Abstract
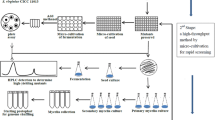
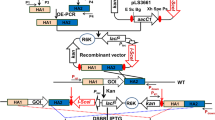
Genome shuffling is a powerful approach for efficiently engineering industrial microbial strains with interested phenotypes. Here we reported a high producer of nuclease P1, Penicillium citrinum G-16, that was bred by the classical physics-mutagenesis and genome shuffling process. The starting populations were generated by 60Co γ-irradiation mutagenesis. The derived two protoplast fractions were inactivated by heat-treatment and ultraviolet radiation respectively, then mixed together and subjected to recursive protoplast fusion. Three recombinants, E-16, F-71, and G-16, were roughly obtained from six cycles of genome shuffling. The activity of nuclease P1 by recombinant G-16 was improved up to 1,980.22 U4/ml in a 5-l fermentor, which was 4.7-fold higher than that of the starting strain. The sporulation of recombinant G-16 was distinguished from the starting strain. Random amplified polymorphic DNA assay revealed genotypic differences between the shuffled strains and the wild type strain. The close similarity among the high producers suggested that the genetic basis of high-yield strains was achieved by genome shuffling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nuclease P1 (E.C.3.1.30.1) is one member of zinc-dependent endonuclease family and produced by the mold Penicillium citrinum [1]. It contains three Zn(II) ions with two mononucleotide-binding sites [2]. The main function of this commercially available enzyme is to cleave single-stranded RNA and DNA into 5-mononucleotide, which is frequently used as an additive in food industry, an auxiliary therapy in hepatitis, nephritis, and muscle diseases [3], and agent in molecular biology research [4]. As a new biocatalyst for direct asymmetric aldol reaction under solvent-free conditions, the enzyme has been developed in biotransformation process [5].
To enhance the utilization efficiency of enzyme and production by P. citrinum, various processes have been investigated. DEAE cellulose [6] and marcoporous absorbent resin [7] as supports were evaluated for RNA hydrolysis in the continuous process. Gangadhara et al. [2] studied the effects of denaturants (GuHCl and urea) on nuclease P1 structure and activity, suggesting that the increased catalytic efficiency of nuclease P1 is due to a more open and flexible conformation of the activated enzyme. To the best of our knowledge, there were only a few reports on breeding of high nuclease P1 producers by P. citrinum. By the low-energy ion beam mutation treatment, strain N409 with the activity of nuclease P1 up to 421 U/ml was obtained [8]. Based on the means of mutations with 60Co γ-irradiation and nitrosoguanidine (NTG), the high-yield mutant HEP2312 (nuclease P1 at 1,508 U/ml), was screened [9]. Genome shuffling is a powerful approach for evolution of industrial strains with desirable phenotypes [10–12]. Recently, genome shuffling has been used to improve production of the low-temperature alkalophilic lipase by Acinetobacter johnsonii [13], xylanase activity by Apsergillus sp. NRCF5 [14], ε-poly-l-lysine production by Streptomyces graminearus [15], and 1,3-propanediol by Clostridium diolis DSM 15410 [16]. These reports demonstrated that genome shuffling for phenotypic improvement was efficient and effective [17].
The aim of this study is to breed high-yield producers of nuclease P1 by induction of mutations and genome shuffling. The protocols for isolating, regenerating protoplasts, and successive rounds of protoplast fusion in P. citrinum were described. A high producer of nuclease P1 (recombinant G-16) was obtained, and its productivity was confirmed in 5 liter fermentor.
Materials and Methods
Fungal Strains and Culture Conditions
Wild type of P. citrinum CCICC 4011 was obtained from China Center of Industrial Culture Collection (Peking, China), can produce nuclease P1 activity at 226 U/ml. The fungus was maintained on potato dextrose agar (PDA) medium at 28 °C for 3–5 days when spores were formed [8]. For nuclease P1 production, glucose and peptone (GP) medium (glucose 70 g, peptone 5 g, K2HPO4 · 3H2O 0.5 g, MgSO4 · 7H2O 0.4 g, ZnSO4 · 7H2O 0.4 g, CaCl2 0.4 g, pH 6.5, per liter deionized water) was prepared.
