Abstract
Biofertilizers are a sustainable solution for the adverse biogeochemical impacts exerted by synthetic agrochemicals. The application of biofertilizers is an eco-friendly approach to facilitate plant growth and development. Biofertilizers are mainly composed of living or dormant microbes, which are usually applied to the soil. These microbes encourage plant growth by a wide variety of mechanisms which includes phytohormone production, nutrient solubilization, biocontrol ability, etc. In the present study, bacterial cultures were isolated from nutrient-rich sample sources and screened for plant growth-promoting (PGP) traits. After several PGP trait screenings; two bacterial consortia that could aid the growth and development of plants were formulated. Further, these screening results were validated by applying selected bacterial strains and the prepared consortia on tomato plants (Solanum lycopersium). The growth parameters were evaluated at the plant's germinative, vegetative, and reproductive stages. The biochemical characteristics under the experimental conditions were also examined at various intervals. The results revealed that both our consortia were able to stimulate the growth and development of the host plant. Hence, the formulated bacterial combinations have real potential to become ecologically sustainable biofertilizers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the years, the use of mineral fertilizers to improve crop growth and productivity has been the trend among cultivators. However, excessive use of such chemicals will negatively impact soil fertility, the soil microbiome, and eventually the environment (Gomiero et al. 2011). To this end, agricultural practice is moving toward a more sustainable and environmentally friendly approach. The use of biostimulants in improving genomic and physiological characteristics is a promising practice that was used over the last decades (Katsenios et al. 2021). The application of plant growth-promoting bacteria (PGPB) as a part of mainstream agricultural practice is an example of a bio-stimulant that is becoming a trend.
The bio-stimulation ability of PGPBs is characterized by the production of various substances (for example; IAA, HCN, etc.). These substances participate either in the direct or indirect mechanisms of PGPB action. The bioactivities such as phytohormone production, nutrient solubilization, and nitrogen fixation along with the production of other bioactive compounds that involve in facilitating the resources for plant growth are categorized as direct mechanisms (Glick 2012). Indirect mechanisms are mainly involved in stimulating induced systemic resistance (ISR). The production of antimicrobial compounds such as hydrogen cyanide, phenazines, lipopeptide biosurfactants, and siderophores can all elevate the plant’s immune response toward biotic and abiotic stress. In addition, the production of certain hydrolytic enzymes like protease, chitinases, and glucanases to degrade fungal cell walls or form biofilms on root surfaces that may protect the plant root against soil-borne infections (Vaikuntapu et al. 2014; García-Fraile et al. 2015).
Tomato (Solanum lycopersicum) is one of the most cultivated as well as utilized crops worldwide (Bihon et al. 2022). It is widely cultivated across the tropics and sub-tropic regions around the world (Sadashiva et al. 2017). Currently, global warming along with other associated geochemical problems is resulting in the reduction of more than 50% average yields of major crops including tomatoes (Singh et al. 2018). Therefore, sustainable agricultural practices such as the PGPB application possess greater importance in the current scenario of food security.
In the present study, the biochemical ability of several bacterial strains isolated from nutrient-rich environment samples in improving plant growth and development has been elucidated. These results were further validated by conducting seed germination and a pot culture study with Solanum lycopersicum. Multiple bacterial isolates were found to be showcasing various PGP traits such as the production of phytohormones, biocontrol activity, and nutrient mineralizing ability. In addition, the compatibility of the selected combinations of bacterial isolates was evaluated. The compatible isolates were used to prepare bacterial consortia and were subjected to seed germination and pot culture experiment with Solanum lycopersicum. Furthermore, the physiological and biochemical changes in plants treated with each bacterial formulation were monitored.
Materials and method
Bacterial source sample collection
Soil samples were collected from Kanyakumari Seashore (8°07′30.4"N 77°33′57.1" E) and a lake bank in Vellore- Tamil Nadu (12°58′07.6"N 79°09′32.7" E). These two source samples were selected expecting nutrient solubilizing ability in the native microflora. Samples were collected in plastic bags and brought to Vellore Institute of Technology (VIT), Vellore under cold conditions for isolation of bacteria. Samples were stored at 4 °C until bacteria were isolated for further studies (Gaete et al. 2020).
