Abstract
PICALM (phosphatidylinositol-binding clathrin assembly protein) mutations have been linked to a number of human disorders, including leukemia, Alzheimer’s disease, and Parkinson’s disease. Nevertheless, the effect of PICALM on cancer, particularly on prognosis and immune infiltration in individuals with BRCA, is unknown. We obtained the data of breast cancer patients from The Cancer Genome Atlas (TCGA) database, and analyzed the expression of PICALM in breast cancer, its impact on survival’ and its role in tumor immune invasion. Finally, in vitro cellular experiments were performed to validate the results. Research has found that PICALM expression was shown to be downregulated in BRCA and to be substantially linked with clinical stage, histological type, PAM50, and age. PICALM downregulation was linked to a lower overall survival (OS) and disease-specific survival (DSS) in BRCA patients. A multivariate Cox analysis revealed that PICALM is an independent predictor of OS. The enriched pathways revealed by functional enrichment analysis included oxidative phosphorylation, angiogenesis, the TGF signaling pathway, and the IL-6/JAK/STAT3 signaling system. Furthermore, the amount of immune cell infiltration by B cells, eosinophils, mast cells, neutrophils, and T cells was positively linked with PICALM expression. Finally, we experimentally verified that low expression of PICALM can reduce proliferation, migration, and invasion in tumor cells. This evidence shows that PICALM expression impacts prognosis, immune infiltration, and pathway expression in breast cancer patients, and it might be a potential predictive biomarker for the disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BRCA) is one of the most common cancers that threaten women’s health worldwide. The incidence and mortality of breast cancer are increasing rapidly each year, with an estimated 268,600 new cases and an estimated 41,760 deaths occurring globally in 2019 [1, 2]. BRCA native subtypes are luminal A, luminal B, HER2-enriched, and basal-like [3,4,5]. At present, local surgery, conventional chemotherapy, precise radiotherapy, hormone therapy, and monoclonal antibody can be used as clinical treatment for BRCA patients. Despite major advances in BRCA therapy, a substantial proportion of individuals still have BRCA and are resistant or relapse after treatment [6]. Due to tumor heterogeneity and complexity, existing biomarkers are poor at predicting BRCA prognosis. Therefore, new molecular biomarkers have emerged in clinical studies as prognostic indicators that can improve prognosis and guide individualized treatment regimens [7, 8].
PICALM (phosphatidylinositol-binding lectin assembly protein) was identified as a translocation partner of AF10 (MLLT10) [9] in the U937 leukemia cell line. PICALM protein is widely expressed and involved in lectin-mediated endocytosis and autophagy and is an important mechanism for intracellular transport of proteins, lipids, growth factors, and neurotransmitters [10,11,12]. Previous studies have shown that overexpression of PICALM can inhibit endocytosis of the transferrin receptor and epidermal growth factor receptor [13, 14]. In addition, PICALM was found to be a chromosomal translocation in cell lines obtained from lymphoma patients [9]. PICALM mutations are associated with a variety of human diseases, including leukemia, Alzheimer’s disease, and Parkinson’s disease [15,16,17,18]. However, whether the reduction of PICALM accelerates cancer growth has not been investigated, especially in BRCA. To our knowledge, no other study has examined its effect on BRCA gene expression.
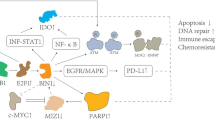
As part of the current work, we used bioinformatics to investigate and understand the relationship between PICALM expression and BRCA, including its clinical, pathological, and prognostic significance, as well as the underlying molecular pathways and immune cell infiltration, as illustrated in Fig. 1. The effect of PICALM on BRCA cells was further investigated in vitro. Our findings highlight a possible oncogenic role of PICALM in BRCA, which may help physicians refine treatment approaches and improve the prognosis of BRCA patients.
PICALM expression levels in various kinds of malignancies and breast cancer. A PICALM expression in various kinds of cancers compared to normal tissues in the TCGA and GTEx databases. B BRCA in TCGA and GTEx databases and in non-matched normal tissues. C ROC curves from the TCGA database for diagnosing breast cancer vs normal breast tissues. D Representative images of PICALM expression in breast cancer tissues and their normal controls. TCGA, The Cancer Genome Atlas; GTEx, Genotype-Tissue Expression Project; ROC, receiver operating characteristic.*P< 0.05, **P< 0.01, ***P< 0.001
Materials and Methods
Expression Analysis of PICALM
Data were from the Cancer Genome Atlas (Cancer Genome Atlas, TCGA, http://portal.gdc.cancer.gov/) and Genotype-Tissue Expression (https://www.commonfund.nih.gov/GTEx) [19]. Transcripts per million reads (TPM) were normalized based on HTSeq FPKM format data. In order to assess the level of protein immunohistochemical platform, HPA (http://www.proteinatlas.org), the database can be in [20] to get online. This public database was used to confirm PICALM protein levels in BRCA.
