Abstract
Progression, prognosis, and therapeutic strategy of stomach adenocarcinoma (STAD) have a close connection with tumor microenvironment (TME). Thus, it is pivotal to delve into the TME and immune-related genes, which may bring possibilities for improving patient’s prognosis. TCGA-STAD dataset was analyzed to acquire differentially expressed lncRNAs in tumor samples, which were overlapped with the immune-related lncRNA datasets in the ImmLnc database. Twenty-six lncRNAs related to STAD immunity and patient’s prognosis were acquired by univariate Cox analysis. Following lncRNA expression patterns, STAD samples could be classified into two clusters with completely different immune patterns. We performed multivariate Cox regression analysis on lncRNAs to identify 7-feature lncRNAs and constructed a corresponding prognostic model. The model validity was verified by survival analysis and ROC curve in validation and training sets. To explore connection between model and TME and tumor drug resistance, this study analyzed differences in immune cell infiltration between samples from high- and low-risk groups and then revealed immune cells follicular helper with significant differences in tumor tissue infiltration. Analysis of resistance to chemotherapeutic drugs revealed that samples in the high-risk group had resistance to cisplatin, doxorubicin, bleomycin, and gemcitabine. Through univariate and multivariate Cox analyses, we manifested that risk score could be an independent prognostic factor. Combining risk score and clinical factors, a nomogram was constructed to accurately predict patient’s prognosis. This model can effectively predict prognosis, TME, and drug resistance of STAD patients, which may provide a reference for tumor development evaluation and precise treatment for clinical STAD.
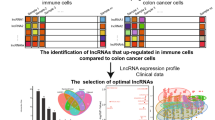
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stomach adenocarcinoma (STAD) is a main histological type of gastric cancer, accounting for 95% of all gastric cancers [1]. Currently, the mainstay of treatment for STAD is surgery combined with chemotherapy, but drug resistance, metastasis, or recurrence in STAD patients make the prognosis still poor [2]. In particular, the high heterogeneity of STAD greatly aggravates the difficulty of treatment. Its heterogeneity is manifested in the differentially activated pathways, immune characteristics, disease susceptibility, etc. and also predicts the need for more accurate prognostic prediction and precise treatment options for patients [3]. Therefore, new prognostic biomarkers for STAD are urgently needed to precisely assess prognosis.
Tumor immune features have attracted much attention because they can assess the clinical benefits of a variety of treatments, and the corresponding immunotherapy has also been used as a novel therapy to achieve good efficacy in the treatment of multiple cancers such as lung cancer, breast cancer, and pancreatic cancer [4,5,6]. The key to the development of immunotherapy lies in the in-depth study of the tumor immune microenvironment [7]. The tumor microenvironment (TME) of STAD similarly contains abundant infiltrating immune cells. Among them, tumor-associated macrophages were remarkably associated with STAD tumor angiogenesis, tumor invasion depth, lymph node metastasis status, and clinical stage [8]. Tumor-infiltrating immune cells act on the regulation of tumor immunity by releasing a variety of cytokines and growth factors, so immune-related gene expression and immune cell infiltration are closely related to tumor development [9]. There are several studies evaluating the prognosis of cancer patients based on the feature genes in the TME. For example, Ren et al. [10] mined the prognostic marker genes in the STAD TME, which can be utilized to evaluate the clinicopathological features and overall survival (OS) rate of patients. The genes associated with inflammation in STAD can represent different dendritic cell and degrees of macrophage infiltration. Wu et al. [11] revealed the potential value of these genes for assessing patient prognosis. Comprehensive analysis, immune-related genes are closely related to OS of patients, which can offer a new reference for the treatment and prognosis prediction of STAD.
We combined TCGA and ImmLnc databases to obtain STAD immune-related lncRNAs and obtained STAD immune-related lncRNAs with prognostic significance by univariate Cox analysis. According to the expression patterns of these lncRNAs, STAD patients can be classified into two subtypes, and ssGSEA analysis reveals the different immune patterns represented by the two subtypes. Multivariate Cox analysis of these immune-related lncRNAs with prognostic significance constructed the STAD prognostic model. The model can accurately predict the OS rate of patients, and the prognostic model can assess immune cell infiltration and resistance of samples to chemotherapeutic drugs. Immune-related feature lncRNAs not only have the ability to predict the prognosis of STAD patients, but also can explain the potential mechanisms affecting the prognosis of patients. This will provide guidance for the precise diagnosis of clinical STAD and the development of targeted therapeutic strategies.