Isolation and selection of bacteria
Each soil sample was weighed for 1 g and was serially diluted in 0.85% NaCl (saline). 100 µL of each 10–3, 10–4, and 10–5 dilutions were used to spread the plate on Luria–Bertani (LB) agar plates. The plates were incubated at 37 °C. Isolates with dissimilar morphology were picked from the plates and were used to prepare pure culture (Vaikuntapu et al. 2014).
Screening for plant growth-promoting traits
Indole-3- acetic acid (IAA) production: quantitative microplate assay
A total of 15 distinctive pure isolates were subjected to quantifying IAA production as described by Sarwar and Kremer (Sarwar and Kremer 1995). Bacterial isolates were incubated at 28 °C in LB broth containing 0.1% tryptophan. After 48 h the broth was centrifuged at 6000 × g for 10 min. To 150 µL supernatant, 100 µL Salkowski reagent (57 mL of 60% of perchloric acid, 41 mL distilled water, and 2 ml of 0.5 M ferric chloride) were added in a 96-well microplate and was allowed to react for 30 min. The color intensity was measured at 530 nm (Sarker and Al-Rashid 2013).
Siderophore production
The siderophore production ability of 15 individual strains was determined using a method described by Schwyn and Neilands (Schwyn and Neilands 1987) with slight modifications. Chrome Azurol S reagent was prepared by mixing 100 ml CAS (121 mg in 100 mL), 20 mL 1 mM FeCl3 (prepared using 10 mM HCl), and 80 mL hexadecyl trimethyl ammonium bromide (HDTMA) solution (729 mg HDTMA in 400 mL distilled water). The LB agar media was autoclaved and mixed with CAS reagent (10 mL per 100 mL LB agar) and poured into the plates. 4 wells were created in each plate. 20 µL of overnight cultures were injected into the well. The formation of a yellow-colored halo zone around the inoculated wells indicates the bacterial production of siderophore.
ACC Deaminase production: quantitative microplate assay
1-aminocyclopropane- 1-carboxylate deaminase (ACC deaminase), an enzyme that actively participates in lowering ethylene levels in the plant is quantified according to Li et al. (Li et al. 2011) protocol. The 15 overnight bacterial cultures were inoculated to Dworkin and Foster (DF) minimal salt medium with 0.05 ML−1 ACC (aminocyclopropane-1-carboxylic acid) and without ACC as control. The broth cultures were centrifuged at 8000 × g for 5 min. 100 µL supernatant was diluted with 900 µL DF medium; from which 60 µL was mixed with 120 µL ninhydrin reagent (500 mg ninhydrin, 15 mg ascorbic acid in 60 mL of ethylene glycol, 60 mL of citrate buffer pH 6) in a PCR tube and mixed by pipetting. These PCR tubes were then heated at 100 °C in the water bath for 30 min. After allowing the samples to cool, it was transferred to a 96-well microplate. The optical density was measured at 530 nm.
Ammonia production
Bacterial strains were inoculated in 3 mL of peptone incubated overnight at 37 °C for 24 h. After incubation 600 µL Nessler’s reagent was added to each test tube. The production of yellow to dark brown color was considered to be positive for ammonia production. The bacterial ammonia production was then quantified using a spectrophotometer at 450 nm. Different concentrations of ammonium sulfate ranging from 100 µM to 500 µM were used to prepare the standard curve (Abdelwahed et al. 2022).
Phosphate solubilization
All 15 bacterial isolates were subjected to a phosphate solubilization assay. 10 µL of each bacterial culture was spot inoculated in National Botanical Research Institutes’-bromophenol blue (NBRIP-BPB) growth medium plate (D-glucose,10 g; Ca3(PO4)2, 5 g; MgCl2.6H2O, 5 g; MgSO4.7H2O, 0.25 g; KCl, 0.2 g; (NH4)2SO4, 0.1 g; bromophenol blue, 0.025 g; agar, 15 g; and distilled water, 1L, pH-7.0) and incubated at 37 °C for 7 days. The formation of a clear halo zone around each colony was closely monitored. The halo diameter (HD) and colony diameter (CD) of each colony was measured and the ratio of HD/CD was calculated. A bacterial culture with an HD/CD ratio greater than or equal to 1.5 is considered to have a strong ability to solubilize phosphate, whereas those cultures having a value between 1.0 to 1.5 are considered to be weak phosphate solubilizers (Li et al. 2019).