Differentially Expressed Gene Analysis
The median PICALM expression in TCGA breast cancer patients was used to separate patients into low and high PICALM expression groups. Using the “limma” package in R software, differential expression analysis was performed before the two groups, and relevant differential genes (DEGs) were screened using the criteria of “|log FC|>1 and P<0.05.” PICALM expression was correlated with the expression of the top ten DEGs through Spearman correlation analysis.
Functional Enrichment Analysis
The R package “clusterProfiler” was used to perform gene set enrichment analysis (GSEA) of the linked DEGs [21,22,23]. Adjusted P<0.05 and false discovery rate (FDR) <0.25 were used as screening criteria to search for significantly enriched functions and pathways.
Immuno-infiltration Analysis
A single sample containing 24 immune cells was subjected to GSEA analysis using the “GSEA” program in R software [24] to determine the relative enrichment score of immune cells in BRCA. Spearman correlation analysis and Wilcoxon rank sum test were used to analyze the difference between high and low PICALM expression in immune infiltration.
Survival Analysis
We employed the Kaplan-Meier plotter (http://kmplot.com/analysis/) [25] to examine the relationship between PICALM expression in BRCA and OS and disease-specific survival (DSS). PICALM’s median expression level was determined as the critical value, and the P-value (P<0.05) was calculated using the log-rank test. A multivariate and univariate analysis of Cox regression was performed to evaluate the effect of clinical factors on the prognosis of BRCA patients. The R software tool “ggplot2” was used to show forest plots.
Quantitative Real-time Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated and reverse transcribed into cDNA for logarithmic growth phase cells of MCF-7 according to RNA reagent instructions, and PICALM expression in each set of cells was measured by qRT-PCR. The primers are as follows: PICALM forward, 5′-GCAGCTGCCTGTTCCTCTTA-3′; PICALM reverse, 5′-TGGCCTTGCATACTGTCTTG-3′; GAPDH forward, 5′-GCACCGTCAAGGCTGAGAAC-3′; GAPDH reverse, 5′-TGGTGAAGACGCCAGTGGA-3′. The reaction conditions were as follows: pre-denaturation at 95 °C for 30 s, denaturation at 95 °C for 5 s, extension at 60 °C for 30 s, and a total of 40 cycles. Each series of studies was performed three times, and the relative expression of PICALM mRNA was determined using the 2−ΔΔCttechnique.
CCK8 Assay and Cell Invasion
Human breast cancer MCF-7 cells were seeded in 96-well plates with a 100:l inoculum per well (cell density of 3103 cells/mL), with six duplicate wells set up for each group. CCK-8 detection reagent (10 μL/well) was added at 24, 48, 72, and 96 h after transfection, the absorbance at 450 nm was measured, and the growth curve was plotted using an enzyme marker after 2 h of incubation.
Cell Scoring and Transwell Invasion Assay
One copy of each cell group of human breast cancer (MCF-7) cells was collected, inoculated, and cultivated in 12-well cell culture plates, a straight line was scratched in the middle of the well, the scratched cells were rinsed out, and the number of migrating cells was counted after 24 h under microscope observation and image. In a 37 °C incubator, add 500 μL of serum-free media to each of the top and bottom chambers with stromal gel chambers and incubate for 2 h. In the upper chamber, 2×105 cells (approximately 300 μL) were injected, and 500 μL of complete media was introduced to the bottom chamber to continue the incubation. Remove the tiny chamber, discard the upper chamber’s medium, clean the upper chamber’s filter membrane with a cotton swab, fix the membrane with 4% formaldehyde for 15 min, stain with crystalline violet for 25 min, wash with PBS, and count and photograph under the microscope (five fields of view were taken randomly for counting, and the mean value represents the number of invasive cells).
Statistical Analysis
This study used R (version 4.0.1) for all statistical analyses. PICALM expression in unpaired and paired tissues was analyzed using the Wilcoxon rank sum test and the paired samples t-test, respectively. To examine the connection between clinical characteristics and PICALM expression, the Wilcoxon rank sum test and logistic regression were performed. P-values less than 0.05 were deemed statistically significant.
