Opinion statement
Leukemia is the most common pediatric cancer and accounts for approximately one third of childhood malignancies. There are germline genetic alterations that significantly increase the risk of developing hematopoietic malignancies in childhood. In this review, we describe a number of these hereditary disorders and their clinical features. These predispositions to cancer syndromes can be attributed to DNA repair/genetic instability, RAS pathway dysfunction, bone marrow failure, telomeropathies, immunodeficiencies, transcription factor abnormalities, pure familial leukemia, and aneuploidy. We focus especially on acute myeloid leukemia associated with Down syndrome, but also include other hereditary syndromes in this review. Recent advances in high-throughput genotyping technology have identified new genetic variations prone to human leukemia. Understanding of the mechanism of leukemia development in these hereditary syndromes allows us to gain insight into leukemogenesis in general and suggests therapeutic strategies based on these findings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute leukemia is the most common pediatric cancer and accounts for approximately one third of childhood cancers, and roughly 80% are lymphoid (acute lymphoblastic leukemia (ALL)) and 20% are myeloid (acute myeloid leukemia (AML)) [1]. While the majority of childhood leukemia cases occur in the absence of any known predisposing factor, a small proportion are truly familial or caused by known hereditary cancer syndromes [2]. Both ALL and AML can be seen with a variety of hereditary cancer syndromes. According to the magnificent past review articles, the categories of leukemia-associated hereditary cancer syndrome can be divided into some groups in several ways [3,4,5,6]. Moreover, recent advances in high-throughput genotyping technology enable comprehensive screens of genetic variation, revealing new genetic variations which are prone to develop leukemia [7]. This review focuses on describing the clinical features of multiple germline syndromes that confer an increased risk of leukemia, and will specifically highlight the process of leukemia development in Down syndrome.
Hereditary cancer syndrome associated with leukemia
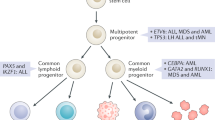
Inherited cancer syndromes associated with leukemia like familial myelodysplastic syndromes (MDSs) and AML syndromes were once considered rare, but are more frequent than previously expected. The number of genes involved in inherited MDS/AML has grown recently with the advance of genomic sequencing technologies, and the known hereditary cancer syndromes associated with leukemia risk account for more than 60 different genes. Each syndrome has a different risk for ALL or AML, sometimes bearing other hematological abnormalities [6]. Table 1 is a modified comprehensive list of the leukemia-associated syndromes from the excellent reviews by Seif [2], Malkin [3], and Stieglitz [4]. To discuss the leukemia-associated inherited cancer syndromes, we divide these syndromes into the following eight main categories based on clinical features, biological functions, and affected pathways: (1) genetic instability/DNA repair syndromes, (2) RAS pathway dysfunction, (3) bone marrow failure syndromes, (4) telomeropathies, (5) immunodeficiency, (6) transcription factor abnormalities, (7) pure familial leukemia, and (8) aneuploidy. Although some of these syndromes belong to multiple categories, we have classified each syndrome to the most representative category by its clinical features. For the purpose of this review, we will briefly describe each category, pick up some hereditary syndromes, and discuss about the process of leukemia development in some of these hereditary cancer syndromes.
DNA repair/genetic instability
Genomic instability is a characteristic of most cancers and leukemia [29]. In hereditary cancer syndromes, genomic instability results from mutations in DNA repair genes and drives malignancy development. A most representative syndrome in this category is Li-Fraumeni syndrome (LFS) [30, 31], caused by p53 mutation [32, 33]. The p53 protein is a transcription factor, upregulating the transcription of target genes involved in cell cycle arrest, DNA repair, apoptosis, and senescence, in response to DNA damage. Mutations in p53 can accumulate additional genetic mutations in hematopoietic progenitor cells. Patients with LFS develop multiple tumors, and also develop leukemia (mostly ALL, and to a lesser extent AML) in 4% of affected mutation carriers in an excellent past review which also found a median age of onset of 12 years [8•]. The newly described constitutional mismatch repair-deficiency syndrome (CMMRD) is caused by bi-allelic (homozygous) alterations in the mismatch repair (MMR) genes, including MLH1, MSH2, MSH6, and PMS2. Some of the manifestations of the syndrome resemble those seen in neurofibromatosis type I, and are associated with multiple café-au-lait spots, pediatric brain tumors, colorectal cancers, and pediatric hematologic malignancies including both ALL and AML [9]. A third of the patients develop hematological malignancies in CMMRD [34]. These MMR gene mutations also cause the adult-onset autosomal dominant Lynch syndrome, previously referred to as hereditary non-polyposis colorectal cancer (HNPCC) [35]. The category of DNA repair/genetic instability also includes Werner syndrome (caused by WRN), Rothmund-Thomson syndrome (RECQL4), Bloom syndrome (BLM), Fanconi anemia (FANC genes), ataxia telangiectasia (ATM), and Nijmegen breakage syndrome (NBS1). Fanconi anemia (FA) is an autosomal recessive (AR) disorder that leads to increased chromosomal breakage through defects in DNA repair [36]. Most FA patients exhibit developmental abnormalities, developing bone marrow failure, AML, and solid malignancies. Bone marrow failure often occurs in childhood, more than a third will develop leukemia, and nearly half will develop MDS [13]. In recent decades, 19 human genes have been found in the cause of FA [14]. These genes code for a group of associated FA, which function cooperatively in a DNA damage recognition and repair. Dysfunction of these protein leads to genomic instability and results in development of MDS and AML.
