescu, O. (2015). α‐Defensins partially protect human neutrophils against Panton‐Valentine leukocidin produced by Staphylococcus aureus. Letters in Applied Microbiology, 61(2), 158–164.
Carlisle, M. D., Srikantha, R. N., & Brogden, K. A. (2009). Degradation of human alpha- and beta-defensins by culture supernatants of Porphyromonas gingivalis strain 381. Journal of Innate Immunity, 1(2), 118–122. https://doi.org/10.1159/000181015
Cash, H. L., Whitham, C. V., Behrendt, C. L., & Hooper, L. V. (2006). Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science, 313(5790), 1126–1130.
Catrysse, L., Vereecke, L., Beyaert, R., & van Loo, G. (2014). A20 in inflammation and autoimmunity. Trends in Immunology, 35, 22–31. https://doi.org/10.1016/j.it.2013.10.005
Chakraborty, K., Ghosh, S., Koley, H., Mukhopadhyay, A. K., Ramamurthy, T., Saha, D. R., Mukhopadhyay, D., Roychowdhury, S., Hamabata, T., Takeda, Y., & Das, S. (2008). Bacterial exotoxins downregulate cathelicidin (hCAP-18/LL-37) and human beta-defensin 1 (HBD-1) expression in the intestinal epithelial cells. Cellular Microbiology, 10(12), 2520–2537. https://doi.org/10.1111/j.1462-5822.2008.01227.x
Chang, T. L., Vargas, J., Jr., DelPortillo, A., & Klotman, M. E. (2005). Dual role of alpha- defensin-1 in anti-HIV-1 innate immunity. Journal of Clinical Investigation, 115, 765–773.
Charo, I. F., & Ransohoff, R. M. (2006). The many roles of chemokines and chemokine receptors in inflammation. New England Journal of Medicine, 354, 610–621.
Charp, P. A., Rice, W. G., Raynor, R. L., Reimund, E., Kinkade Jr, J. M., Ganz, T., Selsted, M. E., Lehrer, R. I., & Kuo, J. F. (1988). Inhibition of protein kinase C by defensins, antibiotic peptides from human neutrophils. Biochemical Pharmacology, 37(5), 951–956.
Chen, X., Niyonsaba, F., Ushio, H., Okuda, D., Nagaoka, I., Ikeda, S., Okumura, K., & Ogawa, H. (2005). Synergistic effect of antibacterial agents human beta-defensins, cathelicidin LL-37 and lysozyme against Staphylococcus aureus and Escherichia coli. Journal of Dermatological Science, 40(2), 123–132. https://doi.org/10.1016/j.jdermsci.2005.03.014
Chessa, M., Van De Bruaene, A., Farooqi, K., Valverde, I., Jung, C., Votta, E., & Gatzoulis, M. A. (2022). Three-dimensional printing, holograms, computational modelling, and artificial intelligence for adult congenital heart disease care: An exciting future. European Heart Journal, 43(28), 2672–2684.
Choe, H., Farzan, M., Konkel, M., Martin, K., Sun, Y., Marcon, L., Cayabyab, M., Berman, M., Dorf, M.E., Gerard, N. & Sodroski, J. (1998). The orphan seven-transmembrane receptor apj supports the entry of primary T-cell-line- tropic and dualtropic human immunodeficiency virus type 1. Journal of Virology, 72, 6113–6118.
Chong, K. T., Xiang, L., Wang, X., Jun, E. L., Xi, L. F., & Schweinfurth, J. M. (2006). High level expression of human epithelial β-defensins (hBD-1, 2 and 3) in papillomavirus induced lesions. Virology Journal, 3, 1–8. https://doi.org/10.1186/1743-422X-3-75
Chu, H., Pazgier, M., Jung, G., Nuccio, S. P., Castillo, P. A., De Jong, M. F., Winter, M.G., Winter, S.E., Wehkamp, J., Shen, B. & Bevins, C. L. (2012). Human alpha-defensin 6 promotes mucosal innate immunity through self-assembled peptide nanonets. Science, 337(6093), 477–481.
Clevers, H. C., & Bevins, C. L. (2013). Paneth cells: Maestros of the small intestinal crypts. Annual Review of Physiology, 75, 289–311.
Cole, J. N., & Nizet, V. (2016). Bacterial evasion of host antimicrobial peptide defenses. Microbiology Spectrum, 4(1). https://doi.org/10.1128/microbiolspec
Cox, A. D., & Wilkinson, S. G. (1991). Ionizing groups in lipopolysaccharides of Pseudomonas cepacia in relation to antibiotic resistance. Molecular Microbiology, 5(3), 641–646. https://doi.org/10.1111/j.1365-2958.1991.tb00735.x
Crandall, A. D., & Montville, T. J. (1998). Nisin resistance in Listeria monocytogenes ATCC 700302 is a complex phenotype. Applied and Environmental Microbiology, 64(1), 231–237. https://doi.org/10.1128/AEM.64.1.231-237.1998
Cullen, T. W., Giles, D. K., Wolf, L. N., Ecobichon, C., Boneca, I. G., & Trent, M. S. (2011). Helicobacter pylori versus the host: Remodeling of the bacterial outer membrane is required for survival in the gastric mucosa. PLoS Pathogens, 7(12), e1002454. https://doi.org/10.1371/journal.ppat.1002454
Dabirian, S., Taslimi, Y., Zahedifard, F., Gholami, E., Doustdari, F., Motamedirad, M., Khatami, S., Azadmanesh, K., Nylen, S., & Rafati, S. (2013). Human neutrophil peptide-1 (HNP-1): a new anti-leishmanial drug candidate. PLOS Neglected Tropical Diseases, 7(10), e2491.
DasGupta, T., Nweze, E. I., Yue, H., Wang, L., Jin, J., Ghosh, S. K., Kawsar, H. I., Zender, C., Androphy, E. J., Weinberg, A., McCormick, T. S., & Jin, G. (2016). Human papillomavirus oncogenic E6 protein regulates human β-defensin 3 (hBD3) expression via the tumor suppressor protein p53. Oncotarget, 7(19), 27430–27444. https://doi.org/10.18632/oncotarget.8443
Demirkhanyan, L. H., Marin, M., Padilla-Parra, S., Zhan, C., Miyauchi, K., Jean-Baptiste, M., Novitskiy, G., Lu, W., & Melikyan, G. B. (2012). Multifaceted mechanisms of HIV-1 entry inhibition by human alpha-defensin. Journal of Biological Chemistry, 287, 28821–38.
Ding, J., Rapista, A., Teleshova, N., Lu, W., Klotman, M. E., & Chang, T. L. (2011). Mucosal human defensins 5 and 6 antagonise the anti-HIV activity of candidate polyanion microbicides. Journal of Innate Immunity, 3, 208–212.