Results
Low Expression of PICALM in BRCA
We undertook a pan-cancer investigation to better understand PICALM expression in malignancies. PICALM expression was shown to be decreased in the majority of malignancies, including ACC, according to the findings (adrenocortical carcinoma), BLCA (uroepithelial carcinoma of the bladder), BRCA (breast cancer), COAD (colon cancer), DLBC (diffuse large B-cell lymphoma), LAML (acute myeloid leukemia), LUAD (lung adenocarcinoma), LUSC (squamous cell carcinoma of the lung) OV (ovarian cancer), PCPG (pheochromocytoma and paraganglioma), SKCM (cutaneous melanoma), TGCT (testicular cancer), THCA (thyroid cancer), THYM (thymoma), UCEC (endometrioid carcinoma), and UCS (uterine carcinosarcoma). Prominently, PICALM expression was greater in CHOL (cholangiocarcinoma), ESCA (esophageal cancer), GBM (glioblastoma multiforme), HNSC (head and neck cancer), KICH (renal suspicious cell carcinoma), LGG (low-grade glioma of the brain), LIHC (liver cancer), PAAD (pancreatic cancer), and STAD (gastric cancer) than in normal tissues (Fig. 1A). PICALM expression was considerably decreased in BRCA samples compared to normal breast tissues (P<0.001, Fig. 1B). Furthermore, we employed ROC curves to determine the predictive value of PICALM expression, with an AUC of 0.825 (95% CI=0.800–0.850) to discriminate BRCA tissues from normal tissues (Fig. 1C).
Lastly, to better describe PICALM expression, we used the HPA database to examine PICALM protein levels in BRCA and normal tissues (Fig. 1D). PICALM protein levels in BRCA tissues were lower than in normal tissues. All of these findings imply that PICALM expression is involved in the formation and advancement of BRCA.
Clinicopathological Features of BRCA and PICALM Expression
We compared the clinical characteristics of BRCA patients with high and low PICALM expression (Table 1) and found that BRCA patients with high PICALM expression had a higher proportion of whites, worse t stage, and more invasive ductal carcinoma tissue types, and patients with basal PAM50 stage of breast cancer had more and worse clinical overall survival and disease-free survival. However, there was no significant difference in age, N stage, M stage, PR status, and ER status between the two groups.
At the same time, we also analyzed the differences in the expression level of PICALM among different clinical features. We found that from the perspective of histological type, the expression of PLCALM was significantly higher in the invasive ductal carcinoma group (LDC) than in the invasive lobular carcinoma group (ILC) (P<0.001). In the PAM50 classification, the expression of PICALM in the basal group was higher than that in the LumA and Her2 groups. The expression of PICALM in the negative ER group was higher than that in the positive group. In addition, in both the OS and DSS groups, the expression of PICALM was significantly higher in deceased patients than in survivors (Fig. 2). In addition, the results of single-gene logistic regression showed an association between PICALM expression and clinicopathological group characteristics (Table 2), and PICALM expression was positively associated with white race, invasive ductal carcinoma, and menopausal status.
PICALM expression correlates with different clinicopathological features of BRCA patients. Data are shown for (A) pathological stage; (B) histological type; (C) age; (D) T stage; (E) N stage; (F) M stage; (G) PAM50; (H) OS event; (I) DSS event; (J) ER status; (K) PR status; and (L) HER2 status. IDC, infiltrating ductal carcinoma; ILC, infiltrating lobular carcinoma; LumA, luminal A; LumB, luminal B; OS, overall survival; DSS, disease-specific survival; ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor receptor 2.*P < 0.05, **P < 0.01, ***P < 0.001
Enrichment Analysis of DEGs in BRCA
There were 1475 genes with differential expression between PICALM expression levels, of which 132 were up-regulated and 1343 were downregulated (FDR<0.05, |log2 FC|>1) (Table 3). We performed GSEA enrichment analysis for differential genes. The results showed that differential genes were significantly enriched in HALLMARK TGF BETA SIGNALING, HALLMARK ANGIOGENESIS, and HALLMARK IL6-JAK-STAT3 SIGNALING pathways (Fig. 3A). Meanwhile, we analyzed the Gene Ontology from three aspects (molecular function, cellular component, and biological process). BP of Gene Ontology gene sets for GSEA analysis results showed GO RIBONUCLEOPROTEIN COMPLEX BIOGENESIS, GO ESTABLISHMENT OF PROTEIN LOCALIZATION TO MEMBRANE, GO RIBOSOME BIOGENESIS, GO ATP METABOLIC PROCESS, and GO PROTEIN DNA COMPLEX SUBUNIT ORGANIZATION were enriched (Fig. 3B). CC of Gene Ontology gene sets for GSEA analysis indicates GO MITOCHONDRIAL MATRIX, GO MITOCHONDRIAL PROTEIN COMPLEX, GO PROTEIN DNA COMPLEX, GO RIBOSOME, and GO RIBOSOMAL subunits were enriched (Fig. 3C). In addition, MF of the Gene Ontology gene sets for GSEA analysis showed GO STRUCTURAL CONSTITUENT OF RIBOSOME, GO HORMONE ACTIVITY and GO RRNA BINDING, GO PRE MRNA BINDING, and GO OXIDOREDUCTASE ACTIVITY ACTING ON NAD PH QUINONE OR SIMILAR COMPOUND AS ACCEPTOR are enriched (Fig. 3D).
