Abstract
A novel gene encoding aspartate dehydrogenase (ASPDH) has been discovered in Achromobacter denitrificans. The product of this gene has a strict dependence on NADH and demonstrated significant reductive activity towards not only oxaloacetate (OAA) but also 2-ketobutyric acid. Further enzymatic characterization revealed the kinetic parameters of ASPDH for OAA and 2-ketobutyric acid were as follows: Km values of 4.25 mM and 0.89 mM, Vmax values of 10.67 U mg−1 and 2.10 U mg−1, and Kcat values of 3.70 s−1 and 0.72 s−1, respectively. The enzyme also showed a dependency on metal ions, with EDTA and Cu2+ exerting strong inhibitory effects, while Ca2+ and Fe2+ exhibited pronounced enhancing effects. By utilizing a whole-cell biocatalyst system comprising glucose dehydrogenase (GDH) and ASPDH as a coupled system to replenish cofactors by oxidizing glucose, enabling the effective conversion of 2-ketobutyric acid to L-2-aminobutyric acid (L-2-ABA) with 97.2% yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
L-amino acid dehydrogenase catalyzes the reversible oxidative deamination of L-amino acids, utilizing either nicotinamide adenine dinucleotide (NAD) or nicotinamide adenine dinucleotide phosphate (NADP) as cofactors, producing the corresponding α-keto acids [1]. The enzymes are involved in the regulation of amino acid metabolism and glucose metabolism. In comparison to previously reported amino acid dehydrogenases (such as alanine dehydrogenase, leucine dehydrogenase, glutamate dehydrogenase, isoleucine dehydrogenase, and phenylalanine dehydrogenase) [1], the research on L-aspartate dehydrogenase (L-ASPDH) is relatively limited [1,2,3,4,5,6,7,8,9]. L-ASPDH has the capability to directly catalyze the reversible oxidative deaminating reaction of L-aspartate, generating oxaloacetate [2,3,4, 8], which can be applied in the production of aspartate.
Yang Z. et al. initially discovered an original L-aspartate dehydrogenase (L-ASPDH) from the thermophilic bacterium Thermotoga maritima [9]. Subsequently, Kazunari Yoneda et al. identified a gene (AF1838) with 38% similarity to the discovered L-ASPDH in the thermophilic archaeon Archaeoglobus fulgidus, and this L-ASPDH exhibited high tolerance to temperatures up to 80 ℃ [6]. Li Y. et al. identified the first bacterial NAD-dependent L-ASPDH from the mesophilic bacterium Pseudomonas aeruginosa PAO1 [4]. Later, T. M. Kuvaeva et al. made the first discovery of NADP-dependent L-ASPDH from the Rhodopseudomonas palustris and Bradyrhizobium japonicum [8]. These studies indicate that L-ASPDH can be widely present in various microorganisms.
In our research on Achromobacter denitrificans, a denitrifying bacterium, we identified a gene in its genome that potentially encodes L-ASPDH. This discovery marks the first identification of this enzyme from the Alcaligenaceae family within the Burkholderiales order. We performed heterologous expression of the gene and investigated its enzymatic properties. The results revealed that the enzyme exhibited activity in both reducing oxaloacetate (OAA) and 2-ketobutyric acid, and this is also the first natural occurrence of L-ASPDH capable of catalyzing the production of L-2-aminobutyrate (L-2-ABA) from 2-ketobutyric acid.
Materials and Methods
Plasmids, Strains, and Media
Yi Xin Biotechnology (Shanghai) Co. synthesized the aspartate dehydrogenase (ASPDH, accession numbers: WP_088148634) gene from Achromobacter Denitrificans. The ASPDH gene was successfully inserted into plasmid pET-28a at the Hind III and EcoR I restriction sites (primers shown in Table S1). E. coli DH5α served as an intermediate host for both amplifying and preserving recombinant plasmids. Subsequently, the ASPDH gene-carrying plasmid was introduced into E. coli BL21 (DE3) for transformation. LB medium was utilized for culturing E. coli bacteria.
