Abstract
Purpose of Review
Advances in the understanding of germline predisposition to pediatric cancers, particularly myeloid neoplasms, have increased rapidly over the last 20 years. Here, we highlight the most up-to-date knowledge regarding known pathogenic germline variants that contribute to the development of myeloid neoplasms in children.
Recent Findings
This discussion enumerates the most notable myeloid neoplasm-causing germline mutations. These mutations may be organized based on their molecular underpinnings—transcriptional control, splicing and signal transduction control, and a group of heterogeneous bone marrow failure syndromes. We review recent findings related to the biochemical mechanisms that predispose to malignant transformation in each condition. Key genetic discoveries such as novel mutations, degrees of penetrance, principles of the two-hit hypothesis, and co-occurrence of multiple mutations are shared. Clinical pearls, such as information regarding epidemiology, natural history, or prognosis, are also discussed.
Summary
Germline mutations predisposing to pediatric myeloid neoplasms are frequent, but underrecognized. They hold major clinical implications regarding prognosis, treatment strategies, and screening for other malignancies. Further research is warranted to better characterize each of these conditions, as well as identify additional novel germline pathogenic variants of interest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Historically, greater attention has been paid to cancer predisposition syndromes in the setting of adult-onset malignancies as compared to pediatric patients. Further recognition is needed regarding the role of predisposition syndromes in the development of pediatric malignancies, as rates of pathogenic germline mutations have been estimated to be similar among pediatric and adult patients with cancer (8.5% vs. 8%) [1]. The first pediatric germline predisposition syndrome for bone marrow failure (BMF) and myeloid neoplasms (MNs) was described nearly a century ago, with the discovery of Fanconi anemia [2]. However, the acquisition of new knowledge regarding other predisposing germline mutations for MNs remained relatively stalled until the last 20 years, during which the discovery of causative mutations has rapidly increased.
Myeloid malignancies occur with lower frequency in children as compared to lymphoid neoplasms, with approximately only 20% of childhood leukemias being of myeloid origin [3]. Of those, up to 25% are estimated to carry pathogenic germline mutations [4]. In light of this growing population, the WHO classification was updated in 2016 to include acute myeloid leukemia (AML) and other MNs associated with germline mutations or a known cancer predisposition syndrome as a distinct entity [5].
Without a heightened index of suspicion, germline mutations may go unrecognized, as many of these pathogenic variants may also be identified as somatic mutations in the neoplasm. Therefore, testing of healthy tissue for germline mutations is of high importance. Care must be taken to ensure that true germline material is utilized for testing. Blood and bone marrow are not viable sources, even if the patient is in remission, as somatic mutations may persist in these sites. Tissues that are easily contaminated also should not be used, such as saliva and nails. The gold standard sample for germline testing is cultured skin fibroblasts via skin biopsy [6]. The diagnosis of a cancer predisposition syndrome often holds major implications for the individual patient, as well as their family members. The identification of a germline pathogenic variant may influence prognosis, treatment recommendations, and monitoring for additional malignancies. Family members may benefit from enhanced screening for various malignancies, as well as knowledge of inheritance patterns for reproductive planning.
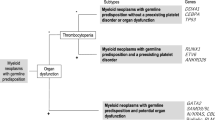
Implicated genes of the germline predisposition mutations for MNs have a wide array of molecular effects. Many are involved in transcriptional control, as well as splicing and signal transduction control. Additionally, syndromes have been identified with defects rooted in DNA repair, telomere biology, ribosome synthesis, and tumor suppression mechanisms. Each of these conditions will be discussed separately here, with an emphasis on current knowledge regarding the genetic and biological mechanisms that predispose to malignancy, as well as key clinical takeaways. Each germline mutation lends itself to a different clinical phenotype, which is partially characterized by age of onset for the MNs depicted in Fig. 1.
Timeline of age of MN onset by mutation/syndrome, averages, and ranges [32]. Depiction of the average (indicated by arrowhead) and range (indicated by bracket) of age of onset for myeloid neoplasms by specific germline mutation or syndrome. Conditions listed in upper figure portion are predominately adult age of onset; conditions listed in the lower figure portion are predominately childhood age of onset
Transcriptional Control
Transcription factors are the proteins which convert DNA into RNA. They enable appropriate hematopoiesis by regulating downstream gene expression through binding to specific consensus sequence elements. Disruptions in transcription factor control over gene expression are critical in the evolution of cancer and can occur through germline predisposition or somatic mutations. Transcription factor mutations that can lead to these disruptions and induce myeloid neoplasms include CEBPA, RUNX1, and GATA2.