Ernst, C. M., Staubitz, P., Mishra, N. N., Yang, S. J., Hornig, G., Kalbacher, H., Bayer, A. S., Kraus, D., & Peschel, A. (2009). The bacterial defensin resistance protein MprF consists of separable domains for lipid lysinylation and antimicrobial peptide repulsion. PLoS Pathogens, 5(11), e1000660. https://doi.org/10.1371/journal.ppat.1000660
Ernst, R. K., Guina, T., & Miller, S. I. (2001). Salmonella typhimurium outer membrane remodeling: Role in resistance to host innate immunity. Microbes and Infection, 3(14–15), 1327–1334. https://doi.org/10.1016/s1286-4579(01)01494-0
Faurschou, M., Sorensen, O. E., Johnsen, A. H., Askaa, J., & Borregaard, N. (2002). Defensinrich granules ofhuman neutrophils: Characterisation of secretory properties. Biochimica Et Biophysica Acta, 1591, 29–35.
Frick, I. M., Akesson, P., Rasmussen, M., Schmidtchen, A., & Björck, L. (2003). SIC, a secreted protein of Streptococcus pyogenes that inactivates antibacterial peptides. The Journal of Biological Chemistry, 278(19), 16561–16566. https://doi.org/10.1074/jbc.M301995200
40. Gabay, J. E., Scott, R. W., Campanelli, D., Griffith, J., Wilde, C., Marra, M. N., Seeger, M. & Nathan, C. F. (1989). Antibiotic proteins ofhuman polymorphonuclear leukocytes. Proceedings of The National Academy of Sciences of the United States of America, 86(14), 5610–5614.
Ganesan, S., Aggarwal, K., Paquette, N., & Silverman, N. (2011). NF-κB/Rel proteins and the humoral immune responses of Drosophila melanogaster. Current Topics in Microbiology and Immunology, 349, 25–60. https://doi.org/10.1007/82_2010_107
Ganz, T. (2003). Defensins: Antimicrobial peptides of innate immunity. Nature Reviews Immunology, 3, 710–720.
Ganz, T. (1987). Extracellular release of antimicrobial defensins by human polymorphonuclear leukocytes. Infection and Immunity, 55, 568–571.
Ganz, T., Selsted, M. E., Szklarek, D., Harwig, S. S., Daher, K., Bainton, D. F., & Lehrer, R. I. (1985). Defensins natural peptide antibiotics of human neutrophils. J Clin Invest, 76I, 1427–1435.
García, J. R., Jaumann, F., Schulz, S., Krause, A., Rodríguez-Jiménez, J., Forssmann, U., Adermann, K., Klüver, E., Vogelmeier, C., Becker, D., & Hedrich, R. (2001). Identification of a novel, multifunctional β-defensin (human β-defensin 3) with specific antimicrobial activity: Its interaction with plasma membranes of Xenopus oocytes and the induction of macrophage chemoattraction. Cell and Tissue Research, 306, 257–264.
Garg, A. V., Ahmed, M., Vallejo, A. N., Ma, A., & Gaffen, S. L. (2013). The deubiquitinase A20 mediates feedback inhibition of interleukin-17 receptor signaling. Science Signaling, 6, ra44. https://doi.org/10.1126/scisignal.2003699
Ghosh, D., Porter, E., Shen, B. O., Lee, S. K., Wilk, D., Drazba, J., Yadav, S.P., Crabb, J.W., Ganz, T. & Bevins, C. L. (2002). Paneth cell trypsin is the processing enzyme for human defensin-5. Nature Immunology, 3(6), 583–590.
Gläser, R., Meyer-Hoffert, U., Harder, J., Cordes, J., Wittersheim, M., Kobliakova, J., Fölster-Holst, R., Proksch, E., Schröder, J. M., & Schwarz, T. (2009). The antimicrobial protein psoriasin (S100A7) is upregulated in atopic dermatitis and after experimental skin barrier disruption. The Journal of Investigative Dermatology, 129, 641–649.
Gombart, A. F., Borregaard, N., & Koeffler, H. P. (2005). Human cathelicidin antimicrobial peptide (CAMP) gene is a direct target of the vitamin D receptor and is strongly up-regulated in myeloid cells by 1,25-dihydroxyvitamin D3. FASEB Journal: Official Publication of the Federation of American Societies for Experimental Biology, 19(9), 1067–1077. https://doi.org/10.1096/fj.04-3284com
Gruenheid, S., & Le Moual, H. (2012). Resistance to antimicrobial peptides in Gram-negative bacteria. FEMS Microbiology Letters, 330(2), 81–89. https://doi.org/10.1111/j.1574-6968.2012.02528.x
Gschwandtner, M., Zhong, S., Tschachler, A., Mlitz, V., Karner, S., Elbe-Bürger, A., & Mildner, M. (2014). Fetal human keratinocytes produce large amounts of antimicrobial peptides: Involvement of histone-methylation processes. The Journal of Investigative Dermatology. https://doi.org/10.1038/jid.2014.165
Guilhelmelli, F., Vilela, N., Albuquerque, P., Derengowski, L. S., Silva-Pereira, I., & Kyaw, C. M. (2013). Antibiotic development challenges: The various mechanisms of action of antimicrobial peptides and of bacterial resistance. Frontiers in Microbiology, 4, 353. https://doi.org/10.3389/fmicb.2013.00353
Guina, T., Yi, E. C., Wang, H., Hackett, M., & Miller, S. I. (2000). A PhoP-regulated outer membrane protease of Salmonella enterica serovar typhimurium promotes resistance to alpha-helical antimicrobial peptides. Journal of Bacteriology, 182(14), 4077–4086. https://doi.org/10.1128/JB.182.14.4077-4086.2000
Gulati, N. M., Miyagi, M., Wiens, M. E., Smith, J. G., & Stewart, P. L. (2019). Alpha-defensin HD5 stabilises human papillomavirus 16 capsid/core interactions. Pathog Immun, 4, 196–234.
Gutsmann, T., Hagge, S. O., Larrick, J. W., Seydel, U., & Wiese, A. (2001). Interaction of CAP18-derived peptides with membranes made from endotoxins or phospholipids. Biophysical Journal, 80(6), 2935–2945. https://doi.org/10.1016/S0006-3495(01)76259-5
Harder, J., Bartels, J., Christophers, E., & Schroder, J. M. (1997). A peptide antibiotic from human skin. Nature, 387, 861.
Harder, J., Bartels, J., Christophers, E., & Schroder, J. M. (2001). Isolation and characterisation of human beta -defensin-3, a novel human inducible peptide antibiotic. Journal of Biological Chemistry, 276, 5707–5713.