Enzymes and Chemical Reagents
The following enzymes were used: QuickCut EcoR I, QuickCut Hind III (TaKaRa), ClonExpress II One Step Clone Kit (Vazyme), 2 × FastPfu SuperMix (Miozyme). All chemical reagents, except for the 3, 4-dihydroxyphenylpyruvic acid synthesized in our laboratory, were purchased from Sigma–Aldrich (Shanghai, China). These chemical reagents were domestically sourced and were of analytical purity.
Expression and Purification for ASPDH
Inoculate 3 μL of E. coli BL21 (DE3) cells carrying the aspartate dehydrogenase (ASPDH) gene into fresh LB liquid medium added to 50 μg ml−1 kanamycin and incubated the culture at 200 rpm and 37 °C for 12 h. An overnight culture of 1 mL was transferred to 50 mL of fresh LB liquid medium added to kanamycin at a concentration of 50 μg mL−1. The culture was cultured at 37 °C and 200 rpm until it attained an approximate OD600 of 0.6. Afterward, an ultimate concentration of 0.3 mM IPTG was supplemented with the medium to induce protein. The medium was further incubated at 20 °C, 200 rpm for an additional duration of 24 h; centrifuged the expressed medium at high speed (8000 rpm) and 4 °C for 10 min; collected the bacteria from the sediment and resuspended them in 0.1 M Tris–HCl buffer (pH 7.0); then, sonicated the resuspended cells in a cold environment (0 °C); and centrifuged the disrupted solution at 8000 rpm and 4 °C for 10 min: the crude enzyme extract is obtained by harvesting the liquid supernatant.
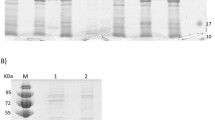
The crude enzyme extract underwent filtration through a membrane and subsequent purification via a 1 mL His TrapHP affinity chromatography column. The column was balanced with a binding buffer (500 mM NaCl, 20 mM imidazole, 20 mM phosphate buffer, pH 7.4) at twice the column volume during purification. The sample was loaded onto the column and then subjected to linear elution using an elution buffer (500 mM NaCl, 500 mM imidazole, 20 mM phosphate buffer, pH 7.4) at five times the column volume. The enzyme sample corresponding to the target elution peak was collected. The collected enzyme sample was additionally desalted using a 5 mL HisTrap desalting column to reduce excessive imidazole and sodium chloride concentrations. Then crude ASPDH extract as well as ASPDH with purification were examined on 12% SDS-PAGE protein electrophoresis [10,11,12]. We used the original strain of E. coli BL21 (DE3) as a blank control. The concentration of protein was assessed by the Bradford method [13].
Assays of Enzyme Activity
We utilized a spectrophotometer to measure the changes in NADH absorption at 340 nm, which is its maximum absorbance wavelength, to assess the enzyme activity [14]. The reaction mixture (3 mL) comprised 10 mM substrate, 0.13 mM NADH, 0.1 M Tris–HCl buffer (pH 7.0), and a right quantity of purified aspartate dehydrogenase (ASPDH). Firstly, the substrate and 0.1 M Tris–HCl buffer (pH 7.0) were mixed. Subsequently, the reaction solution was incubated at 37 °C for 5 min. Next, NADH was added, followed by the addition of the right quantity of purified ASPDH. The decline in NADH absorbance was assessed at 340 nm within a 60-s interval. We designated one ASPDH unit of activity as the quantity of ASPDH capable of utilizing 1 μmol of NADH per minute under optimum environments.
Analysis of ASPDH Substrate Specificity
Assessing the reduction substrate specificity of aspartate dehydrogenase (ASPDH) was a part of our investigation. The relative activities were defined based on the maximal activity [15]. The substrates used in the assay include oxaloacetate, glyoxylic acid, α-ketoglutarate, pyruvate, 2-ketobutyric acid, phenylpyruvate, 4-methyl-2-oxopentanoic acid, sodium 3-methyl-2-oxobutanoate, 4-hydroxyphenylpyruvate, benzoylformic acid, and 3,4-dihydroxyphenylpyruvate. The reaction system (3 mL) consists of 10 mM substrate, 0.13 mM NADH, 0.1 M Tris–HCl buffer (pH 7.0), and a right quantity of purified ASPDH. Please refer to the “Assays of Enzyme Activity” section for detailed experimental procedures. Triplicate assays were conducted for all experiments.