CEBPA
CCAAT enhancer binding protein alpha (CEBPA) is a critical transcription factor identified as a loci of interest in MNs in 2004 [7]. CEBPA encodes the CCAAT enhancer binding protein (C/EBP), which is involved in lineage-specific myeloid differentiation. Mutations in CEBPA confer a high risk of myeloid neoplasms and may be found in 5–10% of AML cases. In de novo AML, up to 15% involve biallelic mutations in CEBPA [8]. De novo AML refers to AML in patients with no clinical history of prior myelodysplastic syndrome (MDS), myeloproliferative disorder, or exposure to potentially leukemogenic therapies or agents [9]. Secondary AML (s-AML) refers to a leukemic process: (1) evolving from prior myelodysplasia (MDS), myeloproliferative disorder (MPN), or aplastic anemia (AA) with or without treatment or (2) as a product of previous exposure to a proven leukemogenic chemotherapeutic agent (therapy-related AML (t-AML)) [10]. Wild-type CEBPA usually encodes for a 42 kDa isoform. In most familial cases of AML involving CEBPA mutations, biallelic mutations are observed. Biallelic mutations prevent DNA binding or C/EBP dimerization and occur when both a C-terminal and an N-terminal mutation are present. The germline CEBPA mutation results in a 30 kDa isoform usually from variants leading to stop-gain frameshifts in the N terminus. A further second mutation occurring in the C-terminal is often acquired as either an in-frame insertion or deletion or a missense variant [11]. Also, possibly contributing to the pathogenesis of AML is the epigenetic control of gene expression. CEBPA promoter methylation has recently been found to silence the gene [12••, 13]. Some variants in biallelic CEBPA mutated AML tend to co-occur such as GATA2, WT1 (WT1 transcription factor), TET2 (Tet methylcytosine dioxygenase 2), and CSF3R (colony-stimulating factor 3 receptor). Others are rare and often occur exclusively such as FLT3 (fms-like tyrosine kinase 3), DNMT3A (DNA methyltransferase 3 alpha), IDH1/2 (isocitrate dehydrogenase (NADP( +)) 1/2), NPM1 (nucleophosmin 1), and RUNX1 [14]. Compared to other cancer predisposition genes, germline CEBPA mutations demonstrate significant penetrance and confer an almost 100% risk of developing AML [15]. Notably biallelic CEBPA gene mutations confer both an increased survival and event-free survival compared to those with monoallelic CEBPA mutation [16].
RUNX1
The RUNX1 gene encodes a protein which is a key transcription factor for hematopoiesis called runt-related transcription factor (RUNX1). Approximately 10% of AML cases involve RUNX1 mutations and a further 10% of these RUNX1-mutated AML cases involve germline variants [17,18,19
GATA2
GATA2 is a transcription factor involved in stem cell maintenance with key roles in regulating gene expression in hematopoietic cells. It belongs to the GATA family of transcription factors which contain zinc fingers in their DNA binding domain. GATA2 is expressed in hematopoietic progenitors as well as nonhematopoietic embryonic stem cells and is involved in the development of the lymphatic system [21]. Germline GATA2 mutations demonstrate high penetrance with up to 70% risk of developing a myeloid neoplasm and a 90% risk of developing clinical symptoms [22, 23]. This topic is further discussed in another chapter.
Splicing and Signal Transduction Control
Splicing describes the process where the noncoding regions of genes are removed from the primary messenger RNA transcript leaving the coding regions which are then joined together to generate mature messenger RNA. Alternative pre-mRNA splicing occurs in myeloid neoplasms with higher frequencies seen in somatic mutations. There are rare germline variants such as mutations in DDX41, SAMD9/SAMD9L, and the RAS/MAPK pathway which predispose to malignancy.
SAMD9 and SAMD9L
Sterile alpha motif domain-containing protein 9 (SAMD9) and sterile alpha motif domain-containing protein 9-like (SAMD9L) genes are located on chromosome 7 and are derived from the same ancestral gene [24••]. They are both interferon TNF-alpha inducible and were discovered due to studies in patients with MNs and acquired 7q21 microdeletions [25]. While they appear to be involved in cytokine signaling and endocytosis, their hematopoietic function is unclear [26].
In pediatric patients with MDS, 17% have germline variants of SAMD9/9L and most are missense mutations which occur in the second half or C-terminus of the SAMD9/9L proteins leading to autosomal-dominant syndromes [27]. Deletions involving the long arm of chromosome 7 as well as monosomy of chromosome 7 are common abnormalities seen in AML and MDS as haploinsufficiency of SAMD9/9L contributes to genetic neoplasms. Conversely, germline mutations which cause gain-of-function mutations are associated with a spectrum of disorders such including ataxia-pancytopenia syndrome (ATXPC), myelodysplasia, and leukemia syndrome with monosomy 7 syndrome and MIRAGE (myelodysplasia, infection, restriction of growth, adrenal hypoplasia, genital phenotypes, and enteropathy) syndrome [28]. An autoinflammatory panniculitis similar to Chronic Atypical Neutrophilic Dermatosis with Lipodystrophy and Elevated Temperatures (CANDLE) syndrome was recently found to be caused by frameshift and truncating germline mutations in SAMD9L [29••]. Separately, SAMD9 mutations demonstrate high penetrance with a high rate of de novo cases as observed in MIRAGE syndrome [28]. Conversely, in SAMD9L syndromes, there is incomplete penetrance which is approximated to 70% for most hematological diseases [30].
Normally, SAMD9/9L inhibits cell cycle progression and suppress myeloid proliferation; thus, activating mutations leads to growth restrictions with organ hypoplasia and pancytopenias [31]. Acquired loss-of-function mutations in SAMD9 or SAMD9L have been shown in studies to balance the gain-of-function germline effect [31]. Another proposition is that an adaptive mechanism to germline SAMD9/9L mutations may be the selective loss of chromosome 7 [27, 28]. The resultant haploinsufficiency of genes which co-occur in this region such as KMT2C/MLL3 (lysine methyltransferase 2C/mixed-lineage leukemia 3), EZH2, and CUX1 can also influence the development of AML or MDS [32].