Nuclease P1 production process in a 5-l fermentor: a loop of PDA culture was inoculated into 100 ml of GP medium in a 500-ml Erlenmeyer flask. After incubation at 28 °C, 200 rpm for 30 h, 100 ml seed broth was transferred into a 3.5-l GP in a 5-l stirred-tank fermentor (B. Braun Inc., German) and incubated at 28 °C, 300 rpm, and 1.0 vvm aeration rate.
Preparation of Starting Strains for Genome Shuffling
The starting strains for genome shuffling were obtained through subtle improvements of the wild-type P. citrinum 4011 by 60Co γ-irradiation with 300 Gy dosage treatment [9, 18]. Briefly, 5 ml of spore suspension from a slant culture were transferred to an aseptic tube. 60Co γ-irradiation was used for mutation treatment. After appropriately diluting, the suspension of spores was spread on the PDA medium. A number of colonies were selected, and their nuclease P1 activity was evaluated by fermentation tests. The mutants with higher production were preserved and used as the starting strains for genome shuffling.
Protoplast Preparation and Inactivation Treatment
Protoplasts were prepared according to a previously reported method by a few modification [19]. Briefly, the uniform conidial suspensions of selected strains were inoculated in 300 ml Erlenmeyer flasks containing 30 ml PDA medium. The flasks were cultivated on a rotary shaker 100 rpm overnight at 28 °C. Mycelium was collected by filtratin through sterile Myracloth, rinsed with and resuspended in 0.6 M NaCl.
DL-dithiothreitol (DTT) solution was used to pretreat the pellets in 100 ml flask on a rotary shaker (100 rpm) for 30 min. Rinsed with 0.6 M NaCl to discard DTT, 250 mg (wet weight) of the pellets was resuspended in 10 ml of the solution of cell wall-lytic enzymes (cellulase, snailase, and lyticase, 2 mg/ml), cultured at 30 °C on a shaker incubator operating at 80 rpm in 50 ml Erlenmeyer flask until maximum protoplastation. Protoplasts were separated from the mixtures by filtration through sterile Myracloth, and the number of protoplasts in the filtered solutions was counted under a microscope using hemacytometer. After centrifuged at 800×g for 15 min in 0.6 M NaCl, protoplasts were washed and recovered.
To improve the efficiency of hybrid screening, parental starting protoplasts were applied to inactivation treatment, then the parental protoplasts cannot grow on regeneration plates. The equal number of protoplasts from starting populations was mixed and then divided equally into two aliquots [15]. One aliquot was inactivated by heat at 50 °C for 30 min, the other was treated by UV inactivation under a 30 W UV lamp for 15 min.
Genome Shuffling
Protoplasts were fused using a modification of the method of Lv et al. [18]. Aliquots of inactivated protoplasts were mixed in a ratio of 1:1, collected by centrifugation and then resuspended in PN solution (40 % PEG-6000 in 0.6 M NaCl buffer, pH 7.8). The mixture was incubated at 30 °C, 80 rpm for 30 min. The pellets were centrifuged, washed twice with 0.6 M NaCl buffer (pH 7.8), and resuspended in 0.6 M NaCl buffer (pH 7.8). It was serially diluted and regenerated on regeneration medium (PDA medium supplemented with 0.6 M NaCl as osmotic stabilizer) plates for 3–7 d at 28 °C. The colonies were isolated and the productivity was evaluated with fermentation test in shaking flasks. The selected high-yield fusants were used as the starting strains for subsequent rounds of genome shuffling. Six successive rounds of genome shuffling were performed. Stable segregants, suggested to be haploid, were obtained by growing the hybrids on PDA containing benlate at a concentration of 0.5 μg/ml. The segregants were checked for stability on fresh medium containing benlate and subsequently phenotypically characterized by replication onto appropriately PDA media [20].