Impact of Temperature on ASPDH Activity
A UV spectrophotometer was employed to identify the optimal temperature of aspartate dehydrogenase (ASPDH) within a range of 20 to 70 °C under pH 7.0 conditions. The necessary quantity of purified ASPDH, along with 10 mM substrate, 0.13 mM NADH, and 0.1 M Tris–HCl buffer (pH 7.0), constituted the reaction mixture (3 mL). To determine the enzyme’s optimal temperature, we measured the decline in absorbance of NADH at 340 nm. Triplicate assays were conducted for all experiments.
Impact of pH on ASPDH Activity
We determined the optimal pH of aspartate dehydrogenase (ASPDH) using a UV spectrophotometer at the enzyme’s optimal temperature within the pH range of 3 to 10. The necessary quantity of purified ASPDH, along with 10 mM substrate, 0.13 mM NADH, and 0.1 M buffer (pH 7.0), constituted the reaction mixture (3 mL). Buffer compositions corresponding to different pH ranges were utilized: citric acid-sodium citrate buffer for pH 3–6, sodium dihydrogen phosphate-disodium hydrogen phosphate buffer for pH 6–8, and Tris–HCl buffer for pH 8–10 [2, 4, 16]. After incubating the enzyme with the corresponding buffer at the optimal temperature for a specified duration, we added the substrate and NADH. The decrease in NADH absorbance at 340 nm was then measured during a defined time period. The optimal pH of the enzyme was determined by identifying the pH value at which the highest enzyme activity was exhibited. Triplicate assays were conducted for all experiments.
Impact of ASPDH Thermal and pH Stability
The thermal stability and pH stability of aspartate dehydrogenase (ASPDH) were assessed by subjecting the enzyme to incubation at temperatures ranging from 20 to 70 °C and at pH values ranging from 3 to 9 for a duration of 1 h. The untreated group (100%) served as the control in the experiment. The remaining enzyme activity was assessed using the aforementioned method to assess the enzyme’s stability under varied temperature and pH conditions. Triplicate assays were conducted for all experiments.
Impact of Metal Ions on ASPDH
The impact of metal ions on the aspartate dehydrogenase (ASPDH) activity was investigated by performing measurements in the previously mentioned reaction mixture. Metal ions (Zn2+, Mg2+, Ca2+, Cu2+, Ni+, Fe2+, Co2+, Mn2+) and EDTA were supplemented at final concentrations of 0.1 mM and 1 mM. Optimal reaction conditions were employed for conducting the determination. Triplicate assays were conducted for all experiments.
Kinetic Study of ASPDH
The determination of the kinetic parameters was performed using a standard assay, wherein the concentration of one substrate was adjusted while maintaining the concentrations of the other substrates constant [17]. Nonlinear fitting of the kinetic data was performed using GraphPad Prism 8.0 software [18, 19]. Triplicate assays were conducted for all experiments.
Construction of Coexpression Plasmid
The construction of the plasmid pET-28a-aspdh-T7-gdh [20, 21] involved several steps. First, the glucose dehydrogenase gene from Bacillus subtilis, along with a RBS and T7 promoter, was synthesized by Synbio Technologies (Suzhou, China). Subsequently, this gene was amplified using primers (Table S2) that included EcoR I and Hind III restriction sites at the forward and reverse ends, respectively. After the amplification of the gene, it was ligated to the pET-28a-aspdh vector that had already undergone digestion with the same restriction enzymes. Following this step, the resulting recombinant plasmids (pET-28a-aspdh-T7-gdh) were transformed into E. coli BL21 (DE3) to facilitate protein expression and then harvested for biocatalyst.
Whole-Cell Biocatalyst and Transformation of L-2-ABA
The procedure for inducing and expressing the coexpression strain followed the previously mentioned protocol. In succession, following a 24-h induction at 15 °C, cells were gathered from a 50-ml fermentation culture, rinsed with a 0.1 M Tris–HCl buffer (pH 7.0), and then centrifuged at 8000 rpm and 4 °C for 10 min. The whole-cell was resuspended in 20 ml Tris–HCl buffer and stored at 4 °C for further studies.