Hayden, M. S., & Ghosh, S. (2011). NF-κB in immunobiology. Cell Research, 21(2), 223–244.
Holly, M. K., Diaz, K., & Smith, J. G. (2017). Defensins in viral infection and pathogenesis. In: Enquist L editor. Annual review of virology (Vol. 4, pp. 369–91). Department of Microbiology, University of Washington.
Hu, H., Di, B., Tolbert, W. D., Gohain, N., Yuan, W., Gao, P., Ma, B., He, Q., Pazgier, M., & Zhao, L. (2019). Systematic mutational analysis of human neutrophil α-defensin HNP4. Biochim Biophys Acta Biomembr, 1861, 835–844. https://doi.org/10.1016/j.bbamem.2019.01.007
Huan, Y., Kong, Q., Mou, H., & Yi, H. (2020). Antimicrobial peptides: Classification, design, application and research progress in multiple fields. Frontiers in Microbiology, 11(October), 1–21. https://doi.org/10.3389/fmicb.2020.582779
Islam, D., Bandholtz, L., Nilsson, J., Wigzell, H., Christensson, B., Agerberth, B., & Gudmundsson, G. (2001). Downregulation of bactericidal peptides in enteric infections: A novel immune escape mechanism with bacterial DNA as a potential regulator. Nature Medicine, 7(2), 180–185. https://doi.org/10.1038/84627
Jin, G., & Weinberg, A. (2019). Human antimicrobial peptides and cancer. Seminars in Cell & Developmental Biology, 88, 156–162.
Jin, T., Bokarewa, M., Foster, T., Mitchell, J., Higgins, J., & Tarkowski, A. (2004). Staphylococcus aureus resists human defensins by production of staphylokinase, a novel bacterial evasion mechanism. Journal of immunology (Baltimore, Md.: 1950), 172(2), 1169–1176. https://doi.org/10.4049/jimmunol.172.2.1169
Kamli, M. R., Alzahrani, E. A., Albukhari, S. M., Ahmad, A., Sabir, J. S., & Malik, M. A. (2022). Combination effect of novel bimetallic Ag-Ni nanoparticles with fluconazole against Candida albicans. Journal of Fungi, 8(7), 733.
Kaur, G., Poljak, A., Ali, S. A., Zhong, L., Raftery, M. J., & Sachdev, P. (2021). Extending the depth of human plasma proteome coverage using simple fractionation techniques. Journal of Proteome Research, 20(2), 1261–1279. https://doi.org/10.1021/acs.jproteome.0c00670
Kim, C. H., & Broxmeyer, H. E. (1999). Chemokines: Signal lamps for trafficking of T and B cells for development and effector function. Journal of Leukocyte Biology, 65, 6–15.
Kim, C., Gajendran, N., Mittrücker, H. W., Weiwad, M., Song, Y. H., Hurwitz, R., Wilmanns, M., Fischer, G. & Kaufmann, S. H. (2005). Human α-defensins neutralize anthrax lethal toxin and protect against its fatal consequences. Proceedings of the National Academy of Sciences, 102(13), 4830–4835.
Klotman, M. E., & Chang, T. L. (2006). Defensins in innate antiviral immunity. Nature Reviews Immunology, 6, 447–456.
Klotman, M. E., Rapista, A., Teleshova, N., Micsenyi, A., Jarvis, G. A., Lu, W., Porter, E. & Chang, T. L. (2008). Neisseria gonorrhoeae-induced human defensins 5 and 6 increase HIV infectivity: Role in enhanced transmission. The Journal of Immunology, 180(9), 6176–6185.
Kolls, J. K., & Chan, Y. R. (2008). Cytokine-mediated regulation of antimicrobial proteins. Nature Reviews Immunology, 8(11), 829. https://doi.org/10.1038/nri2433
Kooi, C., & Sokol, P. A. (2009). Burkholderia cenocepacia zinc metalloproteases influence resistance to antimicrobial peptides. Microbiology (Reading England), 155(Pt, 9), 2818–2825. https://doi.org/10.1099/mic.0.028969-0
Koprivnjak, T., & Peschel, A. (2011). Bacterial resistance mechanisms against host defense peptides. Cellular and Molecular Life Sciences: CMLS, 68(13), 2243–2254. https://doi.org/10.1007/s00018-011-0716-4
Kovács, M., Halfmann, A., Fedtke, I., Heintz, M., Peschel, A., Vollmer, W., Hakenbeck, R., & Brückner, R. (2006). A functional dlt operon, encoding proteins required for incorporation of d-alanine in teichoic acids in gram-positive bacteria, confers resistance to cationic antimicrobial peptides in Streptococcus pneumoniae. Journal of Bacteriology, 188(16), 5797–5805. https://doi.org/10.1128/JB.00336-06
Kraemer, B. F., Campbell, R. A., Schwertz, H., Cody, M. J., Franks, Z., Tolley, N. D., Kahr, W. H., Lindemann, S., Seizer, P., Yost, C. C., & Zimmerman, G. A. (2011). Novel anti-bacterial activities of β-defensin 1 in human platelets: Suppression of pathogen growth and signaling of neutrophil extracellular trap formation. PLoS Pathogens, 7(11), e1002355.
Krikos, A., Laherty, C. D., & Dixit, V. M. (1992). Transcriptional activation of the tumor necrosis factor alpha-inducible zinc finger protein, A20, is mediated by kappa B elements. Journal of Biological Chemistry, 267, 17971–17976. https://doi.org/10.1016/S0021-9258(19)37138-8
Kristian, S. A., Lauth, X., Nizet, V., Goetz, F., Neumeister, B., Peschel, A., & Landmann, R. (2003). Alanylation of teichoic acids protects Staphylococcus aureus against Toll-like receptor 2-dependent host defense in a mouse tissue cage infection model. The Journal of Infectious Diseases, 188(3), 414–423. https://doi.org/10.1086/376533
Kudryashova, E., Quintyn, R., Seveau, S., Lu, W., Wysocki, V. H., & Kudryashov, D. S. (2014). Human defensins facilitate local unfolding of thermodynamically unstable regions of bacterial protein toxins. Immunity, 41, 709–721.
Kurpe, S. R., Grishin, S. Y., Surin, A. K., Panfilov, A. V., Slizen, M. V., Chowdhury, S. D., & Galzitskaya, O. V. (2020). Antimicrobial and amyloidogenic activity of peptides. Can antimicrobial peptides be used against SARS-Antimicrobial and amyloidogenic activity of peptides. Can antimicrobial peptides be used against SARSCoV-2. International Journal of Molecular Sciences, 21(24), 9552. https://doi.org/10.3390/ijms21249552
Lafferty, M. K., Sun, L., DeMasi, L., Lu, W., & Garzino-Demo, A. (2010). CCR6 ligands inhibit HIV by inducing APOBEC3G. Blood, 115, 1564–1571.