Random Amplification Polymorphic DNA Analysis
Genomic DNA extraction and random amplified polymorphic DNA (RAPD) analysis were performed as described by Abe et al. [18, 21, 22], respectively. DNA was extracted from mycelia of P. citrinum. The mycelia were grinded to be powder under liquid nitrogen condition and suspended in liquid (0.1 M NaCl, 30 mM Tris–HCl pH 7.8, 10 mM EDTA). For the RAPD analysis, the primers shown in Table 1 were used. Amplification was achieved using a thermal cycler set at the following program: 94 °C for 4 min, followed by 40 cycles of 94 °C for 30 s, 34 °C for 1 min, and 72 °C for 2 min and extended by 72 °C for 5 min, with the reaction system: 5 μl 10 × PCR buffer, 3 μl MgCl2 (25 mM), 1 μl dNTP(10 mM), 2 μl primers, 2 μl DNA template, 1 μl Taq polymerase, and 36 μl sterile water. The products were separated by electrophoresis in 2.0 % agarose gels. The presence of amplified bands with different intensities and locations were detected and analyzed with the Quantity One 4.1 (BioRad, Hercules, CA, USA) software. The RAPD band patterns were analyzed by the unweighted pair group method using arithmetic average (UPGMA) method.
Nuclease P1 Assay
The activity of nuclease P1 was determined by the method of Ying et al. [23]. In brief, 1.9 ml substrate (containing 1 % RNA (w/v), 125 mM acetic acid buffer (pH 5.1), 1 mM Zn(II)) was heated at 67 °C in the constant temperature water-bath and keep warm, then 0.1 ml diluted enzyme was added and incubated for 15 min. The test tubes were transferred into ice water, 2.0 ml precooling of nucleic acid precipitant (0.25 % (NH4)6Mo7O24 · 4H2O–2.5 % HClO4 (w/v)) was added to stop the reaction, and kept in the ice bath for 20 min. The precipitated RNA was removed by centrifugation. The absorbance at 260 nm of the supernatant was recorded. The enzyme activity (U) was calculated as the below:
Where a is the dilution times of the enzyme before enzyme activity determination.
Statistical Analysis
All the data were subjected to statistical analysis by one-way ANOVA at p < 0.05 and p < 0.01.
Results
Strains Mutagenesis and Starting Strains Selection
The feature of natural evolution is practically mimicked by genome shuffling on recursive genetic recombination. The starting population with an improvement of the desired phenotype from the wild-type is required before genome shuffling. 60Co γ-irradiation was used as a mutagenic agent to improve the enzyme productivity of the wild-type strain. Mutant selection was based on the evaluation of flask tests. Six mutants with an improved activity, named as CM-11, CM-42, CM-33, CM-97, CM-158, and CM-227, were selected from a mutant library of 306 colonies respectively (Fig. 1). In the flask tests, the nuclease activity by mutant CM-11 was up to 420.22 U/ml and 0.86-fold increase over P. citrinum 4011 (nuclease activity 226 U/ml). Consequently, CM-11, CM-42, CM-33, CM-97, CM-158, and CM-227 were used as the starting population for genome shuffling.
Optimization of Protoplast Preparation and Regeneration Conditions
To conduct the protoplast fusion, various factors of protoplast formation and regeneration were investigated (Table 2). Mycelia pretreated with DTT for 45 min was helpful to the rate of protoplast formation by 54 %. Four osmotic stabilizers were tested for their efficacy in releasing protoplasts. When 0.6 M NaCl was used as osmotic stabilizer, the maximum numbers of protoplasts were obtained. For the release of protoplasts, three commercially available enzymes were investigated for their lytic activity against P. citrinum mycelia. Enzyme mixture consisting of cellulose/snailase/lyticase (in a ratio 2:1:1 for 140 min) were able to produce the maximum rate of protoplasts formation (Table 2 and Fig. 2). The process of protoplast formation from mycelia was clearly shown in Fig. 3.