To facilitate further analysis, a bioconversion reaction was prepared by combining 50 ml of 0.1 M Tris–HCl buffer solution with 10 g L−1 wet cells, along with 70 mM 2-ketobutyric acid, 70 mM glucose, and 0.3 g L−1 NAD+. The reaction solution was cultured at 30 °C, 150 rpm. Quantitative determination of the L-2-ABA concentration was performed at 2-h intervals by employing HPLC [22,23,24].
L-2-ABA were Determined by HPLC
The supernatant required for quantifying L-2-ABA levels was obtained by centrifuging an 800 µL culture for 20 min at a speed of 13,800 g. Subsequently, the supernatant was appropriately diluted to the desired concentration to facilitate L-2-ABA quantification via the utilization of the 1260 High-Performance Liquid Chromatography (HPLC) system from Agilent Technology (USA). For separation and quantification of L-2-ABA, the OPA precolumn derivatization method [25] was applied using a Thermo ODS-2HYPERSIL C18 column (250 mm × 4.0 mm, USA). Solvent A, composed of 3.01 g sodium acetate, 200 µL triethylamine, and 5 mL tetrahydrofuran per liter, was used. Solvent B was prepared by dissolving 3.01 g sodium acetate in a mixture of 400 mL acetonitrile, 400 mL methanol, and 200 mL deionized water. The flow rate of both solvents was set at 1.0 mL min−1. The gradient started at 8% of solvent B and reached 100% within a duration of 35.5 min (Table 1). A sample of 1 µL was subjected to derivatization, injection, and subsequent monitoring at 338 nm [26].
Sequence Analysis
Nine protein sequences of the reported ASPDH family were selected from the NCBI database. The secondary structure alignment of ASPDH was carried out using ESPript 3.0. The phylogenetic tree was constructed using MEGA7.0 software [16, 18, 19]. We utilized the Neighbor-Joining algorithm and conducted the bootstrap analysis with 1000 iterations. Subsequently, we selected the consensus tree with the utmost reliability, determined by its bootstrap values, for further analysis.
Results
Expression and Purification of ASPDH
The recombinant plasmid with the ASPDH gene was favorably integrated into E. coli BL21 (DE3) bacteria. The crude ASPDH solution was purified using the AKTA (GE Healthcare) system to obtain the pure ASPDH solutions. Analysis of the SDS-PAGE gel revealed the expected bands of ASPDH at around 31 kDa (Fig. 1), consistent with the theoretical value.
ASPDH Optimal Temperature
ASPDH exhibits optimal activity at a temperature of 30 °C (Fig. 2). Its reducing activity noteworthily ascending from 20 °C towards its optimum temperature. Nevertheless, surpassing the optimum temperature resulted in a gradual decline in enzyme activity, ultimately leading to full inactivation at 60 °C.
ASPDH Optimal pH
The optimal pH for ASPDH is 7.0. ASPDH displayed minimal activity that was nearly undetectable at a pH level of 3.0. At pH 8.0, ASPDH retained 65.42% of its reducing activity. However, at pH 8.5, ASPDH became completely inactive (Fig. 3).
Thermal and pH Stability of ASPDH
ASPDH displayed excellent thermal stability at low temperatures (Fig. 4A). The stability of the enzyme gradually decreased with increasing temperature. When ASPDH is maintained above 25 °C, its enzymatic activity decreased gradually, with a noteworthy decline observed when stored above 45 °C.
In terms of pH stability, ASPDH demonstrated optimal stability at pH 7.0 (Fig. 4B). It maintained over 65% of its enzymatic activity in the pH range of 4.5 to 6.5 and retained 80% activity in the pH range of 7.5 to 8.5. At pH 3.5, it retained 40% activity, indicating that ASPDH had good stability in environments with varying pH, and it was suitable for storage in neutral to weak alkaline conditions.