Zinc solubilization
The selected bacterial isolates were screened for their ability to solubilize the insoluble zinc compound ZnO. Overnight cultures were spot inoculated in tris minimal media (per liter: Tris–HCl 6.06 g; NaCl 4.68 g; KCl 1.49 g; NH4Cl 1.07 g; Na2SO4 0.43 g; MgCl2.2H2O 0.2 g; CaCl2.2H2O, 30 mg, pH 7.0; added 0.1% ZnO (w/v) and 1.5% agar. After inoculation, the plates were incubated in dark conditions at 30 °C for 14 days (Sharma et al. 2011; Kamran et al. 2017). The formation of a clear halo zone indicates zinc solubilization.
Potassium solubilization
The ability of the strains to solubilize potassium was analyzed by inoculating them in the Aleksandrov agar medium (Himedia). Spot inoculation of 10 µL of each bacterial culture into Aleksandrov agar was done and incubated at 30 °C for 3–4 days (RAJAWAT et al. 2016). The formation of the halo zone indicated potassium solubilization.
Hydrogen Cyanide (HCN) production
All the 15 isolates were screened for hydrogen cyanide production following the method described by Lorck (1948). Each isolate was streaked on nutrient agar medium supplemented with glycine (4.4 g/L). The lid of the agar plate was covered with a Whatman number 1 filter paper (Previously soaked in a 0.5% picric acid and 2% sodium carbonate w/v solution). Plates were sealed with parafilm and incubated at room temperature for 4 days. The appearance of orange or red coloration in the paper indicates the production of hydrogen cyanide.
Protease production
Skim milk agar plates prepared by adding 1% sterile skim milk powder to autoclaved nutrient agar were used to detect the bacterial production of proteases. The spot-inoculated plates were incubated overnight at room temperature. The formation of the halo zone around the spot inoculation indicated the bacterial ability to produce protease (Yasmin et al. 2020).
Compatibility test and consortia preparation
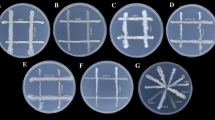
The isolates KC-5, KC-6, VLC-4, and VLC-6 showed superior qualities in PGP screening. Therefore, these isolates were selected for further consortia preparation and plant study. These isolates were streaked on nutrient agar plates in such a way that every culture will cross each other. The plates were incubated at 37 °C for 48 h and the zone of inhibition around each bacterial colony meeting point was observed and recorded (Prasad and Babu 2017).
Biochemical and molecular identification of the bacterial isolates
The DNA of each selected strain was isolated using G. HCl (Guanidine Hydrochloride) method (Gupta et al. 2013). Later, 16 s rRNA gene was amplified using a universal primer sequence set (27F and 1492R). The polymerase chain (PCR) reaction was carried out under the condition of initial denaturation 95° C for 5 min, 35 cycles of 95° C for 30 s, 50° C for 30 s, 72° C for 30 s and final extension of 72° C for 10 min. The PCR product was subjected to Sanger sequencing, and obtained sequences were blasted for retrieving similar sequences (Senthilraj et al. 2016). These similar sequences were used for constructing a phylogenetic tree and further identification of the isolates (Ayyaz et al. 2016).
Biochemical tests such as indole, Methyl Red (MR), Voges Proskauer (VP), citrate, catalase, and oxidase tests along with Gram’s staining were performed to conform to the lineage of the selected isolates (Chauhan and Jindal 2020).
Biofilm formation
The ability of each isolate to form biofilm was tested following the method described by O'Toole and Kolter (O’Toole and Kolter 1998) with slight modifications. The 1% of overnight bacterial cultures were grown in a 24-well microtiter plate containing LB broth media. After 72 h the broth was removed and the adhered bacterial cells were washed with sterilized LB media. The washed bacterial cells were stained using 1% crystal violet and incubated for 15 min. The dye was discarded and the cells were rinsed with distilled water. The dye absorbed by the biofilm formed was then dissolved with 70% ethanol and subsequently quantified with a UV spectrophotometer at 590 nm (Yousef-Coronado et al. 2008; Santiago et al. 2017).