Materials and Methods
Data Sources
The gene transcriptome data of STAD as well as the clinical information of the patient (TCGA-STAD) were obtained from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). The gene transcriptome dataset was annotated to finally obtain the expression information of lncRNAs and mRNAs.
Differentially Expressed Immune-Related lncRNA Screening
Differential analysis of lncRNA data obtained in TCGA-STAD was performed using the R package “edgeR” to screen differentially expressed lncRNAs (DElncRNAs) with logFC > 1.5 and Padj < 0.05 [12]. Through the website ImmLnc (http://bio-bigdata.hrbmu.edu.cn/ImmLnc/index.jsp), the immune-related lncRNA set was retrieved, and the immune-related lncRNA set was overlapped with STAD DElncRNAs to obtain differentially expressed immune-related lncRNAs.
Consensus Clustering Analysis of Samples
The R package “survival” was used to perform univariate Cox regression analysis of immune-related lncRNAs that were differentially expressed in STAD, using Padj < 0.05 as a screening criterion to obtain a lncRNA set with prognostic significance for STAD (https://cran.r-project.org/web/packages/survival/survival.pdf). The R package “ConsensusClusterPlus” was used to perform k-means consensus cluster analysis of STAD samples based on the expression of prognostically relevant lncRNAs, and the optimal cluster number K value was determined using a double sampling scheme [13]. The samples were divided into different subtypes according to the optimal number of clusters, and the sample differences between subtypes were verified using PCA dimensionality reduction. To determine differences in immune profiles of the subtypes, 29 immune-related phenotypes were selected and analyzed by ssGSEA enrichment based on mRNA expression profiles.
Establishment of the Prognostic Model
The samples in TCGA-STAD dataset were divided into training and validation sets in a 7:3 ratio using the R package “caret” (https://cran.r-project.org/web/packages/caret/caret.pdf). Multivariate Cox regression analysis was performed with prognostically relevant lncRNAs to obtain information on feature lncRNAs, their coefficients, and hazard ratios, and a risk assessment model was built based on the coefficients.
Validation of the Prognostic Model
The risk score was calculated based on the expression level and risk coefficient of each feature lncRNA, and the median was used as a cut-off value to distinguish the high- and low-risk groups of the training set and the validation set. PCA dimensionality reduction was performed on samples from the two risk groups by the R package “factoextra” (https://cran.microsoft.com/snapshot/2016-11-30/web/packages/factoextra/factoextra.pdf). Survival curves were plotted using the R package “survival” to verify the model grouping effect. Finally, ROC curves were plotted by the R package “timeROC” to calculate the AUC values corresponding to the risk assessment model predicting the 3- and 5-year survival rates of STAD patients, respectively [14].
Analysis of Immune Microenvironment and Chemotherapy Response of Samples
The R package “CIBERSORT” was used to score the abundance of immune cell infiltration in samples from the high- and low-risk groups of the training set, and a permutation test was used to set p value < 0.05 as the standard to screen immune cells with notable differences in infiltration levels between the two risk groups (https://doi.org/10.1038/nmeth.3337). The R package “pRRophetic” was used to predict the response of high- and low-risk group samples in the training set to chemotherapy with the drugs cisplatin, doxorubicin, bleomycin, and gemcitabine, and the Wilcoxon test was performed on the predicted drug IC50 values of two risk groups to analyze the differences in drug resistance [15].
Construction and Assessment of the Nomogram
Univariate and multivariate Cox regression analyses of risk score, gender, age, and TNM tumor stage were performed in TCGA-STAD training set to assess the independence of the multiple prognostic models. A nomogram was drawn based on the risk score and patient clinical information by the R package “rms,” and a calibration curve was drawn to predict the 3-year and 5-year survival rates of STAD patients [16].