RAS pathway dysfunction
This category of hereditary syndromes includes Noonan syndrome (PTPN11, K-RAS, N-RAS, etc.), CBL syndrome (CBL), and nurofibromatosis type 1 (NF1). Cancer and leukemia are diseases of uncontrolled cell division and usually linked to a series of dysfunctions in the activity of cell cycle regulators. RAS signaling affects many cellular functions, which includes cell proliferation and differentiation in both normal and malignant cells [37]. Juvenile myelomonocytic leukemia (JMML) is a unique hematopoietic disorder of infancy caused by excessive proliferation of cells of mono and granulocytic lineages [38]. Approximately 90% of patients with JMML carry either somatic or germline mutations of PTPN-11, K-RAS, N-RAS, CBL, or NF1 in their leukemic cells. Allogeneic hematopoietic stem cell transplantation (HSCT) remains the therapy of choice for most patients with JMML. A recent report recommends that HSCT be promptly offered to any child with PTPN-11, K-RAS, or NF1-mutated JMML and to the majority of those with N-RAS mutations but not CBL mutations [39]. Because JMML patients with CBL mutations and few of those with N-RAS mutations may have spontaneous resolution of hematologic abnormalities, the decision to proceed to HSCT in these patients must be weighed carefully.
Bone marrow failure
This category contains a variety of clinical syndromes with many different mutated genes, including Diamond Blackfan anemia (DBA), Shwachman-Diamond syndrome (SDS), familial aplastic anemia with SRP72 mutation, congenital amegakaryocytic thrombocytopenia (CAMT), thrombocytopenia and absent radii (TAR) syndrome, thrombocytopenia with ANKRD26 mutation, and Kostmann syndrome. FA also can be included this category. These bone marrow failure syndromes are considered as a risk factor for clonal evolution [40]. In these syndromes, MDS and AML seem to be the most common hematological malignancies with risks ranging from 5 to 25% [18, 41,42,43,44,45,46]. Careful observation of any leukemia-related signs or symptoms has been offered as management for these syndromes. Recently, RPS29, RPS27, and RPL27 genes are reported to be corresponding for DBA by whole exome sequencing [47, 48].
Telomeropathies and immunodeficiency
Table 1 shows a list of telomeropathies and immunodeficiency syndromes associated with leukemia risk. Dyskeratosis congenita (DC) is the representative disease of telomeropathies caused by dysfunction in TERT and TERC, leading to abnormal telomere maintenance [20]. Patients with DC are at increased risk for bone marrow failure (BMF), MDS or AML, solid tumors (usually squamous cell carcinoma of the head/neck or anogenital cancer), and pulmonary fibrosis. To date, ACD, CTC1, DKC1, NHP2, NOP10, PARN, RTEL1, TERC, TERT, TINF2, and WRAP53 are the genes in which pathogenic variants are known to cause DC. HSCT is the only curative treatment for hematological malignancies but has had poor long-term efficacy in DC patients, because of their high incidence of treatment-related toxicities. Therefore, reduced-intensity preparative regimens being studied in a few institutions may improve long-term outcomes [49].