Impact of Metal Ions
This study investigated the impact of metal ions on ASPDH activity. The results revealed a noteworthily inhibitory effect of EDTA and Cu2+ on the enzyme; this may be because Cu2+ can bind to the cysteine residue sulfhydryl group in the center of enzyme activity, inhibiting the activity of the enzyme [27], while all other tested ions (excluding Co2+) demonstrated diverse degrees of enhancement. These findings indicate that ASPDH is dependent on metal ions (Table 2).
Substrate Specificity
To determine the substrate specificity of ASPDH, the substrate spectra were measured using different α-keto acids. NADH was used as the cofactor, and no activity with NADPH was observed. Except for the optimal substrate, oxaloacetate (OAA), ASPDH exhibited significant reducing activity towards other tested α-keto acids such as pyruvate and 2-ketobutyric acid, with reduction activity of 13.9% and 9.6%, respectively. However, it showed only low reducing activity towards sodium 3-methyl-2-oxobutanoate, benzoylformic acid, 4-methyl-2-oxopentanoic acid, α-ketoglutarate, and 4-hydroxyphenylpyruvate. No reducing activity was detected for phenylpyruvate and 3,4-dihydroxyphenylpyruvate (Table 3).
Kinetic Parameters
Using OAA and 2-ketobutyric acid as substrates, along with NADH, the kinetic parameters of ASPDH were calculated under the specified optimal conditions of reaction temperature and pH. The Km value of ASPDH for OAA was 4.25 mM, Vmax was 10.67 U mg−1 and kcat was 3.70 s−1. For 2-ketobutyric acid, ASPDH exhibited a Km value of 0.89 mM, a Vmax of 2.10 U mg−1, and a kcat of 0.72 s−1.
Utilizing the Coupling System Enables the Synthesis of L-2-ABA
Coexpression of ASPDH and GDH in E. coli BL21 (DE3) resulted in the formation of an efficient coupling system. Following 12 h of whole-cell biocatalysis, a nearly complete reduction of 2-ketobutyric acid to L-2-ABA was achieved, resulting in a high conversion rate of 97.2% (Fig. 5).
Sequence Analysis
Based on the phylogenetic tree (Fig. 6), ASPDH from A. Denitrificans shows a high homology with other ASPDHs. The alignment of the amino acid sequence (Fig. 7) revealed that A. denitrificans ASPDH shares similarities with other ASPDHs.
Phylogenetic analysis of aspartate dehydrogenase (ASPDH) from A. Denitrificans with other nine ASPDHs that were reported: Achnromobacter denitrificaris (WP 088148634.1), Achromobacter spanius (WP 050448281.1), Achromobacter sp. MY14(WP 230693706.1), Delftia sp. Cs1-4(WP 013804514.1), Pseudomonas aeruginosa PAO1 (NP 252195.1), Klebsiella pneumoniae (ODM39893.1), Rhodopseudomonas palustris (RJF77029.1), Bradyrhizobium japonicum (KGTS81542.1), Thermotoga maritima (AKE31272.1), Archaeoglobus fulgidus (WP 010879332.1)
Discussion
Aspartic acid, a natural amino acid, participates in energy production by undergoing transformation into various intermediate metabolites through the tricarboxylic acid cycle [1], Aspartic acid also serves as a central hub for the synthesis of other amino acids [2]. It serves as a neurotransmitter and plays a crucial role in the central nervous system of humans and animals, participating in learning and memory processes. Aspartic acid also contributes to maintaining nitrogen balance within cells and participates in transamination reactions along with other amino acids. 2-ketobutyric acid, a metabolic regulator, is involved in the metabolism of carbohydrates, fats, and amino acids. Moreover, during energy deficiency, it serves as an alternative energy source for other tissues through ketone body metabolism. Additionally, 2-ketobutyric acid is a compound with a strong aroma and flavor, making it widely used in the food and beverage industry as a flavoring or flavor additive. Moreover, 2-ketobutyric acid holds potential value in pharmaceutical research due to its ability to synthesize certain antiepileptic drugs through amide formation [28,29,30].