Gene set enrichment analysis (GSEA) of PICALM-associated DEGs. A GSEA analysis of Hallmark gene sets; B BP of Gene Ontology gene sets for GSEA analysis; C CC of Gene Ontology gene sets for GSEA analysis; D MF of the Gene Ontology gene sets for GSEA analysis. GSEA, gene set enrichment analysis; DEGs, differentially expressed genes
Immune Infiltration Correlates with PICALM Expression
To thoroughly understand the connection between PICALM expression and immunity, GSEA was utilized to evaluate immune cell infiltration in BRCA. It was found (Fig. 4A–C) that PICALM expression was shown to be associated with aDC (r=0.126, P<0.001), B cells (r=0.106, P<0.001), DC (r=0.117, P<0.001), eosinophils (r=0.087, P=0.004), iDC (r=0.144, P< 0.001), macrophages (r=0.390, P<0.001), mast cells (r=0.095, P=0.002), neutrophils (r=0.285, P<0.001), NK CD56dim cells (r=0.083, P=0.006), T cells (r=0.119, P<0.001), T helper cells (r=0.370, P<0.001), Tcm (r=0.545, P<0.001), Tem (r=0.160, P<0.001), TFH (r=0.070, P=0.019), Tgd (r=0.313, P<0.001), Th1 cells (r=0.268, P<0.001), and Th2 cells (r=0.129, P<0.001) which were substantially and positively associated with the level of immune cell infiltration. There was a significant inverse relationship between PICALM expression and the degree of immune cell infiltration in NK CD56bright cells (r=−0.262, P<0.001) and pDC (r=−0.234, P<0.001). Further, PICALM high-expression groups showed significantly higher immune enrichment scores compared to PICALM low-expression groups.
Correlation between PICALM expression and the level of immune infiltration of BRCA. A Correlation between PICALM expression and the relative abundance of 24 immune cells. B Immune cell infiltration levels were compared between groups with high and low PICALM expression. C Correlations between the relative enrichment scores of immune cells and the expression of PICALM
Prognostic Value of PICALM in BRCA
PICALM expression was evaluated in relation to the prognosis of BRCA patients using the Kaplan-Meier method. By using the intermediate value of PICALM expression as a threshold value, we classified BRCA patients according to their PICALM expression levels. The findings demonstrate that the PICALM high-expression group had a poorer prognosis for both OS and DSS than the PICALM low-expression group (Fig. 5A, M) (OS, HR= 1.76, 95% CI = 1.26–2.45, P=0.00; DSS, HR= 1.88, 95% CI = 1.21–2.94, P=0.005). Afterward, we examined the relationship between PICALM expression and prognosis in various subgroups of BRCA patients. Both OS (Fig. 5B–L) and DSS (Fig. 5N–X) were significantly worse for patients with high PICALM expression, in many other subgroups, including age >60 years, T1 and T2, N0 and N1, M0, post-menopause, luminal A, ER positive, PR positive, HER2 negative, stages II and III, infiltrating ductal carcinoma (IDC) subgroups (all P<0.05). Furthermore, we investigated prognostic indicators using univariate and multivariate Cox regression. On univariate COX analysis, age >60 years, T4 stage, arbitrary N stage, M1 stage, stage III/IV, LumB, Her2 positive, post-menopause, and PICALM high expression were associated with worse OS, whereas on multivariate analysis, age >60 years, M1 stage, postmenopausal, and PICALM high-expression patients had a worse prognosis (Fig. 6A, B). Furthermore, we examined the connection between clinicopathological characteristics and DSS, and the univariate results revealed that patients with T4 stage, arbitrary N stage, M1 stage, stage III/IV, HER2 positive, basal, and PICALM high expression had worse DSS, whereas the multifactorial analysis revealed that patients with age >60 years, M1 stage, postmenopausal, and PICALM high expression had worse DSS (Fig. 6C, D). Age, M stage, menopausal state, and PICALM expression are clearly connected with the prognosis of breast cancer patients.