Transcription factors
Hereditary syndromes in this category are caused by mutations in genes that code for hematopoeitic transcription factors. Alterations in the associated genes (RUNX1, GATA2, PAX5, SH2B3, and ETV6) are also known to occur in de novo or sporadic leukemia cells. Inherited or sporadic GATA2 gene mutations induce MonoMAC syndrome (monocytopenia with atypical mycobacterial infection), DCML deficiency (loss of dendritic cells, monocytes, and natural killer and B lymphoid cells), Emberger syndrome (lymphedema with MDS), and familial AML/MDS [22•]. Familial AML/MDS caused by GATA2 deficiency is categorized the next group “pure familial MDS/AML” because of without other clinical symptoms. Various types of heterozygous mutations, including substitutions, indel, and deletions, have been detected, scattering among five translated exons of the GATA2 gene [22•]. As a result of these alterations, either a mutant GATA2 protein or no GATA2 protein is produced. Reduced expression of the GATA2 gene in HSCs caused by a heterozygous mutation/deletion appears to play a role in the onset of GATA2-associated hematologic disease [50]. More than half of patients with GATA2 deficiency develop MDS, most of which acquired unfavorable chromosomal abnormalities, and finally progressed to AML [22•]. Recently, we reported a family of GATA2 deficiency treated by reduced intensity stem cell transplantation (RIST) [51]. Incidence of MDS/AML gradually increases with age in GATA2 deficiency, and AML with GATA2 deficiency results in poor outcome unless successfully transplanted. Allowing for these clinical findings, RIST could be a promising treatment option for patients with GATA2 deficiency before progression of advanced MDS and AML.
Other pure familial MDS/AML
We defined familial MDS/AML without other systemic clinical features as “pure familial MDS/AML” in this review. The first genes involved in this category, CEBPA, were identified a decade ago [23]. There have been more genes in this category, exemplified by multiple autosomal dominant, highly penetrant inherited cancer syndromes resulting from germline mutations in GATA2, DDX41 ATG2B, and GSKIP [22•, 24, 52]. Most of these syndromes are inherited in an autosomal dominant manner and maybe grouped by their clinical presentations. Familial mosaic monosomy 7 is also included in this category. Of note, individuals with a family history of monosomy 7 may initially have a normal karyotype in peripheral blood and/or bone marrow and later transition to mosaic monosomy 7 [25].
Aneuploidy
Acquired somatic chromosomal aneuploidies are the most common genetic aberrations in sporadic leukemia. Constitutional aneuploidies also associate with the risk for specific hematological malignancies. Trisomy 21 and trisomy 8 are representatives of constitutional aneuploidies of hereditary cancer syndrome associated with leukemia. Constitutional trisomy 8 is lesser than trisomy 21 [53,54,55], and can be seen mostly as a mosaic in the blood or the skin [28]. Meanwhile, trisomy 8 is the most common among sole cytogenetic abnormalities in both AML and MDS with respective incidences of 6 and 11% [56]; it is sometimes difficult to distinguish constitutional or not [57].
Down syndrome (DS) results from trisomy 21 and occurs in 1 in 700–1000 births [54, 55]. Patients with DS show a 10- to 20-fold higher risk of acute leukemia [58, 59]. The most marked increase in incidence in DS infants is with acute megakaryoblastic leukemia (AMKL). The relative risk of developing AMKL is estimated to be 500 times higher in children with DS than in the general population [60]. In most cases, DS-AMKL is preceded by a temporary form of megakaryoblastic leukemia unique to newborns with DS and known as transient abnormal myelopoiesis (TAM). TAM presents in the fetus or a few days after birth, and in most cases resolves spontaneously within 3 months [61, 62]. After spontaneous remission, 20% of TAM patients develop MDS and DS-AMKL within 4 years [60, 63].