Results
Immune-Related lncRNAs with Prognostic Significance in STAD
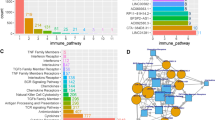
1812 lncRNAs which were remarkably differentially expressed in STAD tumor samples compared with healthy samples were obtained by the differential analysis, of which 1396 were upregulated and 416 were downregulated (Fig. 1A). 199 lncRNAs were obtained after intersection of DElncRNAs with immune-related lncRNAs in the ImmLnc database (Fig. 1B). Univariate Cox analysis of these lncRNAs was performed to obtain 26 lncRNAs that were immune-related in STAD and had prognostic significance (TableS1). In this study, unsupervised consensus clustering was conducted based on prognostically relevant lncRNA expression profiles. The STAD samples were divided into two categories named cluster1 and cluster2 using K = 2 as the optimal number of clusters (Fig. 1C-E). The results of PCA dimensionality reduction showed that this clustering mode made the two groups of samples clearly distinguished (Fig. 1F). To understand the immune patterns of the two types of samples, we obtained the immune cell infiltration level and immune-related gene set enrichment level in the two groups of samples by ssGSEA analysis. The results revealed that the immune cell infiltration level and immune-related gene set enrichment level in cluster2 showed a more remarkable upregulation trend than cluster1. Therefore, we classified cluster1 as the “cold-immune” subtype and cluster2 as the “heat-immune” subtype (Fig. 1G). The above results illustrated that the prognostically relevant lncRNAs obtained in this study had a potential role in distinguishing the immune status of STAD.
Immune-related lncRNAs with prognostic significance in STAD. A Volcano plot of lncRNA differential analysis in STAD tumor samples relative to healthy samples; red represents upregulated genes, and green represents downregulated genes. B Venn diagram of the intersection of differentially expressed lncRNAs with immune-related lncRNAs. C Consensus cluster cumulative distribution function curve with K = 2 ~ 10. D Relative change in the area under the cumulative distribution function curve with K = 2 ~ 10. E Sample agreement matrix for K = 2. F Differences in PCA dimensionality reduction distribution between the 2 cluster subtypes. G Heat map of ssGSEA scores for the two STAD subtypes.
Immune-Related lncRNA Prognostic Model Construction
Combined with multivariate Cox regression analysis of the expression of prognosis-related lncRNAs and OS in STAD samples of the training set, 7-feature lncRNAs were obtained, namely, ZFPM2-AS1, LINC01060, LINC01537, C15orf54, CYMP-AS1, LINC01614, and AC092198.1 (Fig. 2). The risk assessment formula was obtained according to the coefficient and expression level of feature lncRNAs: risk score = 0.146*ZFPM2-AS1 + 0.075*LINC01060 + 0.164*LINC01537 + 0.070*C15orf54 + 0.060*CYMP-AS1 + 0.075*LINC01614-0.071*AC092198.1.
Immune-Related lncRNA Prognostic Model Evaluation
The samples in the training set and validation set were divided into high- and low-risk groups according to the median value of risk score. Seven feature lncRNAs showed notable differential expression in the two risk groups of STAD training set and validation set, of which LINC01060, LINC01537, LINC01614, C15orf54, ZFPM2-AS1, and CYMP-AS1 showed upregulation in the high-risk group of training set, and AC092198.1 showed considerable downregulation in the high-risk group of training set (Fig. 3A). The samples in the training set were arranged according to the risk score from low to high. Then the scatter of sample survival time was plotted, which found that as the risk score increased, the number of deaths gradually increases, accompanying with the gradual shrink of the survival time of patients (Fig. 3B-C). PCA dimensionality reduction of the training set samples revealed a clear distinction between samples from the two risk groups (Fig. 3D). The results of survival analysis indicated that the OS of patients in the high-risk group of the training set was markedly lower than that in the low-risk group (Fig. 3E). These results were equally presented in the validation set (Fig. 4A-E). The ROC curve was plotted to verify the accuracy of the model in predicting the prognosis, and the results showed that the AUC values of the model in predicting the 3- and 5-year survival of STAD patients were 0.69, 0.81 (training set), and 0.75, 0.73 (validation set), respectively (Figs. 3F and 4F). These results indicated that the 7-lncRNA risk assessment model had good predictive performance for the prognosis of STAD patients in TCGA-STAD dataset.