There is a spectrum of clinical presentation in patients affected by SAMD9/9L syndromes ranging from transient cytopenia (most commonly thrombocytopenia) to a much more severe disease often fatal in infancy [24••]. Pancytopenia with hypocellular marrow and MDS with − 7 or del(7q) are also common presentations [24••]. While hematologic abnormalities are shared between both SAMD9/9L syndromes, they each can manifest separately with distinct characteristics. Notably the SAMD9 phenotype presents as MIRAGE syndrome often with intrauterine growth restriction; genital abnormalities such as testicular dysgenesis, cryptorchid testes and clitoromegaly, primary adrenal insufficiency and early onset adrenal hypoplasia, gastrointestinal issues (enteropathy, reflux); severe systemic infections; and cytopenia at birth [33]. Neurological findings are the hallmark in patients affected by SAMD9L mutations can lead to the loss of Purkinje cells, often presenting as cerebellar ataxia or atrophy, nystagmus, and dysmetria [34, 35].
One large cohort of pediatric MDS patients 90 percent of those with a SAMD9/9L mutation showed refractory cytopenia of childhood (RCC) while 10% showed MDS with excess blasts [33]. Furthermore, in pediatric MDS, the median age at diagnosis in those with SAMD9/9L was similar to those with GATA2-related MDS at 9.6 years (Fig. 1) [33].
There is a spectrum of ill-defined immunologic dysfunction in patients with SAMD9/9L mutations [24••]. In some cases, the SAMD9/9L mutation decreased peripheral B/NK-cells, low IgG, and IgM or increased TNF-alpha and IL-6 levels have been recorded [36]. While most of the non-syndromic MDS patients harboring this mutation do not appear to be at increased risk of severe infections, those with syndromic phenotypes such as MIRAGE syndrome and some SAMD9L mutations demonstrate immune dysregulation [28, 37].
The two unique SAMD9/9L disease mechanisms are clonal evolution and somatic revertant mosaicism. Clonal evolution occurs through the non-random loss of chromosome 7 (− 7/del(7q)) which contains mutated SAMD9/9L gene copy and has been termed adaptation by aneuploidy [38]. Chromosome 7 harbors several tumor suppressor genes; therefore, clonal evolution to advanced AML/MDS is a frequent complication in SAMD9/9L-related MDS [39]. This process is usually paired with somatic driver mutations such as RUNX1, PTPN11, KRAS, ETV6, BRAF, SETBP1, ASXL1, CBL, and EZH2 [40].
In somatic revertant mosaicism, the two processes involved include the acquisition of truncating SAMD9/9L mutations or an independent uniparental disomy of 7q (UPD7q) [41]. Somatic SAMD9/9L mutations are thought to exert a loss-of-function effect and cancel the gain-of-function germline mutation as they are acquired on the same allele (in cis) [36].
Currently, due to the novelty of SAMD9/9L with respect to myeloid predisposition, no concrete management guidelines exist. In general, for pediatric patients with MDS, HSCT is recommended as soon as clinically possible especially in those in patients with immunodeficiency, morphologically advanced disease, transfusion dependency, neutropenia, and high-risk cytogenetic and genetic lesions [42].
RAS/MAPK Pathway
The RAS/MAPK pathway is one of the more well-studied signal transduction pathways in cell biology. Its function is to transduce extracellular products to the cell nucleus activating specific genes for cell growth, division, and differentiation [43]. RAS is a superfamily of proteins which are GTPases controlling various cell signaling pathways such as the MAPK pathway and include HRAS, KRAS, and NRAS. RAS controls downstream signaling pathways important for proliferation and differentiation by being active in the GTP-bound state and being inactive in the GDP bound state [44, 45]. Germline pathogenic variants in the RAS/MAPK pathway are associated with a spectrum of congenital disorders such as neurofibromatosis type 1, cardiofaciocutaneous syndrome, Costello syndrome, Noonan syndrome, and Noonan syndrome with multiple lentigines. The somatic variants in KRAS, NRAS, PTPN1, CBL, and NF1 are often seen in various neoplasms. There is a significant risk of myeloproliferative neoplasms conferred by these conditions including a severe pediatric myeloid neoplasm termed juvenile myelomonocytic leukemia (JMML) [46, 47]. In patients with JMML, germline mutations are predominantly located in intron 7 and exons 8 and 9. JMML occurs as the result of dysregulation in RAS/MAPK signaling pathway often caused by either germline mutations in NF1 or CBL tumor suppressors and further biallelic inactivation in hematopoietic cells or heterozygous somatic gain-of-function mutations in KRAS, NRAS, or PTPN1. Copy-neutral loss of heterozygosity (CN-LOH) affecting chromosome 17q, a second NF1 mutation, or somatic interstitial deletions are the main mechanisms by which a second NF1 may occur [48, 49
DDX41
DDX41 (DEAD-Box Helicase 41) encodes an RNA helicase which plays a role in RNA splicing. The gene, located on chromosome 5, may experience mutations which are often mutually exclusive with the spliceosome mutations more frequently seen in myeloid neoplasms [51]. Mutations in this gene modify RNA processing and pre-MRA splicing, thereby causing loss of tumor suppressor function [52]. One of the most commonly occurring postconception alterations found in myeloid tumors is the p.R525H variant in DDX41 [53]. Recent studies show there may be certain locations on DDX41 that are predisposed to germline mutations and are often more common within ethnic groups. For example within the Asian population, the p.A500Cfs*9 germline variant is often found whereas the p.M1I and p.D140Gfs*2 variants are more often found within the Caucasian population [51, 54]. In addition to myeloid neoplasms, these families are also predisposed to the development of non-Hodgkin lymphoma. The germline DDX41 mutations present in myeloid neoplasms are often found in high-risk AML or MDS cases and bone marrow hypocellularity. MDS precedes more than half of reported cases of AML with the DDX41 mutation [55]. Further simultaneous mutations associated with secondary AML include EZH2, ASXL1 (ASXL transcriptional regulator 1), CUX1 (Cut like homeobox 1), SRSF2 (serine- and arginine-rich splicing factor 2), and SETBP1 [56]. Somatic DDX41 mutations are uncommon without predisposing germline DDX41 pathogenic variants and are most often found in older patients with a median age of diagnosis as 69 years. This differs from the RUNX1, GATA2, or CEBPA genes with a high predisposition to familial myeloid neoplasms and usually have a younger age of diagnosis [52].