Multistep leukemogenesis in DS-related myeloid disorders
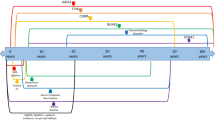
TAM has been considered a preleukemic state and is a suitable pathological condition to analyze the evolutionary process of leukemia. GATA1 is a hematopoietic transcription factor, and GATA1 somatic mutations cause both TAM and subsequent DS-AMKL [64,65,66,67,68]. GATA1 mutation has been reported to be ∼30% of neonates of DS [61, 69], and disappear when TAM/ML-DS enters the remission phase [64]. Although TAM is most commonly seen in infants with DS but barely in those with non-DS, suggesting trisomy 21 also a requirement for development of TAM [70]. Recently, next-generation sequencing (NGS) methodology showed the precise frequency of GATA1 mutation in DS [69]. Besides, NGS studies has also showed that the additional genomic mutations/deletions in the genes coding cohesion components, CTCF, and other epigenetic regulators including EZH2 frequently preceded other driver mutations during progression form TAM to DS-AMKL [26••, 71]. These findings of mutations in epigenetic regulators in DS-AMKL suggest that epigenetics also play a role in the development of DS-AMKL. Mutations are also observed in members of signaling pathways, such as the JAK family of kinases, MPL, SH2B3, and multiple RAS pathway genes [26••, 71]. Importantly, these genomic analyses of DS-AMKL confirm that it evolves from the cells responsible for TAM. Collectively, in the setting of trisomy 21, GATA1 mutations cause TAM (Fig. 1). Although most TAM resolves spontaneously, further mutation in cohesin, CTCF, EZH2, or other epigenetic regulators in residual TAM cells occurs with or without mutation of signal-transducing molecules, leading to AMKL. Considering the age of leukemia onset ranges from childhood to late adulthood in other hereditary leukemia syndromes (often higher than DS-AMKL) [8•, 18, 22•, 23, 72], the fact that most of all DS-AMKL patients develop leukemia within 4 years after birth may indicate that trisomy 21 is a driving event in leukemogenesis by itself [60, 63]. Actually, gain or amplification of chromosome 21 is sometimes observed in poor prognostic childhood leukemia [73, 74], and there are several candidate genes likely to contribute leukemogenesis in chromosome 21 such as RUNX1, APP, ETS2, Dyrk1a, and ERG [75, 76]. Our experimental TAM xenograft model has revealed the existence of multiple genetically distinct subclones in TAM phase [77]. It also enabled the observation of clonal selection and expansion of minor mutant TAM clones, demonstrating the striking genetic heterogeneity and the propagating potential of minor clones in a preleukemic phase [77].
Proposed model of ML-DS pathogenesis. Reprinted from Saida S: Evolution of myeloid leukemia in children with Down syndrome. Int J Hematol 2016, 103:365–372, with permission from the Japanese Society of Hematology. Trisomy 21. vertical lines; GATA1 mutation, black and white triangles; cohesin, CTCF, and other epigenetic regulator mutation, black circle; kinase-signaling molecule mutation, black star.
Conclusions and future directions
Recently, multiple hereditary predisposition syndromes to leukemia have been discovered owing to progression of new genetic technologies including next-generation sequencing, and more genes are likely to be identified in the future. Hereby, patients with these hereditary syndromes are likely not as rare as previously considered. Most of these patients with leukemia need to be treated chemotherapy and/or allogeneic HSCT, and they also need to be accommodated with their own hereditary syndromes because of their fragileness. These issues become especially important when planning a HSCT, because the patients with some of these syndromes are also at increased risk of developing complications by treatments. These genetic abnormalities are also seen as somatic mutations frequently in MDS/AML cells. Therefore, it is important to discriminate between somatic mutations and germline mutations when evaluating an individual for the syndromes associated with germline alterations in these genes. For that reason, physicians should be familiar with these hereditary predisposition syndromes and obtain every clinical symptom and detailed family histories for patients. The identification of these syndromes is a critical step toward individualized follow-up and treatment. The insight provided by these leukemia-associated syndromes will also have diagnostic and possibly therapeutic implication for all patients diagnosed with sporadic leukemia or other hematologic malignancies. Recent advances in genetic research will much more contribute to diagnosis and identification of causative genes of these syndromes [78].
The pathogenesis for leukemia in these syndromes is now beginning to be understood in terms of accumulation of genomic alterations. There would be different mechanisms between each syndrome in the course of leukemia development. In these hereditary syndromes, especially DS-associated myeloid malignancies (TAM/DS-AMKL) are an attractive model to investigate multistep leukemogenesis. The recent findings of mutations in epigenetic regulators in DS-AMKL suggest that epigenetics also play a role in the development of leukemia [27]. Our ongoing research aims to unravel the molecular mechanisms by which these mutations lead to malignancy.
References and Recommended Reading
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Institute NC: SEER (Surveillance, Epidemiology, and End Results) Cancer statistics review, 1975–2007. 2010.
Seif AE. Pediatric leukemia predisposition syndromes: clues to understanding leukemogenesis. Cancer Genet. 2011;204:227–44.
Malkin D, Nichols KE, Zelley K, Schiffman JD. Predisposition to pediatric and hematologic cancers: a moving target. Am Soc Clin Oncol Educ Book. 2014:e44–55.
Stieglitz E, Loh ML. Genetic predispositions to childhood leukemia. Ther Adv Hematol. 2013;4:270–90.