Lee, J. J., Leedale, G. F., & Bradbury, E. M. (Eds) (2002) An Illustrated Guide to the Protozoa. Society of Protozoologists, Lawrence, Kansas, pp 872–951.
Lehrer, R. I., Jung, G., Ruchala, P., Wang, W., Micewicz, E. D., Waring, A. J., Gillespie, E.J., Bradley, K.A., Ratner, A.J., Rest, R.F. & Lu, W. (2009). Human alpha-defensins inhibit hemolysis mediated by cholesterol-dependent cytolysins. Infection and Immunity, 77(9), 4028–4040.
Lehrer, R. I., & Lu, W. (2012). Alpha-defensins in human innate immunity. Immunological Reviews, 245, 84–112.
Lehrer, R. I. (2004). Primate defensins. Nature Reviews Microbiology, 2, 727–738.
Li, C., Wang, S., & He, J. (2019). The two NF-κB pathways regulating bacterial and WSSV infection of shrimp. Frontiers in Immunology, 10, 1785. https://doi.org/10.3389/fimmu.2019.0178
Li, L., Redding, S., & Dongari-Bagtzoglou, A. (2007). Candida glabrata, an emerging oral opportunistic pathogen. Journal of Dental Research, Mar;86(3), 204–215.
Liao, C., Fang, K., Xiao, J., Zhang, W., Zhang, B., Yuan, W., Lu, W. & Xu, D. (2019). Critical determinants of human neutrophil peptide 1 for enhancing host epithelial adhesion of Shigella flexneri Cellular Microbiology, 21(10), e13069.
Lindblad, A., Wu, R., Persson, K., & Demirel, I. (2022). The role of NLRP3 in regulation of antimicrobial peptides and estrogen signaling in UPEC-infected bladder epithelial cells. Cells, 12(18), 2298. https://doi.org/10.3390/cells12182298
Liu, Z. Y., Yang, H. L., Wei, C. Y., Cai, G. H., Ye, J. D., Zhang, C. X., & Sun, Y. Z. (2023). Commensal Bacillus siamensis LF4 induces antimicrobial peptides expression via TLRs and NLRs signaling pathways in intestinal epithelial cells of Lateolabrax maculatus. Fish & Shellfish Immunology, 134, 108634. https://doi.org/10.1016/j.fsi.2023.108634
Llobet, E., Tomás, J. M., & Bengoechea, J. A. (2008). Capsule polysaccharide is a bacterial decoy for antimicrobial peptides. Microbiology (Reading England), 154(Pt 12), 3877–3886. https://doi.org/10.1099/mic.0.2008/022301-0
Lohner, K. (2009). New strategies for novel antibiotics: Peptides targeting bacterial cell membranes. General Physiology and Biophysics, 28(2), 105–116. https://doi.org/10.4149/gpb_2009_02_105
Lough, D., Dai, H., Yang, M., Reichensperger, J., Cox, L., Harrison, C., & Neumeister, M. W. (2013). Stimulation of the follicular bulge LGR5 + and LGR6 + stem cells with the gut-derived human alpha defensin 5 results in decreased bacterial presence, enhanced wound healing, and hair growth from tissues devoid of adnexal structures. Plastic and Reconstructive Surgery, 132(5), 1159–1171.
Loutet, S. A., & Valvano, M. A. (2011). Extreme antimicrobial peptide and polymyxin B resistance in the genus burkholderia. Frontiers in Microbiology, 2, 159. https://doi.org/10.3389/fmicb.2011.00159
Loutet, S. A., Mussen, L. E., Flannagan, R. S., & Valvano, M. A. (2011). A two-tier model of polymyxin B resistance in Burkholderia cenocepacia. Environmental Microbiology Reports, 3(2), 278–285. https://doi.org/10.1111/j.1758-2229.2010.00222.x
Luong, H. X., Thanh, T. T., & Tran, T. H. (2020). Antimicrobial peptides – Advances in Development of Therapeutic Applications. Life Sciences, 260, 118407. https://doi.org/10.1016/j.lfs.2020.118407
Mackewicz, C. E., Yuan, J., Tran, P., Diaz, L., Mack, E., Selsted, M. E., et al. (2003). alpha- Defensins can have anti-HIV activity but are not CD8 cell anti-HIV factors. Aids (London, England), 17, F23-32.
Magana, M., Pushpanathan, M., Santos, A. L., Leanse, L., Fernandez, M., Ioannidis, A., Giulianotti, M.A., Apidianakis, Y., Bradfute, S., Ferguson, A.L. & Tegos, G. P. (2020). The value of antimicrobial peptides in the age of resistance. The Lancet Infectious Diseases, 20(9), e216–e230. https://doi.org/10.1016/S1473-3099
Mahlapuu, M., Björn, C., & Ekblom, J. (2020). Antimicrobial Peptides as Therapeutic Agents: Opportunities and Challenges. Crit. Rev. Biotechnol, 40, 978–992. https://doi.org/10.1080/07388551.2020.1796576
Marchini, G., Lindow, S., Brismar, H., Ståbi, B., Berggren, V., Ulfgren, A. K., Lonne-Rahm, S., Agerberth, B., & Gudmundsson, G. H. (2002). The newborn infant is protected by an innate antimicrobial barrier: Peptide antibiotics are present in the skin and vernix caseosa. British Journal of Dermatology, 147, 1127–1134.
Mascher, T., Margulis, N. G., Wang, T., Ye, R. W., & Helmann, J. D. (2003). Cell wall stress responses in Bacillus subtilis: The regulatory network of the bacitracin stimulon. Molecular Microbiology, 50(5), 1591–1604. https://doi.org/10.1046/j.1365-2958.2003.03786.x
Mason, K. M., Munson, R. S., Jr, & Bakaletz, L. O. (2005). A mutation in the sap operon attenuates survival of nontypeable Haemophilus influenzae in a chinchilla model of otitis media. Infection and Immunity, 73(1), 599–608. https://doi.org/10.1128/IAI.73.1.599-608.2005
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O., & Kasper, D. L. (2005). An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell, 122, 107–118.
McDermott, A. M. (2009). The role of antimicrobial peptides at the ocular surface. Ophthalmic Research, 41(2), 60–75. https://doi.org/10.1159/000187622
McDermott, A. M. (2013). Antimicrobial compounds in tears. Experimental Eye Research, 117, 53–61.