7-lncRNA risk assessment model predicts the survival time and status of patients in the training set. A Heat map of the expression of 7-feature lncRNAs in the high- and low-risk groups. B Distribution of risk scores for STAD patients (green dots, low-risk group; red dots, high-risk group). C Scatterplot showing the survival of STAD patients (green dots, survival; red dots, death). D PCA dimension reduction map of patients in two risk groups. E Kaplan-Meier survival curves showing the difference in patient survival between the high (red) and low (blue)-risk groups. F ROC curve for predicting 3- and 5-year overall survival of patients based on a 7-feature lncRNA risk assessment model.
7-lncRNA risk assessment model predicts the survival time and status of patients in the validation set. A: Heat map of expression and clinicopathological differences of 7-feature lncRNAs in the high- and low-risk groups. B Distribution of risk scores for each STAD patient (green dots, low-risk group; red dots, high-risk group). C Scatterplot showing the survival of STAD patients according to the risk score (green dots, survival; red dots, death). D PCA dimensionality reduction map of patients in two risk groups. E Kaplan-Meier survival curves showing the difference in patient survival between the high (red) and low (blue)-risk groups. F ROC curve for predicting 3- and 5-year overall survival of patients based on a 7-feature lncRNA risk assessment model.
Assessment of the Prognostic Model on the Immune Microenvironment and Chemotherapy Response of STAD
To explore the association between the prognostic model and the immune microenvironment of STAD tumors, this study assessed the abundance of each immune cell infiltration in samples from TCGA-STAD training set (Fig. 5A). There was some difference in immune cell infiltration level in the tumor samples of patients in the two risk groups (Fig. 5B). Correlation analysis between the abundance of each immune cell infiltration indicated a negative correlation between most immune cells (Fig. 5C). These results suggested that cell–cell infiltration may be affected by antagonism, and TME-based assessment of immune levels required comprehensive consideration of multiple cell infiltration levels. The abundance of T cells follicular helper, T cells CD8, and T cells CD4 memory activated cell infiltration, a variety of cells related to anti-tumor immune activation, was prominently downregulated in the high-risk group relative to the low-risk group (Fig. 5D). Afterwards, this study predicted the sensitivity of two risk groups to common chemotherapeutic drugs for STAD. The results showed that the chemotherapeutic drugs cisplatin, doxorubicin, bleomycin, and gemcitabine had higher IC50 values in patients in the high-risk group (Fig. 5E-H), indicating that the high-risk group had favorable chemotherapy drug tolerance. The above results indicated that the 7-lncRNA prognostic model was capable of assessing both immune cell infiltration and chemotherapeutic drug tolerance in STAD samples.
Analysis of immune cell infiltration and drug resistance differences between samples in two risk groups of TCGA-STAD training set. A Relative infiltration abundance of immune cells in STAD samples. B Heat map of relative infiltration abundance of immune cells in two risk groups. C Heat map of correlation between the infiltration abundance of each immune cell. Red, positive correlation; blue, negative correlation; darker color, higher correlation; × indicates p > 0.05. D Difference in infiltration of each immune cell between the high (red) and low (blue)-risk groups. E–H Predicted IC50 difference of chemotherapeutic drugs cisplatin (E), doxorubicin (F), bleomycin (G), and Gemcitabine (H) in samples in two risk groups.
Construction and Evaluation of Nomograms
Univariate Cox regression analysis was performed using the risk score given by the 7-feature lncRNA prognostic model as an independent factor in combination with patient clinical factors. The results exhibited that age (HR = 1.815, p = 0.007), N (HR = 1.761, p = 0.032), M (HR = 2.022, p = 0.029), and risk score (HR = 1.406, p < 0.001) were all significant for patient prognosis and used as prognostic risk factors (Fig. 6A), and multivariate regression analysis indicated that age as well as risk score was closely related to prognosis of patients (Fig. 6B). This result suggested that both age and risk score could be used as independent prognostic risk factors for STAD. Therefore, two indicators, age and risk score, were combined to optimize the risk stratification in this study: the group with risk score lower than the median was selected as the low-risk group, the group with risk score higher than the median and age ≤ 65 was selected as the intermediate risk group, and the group with age > 65 and risk score higher than the median was selected as the high-risk group. Among these three risk groups, there was a remarkable difference in the OS of patients, with patients in the high-risk group having the worst prognosis, and those in the low-risk group having the best prognosis (Fig. 6C). The contribution of this optimized stratification to the accuracy of the model was verified. Finally, considering that the multivariate prediction model was superior to the univariate prediction model, this study plotted a nomogram predicting the 3-year and 5-year OS of STAD patients in combination with the 7-feature lncRNA risk score and optimized age stratification and other clinical factors (Fig. 6D). The corresponding calibration curve showed that the nomogram had good predictive ability (Fig. 6E-F).