Bone Marrow Failure Syndromes and Other Inherited Disorders
Fanconi Anemia (FA)
Bone marrow failure syndromes such as Fanconi Anemia, Dyskeratosis Congenita, Shwachman-Diamond syndrome, Severe congenital Neutropenia, and Diamond–Blackfan anemia significantly increase the likelihood of developing a myeloid neoplasm.
FA is understood to be caused by dysfunctional DNA repair, due to hypersensitivity to mitomycin C (MMC) and diepoxybutane (DEB), which causes DNA cross-linking [57]. There is subsequently a higher rate of chromosomal breakage seen in these cells, which is the basis of diagnostic testing for clinical evaluation of FA. MDS and AML are thought to develop in these patients due to the presence of mutations in these cells which allow for evasion of cell cycle regulation and apoptosis [58].
The general categories of FA genes include the FA core complex which makes proteins which then then activate ID2 complex proteins and a group of proteins in the downstream functional units which work to recruit DNA repair proteins. When genes in this pathway are mutated, there is subsequent impairment in DNA repair, particularly double-strand DNA damage. The risk of developing MDS or AML before the age of 20 is 27% and increases to 52% by the age of 40 as per the International Fanconi Anemia Registry Study [59].
Dyskeratosis Congenita (DC)
Dyskeratosis congenita is a bone marrow failure disorder which is part of the telomere biology disorders. In two-thirds of the patients with DC, mutations in genes encoding core telomerase components and others involved in telomere maintenance have been identified [60]. As shortened telomeres leads to senescence, apoptosis, or oncogenic transformation of somatic cells and thus cause subsequent genetic instability, DC is therefore also a myeloid predisposition syndrome. This is thought to play a critical role in the malignant potential and transformation [61].
Diamond–Blackfan Anemia (DBA)
Diamond–Blackfan anemia is a cancer predisposition syndrome associated with defects in ribosome synthesis. These gene defects, known as ribosomopathies, lead to haploinsufficiency by hosting heterozygous loss-of-function mutations. Numerous studies show that these ribosomopathies can stimulate malignant transformation from the upregulation of protein synthesis [62]. Further evidence has been found showing that ribosomal stress signaling through p53 activation may play an additional role in malignant transformation. This mechanism conveys a higher risk of developing solid tumors, AML, and MDS among DBA in some studies [63].
Li–Fraumeni Syndrome
Li–Fraumeni syndrome is a cancer predisposition disorder linked to p53 tumor suppressor gene germline mutations. The downstream effects of this mutation include dysregulation of the cell cycle, cell proliferation, DNA repair, genomic stability, and homeostasis. In patients with Li–Fraumeni syndrome, the most frequent leukemia is hypodiploid acute lymphoblastic leukemia however myeloid neoplasms are also common. These often occur as a complication from therapy given for a primary malignancy. De novo germline p53 mutations in AML are rare [64, 65].
Trisomy 21
Transient myeloproliferative disorder is a self-resolving abnormality in myelopoiesis that is uniquely seen in neonates with complete or mosaic trisomy 21. This condition arises due to clonal proliferation of immature megakaryoblasts, as a result of cooperation between the chromosome 21 trisomy and somatic mutations in the GATA1 gene [66]. The exact genes on chromosome 21 responsible for this disorder have not yet been identified, but may include RUNX1, ETS2, and ERG [67, 68]. GATA1 mutations in isolation are not thought to be sufficient to cause transient myeloproliferative disorder [69]. Clinically, transient myeloproliferative disorder may range from asymptomatic to life threatening, with hyperleukocytosis, hydrops fetalis, infiltration of solid organs, liver failure, and cardiopulmonary failure [70]. While most patients improve without intervention, 20–30% of patients with transient myeloproliferative disorder will go on to develop acute megakaryoblastic leukemia within 3 years [71].
Conclusion
Knowledge regarding the role of germline mutations in MNs has substantially grown in recent years, with far more patients carrying pathogenic variants than previously suspected. And yet, germline predisposition among pediatric cancer patients remains underdiagnosed. Lack of associated syndromic features and/or family history can make identifying these conditions quite challenging by clinical history. As comprehensive genetic testing becomes more accurate, accessible, and affordable, testing for underlying mutations should be integrated into the care of pediatric patients with newly diagnosed MNs. Spectrum of penetrance and clinical illness may vary widely based on the genetic underpinning. Therefore, identification of a germline mutation can profoundly influence medical decision-making. Further research is warranted to better understand frequency, penetrance, and clinical outcomes among these patients.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Zhang J, et al. Germline mutations in predisposition genes in pediatric cancer. N Engl J Med. 2015;373:2336–46.
Böhles H. Die perniziöse Fanconi-Anämie, 1927. In: Historische Fälle aus der Medizin. Springer, Berlin, Heidelberg. (2020). https://doi.org/10.1007/978-3-662-59833-7_27.
Smith MA, Ries LA, Gurney JG, et al. Leukemia. In: Ries LA, Smith MA, Gurney JG, et al., eds.: Cancer incidence and survival among children and adolescents: United States SEER Program 1975–1995. National Cancer Institute, SEER Program, 1999;NIH 17–34.