Kaplan-Meier method to assess the prognostic value of PICALM expression in BRCA patients. Comparison of A overall survival and M disease-specific survival for breast cancer patients with high and low PICALM. B–L OS survival curves of age >60 years, T1 and T2, N0 and N1, M0, post-menopause, luminal A, ER positive, PR positive, HER2 negative, stages II and III, and IDC subgroups in breast cancer patients with high and low PICALM; N–X DSS survival curves of age >60 years, T1 and T2, N0 and N1, M0, post-menopause, luminal A, ER positive, PR positive, HER2 negative, stages II and III, and IDC subgroups in breast cancer patients with high and low PICALM. OS, overall survival; DSS, disease-specific survival; ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor receptor 2; LumA, luminal A; IDC, infiltrating ductal carcinoma
Low Expression of PICALM Inhibits Proliferation, Migration, and Invasion of BRCA Cells
In order to further understand the significance of the PICALM gene in BRCA, we used in vitro cellular experiments to verify it. The expression of the si-PICALM group was considerably decreased in MCF-7 cells than that of the NC group, according to qRT-PCR data (P<0.01, Fig. 7A). CCK-8 data revealed that si-PICALM might reduce BRCA cell proliferation (P<0.01, Fig. 7B). Depletion of si-PICALM resulted in drastic reductions in proliferation, migration, and invasion of BRCA cells, as shown by the growth curve and transwell experiments (both P<0.01; Fig. 7C, D). Collectively, these results demonstrated that PICALM was highly expressed in BRCA and significantly affected their proliferation, cell migration, and cell invasion.
Downregulation of PICALM reduced the proliferation and migration of MCF-7 breast cancer cells. A qRT-PCR identification of PICALM expression in MCF-7 cells; B the proliferation of MCF-7 cells was examined by CCK-8 assay; C, D the migration of MCF-7 cells was examined by transwell and wound healing/scratch. Image magnification transwell (20×), wound healing/scratches (10×) NC, negative control; ***P < 0.001, **P < 0.01
Discussion
BRCA, as the most common cancer among women worldwide, is the leading cause of cancer-related death among women. Although the survival rate of BRCA patients has improved with the emergence of new radiochemotherapy and immunotherapy methods, unfortunately, only about 20% of metastatic BRCA patients survive for more than 5 years [12, 26, 27]. Traditional markers such as ER and HER2 have important guiding values for the prognosis and treatment of BRCA patients, but they also have obvious limitations and cannot benefit every patient. Therefore, it is particularly important to continuously explore new biomarkers in BRCA and obtain more comprehensive and appropriate evaluation methods.
PICALM is involved in lectin-mediated endocytosis and autophagy, and affects the key processes of protein, lipids, growth factors, and neurotransmitters in the cell [10]. Studies have shown that PICALM is essential for the internalization and localization of cell surface proteins such as transferrin receptor (TfR) and epidermal growth factor receptor (EGFR) [13, 14, 28]. The lack of overexpression of PICALM will destroy the internalization of TfR and EGFR, and receptor internalization is crucial in regulating the conduction of intracellular and extracellular factors, as well as intracellular metabolic activity [29]. Existing studies have found that when t (10; 11), during chromosomal translocation, PICALM fuses with AF10 in the U937 cell line, affecting the progression of various hematological malignancies such as acute lymphoblastic leukemia and myeloid leukemia [30, 31]. Similarly, PICALM-interacting mitotic regulators (PIMREG) that interact with PICALM have also been found to play an important role in maintaining NF-B activation in breast cancer [32, 33]. However, whether PICALM expression promotes tumor growth, especially in BRCA, has not been studied. Open data opens up a feasible way to explore new markers. We used TCGA and GTEx databases to detect the expression of PICALM in various tumors, and found that PICALM was downregulated in a variety of malignant tumors, such as BRCA, ACC, BLCA, COAD, and DLBC. We investigated the relationship between PICALM expression and clinicopathological features of breast cancer patients and found that high PICALM expression was associated with poor clinicopathological parameters such as age, stage, and menopausal status. High expression of PICALM is an independent predictor of poor OS and DSS in BRCA patients. In addition, our experimental study found that knocking down PICALM can inhibit the proliferation, migration, and invasion of BRCA cells. These findings suggest that PICALM plays an important role in the progression of BRCA disease and that PICALM may be an attractive new biological target for the successful treatment of cancer. In order to further explore the potential mechanism of action of PICALM, we analyzed the functions of differential genes in the high-low-expression group of PICALM. Through GSEA analysis, we found that the pathways with high differential gene enrichment in the high-expression and low-expression groups of PICALM included oxidative