In this study, we comprehensively analyzed the enzymatic properties and substrate specificity of ASPDH from Achromobacter Denitrificans. Regarding temperature, ASPDH exhibited optimal activity at 30 °C, with a significant decrease in enzyme activity above 55 °C and complete inactivation at 60 °C. This indicates that ASPDH is sensitive to temperature, and both excessively high and low temperatures can affect its enzymatic activity. In terms of pH stability, ASPDH showed the highest stability at pH 7.0. It maintained over 65% enzyme activity between pH 4.5 and 6.5 and 80% activity between pH 7.5 and 8.5. These findings suggest that ASPDH maintains good stability in pH-changing environments and is suitable for storage in neutral to weak alkaline conditions. This is similar to the enzymes KpnAspDH (ASPDH from Klebsiella pneumoniae) and DelAspDH (ASPDH from Delftia sp. Cs1–4) reported by Li et al. [2].
Kinetic parameters showed that the ASPDH showed strict dependence on NADH, unlike some NADP-dependent ASPDH enzymes reported, such as ASPDH from Thermotoga maritima [9], Archaeoglobus fulgidus [6], and Rhodopseudomonas palustris and Bradyrhizobium japonicum [8]. Additionally, the impact of metal ions on ASPDH activity was researched. The results demonstrated that EDTA and Cu2+ had inhibitory effects on the enzyme, while Zn2+, Mg2+, Ca2+, Ni+, Mn2+, and Fe2+ showed varying degrees of enhancement. These findings indicated that ASPDH was dependent on metal ions.
The substrate specificity of ASPDH was analyzed by testing different α-keto acids. The results showed that ASPDH not only utilized oxaloacetate as a substrate but also exhibited significant reduction activity with pyruvate and 2-ketobutyric acid, reaching 13.9% and 9.6% of the activity compared to oxaloacetate, respectively. Since there are no known enzymes in nature that can directly reduce 2-ketobutyric acid, typically, modification of leucine dehydrogenase is required for the reduction of 2-ketobutyric acid to L-2-aminobutyric acid (L-2-ABA) [23, 24, 31, 32]. However, this study reveals that Achromobacter denitrificans ASPDH possesses an inherent ability to catalyze the synthesis of L-2-ABA through the conversion of 2-ketobutyric acid. The utilization of a whole-cell biocatalyst system incorporating GDH and ASPDH [20,21,22,23] allows for the full utilization of the cofactor NAD+ within the cells, resulting in a more robust catalytic environment conducive to the industrial-scale production of L-2-ABA. These findings indicate the potential for pursuing rational molecular engineering of ASPDH as it lays a crucial groundwork and provides valuable insights for future research and application of ASPDH.
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Li, Y., Ogola, H. J. O., & Sawa, Y. (2012). l-Aspartate dehydrogenase: Features and applications. Applied microbiology and biotechnology, 93, 503–516.
Li, H., Zhu, T., Miao, L., Zhang, D., Li, Y., Li, Q., & Li, Y. (2017). Discovery of novel highly active and stable aspartate dehydrogenases. Scientific Reports, 7(1), 7881.
Li, Y., Ishida, M., Ashida, H., Ishikawa, T., Shibata, H., & Sawa, Y. (2011). A non-NadB type L-aspartate dehydrogenase from Ralstonia eutropha strain JMP134: Molecular characterization and physiological functions. Bioscience, biotechnology, and biochemistry, 75(8), 1524–1532.
Li, Y., Kawakami, N., Ogola, H. J. O., Ashida, H., Ishikawa, T., Shibata, H., & Sawa, Y. (2011). A novel L-aspartate dehydrogenase from the mesophilic bacterium Pseudomonas aeruginosa PAO1: Molecular characterization and application for L-aspartate production. Applied Microbiology and Biotechnology, 90, 1953–1962.
Okamura, T., Noda, H., Fukuda, S., & Ohsugi, M. (1998). Aspartate dehydrogenase in vitamin B12-producing Klebsiella pneumoniae IFO 13541. Journal of Nutritional Science and Vitaminology, 44(4), 483–490.