Meehl, M., Herbert, S., Götz, F., & Cheung, A. (2007). Interaction of the GraRS two-component system with the VraFG ABC transporter to support vancomycin-intermediate resistance in Staphylococcus aureus. Antimicrobial Agents and Chemotherapy, 51(8), 2679–2689. https://doi.org/10.1128/AAC.00209-07
Mittal, D., Kaur, G., Singh, P., Yadav, K., & Ali, S. A. (2020). Nanoparticle-based sustainable agriculture and food science: Recent advances and future outlook. Frontiers in Nanotechnology, 2, 579954. https://doi.org/10.3389/fnano.2020.579954
Mogensen, T. H. (2009). Pathogen recognition and inflammatory signaling in innate immune defenses. Clinical Microbiology Reviews, 22(2), 240–273. https://doi.org/10.1128/CMR.00046-08
Moretta, A., Scieuzo, C., Petrone, A. M., Salvia, R., Manniello, M. D., Franco, A., Lucchetti, D., Vassallo, A., Vogel, H., Sgambato, A., & Falabella, P. (2021). Antimicrobial peptides: A new hope in biomedical and pharmaceutical fields. Frontiers in Cellular and Infection Microbiology, 11(June), 1–26. https://doi.org/10.3389/fcimb.2021.668632
Morio, K. A., Sternowski, R. H., & Brogden, K. A. (2023). Induction of endogenous antimicrobial peptides to prevent or treat oral infection and inflammation. Antibiotics (Basel Switzerland), 12(2), 361. https://doi.org/10.3390/antibiotics12020361
Mount, K. L., Townsend, C. A., Rinker, S. D., Gu, X., Fortney, K. R., Zwickl, B. W., Janowicz, D. M., Spinola, S. M., Katz, B. P., & Bauer, M. E. (2010). Haemophilus ducreyi SapA contributes to cathelicidin resistance and virulence in humans. Infection and Immunity, 78(3), 1176–1184. https://doi.org/10.1128/IAI.01014-09
Munch, D., & Sahl, H. G. (2015). Structural variations of the cell wall precursor lipid II in Gram-positive bacteria - Impact on binding and efficacy of antimicrobial peptides. Biochimica Et Biophysica Acta, 1848(11 Pt B), 3062–3071.
Naik, S., Bouladoux, N., Linehan, J. L., Han, S. J., Harrison, O. J., Wilhelm, C., Conlan, S., Himmelfarb, S., Byrd, A.L., Deming, C. & Belkaid, Y. (2015). Commensal-dendritic-cell interaction specifies a unique protective skin immune signature. Nature, 520(7545), 104–108.
Neuhaus, F. C., & Baddiley, J. (2003). A continuum of anionic charge: Structures and functions of D-alanyl-teichoic acids in gram-positive bacteria. Microbiology and Molecular Biology Reviews: MMBR, 67(4), 686–723. https://doi.org/10.1128/MMBR.67.4.686-723.2003
O’Neil, D. A. (2003). Regulation of expression of beta-defensins: Endogenous enteric peptide antibiotics. Molecular Immunology, 40, 445–450.
Olson, T. S., & Ley, K. (2002). Chemokines and chemokine receptors in leukocyte trafficking. American Journal of Physiology: Regulatory, Integrative and Comparative Physiology, 283, R7-28.
Ordonez, S. R., Veldhuizen, E. J. A., van Eijk, M., & Haagsman, H. P. (2017). Role of soluble innate effector molecules in pulmonary defense against fungal pathogens. Frontiers in Microbiology, 8, 2098.
Oren, Z., Lerman, J. C., Gudmundsson, G. H., Agerberth, B., & Shai, Y. (1999). Structure and organization of the human antimicrobial peptide LL-37 in phospholipid membranes: Relevance to the molecular basis for its non-cell-selective activity. The Biochemical Journal, 341(Pt 3), 501–513
Otte, J. M., Neumann, H. M., Brand, S., Schrader, H., Schmidt, W. E., & Schmitz, F. (2009). Expression of beta-defensin 4 is increased in human gastritis. European Journal of Clinical Investigation, 39(2), 126–138.
Parisi, K., Shafee, T. M. A., Quimbar, P., van der Weerden, N. L., Bleackley, M. R., & Anderson, M. A. (2019). The evolution, function and mechanisms of action for plant defensins. Seminars in Cell & Developmental Biology, 88, 107–118.
Patterson-Delafield, J., Martinez, R. J., & Lehrer, R. I. (1980). Microbicidal cationic proteins in rabbit alveolar macrophages: A potential host defense mechanism. Infection and Immunity, 30, 180–192.
Patterson-Delafield, J., Szklarek, D., Martinez, R. J., & Lehrer, R. I. (1981). Microbicidal cationic proteins of rabbit alveolar macrophages: Amino acid composition and functional attributes. Infection and Immunity, 31, 723–731.
Paulsen, F., Pufe, T., Conradi, L., Varoga, D., Tsokos, M., Papendieck, J., & Petersen, W. (2002). Antimicrobial peptides are expressed and produced in healthy and inflamed human synovial membranes. The Journal of Pathology, 198, 369–377.
Pazgier, M., Hoover, D. M., Yang, D., Lu, W., & Lubkowski, J. (2006). Human beta-defensins. Cellular and Molecular Life Sciences, 63, 1294–1313.
Pence, M. A., Rooijakkers, S. H., Cogen, A. L., Cole, J. N., Hollands, A., Gallo, R. L., & Nizet, V. (2010). Streptococcal inhibitor of complement promotes innate immune resistance phenotypes of invasive M1T1 group A Streptococcus. Journal of Innate Immunity, 2(6), 587–595. https://doi.org/10.1159/000317672
Peschel, A., Jack, R. W., Otto, M., Collins, L. V., Staubitz, P., Nicholson, G., Kalbacher, H., Nieuwenhuizen, W. F., Jung, G., Tarkowski, A., van Kessel, K. P., & van Strijp, J. A. (2001). Staphylococcus aureus resistance to human defensins and evasion of neutrophil killing via the novel virulence factor MprF is based on modification of membrane lipids with l-lysine. The Journal of Experimental Medicine, 193(9), 1067–1076. https://doi.org/10.1084/jem.193.9.1067
Piddock, L. J. (2006). Multidrug-resistance efflux pumps - Not just for resistance. Nature Reviews Microbiology, 4(8), 629–636. https://doi.org/10.1038/nrmicro1464
Porter, E., Yang, H., Yavagal, S., Preza, G. C., Murillo, O., Lima, H., Greene, S., Mahoozi, L., Klein-Patel, M., Diamond, G. & Quayle, A. J. (2005). Distinct defensin profiles in Neisseria gonorrhoeae and Chlamydia trachomatis urethritis reveal novel epithelial cell-neutrophil interactions. Infection and Immunity, 73(8), 4823–4833.