Antibiotic resistance
The selected bacterial isolates were tested for antibiotic resistance against antibiotics namely Rifampicin (10 mcg), Ampicillin (10 mcg), Erythromycin (15 mcg), Kanamycin (30 mcg), Tetracycline (30 mcg), and Chloramphenicol (30 mcg) by the antibiotic sensitivity assay. The bacterial cultures were swabbed on Mueller Hinton agar (MHA) media plates. The antibiotic discs (Himedia, India) were placed above the media surface. The plates were incubated at 37 °C for 24 h. A zone of bacterial growth inhibition around the disc implies the sensitivity of the bacteria toward the respective antibiotic. The diameter of the zone was measured subsequently (Prakash Singh and Nath Jha 2015).
Seed germination study
The seed germination study was carried out following the procedure described by Vasseur-Coronado et al. (Vasseur-Coronado et al. 2021) with slight modifications. The isolates namely, KC-5, KC-6, and VLC-4 were used for this study as individual and as consortia along with commercially available Pseudomonas sp. based biofertilizer control- bought from farmer’s biofertilizers (https://www.farmersbiofertilizers.com/). The bacterial cultures for the seed treatment were prepared by inoculating the individual cultures and the consortia in nutrient broth. Overnight cultures were pelleted out by centrifugation. These pellets were dissolved in PBS in such a way that it contains 0.5 OD cells @ 630 nm. Subsequently, the PKM-1tomato (Solanum lycopersicum) variety seeds were disinfected by submerging them in the following solutions: 70% ethanol for 2 min, and 1% sodium hypochlorite (NaOCl) (SDfine, India) for 5 min. These seeds were then subjected to washing with sterile distilled water multiple times. The disinfected seeds were then soaked in either the respective bacterial formulations or in PBS (control group) for 30 min under shaking. The treated seeds are then placed in germination paper moistened with sterile distilled water (25 times its weight). A total of 12 seeds were placed in each germination paper in triplicates for each treatment group. Petri dishes were incubated at 25–30 °C. After 12 days, the physiological parameters were measured and the vigor index was calculated. The seed vigor index was calculated by applying the following formula.
Pot culture study- physiology and biochemical analysis
The bacterial cultures were prepared by inoculating the individual cultures and the consortia in nutrient broth. After 72 h at 30° C, the obtained cells were pelleted out by centrifuging. These pellets were then washed 3 times in sterile 0.85% NaCl and the concentration of the cells was appropriated to 0.6 OD at 600 nm using the same saline solution. The prepared bacterial suspensions were used to treat the tomato seedlings (Helal et al. 2022).
PKM-1 tomato (Solanum lycopersicum) variety seeds were sown and allowed to grow in the seedling tray for one month. The one-month-old seedlings were transplanted into polythene grow bags. Before transplanting, the roots of the seedlings were washed with water to remove the soil around them. These seedlings were then treated with bacterial cultures (control-no treatment, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and commercially available Pseudomonas sp. based biofertilizer control- bought from farmer’s biofertilizers (https://www.farmersbiofertilizers.com/) by soaking the roots in the respective bacterial suspension. After transplantation, the plant grow bags was kept in a completely randomized design with 5 replications in each treatment including control. The transplanted plants were again treated with the respective bacterial culture treatments after 15 days of treatment.
Plant-Physiological analysis
Physiological parameters like the shoot length, number of branches, and the number of leaves, per plant, were measured in different intervals after transplantation until the stage of reproduction.
Plant-Biochemical analysis
Biochemical analyses such as the quantification of carbohydrates, protein, phenol, and pigments were done on the 30th, 45th, and 60th day after transplantation.
Estimation of carbohydrates
Fresh tomato leaves weighing 40 mg were picked from each treatment group, washed with distilled water, and squashed in a mortar and pestle with 2 mL distilled water. The mixture was heated at 70° C for 20 min followed by centrifugation at 5000 × g at 4o C for 15 min (Karthik et al. 2020). The resulting supernatant was then used for the quantification of carbohydrates by using a phenol–sulphuric acid assay (Chow and Landhäusser 2004) in a 96-well microtiter plate.