Czuchlewski DR, Peterson LC. Myeloid neoplasms with germline predisposition: a new provisional entity within the World Health Organization classification. Surg Pathol Clin. 2016;9:165–76.
Kratz CP, Stanulla M, Cave H. Genetic predisposition to acute lymphoblastic leukemia: overview on behalf of the I-BFM ALL host genetic variation working group. Eur J Med Genet. 2016;59:111–5.
Moriyama T, Relling MV, Yang JJ. Inherited genetic variation in childhood acute lymphoblastic leukemia. Blood. 2015;125:3988–95.
• Bougeard G, Renaux-Petel M, Flaman JM, Charbonnier C, Fermey P, Belotti M, Gauthier-Villars M, Stoppa-Lyonnet D, Consolino E, Brugieres L, et al. Revisiting Li-Fraumeni syndrome from TP53 mutation carriers. J Clin Oncol. 2015;33:2345–52. Large clinical study of “Li-Fraumeni syndrome”
Wimmer K, Kratz CP. Constitutional mismatch repair-deficiency syndrome. Haematologica. 2010;95:699–701.
Lauper JM, Krause A, Vaughan TL, Monnat RJ Jr. Spectrum and risk of neoplasia in Werner syndrome: a systematic review. PLoS One. 2013;8:e59709.
Stinco G, Governatori G, Mattighello P, Patrone P. Multiple cutaneous neoplasms in a patient with Rothmund-Thomson syndrome: case report and published work review. J Dermatol. 2008;35:154–61.
German J. Bloom’s syndrome. XX. The first 100 cancers. Cancer Genet Cytogenet. 1997;93:100–6.
Alter BP. Cancer in Fanconi anemia, 1927–2001. Cancer. 2003;97:425–40.
Dong H, Nebert DW, Bruford EA, Thompson DC, Joenje H, Vasiliou V. Update of the human and mouse Fanconi anemia genes. Human Genomics. 2015;9:32.
Rosenberg PS, Greene MH, Alter BP. Cancer incidence in persons with Fanconi anemia. Blood. 2003;101:822–6.
Taylor AM, Metcalfe JA, Thick J, Mak YF. Leukemia and lymphoma in ataxia telangiectasia. Blood. 1996;87:423–38.
Stiller CA, Chessells JM, Fitchett M. Neurofibromatosis and childhood leukaemia/lymphoma: a population-based UKCCSG study. Br J Cancer. 1994;70:969–72.
Vlachos A, Rosenberg PS, Atsidaftos E, Alter BP, Lipton JM. Incidence of neoplasia in diamond Blackfan anemia: a report from the diamond Blackfan anemia registry. Blood. 2012;119:3815–9.
Noris P, Favier R, Alessi MC, Geddis AE, Kunishima S, Heller PG, Giordano P, Niederhoffer KY, Bussel JB, Podda GM, et al. ANKRD26-related thrombocytopenia and myeloid malignancies. Blood. 2013;122:1987–9.
Savage SA. Dyskeratosis congenita. In GeneReviews(R). Edited by Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, Bird TD, Ledbetter N, Mefford HC, Smith RJH, et al.: University of Washington, Seattle. GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved.; 1993.
Godley LA. Inherited predisposition to acute myeloid leukemia. Semin Hematol. 2014;51:306–21.
• Spinner MA, Sanchez LA, Hsu AP, Shaw PA, Zerbe CS, Calvo KR, Arthur DC, Gu W, Gould CM, Brewer CC, et al. GATA2 deficiency: a protean disorder of hematopoiesis, lymphatics, and immunity. Blood. 2014;123:809–21. Excellent review article of “GATA2 deficiency”
Smith ML, Cavenagh JD, Lister TA, Fitzgibbon J. Mutation of CEBPA in familial acute myeloid leukemia. N Engl J Med. 2004;351:2403–7.
Saliba J, Saint-Martin C, Di Stefano A, Lenglet G, Marty C, Keren B, Pasquier F, Valle VD, Secardin L, Leroy G, et al. Germline duplication of ATG2B and GSKIP predisposes to familial myeloid malignancies. Nat Genet. 2015;47:1131–40.
Morrissette JJD, Wertheim G, Olson T. Familial monosomy 7 syndrome. In GeneReviews(R). Edited by Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, Bird TD, Ledbetter N, Mefford HC, Smith RJH, et al.: University of Washington, Seattle. GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved.; 1993.