phosphorylation, angiogenesis, TGF-β signaling pathway, and IL-6/JAK/STAT3 signaling pathway. These pathways play an important role in tumors. Tumor angiogenesis refers to the process in which endothelial cells proliferate and migrate in differentiated arteries in the presence of vascular growth factor, and rebuild vascular networks through degradation of the extravascular matrix and basement membrane. The proliferation, invasion, and migration of tumor cells are closely related to angiogenesis. Studies have shown that in the absence of angiogenesis, endothelial cell proliferation and migration are closely related to angiogenesis. The diameter of tumor tissue does not exceed 2 mm [34,35,36,37,38]. Transforming growth factor β (TGF-β) plays a variety of roles in the occurrence and development of BRCA, including promoting carcinogenesis and apoptosis in the early stage of BRCA and promoting tumor invasion in the late stage of BRCA. Abnormal regulation of TGF-β signaling and mutations in TGF-β receptors or downstream complexes have been shown to lead to tumor growth. In BRCA tissues, TGF-β is positively correlated with EGFR expression, and TGF-β signaling pathway is systematically correlated with EGFR deactivation [39]. The co-expression of PICALM may directly or indirectly participate in the pathogenesis and development of BRCA by affecting the oxidative phosphorylation, angiogenesis, and TGF signaling pathway of BRCA. Obviously, more experimental verification is needed to enrich the biological functions of PICALM in BRCA.
In addition, we also studied the relationship between immune cells and PICALM expression in BRCA. Immune cell infiltration, which is known to be associated with tumor growth and recurrence, may provide insights into neoadjuvant chemotherapy and immune checkpoint suppression (ICI) responses and is considered a key driver of clinical outcomes and immunotherapy reactivity [40,41,42]. Therefore, monitoring BRCA-infiltrating immune cells can not only be used in combination with ICI treatment, but also has prognostic significance for ICI treatment. We found that PICALM expression was negatively correlated with the infiltration of several immune cells, including B cells, eosinophils, mast cells, neutrophils, and T cells. B cells are important immune cells that perform a variety of activities in the immune response. Some studies have found that B-cell infiltration is related to the poor survival rate of BRCA individuals, and tumor-infiltrated B-lymphocytes can slow down tumor development by secreting antibodies, stimulating T cell response, and directly destroying tumor cells [43, 44]. Mast cells are immune cells that release a variety of active chemicals that help the immune system respond [45]. Giuseppe et al. believe that mast cell density is enhanced in gastric cancer and plays a tumorigenic role through the secretion of angiogenic factors and lymphangiogenic factors [46]. In addition, Reddy et al. [47] found that mast cell infiltration plays a role in the harmful consequences of inflammatory BRCA neoadjuvant chemotherapy. These findings suggest that inhibiting PICALM expression may affect BRCA progression and survival by controlling the number of invading immune cells.
Although our recent work increases our understanding of the association between PICALM expression and prognosis in BRCA patients, there are some limitations that need to be considered. Firstly, because this study relied on data from a database, we did not have access to some important clinical information, such as chemotherapy regimens. Secondly, while our in vitro studies and experimental results of PICALM expression in BRCA cells are consistent with our predictions, further studies and rigorous validation in animal experiments or clinical sample systems are needed to elucidate the carcinogenesis and progression of PICALM in BRCA.
Conclusion
To conclude, we found that PICALM expression was downregulated in many tumors, and importantly, that low PICALM expression was an independent poor predictive marker in BRCA patients, inexorably connected to aggressive clinical features and poor immune cell infiltration in BRCA patients. Next, in vitro cellular experiments were used to confirm our predictions. Finally, PICALM can be utilized to forecast the prognosis of BRCA patients. Our results will aid in the understanding of the processes behind human BRCA initiation and progression, as well as the development of more effective medical therapy in the future.
Data Availability
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
References
Sung, H., Ferlay, J., Siegel, R. L., et al. (2021). Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a Cancer Journal for Clinicians, 71(3), 209–249.
Siegel, R. L., Miller, K. D., & Jemal, A. (2019). Cancer statistics, 2019. CA: a Cancer Journal for Clinicians, 69(1), 7–34.
Zuo, Y., Li, Y., Zhou, Z., et al. (2017). Long non-coding RNA MALAT1 promotes proliferation and invasion via targeting miR-129-5p in triple-negative breast cancer. Biomedicine & Pharmacotherapy, 95, 922–928.
Hassan, M. K., Kumar, D., Naik, M., et al. (2018). The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS One, 13(1), e0191377.