Genome Shuffling of Improved Mutant Populations
Genome shuffling is depended on the high frequency of protoplast formation and fusion, while the recursive fusion of protoplasts that permits obtaining quickly the phenotype of interest is the basis of genome shuffling [24, 25]. To bleach parental populations from fusants, UV radiation and heat treatment were utilized [26]. After UV radiation for 15 min or heating at 55 °C for 30 min, the inactivation rate of protoplasts of P. citrinum was up to 100 % (Table 3). The temperature and time in the fusion process were further evaluated too. Good fusion rate would be produced at 30 °C for 15 min (Fig. 4).
Genome shuffling was applied to improve nuclease P1 production by the six mutants, CM-11, CM-42, CM-33, CM-97, CM-158, and CM-227, obtained by 60Co mutagenesis. After the first round of shuffling, 200 colonies were selected. Through the enzyme activity assay by flask tests, five recombinants, G1-7, G1-19, G1-163, G1-192, and G1-195, showed high enzyme activities and were pooled as starting populations for the next shuffling process. After six successive rounds of genome shuffling, from the 1183 shuffling colonies three recombinants (E-16, F-71, and G-16) were selected by their nuclease P1 activity (Fig. 1). The enzyme activity by recombinant G-16 was up to 1762.49 U/ml, 4.2 times higher than that of the starting strain CM-11 (420.22 U/ml). The genetic stability of G-16 was evaluated by five successive subcultivation tests on PDA containing benlate at a concentration of 0.5 μg/ml. The rang of production levels among five generations was 1,684.3 to 1,792.7 U/ml, indicating that the hereditary characters of the high nuclease P1 producing recombinant, P. citrinum G-16, was stable.
Morphological Characteristics of Parental and the Shuffled Strains
The size of mycelium was measured at the beginning of the cultivation on PDA agar. Figure 5 showed the comparison of the morphological characters of mycelia, pycnidia, and conidiospores development between the recombinant G-16 with the parental strain 4011. Cultured on PDA agar surface, the recombinant G-16 achieved a quick development process of mycelium into sporulation and produced a great deal of conidiospores (within 2 days, 28 °C). However, strain 4011 exhibited good growth of mycelia and little formation of conidiospore. This result suggested that improved spore productivity may be correlated with metabolite synthesis, such as nuclease P1.
Characterization of Nuclease P1 Production Profiles of the Shuffled Strain
Nuclease P1 production and cell growth of the recombinant G-16 was further examined in a 5 liter fermentor (Fig. 6). Consistent with the results obtained in flask tests, recombinant G-16 produced more than 4.7-fold nuclease P1 yield (1,980.22 U/ml), compared with that of the starting strain after 84 h. As shown in Fig. 6 nuclease P1 was produced along with exponential growth in the fermentation process. Sugar was consumed rapidly in the exponential growth phase. The mycelium growth was almost consistent with the sugar consumption and stop in the stationary phase. These results implied that there were two rapid growth phases, M1 (12–36 h) and M2 (48–72 h). Corresponded to M1 transition into M2 phases, the producing rate of nuclease P1 was changed to a higher level, which was consistent with the sporulation time (Fig. 5). Although mycelium biomass growth stop at 72 h, nuclease P1 production continued and reached the peak at 84 h (Fig. 6). The amino nitrogen was consumed fast in the first 24 h of cultivation, which corresponded with sugar consumption and pH decreased. After that, amino nitrogen concentration increased sharply with 50 h and then decreased slowly again.
Polymorphism of Genomic Population by RAPD Analysis
RAPD analysis was used to examine the genetic diversity among 6 shuffled recombinants with different enzyme productivity and the parental strain 4011. Six of the pre-screened primers (s199, s197, s10, s155, s104, and s156) successfully amplified polymorphic DNA bands. Amplified bands were characterized based on their size, which ranged from approximately 150 to 2,500 kb. RAPD band pattern was analyzed using the Nei similarity index, and a dendrogram was constructed based on the similarity matrix data by applying the UPGMA cluster analysis method (Fig. 7). The shuffled recombinants were segregated and high producers (E-16, F-71, and G-16) were clustered into a subgroup. Based on the dendrogram, close homology existed in those strains that the similarity coefficient of the seven strains varied from 0.58 to 0.77. The shuffled recombinant G-16 was the closest to F-71 (their genetic similarity coefficient was 0.77, while genetic distance was 0.185), but was distant to the low producer E-9 (their genetic similarity coefficient was 0.58, while genetic distance was 0.462). The specific bands and UPGMA dendrogram of cluster analysis obtained in our study showed that genetic information was transferred from the parental strains to the shuffled strains by genome shuffling and the genetic information of the shuffled strains had been changed.