•• Yoshida K, Toki T, Okuno Y, Kanezaki R, Shiraishi Y, Sato-Otsubo A, Sanada M, Park MJ, Terui K, Suzuki H, et al. The landscape of somatic mutations in Down syndrome-related myeloid disorders. Nat Genet. 2013;45:1293–9. Comprehensive genetic study first reporting the mutations of epigenetic factor in “Down syndrome related myeoid disorders”
Saida S. Evolution of myeloid leukemia in children with Down syndrome. Int J Hematol. 2016;103:365–72.
Hasle H, Clausen N, Pedersen B, Bendix-Hansen K. Myelodysplastic syndrome in a child with constitutional trisomy 8 mosaicism and normal phenotype. Cancer Genet Cytogenet. 1995;79:79–81.
Negrini S, Gorgoulis VG, Halazonetis TD. Genomic instability—an evolving hallmark of cancer. Nat Rev Mol Cell Biol. 2010;11:220–8.
Li FP, Fraumeni JF Jr. Soft-tissue sarcomas, breast cancer, and other neoplasms. A familial syndrome? Ann Intern Med. 1969;71:747–52.
Li FP, Fraumeni JF Jr, Mulvihill JJ, Blattner WA, Dreyfus MG, Tucker MA, Miller RW. A cancer family syndrome in twenty-four kindreds. Cancer Res. 1988;48:5358–62.
Malkin D, Li FP, Strong LC, Fraumeni JF Jr, Nelson CE, Kim DH, Kassel J, Gryka MA, Bischoff FZ, Tainsky MA, et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science. 1990;250:1233–8.
Srivastava S, Zou ZQ, Pirollo K, Blattner W, Chang EH. Germ-line transmission of a mutated p53 gene in a cancer-prone family with Li-Fraumeni syndrome. Nature. 1990;348:747–9.
Vasen HF, Ghorbanoghli Z, Bourdeaut F, Cabaret O, Caron O, Duval A, Entz-Werle N, Goldberg Y, Ilencikova D, Kratz CP, et al. Guidelines for surveillance of individuals with constitutional mismatch repair-deficiency proposed by the European consortium “care for CMMR-D” (C4CMMR-D). J Med Genet. 2014;51:283–93.
Sijmons RH, Hofstra RM. Review: clinical aspects of hereditary DNA mismatch repair gene mutations. DNA Repair (Amst). 2016;38:155–62.
Mehta PA, Tolar J. Fanconi anemia. In GeneReviews(R). Edited by Pagon RA, Adam MP, Ardinger HH, Wallace SE, Amemiya A, Bean LJH, Bird TD, Ledbetter N, Mefford HC, Smith RJH, et al.: University of Washington, Seattle. GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved.; 1993.
Downward J. Targeting RAS signalling pathways in cancer therapy. Nat Rev Cancer. 2003;3:11–22.
Niemeyer CM. RAS diseases in children. Haematologica. 2014;99:1653–62.
Locatelli F, Niemeyer CM. How I treat juvenile myelomonocytic leukemia. Blood. 2015;125:1083–90.
Bagby GC, Meyers G. Bone marrow failure as a risk factor for clonal evolution: prospects for leukemia prevention. Hematology Am Soc Hematol Educ Program. 2007:40–6.
Dror Y, Freedman MH. Shwachman-diamond syndrome. Br J Haematol. 2002;118:701–13.
Freedman MH, Bonilla MA, Fier C, Bolyard AA, Scarlata D, Boxer LA, Brown S, Cham B, Kannourakis G, Kinsey SE, et al. Myelodysplasia syndrome and acute myeloid leukemia in patients with congenital neutropenia receiving G-CSF therapy. Blood. 2000;96:429–36.
Rosenberg PS, Alter BP, Bolyard AA, Bonilla MA, Boxer LA, Cham B, Fier C, Freedman M, Kannourakis G, Kinsey S, et al. The incidence of leukemia and mortality from sepsis in patients with severe congenital neutropenia receiving long-term G-CSF therapy. Blood. 2006;107:4628–35.
Rosenberg PS, Alter BP, Link DC, Stein S, Rodger E, Bolyard AA, Aprikyan AA, Bonilla MA, Dror Y, Kannourakis G, et al. Neutrophil elastase mutations and risk of leukaemia in severe congenital neutropenia. Br J Haematol. 2008;140:210–3.
Ballmaier M, Germeshausen M. Congenital amegakaryocytic thrombocytopenia: clinical presentation, diagnosis, and treatment. Semin Thromb Hemost. 2011;37:673–81.