To 100 µL of each leaf extract 100 µL of 5% phenol was added along with 500 µL of concentrated sulphuric acid in ice-cold conditions. The mixture was then brought to room temperature by heating it in a hot water bath for 20 min. The samples were then allowed to cool and the absorbance was measured @ 490 nm. Glucose standard concentrations ranging from 0.025 to 2 mg/mL were used to plot the absorbance against the concentration standard curve. Using the standard graph, the unknown carbohydrate concentration in plant samples was estimated. The experiment was done in replications and the mean value was considered.
Estimation of protein
Fresh leaf samples weighing 40 mg were ground with 3 mL water in a mortar and pestle. The mixture was centrifuged at 14000 × g at 4o C for 10 min (Mathur et al. 2015). The resulting supernatant was used for the estimation of protein using the BCA protein quantification assay kit (TAKARA, INDIA) following manufacturer protocol. Bovine Serum Albumin (BSA) concentrations ranging from 0.025 to 2 mg/mL were used to plot absorbance against the concentration standard curve. The experiment was done in triplicates and the mean value was considered (Sun et al. 2007).
Estimation of phenol
Fresh leaf samples weighing 40 mg were crushed with 2 mL of 80% ethanol followed by the centrifugation of the mixture at 14,000 × g at 4o C for 20 min. The supernatant was separated and used as the leaf extract for the estimation of phenol. Phenol in the sample was quantified using Folin Ciocalteau (FC) reagent (Maheswari and Elakkiya 2014). In a 1.5 ml tube, 100 µL of each sample and 200 µL of FC reagent were added and mixed. The mixture was then neutralized with 400 µL 7.5% sodium carbonate and was incubated at room temperature for 30 min with shaking for color development. 200 µL of the reaction mixture was transferred into a 96-well plate. The absorbance of the resulting blue color was measured at 725 nm. Gallic acid concentrations ranging from 0.025 to 2 mg/mL were used to plot absorbance against the concentration standard curve. The experiment was done in triplicates and the mean value was considered.
Estimation of pigments
Leaf samples weighing 100 mg were ground with 10 mL of 80% acetone in a mortar and pestle. The mixture was then centrifuged at 14,000 × g at 4o C for 10 min. The optical density of the supernatant was analyzed at 645 nm, 663 nm, and 470 nm. Based on the OD values obtained the concentrations of chlorophyll a (Chl A), chlorophyll b (Chl B), and carotenoids were calculated (Sarkar et al. 2018). The experiment was done in triplicates and the mean value was considered.
Results
Isolation and selection of bacterial cultures
Each sample source was selected for scientific reasons. A total of 15 morphologically distinctive bacterial isolates were selected from both sample sources for the screening of PGP properties.
Screening for plant growth-promoting traits
Indole-3- acetic acid (IAA) production: quantitative microplate assay
The ability of the selected 15 isolates in producing IAA, siderophore, and ACC deaminase was investigated predominately. Out of the 15 strains; KC-5 (87.93 µg mL−1), and KC-7 (77.18 µg mL−1) showed the best performance in IAA production. However, KC-3 (15.67 µg mL−1), VLC-4 (15.26 µg mL−1), and VLC-8 (11.17 µg mL−1) were found to be moderate IAA producers (Fig. 1).
Siderophore production
The CAS-qualitative agar test for siderophore production revealed a yellow zone around siderophore-producing bacterial strains (Fig. 2). Isolates namely- KC-2 and KC-5 showed the formation of a strong halo after 24 h of incubation. Isolates like KC-1, KC-4, KC-6, VLC-1, VLC-4, and VLC-6 showed gradual zone formation from mild to strong in the 48 h period. The rest of the bacterial isolates showed no or negligible zone formation implying the inability of these strains to produce siderophore.