Quayle, A. J., Porter, E. M., Nussbaum, A. A., Wang, Y. M., Brabec, C., Yip, K. P., & Mok, S. C. (1998). Gene expression, immunolocalisation, and secretion of human defensin-5 in human female reproductive tract. American Journal of Pathology, 152(5), 1247–1258.
Raj, P. A., & Dentino, A. R. (2002). Current status of defensins and their role in innate and adaptive immunity. Fems Microbiology Letters, 206, 9–18.
Rapista, A., Ding, J., Benito, B., Lo, Y. T., Neiditch, M. B., Lu, W., & Chang, T. L. (2011). Human defensins 5 and 6 enhance HIV-1 infectivity through promoting HIV attachment. Retrovirology, 8(1), 1–10.
Salzman, N. H., Hung, K., Haribhai, D., Chu, H., Karlsson-Sjöberg, J., Amir, E., Teggatz, P., Barman, M., Hayward, M., Eastwood, D. & Bos, N. A. (2010). Enteric defensins are essential regulators of intestinal microbial ecology. Nature Immunology, 11(1), 76–83.
Sass, V., Schneider, T., Wilmes, M., Körner, C., Tossi, A., Novikova, N., Shamova, O., & Sahl, H. G. (2010). Human β-defensin 3 inhibits cell wall biosynthesis in Staphylococci. Infection and Immunity, 78(6), 2793–2800.
Schlee, M., Wehkamp, J., Altenhoefer, A., Oelschlaeger, T. A., Stange, E. F., & Fellermann, K. (2007). The induction of human beta-defensin-2 by the probiotic Escherichia coli Nissle 1917 is mediated through flagellin. Infection and Immunity, 75, 2399–2407.
Schlee, M., Harder, J., Köten, B., Stange, E. F., Wehkamp, J., & Fellermann, K. (2008). Probiotic lactobacilli and VSL#3 induce enterocyte beta-defensin 2. Clinical and Experimental Immunology, 151(3), 528–535. https://doi.org/10.1111/j.1365-2249.2007.03587.x
Schmidtchen, A., Frick, I. M., Andersson, E., Tapper, H., & Björck, L. (2002). Proteinases of common pathogenic bacteria degrade and inactivate the antibacterial peptide LL-37. Molecular Microbiology, 46(1), 157–168. https://doi.org/10.1046/j.1365-2958.2002.03146.x
Schmidtchen, A., Pasupuleti, M., & Malmsten, M. (2014). Effect of hydrophobic modifications in antimicrobial peptides. Advances in Colloid and Interface Science, 205, 265–274. https://doi.org/10.1016/j.cis.2013.06.009
Schneider, T., Kruse, T., Wimmer, R., Wiedemann, I., Sass, V., Pag, U., Jansen, A., Nielsen, A.K., Mygind, P.H., Raventós, D.S. & Kristensen, H. H. (2010). Plectasin, a fungal defensin, targets the bacterial cell wall precursor lipid II. Science, 328(5982), 1168–1172.
Schroeder, B. O., Wu, Z., Nuding, S., Groscurth, S., Marcinowski, M., Beisner, J., Buchner, J., Schaller, M., Stange, E.F. & Wehkamp, J. (2011). Reduction of disulphide bonds unmasks potent antimicrobial activity of human beta-defensin 1. Nature, 469(7330), 419–423.
Sekirov, I., Russell, S. L., Antunes, L. C., & Finlay, B. B. (2010). Gut microbiota in health and disease. Physiological Reviews, 90, 859–904.
Selsted, M. E., Harwig, S. S., Ganz, T., Schilling, J. W., & Lehrer, R. I. (1985). Primary structures of three human neutrophil defensins. J Clin Invest, 76, 1436–1439.
Semple, F., & Dorin, J. R. (2012). beta-Defensins: Multifunctional modulators of infection, inflammation and more. Journal of Innate Immunity, 4, 337–348.
Shafer, W. M., Qu, X., Waring, A. J., & Lehrer, R. I. (1998). Modulation of Neisseria gonorrhoeae susceptibility to vertebrate antibacterial peptides due to a member of the resistance/nodulation/division efflux pump family. Proceedings of the National Academy of Sciences of the United States of America, 95(4), 1829–1833. https://doi.org/10.1073/pnas.95.4.1829
Shinnar, A. E., Butler, K. L., & Park, H. J. (2003). Cathelicidin family of antimicrobial peptides: Proteolytic processing and protease resistance. Bioorganic Chemistry, 31(6), 425–436. https://doi.org/10.1016/s0045-2068(03)00080-4
Shireen, T., Singh, M., Das, T., & Mukhopadhyay, K. (2013). Differential adaptive responses of Staphylococcus aureus to in vitro selection with different antimicrobial peptides. Antimicrobial Agents and Chemotherapy, 57(10), 5134–5137. https://doi.org/10.1128/AAC.00780-13
Sieprawska-Lupa, M., Mydel, P., Krawczyk, K., Wójcik, K., Puklo, M., Lupa, B., Suder, P., Silberring, J., Reed, M., Pohl, J., Shafer, W., McAleese, F., Foster, T., Travis, J., & Potempa, J. (2004). Degradation of human antimicrobial peptide LL-37 by Staphylococcus aureus-derived proteinases. Antimicrobial Agents and Chemotherapy, 48(12), 4673–4679. https://doi.org/10.1128/AAC.48.12.4673-4679.2004
Simanski, M., Erkens, A. S., Rademacher, F., & Harder, J. (2019). Staphylococcus epidermidis-induced interleukin-1 beta and human beta-defensin-2 expression in human keratinocytes is regulated by the host molecule A20 (TNFAIP3). Acta Dermato-Venereologica, 99(2), 181–187. https://doi.org/10.2340/00015555-3073
Smith, J. G., Silvestry, M., Lindert, S., Lu, W., Nemerow, G. R., & Stewart, P. L. (2010). Insight into the mechanisms of adenovirus capsid disassembly from studies of defensin neutralisation. Plos Pathogens, 6, e1000959.
Spencer, J. D., Hains, D. S., Porter, E., Bevins, C. L., DiRosario, J., Becknell, B., Wang, H. & Schwaderer, A. L. (2012). Human alpha defensin 5 expression in the human kidney and urinary tract. PLoS One1, 7(2), e31712.