Polymorphism of genomic population by RAPD analysis. a RAPD-PCR profiles of strains using primer s199 and s156. M is the DNA marker. Lanes 1–7 are the parent strain and the high-yielding isolates (from left to right: 4011, F-71, B-21, G-16, C-5, E-16, E-9) by using primer s199, while lanes 8–14 are the parent strain and the high-yielding isolates (from left to right: 4011, F-71, B-21, G-16, C-5, E-16, E-9) by using primer s156. b Dendrogram of the high/low nuclease P1-producing mutants based on UPGMA cluster analysis and similarity index
Discussion
Nuclease P1 is widely produced by various industrial process. Exploring more effective method to increase the enzyme production is the interest for its manufacture. In the present study, we successfully applied genome shuffling method to improve production of nuclease P1. After six successive rounds of genome shuffling, the recombinant G-16 with high production was picked up from 1183 shuffled isolates. The enzyme activity by recombinant G-16 was up to 1,762.49 U/ml, 4.2-fold higher than the starting strain CM-11 (420.22 U/ml; Fig. 1). This improvement was at similar level as that of previous report for S. fradiae (6-fold) [11], Lactobacillus (3-fold) [25], and Streptomyces sp. U121 (5-fold) [27]. In the 5-l fermentor, the enzyme production was increased further to 1,980.22 U/ml (Fig. 6). The production of G-16 was higher compared to other strain development efforts that have been reported [8, 9]. For example, Li et al. (2007) screened a nuclease P1 producer of P. citrinum by ultraviolet mutagenesis, its activity reached 1,508 U/ml [9]. Therefore, recombinant G-16 has a potential application in the industry.
One clue of the improved performance of recombinant G-16 was its differentiations in morphological and genetic characteristics compared to the starting strains (Figs. 5 and 7). Recombinant G-16 had a high productivity of conidiospores in 48 h growth, while the parental strain 4011 was sparse. It has been reported that the ability of producing enzyme is related directly to the ability of sporulation by microbe [28]. In Aspergillus niger, a predictable behavior fungal morphology was correlated closely with productivity [29]. In S. coelicolor, some associations between the sporulation and secondary synthesis were often observed [30]. Therefore, the colonies with abundant conidiospores would be hyper-producers of the enzyme.
Molecular genetic techniques, such as amplified fragment length polymorphism (AFLP) [31] or RAPD [21, 32], have become powerful tools to analyze genetic relationships and diversity. In this study, polymorphism of DNA RAPD was utilized to investigate the genetic variation and evolution between fusants and parental strains. As shown in Fig. 7, the higher producers were clustered together (E-16, F-71, and G-16) and were separated from those low-yield strains. Taken together with the results of morphological and metabolic characteristics, our results implied the production improvements of fusants are derived from genetic variation, not from modification. Hence we hypothesized that some unique DNA fragments correlated with hyper-yielding characteristics was common among the hyper-yielding shuffled strains.
Conclusion
In this study, we described a genome shuffling protocol for isolating, regenerating protoplasts, and successive rounds of protoplast fusion in P. citrinum. Our results demonstrated that rapid improvement of nuclease P1 activity was achieved by this method. The starting mutant population was generated by 60Co γ-irradiation treatments of the spores. After six rounds of protoplast fusion, an improved recombinant G-16 was obtained and its activity yield of nuclease P1 reached 1980.22 U/ml in a 5-l fermentor. The morphological and molecular difference between the parent and shuffled recombinants were observed, and the mechanism of the improved shuffled recombinants was discussed. Our results suggested that G-16 would be a promise candidate strain for industrial application, and genome shuffling is a powerful approach for molecular breeding of industrial strains.