ACC Deaminase production: quantitative microplate assay
The PCR-plate ninhydrin ACC assay determined the ability of the selected 15 cultures in producing ACC deaminase. All 15 isolates were found to be having some ACC deaminase activity as the average final ACC concentration in the broth after 24 h incubation was around 5 mML−1 (Fig. 3). However, certain isolates like VLC-4, VLC-6, KC-2, KC-3, KC-5, and KC-6 showed better ACC utilization (final ACC concentration 1.5mML−1 or below), implying better performance for ACC deaminase production.
Ammonia production
The qualitative test showed that 8 isolates tested were positive for ammonia production (Fig. 4). All isolates except KC-5 and KC-7 showed an average ammonia production of 150 µM.
Phosphate solubilization
Among the 15 isolates that were subjected to phosphate solubilization assay, 5 of the isolates showed superiority in mineralizing phosphate (HD/CD above 1.5). These 5 isolates (namely KC-7, VLC-4, VLC-6, KC-2, and KC-6) were then closely monitored for their solubilizing efficiency (Fig. 5). The isolates CSC-1 and KC-6 showed a steady growth rate of phosphate solubilizing halo (HD/CD) over the 72 h period. VLC-6 showed a third-day spike in the halo, implying the gradual attaining of the mineralization target. KC-7 and KC-2 showed no specific growth rate of halo over time, even though KC-2 showed a better HD/CD ratio. VLC-4 seemed to end its phosphate mineralization on the second day since there was not much change observed in the halo.
Zinc solubilization
Here we screened the selected cultures for their ability to solubilize ZnO. Out of the 15 isolates screened for zinc solubilization in ZnO-incorporated tris minimal media, only VLC-6 was found to show positive zone formation (Fig. 6).
Potassium solubilization
The zone formed around the cultures suggested that 4 (namely, KC-7, VLC-4, VLC-6, and KC-6) out of the 15 isolates were found to have the ability to solubilize the insoluble form of potassium (Potassium aluminum silicate) in the Aleksandrov medium (Fig. 7).
HCN production
The picric acid dye present in the filter paper undergoes molecular rearrangement when it reacts with the HCN produced by the bacteria- changing its coloration from yellow to orange. Among the 15 isolates screened only VLC-6 was found to be able to produce HCN (Fig. 8).
Protease production
Out of the 15 isolates screened for protease production, VLC-4 was found to be the only isolate that produces protease (Fig. 9).
Identification of the bacterial isolates
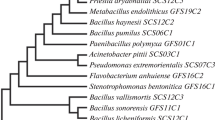
The constructed phylogenetic trees revealed that the bacterial isolate VLC-4, KC-5, KC-6 and VLC-6 were closer to Enterobacter sp., Lysinibacillus sp., Acinetobacter sp. and Pseudomonas aeruginosa respectively (Fig. S1). The sequences of the respective isolate were then submitted to the NCBI database and accession numbers were retrieved as follows; Enterobacter Sp: OP050456, Lysinibacillus sp.: OP070953, Acinetobacter sp.: OP080714 and Pseudomonas aeruginosa: OP055908. The biochemical Characterization of the isolates (Table 1) further validated the molecular identification.
Biofilm formation
Among the isolates KC-5, KC-6, VLC-4, and VLC-6; VLC-6 and KC-5 were seen to be having a greater ability to form a biofilm with 0.4 and 0.15 OD respectively. However, KC-6 and VLC-4 are moderate biofilm producers (Fig. 10).
Antibiotic resistance
VLC-6 was seen to be resistant to most of the antibiotics which were tested. VLC-4 and KC-6 were showing moderate susceptibility to most of the antibiotics. KC-5 was susceptible to all the antibiotics tested (Table 2).
Compatibility test and consortia preparation
KC-5, KC-6, VLC-4, and VLC-6 were subjected to compatibility tests in different combinations, among which VLC-6 was seen inhibiting the growths of KC-5 & KC-6. However, KC-5 + KC-6, KC-5 + VLC-4, and VLC-4 + VLC-6 were compatible with each other (Fig. 11). KC-5 + KC-6 and KC-5 + VLC-4 were selected for further plant study as VLC-6 was identified to be Pseudomonas aeruginosa which is reported to be a human pathogen.