Samaraweera SE, Wang PPS, Li KL, Casolari DA, Feng J, Pinese M, Maung KZY, Leo P, Cowley M, Perkins K, Smith AM, Ellis J, Wee A, Hiwase DK, Scott HS, Schreiber AW, Brown AL, Deans AJ, Ross DM, Moore AS, Gonda TJ, Hahn CN, D’Andrea RJ. Childhood acute myeloid leukemia shows a high level of germline predisposition. Blood. 2021;138(22):2293–8. https://doi.org/10.1182/blood.2021012666.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, Bloomfield CD, Cazzola M, Vardiman JW. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–405. https://doi.org/10.1182/blood-2016-03-643544.
Desai AV, Perpich M, Godley LA. Clinical assessment and diagnosis of germline predisposition to hematopoietic malignancies: the University of Chicago Experience. Front Pediatr. 2017;6(5):252. https://doi.org/10.3389/fped.2017.00252.
Smith ML, Cavenagh JD, Lister TA, Fitzgibbon J. Mutation of CEBPA in familial acute myeloid leukemia. N Engl J Med. 2004;351(23):2403–7. https://doi.org/10.1056/NEJMoa041331.
Mannelli F, Ponziani V, Bencini S, Bonetti MI, Benelli M, Cutini I, Gianfaldoni G, Scappini B, Pancani F, Piccini M, Rondelli T, Caporale R, Gelli AM, Peruzzi B, Chiarini M, Borlenghi E, Spinelli O, Giupponi D, Zanghì P, Bassan R, Bosi A. CEBPA-double-mutated acute myeloid leukemia displays a unique phenotypic profile: a reliable screening method and insight into biological features. Haematologica. 2017;102(3):529–40. https://doi.org/10.3324/haematol.2016.151910.
Cheson BD, Bennett JM, Kopecky KJ, Büchner T, Willman CL, Estey EH, Schiffer CA, Doehner H, Tallman MS, Lister TA, Lo-Coco F, Willemze R, Biondi A, Hiddemann W, Larson RA, Löwenberg B, Sanz MA, Head DR, Ohno R, Bloomfield CD; International Working Group for Diagnosis, Standardization of Response Criteria, Treatment Outcomes, and Reporting Standards for Therapeutic Trials in Acute Myeloid Leukemia. Revised recommendations of the International Working Group for Diagnosis, Standardization of Response Criteria, Treatment Outcomes, and Reporting Standards for Therapeutic Trials in Acute Myeloid Leukemia. J Clin Oncol. 2003 Dec 15;21(24):4642–9. https://doi.org/10.1200/JCO.2003.04.036. Erratum in: J Clin Oncol. 2004;Feb 1;22(3):576. LoCocco, Francesco [corrected to Lo-Coco, Francesco].
Boddu P, Kantarjian HM, Garcia-Manero G, Ravandi F, Verstovsek S, Jabbour E, Borthakur G, Konopleva M, Bhalla KN, Daver N, DiNardo CD, Benton CB, Takahashi K, Estrov Z, Pierce SR, Andreeff M, Cortes JE, Kadia TM. Treated secondary acute myeloid leukemia: a distinct high-risk subset of AML with adverse prognosis. Blood Adv. 2017;1(17):1312–23. https://doi.org/10.1182/bloodadvances.2017008227.
Wurm AA, Zjablovskaja P, Kardosova M, Gerloff D, Bräuer-Hartmann D, Katzerke C, Hartmann JU, Benoukraf T, Fricke S, Hilger N, Müller AM, Bill M, Schwind S, Tenen DG, Niederwieser D, Alberich-Jorda M, Behre G. Disruption of the C/EBPα-miR-182 balance impairs granulocytic differentiation. Nat Commun. 2017;8(1):46. https://doi.org/10.1038/s41467-017-00032-6.
•• Kimura Y, Iwanaga E, Iwanaga K, et al. A regulatory element in the 3′-untranslated region of CEBPA is associated with myeloid/NK/T-cell leukemia. Eur J Haematol. 2021;106:327–39. https://doi.org/10.1111/ejh.13551. DNA methylation in the CEBPA 3′-untranslated region (UTR) is associated with myeloid/NK/T-cell leukemia and CEBPA downregulation. The CEBPA 3′-UTR has the enhancer-like activity that is positively controlled by IKZF1. The methylation testing of CEBPA 3′-UTR helps to classify myeloid/NK/T-cell leukemia.
Wen XM, Hu JB, Yang J, Qian W, Yao DM, Deng ZQ, Zhang YY, Zhu XW, Guo H, Lin J, et al. CEBPA methylation and mutation in myelodysplastic syndrome. Med Oncol. 2015;32:192. https://doi.org/10.1007/s12032-015-0605-z.
Wilhelmson AS, Porse BT. CCAAT enhancer binding protein alpha (CEBPA) biallelic acute myeloid leukaemia: cooperating lesions, molecular mechanisms and clinical relevance. Br J Haematol. 2020;190:495–507. https://doi.org/10.1111/bjh.16534.
Tawana K, Wang J, Renneville A, Bodor C, Hills R, Loveday C, Savic A, Van Delft FW, Treleaven J, Georgiades P, et al. Disease evolution and outcomes in familial AML with germline CEBPA mutations. Blood. 2015;126:1214–23. https://doi.org/10.1182/blood-2015-05-647172.
Swerdlow SH. Who classification of tumours of haematopoietic and lymphoid tissues. International Agency for Research on Cancer; 2017.