Alter BP, Giri N, Savage SA, Rosenberg PS. Cancer in the National Cancer Institute inherited bone marrow failure syndrome cohort after 15 years of follow-up. Blood. 2016;128:334.
Mirabello L, Macari ER, Jessop L, Ellis SR, Myers T, Giri N, Taylor AM, McGrath KE, Humphries JM, Ballew BJ, et al. Whole-exome sequencing and functional studies identify RPS29 as a novel gene mutated in multicase diamond-Blackfan anemia families. Blood. 2014;124:24–32.
Wang R, Yoshida K, Toki T, Sawada T, Uechi T, Okuno Y, Sato-Otsubo A, Kudo K, Kamimaki I, Kanezaki R, et al. Loss of function mutations in RPL27 and RPS27 identified by whole-exome sequencing in diamond-Blackfan anaemia. Br J Haematol. 2015;168:854–64.
Dietz AC, Orchard PJ, Baker KS, Giller RH, Savage SA, Alter BP, Tolar J. Disease-specific hematopoietic cell transplantation: nonmyeloablative conditioning regimen for dyskeratosis congenita. Bone Marrow Transplant. 2011;46:98–104.
Shimizu R, Yamamoto M. GATA-related hematologic disorders. Exp Hematol. 2016;44:696–705.
Saida S, Umeda K, Yasumi T, Matsumoto A, Kato I, Hiramatsu H, Ohara O, Heike T, Adachi S. Successful reduced-intensity stem cell transplantation for GATA2 deficiency before progression of advanced MDS. Pediatr Transplant. 2016;20:333–6.
Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, Singh J, Padgett RA, Gu X, Phillips JG, et al. Inherited and somatic defects in DDX41 in myeloid neoplasms. Cancer Cell. 2015;27:658–70.
Seghezzi L, Maserati E, Minelli A, Dellavecchia C, Addis P, Locatelli F, Angioni A, Balloni P, Miano C, Cavalli P, et al. Constitutional trisomy 8 as first mutation in multistep carcinogenesis: clinical, cytogenetic, and molecular data on three cases. Genes Chromosomes Cancer. 1996;17:94–101.
Parker SE, Mai CT, Canfield MA, Rickard R, Wang Y, Meyer RE, Anderson P, Mason CA, Collins JS, Kirby RS, et al. Updated National Birth Prevalence estimates for selected birth defects in the United States, 2004–2006. Birth Defects Res A Clin Mol Teratol. 2010;88:1008–16.
Roizen NJ, Patterson D. Down’s syndrome. Lancet. 2003;361:1281–9.
Paulsson K, Johansson B. Trisomy 8 as the sole chromosomal aberration in acute myeloid leukemia and myelodysplastic syndromes. Pathol Biol (Paris). 2007;55:37–48.
Saumell S, Sole F, Arenillas L, Montoro J, Valcarcel D, Pedro C, Sanzo C, Luno E, Gimenez T, Arnan M, et al. Trisomy 8, a cytogenetic abnormality in myelodysplastic syndromes, is constitutional or not? PLoS One. 2015;10:e0129375.
Fong CT, Brodeur GM. Down’s syndrome and leukemia: epidemiology, genetics, cytogenetics and mechanisms of leukemogenesis. Cancer Genet Cytogenet. 1987;28:55–76.
Hasle H, Clemmensen IH, Mikkelsen M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet. 2000;355:165–9.
Zipursky A, Poon A, Doyle J. Leukemia in Down syndrome: a review. Pediatr Hematol Oncol. 1992;9:139–49.
Pine SR, Guo Q, Yin C, Jayabose S, Druschel CM, Sandoval C. Incidence and clinical implications of GATA1 mutations in newborns with Down syndrome. Blood. 2007;110:2128–31.
Massey GV, Zipursky A, Chang MN, Doyle JJ, Nasim S, Taub JW, Ravindranath Y, Dahl G, Weinstein HJ. A prospective study of the natural history of transient leukemia (TL) in neonates with Down syndrome (DS): Children’s oncology group (COG) study POG-9481. Blood. 2006;107:4606–13.
Hitzler JK. Acute megakaryoblastic leukemia in Down syndrome. Pediatr Blood Cancer. 2007;49:1066–9.
Ahmed M, Sternberg A, Hall G, Thomas A, Smith O, O’Marcaigh A, Wynn R, Stevens R, Addison M, King D, et al. Natural history of GATA1 mutations in Down syndrome. Blood. 2004;103:2480–9.