Evaluation of independent prognostic factors and nomogram construction. A Forest plot for univariate Cox regression analysis of risk score, age, gender, and T, N, and M stages. B Forest plot of multivariate Cox regression analysis for risk score, age, gender, and T, N, and M stages. C TCGA-STAD patients’ survival curves after the optimized stratification of age and risk score. D Nomogram of risk score combined with clinical information. E–F Calibration curve of nomograms to predict 3- (E) and 5-year (F) survival of STAD patients.
Discussion
STAD is a frequent malignancy of digestive system in the antrum or pylorus with high level of heterogeneity [17]. This heterogeneity occurs not only macroscopically, but also in the heterogeneous TME and gene expression patterns. Just as gene expression differences alone can divide STAD into four clusters representing different phenotypes, mesenchymal-like subtypes, microsatellite-unstable, TP53-activated, and non-activated types, and have different cytokine expression patterns in different clusters [18]. That is, different subtypes have differences in immune profiles. Therefore, we focused on the correlation between STAD tumor immunity and patient prognosis, obtained STAD-related lncRNA expression data and clinical information from TCGA, identified and validated prognostic feature genes related to tumor immunity, and established a prognostic risk assessment model. The study is important to explore STAD prognosis and treatment options.
In this study, we applied an unsupervised consensus clustering algorithm based on prognostically relevant lncRNAs to divide the samples into two groups of subtypes that could represent different immune patterns and classified the overall high-level infiltration and expression groups of immune cells and related genes as “heat-immune” and the low-level infiltration and expression groups as “cold-immune.” Subsequently, differences were found in Th1 and Th2 cell infiltration and APC and immune checkpoint gene expression levels between the two groups. Th1 and Th2 are two helper T cell subtypes that both act on cancer progression by secreting cytokines, especially interleukins [19]. In another therapeutic study for colon cancer, a typical digestive cancer, it was clearly stated that high expression of helper T cells can significantly prolong patient survival [20]. The tumor inhibitor gene APC plays a vital role in regulating tumorigenesis, cell cycle regulation, and apoptosis, especially in digestive system diseases such as gastric cancer and colon cancer, which are involved in regulating the proliferation and migration of tumor cells [21, 22]. High expression of APC in gastric cancer can reduce adverse prognosis of patients by blocking the Wnt/β-catenin pathway, inhibiting epithelial-mesenchymal transition and metastasis of tumors, and reducing the tumor aggressive phenotype [23]. Therefore, the difference in immune patterns reflected by the two subtypes may be closely related to patients’ prognosis.
Several stomach adenocarcinoma prognostic biomarkers have been studied and developed, among which ctDNA was understood for its convenience and reliability. In Maron’s study, gastroesophageal adenocarcinoma prognostic status was evaluated by ctDNA sequencing, in which genetic alterations were analyzed. However, it was admitted that some desired genetic alterations could not be detected for technique limitation. Herein, we identified 7 immune-related feature lncRNAs for constructing a prognostic model of STAD. Among the seven lncRNAs, only AC092198.1 served as a patient prognostic protective factor, and the remaining six (ZFPM2-AS1, LINC01060, LINC01537, C15orf54, CYMP-AS1, LINC01614) served as prognostic risk factors. The drawbacks are that AC092198.1 and C15orf54 have not been reported in relevant studies, and their properties acting on cancer remain to be further studied. In this study, we found these two lncRNAs may be associated with the prognosis of STAD. The majority of the remaining feature lncRNAs are associated with cancer progression and poor patient prognosis. ZFPM2-AS1 is activated by environmental factors in tumor tissues, and its abnormally high expression in gastric cancer samples can promote gastric cancer cell proliferation and inhibit apoptosis by inhibiting the p53 signaling pathway, so the OS and progression-free survival of gastric cancer patients with high expression of this lncRNA are significantly decreased [24]. Studies on LINC01060 acting on digestive system cancers have not been reported, and the current study only shows that this lncRNA worsens disease progression by promoting oncogene c-Myc transcription and accumulation of hypoxia-inducible factor HIF-1 in highly differentiated glioma stem cells [25]. This has reference significance for STAD, which is also highly heterogeneous in this study, and perhaps the upregulation of transcription factors is also an important reason why this lncRNA acts as a prognostic adverse factor. LINC01537 was significantly associated with the prognostic survival of patients with lung adenocarcinoma, and