Yoneda, K., Kawakami, R., Tagashira, Y., Sakuraba, H., Goda, S., & Ohshima, T. (2006). The first archaeal L-aspartate dehydrogenase from the hyperthermophile Archaeoglobus fulgidus: Gene cloning and enzymological characterization. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics, 1764, 1087–1093.
Yoneda, K., Sakuraba, H., Tsuge, H., Katunuma, N., & Ohshima, T. (2007). Crystal structure of archaeal highly thermostable l-aspartate dehydrogenase/NAD/citrate ternary complex. The FEBS Journal, 274(16), 4315–4325.
Kuvaeva, T., Katashkina, J., Kivero, A., & Smirnov, S. (2013). Novel NADPH-dependent L-aspartate dehydrogenases from the mesophilic nitrogen-fixing bacteria Rhodopseudomonas palustris and Bradyrhizobium japonicum. Applied Biochemistry and Microbiology, 49, 136–143.
Yang, Z., Savchenko, A., Yakunin, A., Zhang, R., Edwards, A., Arrowsmith, C., & Tong, L. (2003). Aspartate dehydrogenase, a novel enzyme identified from structural and functional studies of TM1643. Journal of Biological Chemistry, 278(10), 8804–8808.
Zheng, Z., Ma, C., Gao, C., Li, F., Qin, J., Zhang, H., Wang, K., & Xu, P. (2011). Efficient conversion of phenylpyruvic acid to phenyllactic acid by using whole cells of Bacillus coagulans SDM. PLoS ONE, 6(4), e19030.
Hari Prasad, O., Nanda Kumar, Y., Reddy, O., Chaudhary, A., & Sarma, P. (2012). Cloning, expression, purification and characterization of UMP kinase from Staphylococcus aureus. The protein journal, 31, 345–352.
Prasad, U. V., Vasu, D., Kumar, Y. N., Kumar, P. S., Yeswanth, S., Swarupa, V., Phaneendra, B., Chaudhary, A., & Sarma, P. (2013). Cloning, expression and characterization of NADP-dependent isocitrate dehydrogenase from Staphylococcus aureus. Applied Biochemistry and Biotechnology, 169, 862–869.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72(1–2), 248–254.
Reid, M. F., & Fewson, C. A. (1994). Molecular characterization of microbial alcohol dehydrogenases. Critical Reviews in Microbiology, 20(1), 13–56.
Min, K., Yeon, Y. J., Um, Y., & Kim, Y. H. (2016). Novel NAD-independent d-lactate dehydrogenases from Acetobacter aceti and Acidocella species MX-AZ02 as potential candidates for in vitro biocatalytic pyruvate production. Biochemical Engineering Journal, 105, 358–363.
Wang, L., Fan, T.-P., Wang, M., Bai, Y., Zheng, X., & Cai, Y. (2023). Human lactate dehydrogenase and malate dehydrogenase possess the catalytic properties to produce aromatic α-hydroxy acid. Systems Microbiology and Biomanufacturing, 3, 627–633.
Ohshima, T., Misono, H., & Soda, K. (1978). Properties of crystalline leucine dehydrogenase from Bacillus sphaericus. Journal of Biological Chemistry, 253(16), 5719–5725.
Qu, W., Bai, Y., Fan, T.-P., Zheng, X., & Cai, Y. (2022). Characterisation of aldo-keto reductases from Lactobacillus reuteri DSM20016. Process Biochemistry, 122, 172–180.
Zhu, J., Bai, Y., Fan, T.-P., Zheng, X., & Cai, Y. (2023). Characterization of acid-resistant aldo–keto reductases capable of asymmetric synthesis of (R)-CHBE from Lactobacillus plantarum DSM20174. Systems Microbiology and Biomanufacturing, 3, 634–646.
Lu, H., Bai, Y., Fan, T.-P., Zhao, Y., Zheng, X., & Cai, Y. (2018). Identification of a L-Lactate dehydrogenase with 3, 4-dihydroxyphenylpyruvic reduction activity for l-Danshensu production. Process Biochemistry, 72, 119–123.