Ernst MPT, Kavelaars FG, Lowenberg B, Valk PJM, Raaijmakers M. RUNX1 germline variants in RUNX1-mutant AML: how frequent? Blood. 2021;137:1428–31. https://doi.org/10.1182/blood.2020008478.
Mendler JH, Maharry K, Radmacher MD, Mrozek K, Becker H, Metzeler KH, Schwind S, Whitman SP, Khalife J, Kohlschmidt J, et al. RUNX1 mutations are associated with poor outcome in younger and older patients with cytogenetically normal acute myeloid leukemia and with distinct gene and MicroRNA expression signatures. J Clin Oncol. 2012;30:3109–18. https://doi.org/10.1200/JCO.2011.40.6652.
Gaidzik VI, Teleanu V, Papaemmanuil E, Weber D, Paschka P, Hahn J, Wallrabenstein T, Kolbinger B, Kohne CH, Horst HA, et al. RUNX1 mutations in acute myeloid leukemia are associated with distinct clinico-pathologic and genetic features. Leukemia. 2016;30:2160–8. https://doi.org/10.1038/leu.2016.126.
Drazer MW, Kadri S, Sukhanova M, Patil SA, West AH, Feurstein S, Calderon DA, Jones MF, Weipert CM, Daugherty CK, et al. Prognostic tumor sequencing panels frequently identify germ line variants associated with hereditary hematopoietic malignancies. Blood Adv. 2018;2:146–50. https://doi.org/10.1182/bloodadvances.2017013037.
OMIM entry - * 137295 - GATA-binding protein 2; GATA2. (n.d.). Retrieved from https://www.omim.org/entry/137295
Collin M, Dickinson R, Bigley V. Haematopoietic and immune defects associated with GATA2 mutation. Br J Haematol. 2015;169:173–87. https://doi.org/10.1111/bjh.13317.
Spinner MA, Sanchez LA, Hsu AP, Shaw PA, Zerbe CS, Calvo KR, Arthur DC, Gu W, Gould CM, Brewer CC, et al. GATA2 deficiency: A protean disorder of hematopoiesis, lymphatics, and immunity. Blood. 2014;123:809–21. https://doi.org/10.1182/blood-2013-07-515528.
•• Sahoo SS, Kozyra EJ, Wlodarski MW. Germline predisposition in myeloid neoplasms: unique genetic and clinical features of GATA2 deficiency and SAMD9/SAMD9L syndromes. Best Pract Res Clin Haematol. 2020;33(3):101197. https://doi.org/10.1016/j.beha.2020.101197. A great summary highlighting the current knowledge regarding the GATA2 and SAMD9/9L syndromes with all the key references and stakeholders.
Asou H, Matsui H, Ozaki Y, Nagamachi A, Nakamura M, Aki D, et al. Identification of a common microdeletion cluster in 7q21.3 subband among patients with myeloid leukemia and myelodysplastic syndrome. Biochem Biophys Res Commun. 2009;383:245–51.
Inaba T, Nagamachi A. Revertant somatic mosaicism as a cause of cancer. Cancer Sci. 2021;112(4):1383–9. https://doi.org/10.1111/cas.14852.
Schwartz JR, Ma J, Lamprecht T, et al. The genomic landscape of pediatric myelodysplastic syndromes. Nat Commun. 2017;8:1557. https://doi.org/10.1038/s41467-017-01590-5.
Narumi S, Amano N, Ishii T, Katsumata N, Muroya K, Adachi M, Toyoshima K, Tanaka Y, Fukuzawa R, Miyako K, et al. SAMD9 mutations cause a novel multisystem disorder, MIRAGE syndrome, and are associated with loss of chromosome 7. Nat Genet. 2016;48:792–7. https://doi.org/10.1038/ng.3569.
•• de Jesus AA, Hou Y, Brooks S, Malle L, Biancotto A, Huang Y, et al. Distinct interferon signatures and cytokine patterns define additional systemic autoinflammatory diseases. J Clin Invest. 2020;130(4):1669–82. A very interesting paper showing the correlation between immune-dysregulatory/autoinflammatory diseases and myeloid predisposition syndromes.
Pastor VB, Sahoo SS, Boklan J, Schwabe GC, Saribeyoglu E, Strahm B, et al. Constitutional SAMD9L mutations cause familial myelodysplastic syndrome and transient monosomy 7. Haematologica. 2018;103:427–37. https://doi.org/10.3324/haematol.2017.180778.
Davidsson J, Puschmann A, Tedgard U, Bryder D, Nilsson L, Cammenga J. SAMD9 and SAMD9L in inherited predisposition to ataxia, pancytopenia, and myeloid malignancies. Leukemia. 2018;32:1106–15. https://doi.org/10.1038/s41375-018-0074-4.
Sperling AS, Gibson CJ, Ebert BL. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat Rev Cancer. 2017;17(1):5–19. https://doi.org/10.1038/nrc.2016.112.
Sahoo SS, Pastor VP, Panda PK, Szvetnik EK, Kozyra EJ, Voss R, et al. SAMD9 and SAMD9L germline disorders in patients Enrolled in studies of the European working group of MDS in childhood (EWOG-MDS): prevalence, outcome, phenotype and functional characterisation. Blood. 2018;132(1):613. https://doi.org/10.1182/blood-2018-99-118389.
Chen DH, Below JE, Shimamura A, Keel SB, Matsushita M, Wolff J, et al. Ataxia-pancytopenia syndrome is caused by missense mutations in SAMD9L. Am J Hum Genet. 2016;98:1146–58.