interestingly, this lncRNA has the effect of inhibiting the growth and metastasis of lung adenocarcinoma and can be used as a favorable factor for the prognosis of lung adenocarcinoma by regulating tumor metabolic capacity [26]. In this study, on the other hand, LINC01537 served as an adverse factor for the prognosis of STAD, indicating that this lncRNA plays different roles in different cancers. Same as this study, a study by Wu et al. [27] has shown that high expression of LINC01537 in gastric cancer is associated with shorter prognostic survival time of patients. CYMP-AS1 and LINC01614 have been shown to be adverse prognostic factors in gastric cancer, and it is worth mentioning that LINC01614 has a more remarkable promoting effect on cell proliferation, migration, invasion, epithelial-mesenchymal transition, and tumor growth in vivo and accelerates the progression of G2/M phase in the cell cycle [27]. These results indicate that most of the feature lncRNAs in this study are associated with cancer progression and have potential value in predicting patient prognosis.
Immune microenvironment analysis based on prognostic models showed that the abundance of T cells CD4 memory resting cell infiltration was markedly upregulated in the high-risk group, accompanied by a significant downregulation of the abundance of T cells follicular helper, T cells CD4 memory, and T cells CD8 activated cell infiltration. Among them, T cells CD4 memory resting is common in the study of HIV. Generally, the infiltration of this cell has a protective effect on HIV and is not conducive to the clearance of HIV. In other words, the presence of these dormant memory CD4 T cells hinders the clearance of the target pathogen by the drug [28], which may be closely related to the higher chemoresistance of STAD samples in the high-risk group revealed in this study. Both CD8 T cells and activated CD4 memory T cells are associated with upregulated immune levels [29]. T cells follicular helper plays a crucial role in both initiating B-cell antibody responses and maintaining the body humoral immunity, with anti-microbial defense and the ability to inhibit cancer development [30]. These immune cells can produce cytokines such as TNF-α and interleukins [31, 32]. Immune cells in the tumor immune microenvironment can exhibit differences in drug resistance through different accumulation of cytokines[33]. Thus, the downregulation of these immune cell infiltration levels predicts a lower immune level of the tumor immune microenvironment and also shows different drug resistance. Patients with STAD often present poor prognoses. The prognostic model constructed in this study can be used to assess the immune microenvironment and chemotherapeutic drug response in STAD patients, which can provide guidance for the precise treatment of STAD patients. Similarly, Chen and his colleagues displayed a TME-associated prognostic signature for colon cancer. In their study, colon cancer samples were scored in immune and stromal degree, followed by screening DEGs according to the scores, and Cox regression was introduced to the DEGs constructing a TME-associated prognostic model (PMID: 34,497,681). Comparing to their work, our study presented prognostic model with lncRNA signature, which was different from Chen`s study. However, both the similar analytical steps showed reliable.
In conclusion, we obtained 26 lncRNAs associated with immunity and prognosis by univariate Cox analysis. And their close association with STAD immunity was verified by consensus clustering. We then performed multivariate Cox analysis of prognosis-related lncRNAs, by which we had successfully mined the feature lncRNAs and constructed a 7-lncRNA prognostic model whose their effectiveness had been evaluated then. Based on this prognostic model, it can not only predict the OS of STAD patients, but also assess the tumor immune infiltration and sample resistance to chemotherapeutic drugs. Finally, in this study, nomograms were constructed based on prognostic model risk score, age, gender, and TNM tumor stage for accurately predicting the prognosis of STAD patients. The prognostic model constructed in this study can be used as reliable data support for clinical STAD prognosis prediction and precise treatment.
Our study firstly presented immune-related DElncRNAs signature combining K-means clustering and Cox regression analyses to assess the prognosis of stomach adenocarcinoma patients. However, this study still has limitations. First, the performance evaluation of our prognostic model should be validated on more STAD datasets. Second, all data are derived from open databases, and there are systematic errors among the data. Finally, this study is a pure bioinformatics study, and the results of the role of feature lncRNAs on patient prognosis and drug resistance differences in samples should be further confirmed by clinical experiments. Hence, in the follow-up study, we will try to establish our own sample database to further verify the experimental results and explore its potential mechanism in depth.