Sperandio, B., Regnault, B., Guo, J., Zhang, Z., Stanley, S. L., Jr, Sansonetti, P. J., & Pédron, T. (2008). Virulent Shigella flexneri subverts the host innate immune response through manipulation of antimicrobial peptide gene expression. The Journal of Experimental Medicine, 205(5), 1121–1132. https://doi.org/10.1084/jem.20071698
Staubitz, P., Neumann, H., Schneider, T., Wiedemann, I., & Peschel, A. (2004). MprF-mediated biosynthesis of lysylphosphatidylglycerol, an important determinant in staphylococcal defensin resistance. FEMS Microbiology Letters, 231(1), 67–71. https://doi.org/10.1016/S0378-1097(03)00921-2
Stein, J. V., & Nombela-Arrieta, C. (2005). Chemokine control of lymphocyte trafficking: A general overview. Immunology, 116, 1–12.
Stein, T., Heinzmann, S., Düsterhus, S., Borchert, S., & Entian, K. D. (2005). Expression and functional analysis of the subtilin immunity genes spaIFEG in the subtilin-sensitive host Bacillus subtilis MO1099. Journal of Bacteriology, 187(3), 822–828. https://doi.org/10.1128/JB.187.3.822-828.2005
Suárez, J. M., Edwards, A. N., & McBride, S. M. (2013). The Clostridium difficile cpr locus is regulated by a noncontiguous two-component system in response to type A and B lantibiotics. Journal of Bacteriology, 195(11), 2621–2631. https://doi.org/10.1128/JB.00166-13
Sun, L., Finnegan, C. M., Kish-Catalone, T., Blumenthal, R., Garzino-Demo, P., La Terra Maggiore, G. M., Berrone, S., Kleinman, C., Wu, Z., Abdelwahab, S., Lu, W., & Garzino-Demo, A. (2005). Human beta-defensins suppress human immunodeficiency virus infection: potential role in mucosal protection. Journal of Virology, 79(22), 14318–14329.
Szukiewicz, D., Alkhalayla, H., Pyzlak, M., Watroba, M., Szewczyk, G., & Wejman, J. (2016). Human beta-defensin 1, 2 and 3 production by amniotic epithelial cells with respect to human papillomavirus (HPV) infection, HPV oncogenic potential and the mode of delivery. Microbial Pathogenesis, 97, 154–165. https://doi.org/10.1016/j.micpath.2016.06.010
Taggart, C. C., Greene, C. M., Smith, S. G., Levine, R. L., McCray, P. B. Jr, O’Neill, S., & McElvaney, N. G. (2003). Inactivation of human beta-defensins 2 and 3 by elastolytic cathepsins. Journal of immunology (Baltimore, Md.: 1950), 171(2), 931–937.
Tang, Y. Q., Yuan, J., Osapay, G., Osapay, K., Tran, D., Miller, C. J., Ouellette, A.J. & Selsted, M. E. (1999). A cyclic antimicrobial peptide produced in primate leukocytes by the ligation of two truncated alpha-defensins. Science, 286(5439), 498–502.
Tenge, V. R., Gounder, A. P., Wiens, M. E., Lu, W., & Smith, J. G. (2014). Delineation of interfaces on human alpha-defensins critical for human adenovirus and human papillomavirus inhibition. Plos Pathogens, 10, e1004360.
Thomassin, J. L., Brannon, J. R., Gibbs, B. F., Gruenheid, S., & Le Moual, H. (2012). OmpT outer membrane proteases of enterohemorrhagic and enteropathogenic Escherichia coli contribute differently to the degradation of human LL-37. Infection and Immunity, 80(2), 483–492. https://doi.org/10.1128/IAI.05674-11
Thwaite, J. E., Hibbs, S., Titball, R. W., & Atkins, T. P. (2006). Proteolytic degradation of human antimicrobial peptide LL-37 by Bacillus anthracis may contribute to virulence. Antimicrobial Agents and Chemotherapy, 50(7), 2316–2322. https://doi.org/10.1128/AAC.01488-05
Tollner, T. L., Bevins, C. L., & Cherr, G. N. (2012). Multifunctional glycoprotein DEFB126—A curious story of defensin-clad spermatozoa. Nature Reviews. Urology, 9, 365–375.
Tollner, T. L., Yudin, A. I., Tarantal, A. F., Treece, C. A., Overstreet, J. W., & Cherr, G. N. (2008). Beta-defensin 126 on the surface of macaque sperm mediates attachment of sperm to oviductal epithelia. Biology of Reproduction, 78, 400–412.
Tongaonkar, P., Golji, A. E., Tran, P., Ouellette, A. J., & Selsted, M. E. (2012). High fidelity processing and activation of the human alpha-defensin HNP1 precursor by neutrophil elastase and proteinase 3. PLoS One1, 7, e32469.
Tran, A. X., Whittimore, J. D., Wyrick, P. B., McGrath, S. C., Cotter, R. J., & Trent, M. S. (2006). The lipid A 1-phosphatase of Helicobacter pylori is required for resistance to the antimicrobial peptide polymyxin. Journal of bacteriology, 188(12), 4531–4541. https://doi.org/10.1128/JB.00146-06
Tzeng, Y. L., Ambrose, K. D., Zughaier, S., Zhou, X., Miller, Y. K., Shafer, W. M., & Stephens, D. S. (2005). Cationic antimicrobial peptide resistance in Neisseria meningitidis. Journal of Bacteriology, 187(15), 5387–5396. https://doi.org/10.1128/JB.187.15.5387-5396.2005
Valere, K., Lu, W., & Chang, T. L. (2017). Key determinants of human alpha-defensin 5 and 6 for enhancement of HIV infectivity. Viruses, 9, 244.
Valere, K., Rapista, A., Eugenin, E., Lu, W., & Chang, T. L. (2015). Human alpha-defensin HNP1 increases HIV traversal of the epithelial barrier: A potential role in STI-mediated enhancement of HIV transmission. Viral Immunology, 28, 609–615.
Van Cleemput, J., Poelaert, K. C. K., Laval, K., Vanderheijden, N., Dhaenens, M., Daled, S., Boyen, F., Pasmans, F., & Nauwynck, H. J. (2020). An Alphaherpesvirus Exploits Antimicrobial β-Defensins to Initiate Respiratory Tract Infection. Journal of Virology, 94(8), e1676–19. https://doi.org/10.1128/JVI.01676-19
van der Does, A. M., Hiemstra, P. S., & Mookherjee, N. (2019). Antimicrobial host defence peptides: Immunomodulatory functions and translational prospects. Advances in Experimental Medicine And Biology, 1117, 149–171.