Xu, J., Liu, L., Ma, R., et al. (2021). E2F1 Induces KIF26A transcription and promotes cell cycle progression via CDK-RB-E2Fs feedback loop in breast cancer. Frontiers in Oncology, 10, 530933.
Liu, H., Qiu, C., Wang, B., et al. (2021). Evaluating DNA methylation, gene expression, somatic mutation, and their combinations in inferring tumor tissue-of-origin. Frontiers in Cell and Development Biology, 9, 619330.
Hunter, N. B., Kilgore, M. R., & Davidson, N. E. (2020). The long and winding road for breast cancer biomarkers to reach clinical utility. Clinical Cancer Research, 26(21), 5543–5545.
Zhang, Y., Xiang, J., Tang, L., et al. (2021). Identifying breast cancer-related genes based on a novel computational framework involving KEGG pathways and PPI network modularity. Frontiers in Genetics, 12, 596794.
Dreyling, M. H., Martinez-Climent, J. A., Zheng, M., et al. (1996). The t(10;11)(p13;q14) in the U937 cell line results in the fusion of the AF10 gene and CALM, encoding a new member of the AP-3 clathrin assembly protein family. Proceedings of the National Academy of Sciences of the United States of America, 93(10), 4804–4809.
Tebar, F., Bohlander, S. K., & Sorkin, A. (1999). Clathrin assembly lymphoid myeloid leukemia (CALM) protein: Localization in endocytic-coated pits, interactions with clathrin, and the impact of overexpression on clathrin-mediated traffic. Molecular Biology of the Cell, 10(8), 2687–2702.
Caudell, D., & Aplan, P. D. (2008). The role of CALM-AF10 gene fusion in acute leukemia. Leukemia, 22(4), 678–685.
Meyerholz, A., Hinrichsen, L., Groos, S., et al. (2005). Effect of clathrin assembly lymphoid myeloid leukemia protein depletion on clathrin coat formation. Traffic, 6(12), 1225–1234.
Scotland, P. B., Heath, J. L., Conway, A. E., et al. (2012). The PICALM protein plays a key role in iron homeostasis and cell proliferation. PLoS One, 7(8), e44252.
Suzuki, M., Tanaka, H., Tanimura, A., et al. (2012). The clathrin assembly protein PICALM is required for erythroid maturation and transferrin internalization in mice. PLoS One, 7(2), e31854.
Narayan, P., Sienski, G., Bonner, J. M., et al. (2020). PICALM rescues endocytic defects caused by the Alzheimer’s disease risk factor APOE4. Cell Reports, 33(1), 108224.
Periñán, M. T., Macías-García, D., Labrador-Espinosa, M. Á., et al. (2021). Association of PICALM with cognitive impairment in Parkinson’s disease. Movement Disorders, 36(1), 118–123.
Xu, W., Tan, L., & Yu, J. T. (2015). The role of PICALM in Alzheimer’s disease. Molecular Neurobiology, 52(1), 399–413.
Borel, C., Dastugue, N., Cances-Lauwers, V., et al. (2012). PICALM-MLLT10 acute myeloid leukemia: A French cohort of 18 patients. Leukemia Research, 36(11), 1365–1369.
Tang, Z., Kang, B., Li, C., et al. (2019). GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Research, 47(W1), W556–W560.
Uhlen, M., Zhang, C., Lee, S., et al. (2017). A pathology atlas of the human cancer transcriptome. Science., 357(6352), eaan2507.
Love, M. I., Huber, W., & Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology, 15(12), 550.
Zhou, Y., Zhou, B., Pache, L., et al. (2019). Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nature Communications, 10(1), 1523.
Yu, G., Wang, L. G., Han, Y., et al. (2012). clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS, 16(5), 284–287.
Bindea, G., Mlecnik, B., Tosolini, M., et al. (2013). Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity, 39(4), 782–795.
Mayank, J. V. (2014). Drug target strategies in breast cancer treatment: Recent developments. Anti-Cancer Agents in Medicinal Chemistry, 14(10), 1414–1427.
Maruthanila, V. L., Elancheran, R., Kunnumakkara, A. B., et al. (2017). Recent development of targeted approaches for the treatment of breast cancer. Breast Cancer, 24(2), 191–219.
Sun, C. C., Li, S. J., Hu, W., et al. (2022). Retraction notice to: Comprehensive analysis of the expression and prognosis for E2Fs in human breast cancer. Molecular Therapy, 30(7), 2639.
Huang, F., Khvorova, A., Marshall, W., & Sorkin, A. (2004). Analysis of clathrin-mediated endocytosis of epidermal growth factor receptor by RNA interference. The Journal of Biological Chemistry, 279(16), 16657–16661.