Data Availability
The data used to support the findings of this study are included within the article. The data and materials in the current study are available from the corresponding author on reasonable request.
References
Balakrishnan, M., George, R., Sharma, A., & Graham, D. Y. (2017). Changing trends in stomach cancer throughout the world. Current gastroenterology reports, 19, 36. https://doi.org/10.1007/s11894-017-0575-8
Hoshi, H. (2020). Management of Gastric Adenocarcinoma for general surgeons. Surgical Clinics of North America, 100, 523–534. https://doi.org/10.1016/j.suc.2020.02.004
Zhang, M., et al. (2021). Dissecting transcriptional heterogeneity in primary gastric adenocarcinoma by single cell RNA sequencing. Gut, 70, 464–475. https://doi.org/10.1136/gutjnl-2019-320368
Steven, A., Fisher, S. A., & Robinson, B. W. (2016). Immunotherapy for lung cancer. Respirology, 21, 821–833. https://doi.org/10.1111/resp.12789
Sugie, T. (2018). Immunotherapy for metastatic breast cancer. Chin Clin Oncol, 7, 28. https://doi.org/10.21037/cco.2018.05.05
Morrison, A. H., Byrne, K. T., & Vonderheide, R. H. (2018). Immunotherapy and prevention of pancreatic cancer. Trends Cancer, 4, 418–428. https://doi.org/10.1016/j.trecan.2018.04.001
Lei, X., et al. (2020). Immune cells within the tumor microenvironment: Biological functions and roles in cancer immunotherapy. Cancer Letters, 470, 126–133. https://doi.org/10.1016/j.canlet.2019.11.009
Zhang, H., et al. (2015). Infiltration of diametrically polarized macrophages predicts overall survival of patients with gastric cancer after surgical resection. Gastric Cancer, 18, 740–750. https://doi.org/10.1007/s10120-014-0422-7
Hinshaw, D. C., & Shevde, L. A. (2019). The tumor microenvironment innately modulates cancer progression. Cancer Research, 79, 4557–4566. https://doi.org/10.1158/0008-5472.CAN-18-3962
Ren, N., Liang, B. & Li, Y. (2020). Identification of prognosis-related genes in the tumor microenvironment of stomach adenocarcinoma by TCGA and GEO datasets. Biosci Rep 40, 10.1042/BSR20200980.
Wu, M. et al. (2020). Development and validation of an immune-related gene prognostic model for stomach adenocarcinoma. Biosci Rep 40, 10.1042/BSR20201012.
Robinson, M. D., McCarthy, D. J., & Smyth, G. K. (2010). edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616
Wilkerson, M. D., & Hayes, D. N. (2010). ConsensusClusterPlus: A class discovery tool with confidence assessments and item tracking. Bioinformatics, 26, 1572–1573. https://doi.org/10.1093/bioinformatics/btq170
Blanche, P., Dartigues, J. F., & Jacqmin-Gadda, H. (2013). Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Statistics in Medicine, 32, 5381–5397. https://doi.org/10.1002/sim.5958
Hu, X., Wu, L., Liu, B., & Chen, K. (2021). Immune infiltration subtypes characterization and identification of prognosis-related lncrnas in adenocarcinoma of the esophagogastric junction. Frontiers in Immunology, 12, 651056. https://doi.org/10.3389/fimmu.2021.651056
Huang, C., et al. (2019). Clinical significance of serum CA125, CA19-9, CA72-4, and fibrinogen-to-lymphocyte ratio in gastric cancer with peritoneal dissemination. Frontiers in Oncology, 9, 1159. https://doi.org/10.3389/fonc.2019.01159
Cancer Genome Atlas Research, N. (2014). Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209. https://doi.org/10.1038/nature13480.