References
Munetaka, S., Jun, I., Shigenmi, A., & Hiroo, F. (2000). Endonucleases. Plant Molecular Biology, 44, 387–397.
Gangadhara, B. N., Kumar, P. R., & Prakash, V. (2008). Enhancement of nuclease P1 activity in low concentration of denaturants. Enzyme and Microbial Technology, 43, 336–342.
Burdock, G. A., Flamm, W. G., & Carabin, I. G. (2000). Toxicity and mutagenicity studies of DN-50000® and RP-1® enzymes. Food and Chemical Toxicology, 38, 429–442.
Warnecke, J. M., Sontheimer, E. J., Piccirilli, J. A., & Hartmann, R. K. (2000). Active site constraints in the hydrolysis reaction catalyzed by bacterial RNase P: analysis of precursor tRNAs with a single 3′-S-phosphorothiolate internucleotide linkage. Nucleic Acids Research, 28, 720–727.
Li, H. H., He, Y. H., Yuan, Y., & Guan, Z. (2011). Nuclease p1: a new biocatalyst for direct asymmetric aldol reaction under solvent-free conditions. Green Chemistry, 13, 185–189.
Shi, L. E., Yi, Y., Tang, Z. X., Xiong, W. Y., Mei, J. F., & Ying, G. Q. (2010). Nuclease p1 immobilized on deae cellulose. Brazilian Journal of Chemical Engineering, 27, 31–39.
Li, B., Chen, Y., Chen, X., Liu, D., Niu, H., Xiong, J., et al. (2012). A novel immobilization method for nuclease P1 on macroporous absorbent resin with glutaraldehyde cross-linking and determination of its properties. Process Biochemistry, 47, 655–670.
He, Q. T., Li, N., Chen, X. C., Ye, Q., Bai, J. X., Xiong, J., et al. (2011). Mutation breeding of nuclease p1 production in Penicillium citrinum by low-energy ion beam implantation. Korean Journal of Chemical Engineering, 28, 544–549.
Li, K., Zeng, Q., & Xiang, Z. (2007). Selection of Biochemical Mutants that Overproduce Nuclease P1 and Optimization of the Fermentation Conditions. Journal of Yunnan Agricultural University, 6, 898–904.
Matthew, T. W., & Laurence, D. H. (2012). Direct and indirect consequences of meiotic recombination: implications for genome evolution. Trends in Genetics, 28, 101–109.
Zhang, Y. X., Perry, K., Vinci, V. A., Powell, K., Stemmer, W. P., & Del Cardayré, S. B. (2002). Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature, 415, 644–646.
Jin, Z. H., Xu, B., Lin, S. Z., Jin, Q. C., & Cen, P. L. (2009). Enhanced Production of Spinosad in Saccharopolyspora spinosa by Genome Shuffling. Applied Biochemistry and Biotechnology, 159, 655–663.
Wang, H. K., Zhang, J., & Wang, X. J. (2012). Genome shuffling improves production of the low-temperature alkalophilic lipase by Acinetobacter johnsonii. Biotechnology Letters, 34, 145–151.
El-Bondkly, A. M. A. (2012). Molecular identification using ITS sequences and genome shuffling to improve 2-deoxyglucose tolerance and xylanase activity of marine-derived fungus, Aspergillus sp. NRCF5. Applied Biochemistry and Biotechnology, 167, 2160–2173.
Li, S., Li, F., Chen, X. S., Wang, L., Xu, J., Tang, L., et al. (2012). Genome shuffling enhanced ε-poly-l-lysine production by improving glucose tolerance of Streptomyces graminearus. Applied Biochemistry and Biotechnology, 166, 414–423.
Burkhard, O., Eike, G., Osama, M., & Stefan, J. (2009). Genome shuffling in Clostridium diolis DSM 15410 for improved 1,3-propanediol production. Applied and Environmental Microbiology, 75, 7610–7616.
Gregory, S. (2002). Metabolic engineering by genome shuffling. Nature Biotechnology, 20, 666–668.