Wang, Y., Bai, Y., Fan, T. P., Zheng, X., & Cai, Y. (2018). Reducing 3, 4-dihydroxyphenylpyruvic acid to d-3, 4-dihydroxyphenyllactic acid via a coenzyme nonspecific d-lactate dehydrogenase from Lactobacillus reuteri. Journal of Applied Microbiology, 125(6), 1739–1748.
Zhang, L., Xiao, Y., Yang, W., Hua, C., Wang, Y., Li, J., & Yang, T. (2020). Synthesis of L-2-aminobutyric acid by leucine dehydrogenase coupling with an NADH regeneration system. Sheng wu Gong Cheng xue bao= Chinese Journal of Biotechnology, 36(5), 992–1001.
Chen, J., Zhu, R., Zhou, J., Yang, T., Zhang, X., Xu, M., & Rao, Z. (2021). Efficient single whole-cell biotransformation for L-2-aminobutyric acid production through engineering of leucine dehydrogenase combined with expression regulation. Bioresource Technology, 326, 124665.
Fu, Y., Zhang, J., Fu, X., Xie, Y., Ren, H., Liu, J., Chen, X., & Liu, L. (2020). Production of L-2-aminobutyric acid from L-threonine using a trienzyme cascade. Sheng wu Gong Cheng xue bao= Chinese Journal of Biotechnology, 36(4), 782–791.
Yang, J., Fang, Y., Wang, J., Wang, C., Zhao, L., & Wang, X. (2019). Deletion of regulator-encoding genes fadR, fabR and iclR to increase L-threonine production in Escherichia coli. Applied Microbiology and Biotechnology, 103, 4549–4564.
Wang, S., Fang, Y., Wang, Z., Zhang, S., Wang, L., Guo, Y., & Wang, X. (2021). Improving L-threonine production in Escherichia coli by elimination of transporters ProP and ProVWX. Microbial Cell Factories, 20(1), 1–13.
Kim, J.H., Cho, H.J., Ryu, S.E., & choi, M. (2000), Effects of metal ions on the activity of protein tyrosine phosphatase VHR: Highly potent and reversible oxidative inactivation by Cu2+ ion. Archives of Biochemistry and Biophysics 382 1 (2000): 72–80.
Wang, Y., Li, G.-S., Qiao, P., Lin, L., Xue, H.-L., Zhu, L., Wu, M.-B., Lin, J.-P., & Yang, L.-R. (2018). Increased productivity of L-2-aminobutyric acid and total turnover number of NAD+/NADH in a one-pot system through enhanced thermostability of L-threonine deaminase. Biotechnology Letters, 40, 1551–1559.
Sasa, M. (2006). A new frontier in epilepsy: Novel antiepileptogenic drugs. Journal of Pharmacological Sciences, 100(5), 487–494.
Liao, J.C., Zhang, K., Cho, K.M. (2011). Compositions and methods for the production of l-homoalanine. WO2011106696A2.
J. Chen, M. Xu, T. Yang, X. Zhang, M. Shao, H. Li, Z. Rao, Rational design of the C-terminal Loop region of leucine dehydrogenase and cascade biosynthesis L-2-aminobutyric acid, Sheng wu Gong Cheng xue bao= Chinese Journal of Biotechnology 37(12) (2021) 4254–4265.
Zhou, J., Wang, Y., Chen, J., Xu, M., Yang, T., Zheng, J., Zhang, X., & Rao, Z. (2019). Rational engineering of Bacillus cereus leucine dehydrogenase towards α-keto acid reduction for improving unnatural amino acid production. Biotechnology Journal, 14(3), 1800253.
Author information
Authors and Affiliations
Contributions
Authors YC and ZW designed the experiments. Author ZW performed cell culture studies, experiments, and enzymatic assays. Authors ZW, WL, and YY performed data processing. Authors YC and ZW contributed to manuscript preparation. All authors participated in the discussion.
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent for Publication
This article’s publication has been approved by all co-authors.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Zifeng Wang is the first author of this study.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Z., Liu, W., Yan, Y. et al. Characterization and Application of an Aspartate Dehydrogenase from Achromobacter denitrificans. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-024-04867-w
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-024-04867-w