Cristescu, R., et al. (2015). Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nature Medicine, 21, 449–456. https://doi.org/10.1038/nm.3850
Yoshida, N., et al. (2016). A High RORgammaT/CD3 Ratio is a strong prognostic factor for postoperative survival in advanced colorectal cancer: Analysis of helper T cell lymphocytes (Th1, Th2, Th17 and regulatory T cells). Annals of Surgical Oncology, 23, 919–927. https://doi.org/10.1245/s10434-015-4923-3
Tosolini, M., et al. (2011). Clinical impact of different classes of infiltrating T cytotoxic and helper cells (Th1, th2, treg, th17) in patients with colorectal cancer. Cancer Research, 71, 1263–1271. https://doi.org/10.1158/0008-5472.CAN-10-2907
Morin, P. J. (2019). Colorectal cancer: The APC-lncRNA link. The Journal of clinical investigation, 129, 503–505. https://doi.org/10.1172/JCI125985
Wang, L., et al. (2015). Targeting Cdc20 as a novel cancer therapeutic strategy. Pharmacology & Therapeutics, 151, 141–151. https://doi.org/10.1016/j.pharmthera.2015.04.002
Yang, X. Z., et al. (2018). LINC01133 as ceRNA inhibits gastric cancer progression by sponging miR-106a-3p to regulate APC expression and the Wnt/beta-catenin pathway. Molecular Cancer, 17, 126. https://doi.org/10.1186/s12943-018-0874-1
Kong, F., et al. (2018). Correction: ZFPM2-AS1, a novel lncRNA, attenuates the p53 pathway and promotes gastric carcinogenesis by stabilizing MIF. Oncogene, 37, 6010. https://doi.org/10.1038/s41388-018-0412-z
Li, J., et al. (2021). Hypoxic glioma stem cell-derived exosomes containing Linc01060 promote progression of glioma by regulating the MZF1/c-Myc/HIF1alpha axis. Cancer Research, 81, 114–128. https://doi.org/10.1158/0008-5472.CAN-20-2270
Gong, W. et al. (2019). Analysis of survival-related lncRNA landscape identifies a role for LINC01537 in energy metabolism and lung cancer progression. Int J Mol Sci 20, https://doi.org/10.3390/ijms20153713.
Wu, H., et al. (2021). Survival-related lncRNA landscape analysis identifies LINC01614 as an oncogenic lncRNA in gastric cancer. Frontiers in Genetics, 12, 698947. https://doi.org/10.3389/fgene.2021.698947
Campbell, G. R., & Spector, S. A. (2019). DIABLO/SMAC mimetics selectively kill HIV-1-infected resting memory CD4(+) T cells: A potential role in a cure strategy for HIV-1 infection. Autophagy, 15, 744–746. https://doi.org/10.1080/15548627.2019.1569950
Ino, Y., et al. (2013). Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. British Journal of Cancer, 108, 914–923. https://doi.org/10.1038/bjc.2013.32
Vinuesa, C. G., Linterman, M. A., Yu, D., & MacLennan, I. C. (2016). Follicular helper T cells. Annual Review of Immunology, 34, 335–368. https://doi.org/10.1146/annurev-immunol-041015-055605
Wen, Y., et al. (2020). TNF-alpha in T lymphocytes attenuates renal injury and fibrosis during nephrotoxic nephritis. American Journal of Physiology. Renal Physiology, 318, F107–F116. https://doi.org/10.1152/ajprenal.00347.2019
Han, Y., et al. (2019). IL-38 Ameliorates skin inflammation and limits IL-17 Production from gammadelta T Cells. Cell Rep, 27(835–846), 835. https://doi.org/10.1016/j.celrep.2019.03.082
Wu, T., & Dai, Y. (2017). Tumor microenvironment and therapeutic response. Cancer Letters, 387, 61–68. https://doi.org/10.1016/j.canlet.2016.01.043
Author information
Authors and Affiliations
Contributions
Dr. CBX contributed to the study conception and design. Dr. ZWC and Dr. XMP contributed to the material preparation, data collection, and analysis. Dr. ML and Dr. GXC draft the manuscript. Dr. JXL and Dr. YJM revised the manuscript. Then Dr. YJM gave the final approval of the version to be submitted. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
It does not contain any studies with human or animal subjects.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, C., Chen, Z., Pan, X. et al. Construction of a Prognostic Evaluation Model for Stomach Adenocarcinoma on the Basis of Immune-Related lncRNAs. Appl Biochem Biotechnol 194, 6255–6269 (2022). https://doi.org/10.1007/s12010-022-04098-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-04098-x