Bohlander, S. K., Muschinsky, V., Schrader, K., et al. (2000). Molecular analysis of the CALM/AF10 fusion: Identical rearrangements in acute myeloid leukemia, acute lymphoblastic leukemia and malignant lymphoma patients. Leukemia, 14(1), 93–99.
Abdelhaleem, M., Beimnet, K., Kirby-Allen, M., et al. (2007). High incidence of CALM-AF10 fusion and the identification of a novel fusion transcript in acute megakaryoblastic leukemia in children without Down’s syndrome. Leukemia, 21(2), 352–353.
Jiang, L., Ren, L., Zhang, X., et al. (2019). Overexpression of PIMREG promotes breast cancer aggressiveness via constitutive activation of NF-κB signaling. EBioMedicine, 43, 188–200.
Li, T., Kang, G., Wang, T., et al. (2018). Tumor angiogenesis and anti-angiogenic gene therapy for cancer. Oncology Letters, 16(1), 687–702.
Ying, L., Chen, Q., Wang, Y., et al. (2012). Upregulated MALAT-1 contributes to bladder cancer cell migration by inducing epithelial-to-mesenchymal transition. Molecular BioSystems, 8(9), 2289–2294.
Kuol, N., Stojanovska, L., Apostolopoulos, V., et al. (2018). Role of the nervous system in tumor angiogenesis. Cancer Microenvironment, 11(1), 1–11.
Roskoski, R., Jr. (2007). Vascular endothelial growth factor (VEGF) signaling in tumor progression. Critical Reviews in Oncology/Hematology, 62(3), 179–213.
Garcea, G., Lloyd, T. D., Gescher, A., et al. (2004). Angiogenesis of gastrointestinal tumours and their metastases--A target for intervention? European Journal of Cancer, 40(9), 1302–1313.
Syed, V. (2016). TGF-β Signaling in Cancer. Journal of Cellular Biochemistry, 117(6), 1279–1287.
Tang, X., Shi, L., Xie, N., et al. (2017). SIRT7 antagonizes TGF-β signaling and inhibits breast cancer metastasis. Nature Communications, 8(1), 318.
Cao, W. H., Liu, X. P., Meng, S. L., et al. (2016). USP4 promotes invasion of breast cancer cells via Relaxin/TGF-β1/Smad2/MMP-9 signal. European Review for Medical and Pharmacological Sciences, 20(6), 1115–1122.
Li, B., Severson, E., Pignon, J. C., et al. (2016). Comprehensive analyses of tumor immunity: Implications for cancer immunotherapy. Genome Biology, 17(1), 174.
Liu, J., Tan, Z., He, J., et al. (2020). Identification of three molecular subtypes based on immune infiltration in ovarian cancer and its prognostic value. Bioscience Reports, 40(10), BSR20201431.
Havel, J. J., Chowell, D., & Chan, T. A. (2019). The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nature Reviews. Cancer, 19(3), 133–150.
Wang, S. S., Liu, W., Ly, D., et al. (2019). Tumor-infiltrating B cells: Their role and application in anti-tumor immunity in lung cancer. Cellular & Molecular Immunology, 16(1), 6–18.
Yang, C., Lee, H., Jove, V., et al. (2013). Prognostic significance of B-cells and pSTAT3 in patients with ovarian cancer. PLoS One, 8(1), e54029.
Komi, D. E. A., & Redegeld, F. A. (2020). Role of mast cells in shaping the tumor microenvironment. Clinical Reviews in Allergy and Immunology, 58(3), 313–325.
Sammarco, G., Varricchi, G., Ferraro, V., et al. (2019). Mast cells, angiogenesis and lymphangiogenesis in human gastric cancer. International Journal of Molecular Sciences, 20(9), 2106.
Reddy, S. M., Reuben, A., Barua, S., et al. (2019). Poor response to neoadjuvant chemotherapy correlates with mast cell infiltration in inflammatory breast cancer. Cancer Immunology Research, 7(6), 1025–1035.
Acknowledgements
We thank the TCGA database for sharing a large amount of data and the convenience provided by several online database tools. At the same time, thank you to Baidu Translate and DeepL for providing free translation and polishing.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. All authors read and approved the final manuscript. All authors commented on previous versions of the manuscript. All authors agree to become authors of the manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable
Consent for Publication
All authors consent to the publication of this work.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
A, N., Lyu, P., Yu, Y. et al. PICALM as a Novel Prognostic Biomarker and Its Correlation with Immune Infiltration in Breast Cancer. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-023-04840-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-023-04840-z