Lv, X. A., Jin, Y. Y., Li, Y. D., Zhang, H., & Liang, X. L. (2013). Genome shuffling of Streptomyces viridochromogenes for improved production of avilamycin. Applied Microbiology and Biotechnology, 97, 641–648.
Futoshi, N., Ichiro, W., & Nobufusa, S. (1993). Development of transformation system for the filamentous, ML-236B (compactin) - producing fungus Penicilliurn citrinum. Current Genetics, 23, 28–32.
Tahoun, M. K. (1993). Gene manipulation by protoplast fusion and penicillin production by Penicillium chrysogenum. Applied Biochemistry and Biotechnology, 39(40), 445–453.
Savitha, S., Sadhasivam, S., & Swaminathan, K. (2010). Regeneration and molecular characterization of an intergeneric hybrid between Graphium putredinis and Trichoderma harzianum by protoplasmic fusion. Biotechnology Advances, 28, 282–292.
Abe, Y., Baba, S., Suzuke, T., Ono, C., Iwamoto, K., & Hosobuchi, M. (2004). Molecular basis of ML-236B production in the high-producing mutant No. 41520 of Penicillium citrinum. Journal of General Applied Microbiology, 50, 169–176.
Ying, G. Q., Shi, L. E., Yua, Y., Tang, Z. X., & Chen, J. S. (2006). Production, purification and characterization of nuclease p1 from Penicillium citrinum. Process Biochemistry, 41, 1276–1281.
Patnaik, R., Louie, S., Gavrilovic, V., Perry, K., Stemmer, W. P. C., Ryan, C. M., et al. (2002). Genome shuffling of Lactobacillus for improved acid tolerance. Nature, 20, 707–712.
Yu, L., Pei, X., Lei, T., Wang, Y., & Feng, Y. (2008). Genome shuffling enhanced l-lactic acid production by improving glucose tolerance of Lactobacillus rhamnosus. Journal of Biotechnology, 134, 154–159.
Cheng, Y., Song, X., Qin, Y., & Qu, Y. (2009). Genome shuffling improves production of cellulase by Penicillium decumbens JU-A10. Journal of Applied Microbiology, 107, 1738–1846.
Hide, H., Yamada, T., & Yamada, Y. (2007). Genome shuffling of Streptomyces sp. U121 for improved production of hydroxycitic acid. Applied Microbiology and Biotechnology, 73, 1387–1393.
Wucherpfennig, T., Kiep, K. A., Driouch, H., Wittmann, C., & Krull, R. (2010). Chapter 4 - morphology and rheology in filamentous cultivations. Advances in Applied Microbiology, 73, 89–136.
Wucherpfennig, T., Hestler, T., & Krull, R. (2001). Morphology engineering - osmolality and its effect on Aspergillus niger morphology and productivity. Microbial Cell Factories, 10, 58.
Angel, M., & Ruben, A. (2008). Mycelium differentiation and antibiotic production in submerged cultures of Streptomyces coelicolor. Applied and Environmental Microbiology, 12, 3877–3886.
Xu, B., Jin, Z. H., Wang, H. Z., Jin, Q. C., Jin, X., & Cen, P. L. (2008). Evolution of Streptomyces pristinaespiralis for resistance and production of pristinamycin by genome shuffling. Applied Microbiology and Biotechnology, 80, 261–267.
El-Gendy, M. M., & EL-Bondkly, A. M. (2011). Genome shuffling of marine derived bacterium Nocardia sp. ALAA 2000 for improved ayamycin production. Antonie Van Leeuwenhoek, 99, 773–780.
Acknowledgments
The work was financially supported by the National Nature and Science Foundation of China (3117175) and Nature and Science Foundation of Zhejiang Province (Y3100609).
Conflict of interests
The authors declare that they have no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, C., Wu, G., Li, Y. et al. Genome Shuffling of Penicillium citrinum for Enhanced Production of Nuclease P1. Appl Biochem Biotechnol 170, 1533–1545 (2013). https://doi.org/10.1007/s12010-013-0297-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-013-0297-9