Thunstrom S, Axelsson M. Leukoencephalopathia, demyelinating peripheral neuropathy and dural ectasia explained by a not formerly described de novo mutation in the SAMD9L gene, ends 27 years of investigations - a case report. BMC Neurol. 2019;19:89.
Tesi B, Davidsson J, Voss M, Rahikkala E, Holmes TD, Chiang SC, et al. Gain-of-function SAMD9L mutations cause a syndrome of cytopenia, immunodeficiency. MDS and neurological symptoms Blood. 2017;129(16):2266–79.
Bluteau O, Sebert M, Leblanc T, Peffault de Latour R, Quentin S, Lainey E, et al. A landscape of germ line mutations in a cohort of inherited bone marrow failure patients. Blood. 2018;131:717–32.
Perisa MP, Rose MJ, Varga E, Kamboj MK, Spencer JD, Bajwa RPS. A novel SAMD9 variant identified in patient with MIRAGE syndrome: further defining syndromic phenotype and review of previous cases. Pediatr Blood Canc. 2019;66:e27726.
Schwartz JR, Wang S, Ma J, Lamprecht T, Walsh M, Song G, et al. Germline SAMD9 mutation in siblings with monosomy 7 and myelodysplastic syndrome. Leukemia. 2017;31:1827–30.
Wong JC, Bryant V, Lamprecht T, Ma J, Walsh M, Schwartz J, et al. Germline SAMD9 and SAMD9L mutations are associated with extensive genetic evolution and diverse hematologic outcomes. JCI Insight 2018:3
Buonocore F, Kuhnen P, Suntharalingham JP, Del Valle I, Digweed M, Stachelscheid H, et al. Somatic mutations and progressive monosomy modify SAMD9- related phenotypes in humans. J Clin Invest. 2017;127(5):1700–13.
Niemeyer CM. Pediatric MDS including refractory cytopenia and juvenile myelomonocytic leukemia. In: th Carreras E, Dufour C, Mohty M, Kroger N, editors. The EBMT handbook: hematopoietic stem cell transplantation and cellular therapies; 2019 557–60. Cham (CH).
Molina JR, Adjei AA. The Ras/Raf/MAPK pathway. J Thorac Oncol. 2006;1(1):7–9 (PMID: 17409820).
Drosten M, Dhawahir A, Sum EY, Urosevic J, Lechuga CG, Esteban LM, Castellano E, Guerra C, Santos E, Barbacid M. Genetic analysis of Ras signalling pathways in cell proliferation, migration and survival. EMBO J. 2010;29:1091–104. https://doi.org/10.1038/emboj.2010.7.
Shapiro P. Ras-MAP kinase signaling pathways and control of cell proliferation: Relevance to cancer therapy. Crit Rev Clin Lab Sci. 2002;39:285–330. https://doi.org/10.1080/10408360290795538.
Strullu M, Caye A, Lachenaud J, Cassinat B, Gazal S, Fenneteau O, Pouvreau N, Pereira S, Baumann C, Contet A, et al. Juvenile myelomonocytic leukaemia and Noonan syndrome. J Med Genet. 2014;51:689–97. https://doi.org/10.1136/jmedgenet-2014-102611.
Perez B, Mechinaud F, Galambrun C, Ben RN, Isidor B, Philip N, Derain-Court J, Cassinat B, Lachenaud J, Kaltenbach S, et al. Germline mutations of the CBL gene define a new genetic syndrome with predisposition to juvenile myelomonocytic leukaemia. J Med Genet. 2010;47:686–91. https://doi.org/10.1136/jmg.2010.076836.
Loh ML, Sakai DS, Flotho C, Kang M, Fliegauf M, Archambeault S, Mullighan CG, Chen L, Bergstraesser E, Bueso-Ramos CE, et al. Mutations in CBL occur frequently in juvenile myelomonocytic leukemia. Blood. 2009;114:1859–63. https://doi.org/10.1182/blood-2009-01-198416.
Caye A, Strullu M, Guidez F, Cassinat B, Gazal S, Fenneteau O, Lainey E, Nouri K, Nakhaei-Rad S, Dvorsky R, et al. Juvenile myelomonocytic leukemia displays mutations in components of the RAS pathway and the PRC2 network. Nat Genet. 2015;47:1334–40. https://doi.org/10.1038/ng.3420.
Stieglitz E, Taylor-Weiner AN, Chang TY, Gelston LC, Wang YD, Mazor T, Esquivel E, Yu A, Seepo S, Olsen S, et al. The genomic landscape of juvenile myelomonocytic leukemia. Nat Genet. 2015;47:1326–33. https://doi.org/10.1038/ng.3400.
Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, Singh J, Padgett RA, Gu X, Phillips JG, et al. Inherited and somatic defects in DDX41 in myeloid neoplasms. Cancer Cell. 2015;27:658–70. https://doi.org/10.1016/j.ccell.2015.03.017.
Sebert M, Passet M, Raimbault A, Rahme R, Raffoux E, Sicre de Fontbrune F, Cerrano M, Quentin S, Vasquez N, Da Costa M, et al. Germline DDX41 mutations define a significant entity within adult MDS/AML patients. Blood. 2019;134:1441–4. https://doi.org/10.1182/blood.2019000909.
Cheah JJC, Hahn CN, Hiwase DK, et al. Myeloid neoplasms with germline DDX41 mutation. Int J Hematol. 2017;106:163–74. https://doi.org/10.1007/s12185-017-2260-y.