Verheul, A., Russell, N. J., Hof, V. T., Rombouts, R., & Abee, T. (1997). Modifications of membrane phospholipid composition in nisin-resistant Listeria monocytogenes Scott A. Applied and Environmental Microbiology, 63(9), 3451–3457. https://doi.org/10.1128/aem.63.9.3451-3457.1997
Wang, A., Chen, F., Wang, Y., Shen, M., Xu, Y., Hu, J., Wang, S., Geng, F., Wang, C., Ran, X. & Wang, J. (2013). Enhancement of antiviral activity of human alpha-defensin 5 against herpes simplex virus 2 by arginine mutagenesis at adaptive evolution sites. Journal of Virology, 87(5), 2835–2845.
Wassing, G. M., Ilehag, N., Frey, J., & Jonsson, A. B. (2021). Modulation of human beta-defensin 2 expression by pathogenic neisseria meningitidis and commensal Lactobacilli. Antimicrobial Agents and Chemotherapy, 65(4), e02002–e02020. https://doi.org/10.1128/AAC.02002-20
Wehkamp, J., Harder, J., Wehkamp, K., Wehkamp-von Meissner, B., Schlee, M., Enders, C., Sonnenborn, U., Nuding, S., Bengmark, S., Fellermann, K., Schröder, J. M., & Stange, E. F. (2004). NF-kappaB- and AP-1-mediated induction of human beta defensin-2 in intestinal epithelial cells by Escherichia coli Nissle 1917: A novel effect of a probiotic bacterium. Infection and Immunity, 72(10), 5750–5758. https://doi.org/10.1128/IAI.72.10.5750-5758.2004
Wei, G., de Leeuw, E., Pazgier, M., Yuan, W., Zou, G., Wang, J., Ericksen, B., Lu, W.Y., Lehrer, R.I. & Lu, W. (2009). Through the looking glass, mechanistic insights from enantiomeric human defensins. Journal of Biological Chemistry, 284(42), 29180–29192.
Wiens, M. E., & Smith, J. G. (2015). Alpha-Defensin HD5 inhibits furin cleavage of human papillomavirus 16 L2 to block infection. Journal of Virology, 89, 2866–2874.
Wiens, M. E., & Smith, J. G. (2017). Alpha-Defensin HD5 inhibits human papillomavirus 16 infection via capsid stabilisation and redirection to the lysosome. MBio, 8, e2304–e2316.
Wilson, C. L., Ouellette, A. J., Satchell, D. P., Ayabe, T., Lopez-Boado, Y. S., Stratman, J. L., Hultgren, S.J., Matrisian, L.M. & Parks, W. C. (1999). Regulation of intestinal α-defensin activation by the metalloproteinase matrilysin in innate host defense. Science, 286(5437), 113–117.
Wilson, S. S., Bromme, B. A., Holly, M. K., Wiens, M. E., Gounder, A. P., Sul, Y., & Smith, J. G. (2017). Alpha-defensin-dependent enhancement of enteric viral infection. Plos Pathogens, 13(6), e1006446.
Wittersheim, M., Cordes, J., Meyer-Hoffert, U., Harder, J., Hedderich, J., & Gläser, R. (2013). Differential expression and in vivo secretion of the antimicrobial peptides psoriasin (S100A7), RNase 7, human beta-defensin-2 and – 3 in healthy human skin. Experimental Dermatology, 22, 364–366.
Wu, Z., Hoover, D. M., Yang, D., Boulegue, C., Santamaria, F., Oppenheim, J. J., Lubkowski, J. & Lu, W. (2003). Engineering disulfide bridges to dissect antimicrobial and chemotactic activities of human beta-defensin 3. Proceedings of the National Academy of Sciences, 100(15), 8880–8885.
Xu, C., Wang, A., Marin, M., Honnen, W., Ramasamy, S., Porter, E., Subbian, S., Pinter, A., Melikyan, G. B., Lu, W., & Chang, T. L. (2021). Human defensins inhibit SARS-CoV-2 infection by blocking viral entry. Viruses, 13(7), 1246.
Xu, D., Liao, C., Xiao, J., Fang, K., Zhang, W., Yuan, W., & Lu, W. (2019). Human enteric defensin 5 promotes Shigella infection of macrophages. Infection and Immunity, 88(1), 10–1128.
Xu, D., Liao, C., Zhang, B., Tolbert, W. D., He, W., Dai, Z., Zhang, W., Yuan, W., Pazgier, M., Liu, J. & Lu, W. (2018). Human enteric alpha-defensin 5 promotes Shigella infection by enhancing bacterial adhesion and invasion. Immunity, 48(6), 1233–1244.
Yamaguchi, Y., Nagase, T., Makita, R., Fukuhara, S., Tomita, T., Tominaga, T., Kurihara, H., & Ouchi, Y. (2002). Identification of multiple novel epididymis-specific β-defensin isoforms in humans and mice. The Journal of Immunology, 169(5), 2516–2523.
Yoshimura, T., McLean, M. H., Dzutsev, A. K., Yao, X., Chen, K., Huang, J., Gong, W., Zhou, J., Xiang, Y., H Badger, J. & Wang, J. M. (2018). The antimicrobial peptide CRAMP is essential for colon homeostasis by maintaining microbiota balance. The Journal of Immunology, 200(6), 2174–2185.
Yount, N. Y., & Yeaman, M. R. (2013). Peptide antimicrobials: Cell wall as a bacterial target. Annals of the New York Academy of Sciences, 1277, 127–138. https://doi.org/10.1111/nyas.12005
Yu, H., Dong, J., Gu, Y., Liu, H., Xin, A., Shi, H., Sun, F., Zhang, Y., Lin, D., & Diao, H. (2013). The novel human β-defensin 114 regulates lipopolysaccharide (LPS)-mediated inflammation and protects sperm from motility loss. Journal of Biological Chemistry, 288, 12270–12282.
Zugasti, O., & Ewbank, J. J. (2009). Neuroimmune regulation of antimicrobial peptide expression by a noncanonical TGF-beta signaling pathway in Caenorhabditis elegans epidermis. Nature Immunology, 10(3), 249–256. https://doi.org/10.1038/ni.1700
Acknowledgements
The authors acknowledge the Department of Microbiology, Faculty of Science, Annamalai University, for the support during the study.
Author information
Authors and Affiliations
Contributions
VK conceptualized and designed the work, collected the articles, and wrote the manuscript. YK revised the manuscript, prepared graphical illustration, and carried out data analysis and English editing. SS carried out data curation. GP supervised the work and carried out English and grammar editing.
Corresponding author
Ethics declarations
Ethical Approval
Nil.
Consent to Participate
Not applicable
Consent for Publication
No informed consent for publication
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumaresan, V., Kamaraj, Y., Subramaniyan, S. et al. Understanding the Dynamics of Human Defensin Antimicrobial Peptides: Pathogen Resistance and Commensal Induction. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-024-04893-8
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-024-04893-8