Groet J, McElwaine S, Spinelli M, Rinaldi A, Burtscher I, Mulligan C, Mensah A, Cavani S, Dagna-Bricarelli F, Basso G, et al. Acquired mutations in GATA1 in neonates with Down’s syndrome with transient myeloid disorder. Lancet. 2003;361:1617–20.
Wechsler J, Greene M, McDevitt MA, Anastasi J, Karp JE, Le Beau MM, Crispino JD. Acquired mutations in GATA1 in the megakaryoblastic leukemia of Down syndrome. Nat Genet. 2002;32:148–52.
Rainis L, Bercovich D, Strehl S, Teigler-Schlegel A, Stark B, Trka J, Amariglio N, Biondi A, Muler I, Rechavi G, et al. Mutations in exon 2 of GATA1 are early events in megakaryocytic malignancies associated with trisomy 21. Blood. 2003;102:981–6.
Xu G, Nagano M, Kanezaki R, Toki T, Hayashi Y, Taketani T, Taki T, Mitui T, Koike K, Kato K, et al. Frequent mutations in the GATA-1 gene in the transient myeloproliferative disorder of Down syndrome. Blood. 2003;102:2960–8.
Roberts I, Alford K, Hall G, Juban G, Richmond H, Norton A, Vallance G, Perkins K, Marchi E, McGowan S, et al. GATA1-mutant clones are frequent and often unsuspected in babies with Down syndrome: identification of a population at risk of leukemia. Blood. 2013;122:3908–17.
Carpenter E, Valverde-Garduno V, Sternberg A, Mitchell C, Roberts I, Vyas P, Vora A. GATA1 mutation and trisomy 21 are required only in haematopoietic cells for development of transient myeloproliferative disorder. Br J Haematol. 2005;128:548–51.
Nikolaev SI, Santoni F, Vannier A, Falconnet E, Giarin E, Basso G, Hoischen A, Veltman JA, Groet J, Nizetic D, et al. Exome sequencing identifies putative drivers of progression of transient myeloproliferative disorder to AMKL in infants with Down syndrome. Blood. 2013;122:554–61.
Alter BP. Fanconi anemia and the development of leukemia. Best Pract Res Clin Haematol. 2014;27:214–21.
Ma SK, Wan TS, Cheuk AT, Fung LF, Chan GC, Chan SY, Ha SY, Chan LC. Characterization of additional genetic events in childhood acute lymphoblastic leukemia with TEL/AML1 gene fusion: a molecular cytogenetics study. Leukemia. 2001;15:1442–7.
Moorman AV, Richards SM, Robinson HM, Strefford JC, Gibson BE, Kinsey SE, Eden TO, Vora AJ, Mitchell CD, Harrison CJ. Prognosis of children with acute lymphoblastic leukemia (ALL) and intrachromosomal amplification of chromosome 21 (iAMP21). Blood. 2007;109:2327–30.
Antonarakis SE. Down syndrome and the complexity of genome dosage imbalance. Nat Rev Genet. 2017;18:147–63.
Malinge S, Bliss-Moreau M, Kirsammer G, Diebold L, Chlon T, Gurbuxani S, Crispino JD. Increased dosage of the chromosome 21 ortholog Dyrk1a promotes megakaryoblastic leukemia in a murine model of Down syndrome. J Clin Invest. 2012;122:948–62.
Saida S, Watanabe K, Sato-Otsubo A, Terui K, Yoshida K, Okuno Y, Toki T, Wang R, Shiraishi Y, Miyano S, et al. Clonal selection in xenografted TAM recapitulates the evolutionary process of myeloid leukemia in Down syndrome. Blood. 2013;121:4377–87.
Muramatsu H, Okuno Y, Yoshida K, Shiraishi Y, Doisaki S, Narita A, Sakaguchi H, Kawashima N, Wang X, Xu Y, et al. Clinical utility of next-generation sequencing for inherited bone marrow failure syndromes. Genet Med. 2017;
Acknowledgements
This work was supported by Japan Society for the Promotion of Science (JSPS) KAKENHI Grant-in-Aid for Young Scientists (B) Research Project Number 26860795.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The author declares that he has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Leukemia
Rights and permissions
About this article
Cite this article
Saida, S. Predispositions to Leukemia in Down Syndrome and Other Hereditary Disorders. Curr. Treat. Options in Oncol. 18, 41 (2017). https://doi.org/10.1007/s11864-017-0485-x
Published:
DOI: https://doi.org/10.1007/s11864-017-0485-x