Lewinsohn M, Brown AL, Weinel LM, Phung C, Rafidi G, Lee MK, Schreiber AW, Feng J, Babic M, Chong CE, et al. Novel germ line DDX41 mutations define families with a lower age of MDS/AML onset and lymphoid malignancies. Blood. 2016;127:1017–23. https://doi.org/10.1182/blood-2015-10-676098.
Quesada AE, Routbort MJ, DiNardo CD, Bueso-Ramos CE, Kanagal-Shamanna R, Khoury JD, Thakral B, Zuo Z, Yin CC, Loghavi S, et al. DDX41 mutations in myeloid neoplasms are associated with male gender, TP53 mutations and high-risk disease. Am J Hematol. 2019;94:757–66.
Lindsley RC, Mar BG, Mazzola E, Grauman PV, Shareef S, Allen SL, Pigneux A, Wetzler M, Stuart RK, Erba HP, et al. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood. 2015;125:1367–76. https://doi.org/10.1182/blood-2014-11-610543.
Kim H, D’Andrea AD. Regulation of DNA cross-link repair by the Fanconi anemia/BRCA pathway. Genes Dev. 2012;26(13):1393–408. https://doi.org/10.1101/gad.195248.112.
Lensch MW, Rathbun RK, Olson SB, Jones GR, Bagby GC Jr. Selective pressure as an essential force in molecular evolution of myeloid leukemic clones: a view from the window of Fanconi anemia. Leukemia. 1999;13:1784–9. https://doi.org/10.1038/sj.leu.2401586.
Butturini A, Gale RP, Verlander PC, Adler-Brecher B, Gillio AP, Auerbach AD. Hematologic abnormalities in Fanconi anemia: an International Fanconi Anemia Registry study. Blood. 1994;84:1650–5. https://doi.org/10.1182/blood.V84.5.1650.1650.
Savage SA, Bertuch AA. The genetics and clinical manifestations of telomere biology disorders. Genet Med. 2010;12(12):753–64. https://doi.org/10.1097/GIM.0b013e3181f415b5.
Kirschner M, Maurer A, Wlodarski MW, Ventura Ferreira MS, Bouillon AS, Halfmeyer I, Blau W, Kreuter M, Rosewich M, Corbacioglu S, et al. Recurrent somatic mutations are rare in patients with cryptic dyskeratosis congenita. Leukemia. 2018;32:1762–7. https://doi.org/10.1038/s41375-018-0125-x.
Danilova N, Gazda HT. Ribosomopathies: how a common root can cause a tree of pathologies. Dis Model Mech. 2015;8(9):1013–26. https://doi.org/10.1242/dmm.020529.
Aspesi A, Monteleone V, Betti M, et al. Lymphoblastoid cell lines from Diamond Blackfan anaemia patients exhibit a full ribosomal stress phenotype that is rescued by gene therapy. Sci Rep. 2017;7:12010. https://doi.org/10.1038/s41598-017-12307-5.
Zebisch A, Lal R, Muller M, Lind K, Kashofer K, Girschikofsky M, Fuchs D, Wolfler A, Geigl JB, Sill H. Acute myeloid leukemia with TP53 germ line mutations. Blood. 2016;128:2270–2. https://doi.org/10.1182/blood-2016-08-732610.
McNerney ME, Godley LA, Le Beau MM. Therapy-related myeloid neoplasms: when genetics and environment collide. Nat Rev Cancer. 2017;17:513–27. https://doi.org/10.1038/nrc.2017.60.
Mundschau G, Gurbuxani S, Gamis AS, Greene ME, Arceci RJ, Crispino JD. Mutagenesis of GATA1 is an initiating event in Down syndrome leukemogenesis. Blood. 2003;101:4298–300.
Banno K, Omori S, Hirata K, Nawa N, Nakagawa N, Nishimura K, et al. Systematic cellular disease models reveal synergistic interaction of trisomy 21 and GATA1 mutations in hematopoietic abnormalities. Cell Rep. 2016;15:1228–41.
Lukes J, Danek P, Alejo-Valle O, et al. Chromosome 21 gain is dispensable for transient myeloproliferative disorder driven by a novel GATA1 mutation. Leukemia. 2020;34:2503–8. https://doi.org/10.1038/s41375-020-0769-1.
Hollanda LM, Lima CS, Cunha AF, Albuquerque DM, Vassallo J, Ozelo MC, et al. An inherited mutation leading to production of only the short isoform of GATA-1 is associated with impaired erythropoiesis. Nat Genet. 2006;38:807–12.
Gamis AS, Alonzo TA, Gerbing RB, Hilden JM, Sorrell AD, Sharma M, Loew TW, Arceci RJ, Barnard D, Doyle J, Massey G, Perentesis J, Ravindranath Y, Taub J, Smith FO. Natural history of transient myeloproliferative disorder clinically diagnosed in Down syndrome neonates: a report from the Children’s Oncology Group Study A2971. Blood. 2011;118(26):6752–9. https://doi.org/10.1182/blood-2011-04-350017.
Crispino JD. GATA1 mutations in Down syndrome: implications for biology and diagnosis of children with transient myeloproliferative disorder and acute megakaryoblastic leukemia. Pediatr Blood Cancer. 2005;44:40–4. https://doi.org/10.1002/pbc.20066.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Germline Predisposition to Myeloid Neoplasms
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thompson, C., Ariagno, S. & Kohorst, M.A. Pediatric Germline Predisposition to Myeloid Neoplasms. Curr Hematol Malig Rep 17, 266–274 (2022). https://doi.org/10.1007/s11899-022-00681-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11899-022-00681-5