Seed germination study
The measurement of shoot length, root length, and seedling length was noted down (Table 3). Subsequently the vigor index was also calculated to characterize the impact of the bacterial treatments on the germination of tomato seeds. The results showcased that all the bacterial treatments considerably promoted ‘germination’ when compared with the control group (Fig. S2). The highest vigor index was found to be shown by the KC-6 (983.98) and biofertilizer control group (957.27). However, the formulated consortia KC-5 + KC-6 and KC-5 + VLC-4 showed average performance with a vigor index of 785.80 and 700.31, respectively.
Pot culture study- physiology and biochemical analysis
Physiological analysis
The measurement of plant height, number of branches, and number of leaves on the 10th, 20th, 30th, and 50th days were observed to understand the influence of bacterial treatments on plant growth (Table 4). The results suggested that the application of the consortium KC-5 + KC-6 and the biofertilizer control have the highest values with statistically significant (p < 0.05) differences compared to all other treatment groups (Fig. S3). Moreover, all PGPB treatments had statistically significant differences compared to that of the control.
Biochemical analysis
Carbohydrates
The concentrations of carbohydrates were better in all the treatment groups when compared to the control group at each of the time intervals the experiment was conducted (Fig. 12). On the 30th day, to a 0.264 mg/mL of control; 0.544, 0.563, 0.301, 0.574, 0.384, and 0.611 mg/mL carbohydrates were present in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values changed to a 0.38 mg/mL control; 0.38, 0.485, 0.465, 0.47, 0.59, 0.58, and 0.615 mg/mL in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of the 60th day, the values were; to a 0.279 mg/mL control; 0.362, 0.343, 0.376, 0.515, 0.460, and 0.544 mg/mL in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
Protein
The protein content in each treatment group was estimated to be similar on the 30th day (Fig. 12). A considerable change in protein content was observed on days 45 and 60. On the 30th day, the values were 2.043, 1.962, 1.981, 1.908, 2.138, 2.119, and. 2.172 mg/mL in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values were 0.4471, 0.930, 1.456, 0.585, 0.807, 1.031, and 0.982 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of 60th day, the protein was estimated to be 1.742, 1.952, 1.835, 1.843, 1.955, 1.897, and 1.773 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
Phenol
The phenol content steadily increased with each culture treatment when compared to the control group on each of the 3-time intervals the experiment was performed (Fig. 12). On the 30th day, to a 0.105 mg/mL of control; 0.145, 0.115, 0.136, 0.125, 0.144, and 0.131 mg/mL carbohydrates were present in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values changed to a 0.484 mg/mL control; 0.753, 0.832, 0.730, 0.573, 0.572, and 0.869 mg/mL in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of the 60th day, the values were; to a 0.091 mg/mL control; 0.258, 0.291, 0.190, 0.379, 0.255, and 0.402 mg/mL in KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
Pigments
The estimated Chlorophyll a, chlorophyll b, and carotenoid contents in all the treatment groups showed a significant increase influence of the treated isolates in each of the 3-time intervals except on a few occasions (Fig. 13). On the 30th day, the values of chl A were 0.494, 0.762, 0.523, 0.612, 0.502, 0.575, 0.466 mg/mL in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values were 0.348, 0.638, 0.697, 0.527, 0.682, 0.616, and 0.299 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of the 60th day, the chl A was estimated to be 0.355, 0.601, 0.563, 0.456, 0.633, 0.479, and 0.533 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
On the 30th day, the values of chl B were 0.884, 1.180, 0.918, 0.692, 1.050, 1.01, and 0.887 mg/mL in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values were 0.651, 1.183, 1.184, 0.601, 1.377, 1.197, and 0.930 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of the 60th day, the chl B was estimated to be 0.668, 1.107, 1.040, 0.861, 1.134, 0.910, and 0.966 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
On the 30th day, the values of carotenoids were found to be 0.442, 0.657, 0.432, 0.766, 0.587, 0.498, and 0.386 mg/mL in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. On the 45th day, the values were 0.419, 0.538, 0.579, 0.386, 0.684, 0.522, and 0.438 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively. In the case of the 60th day, the carotenoids were estimated to be 0.415, 0.542, 0.535, 0.455, 0.567, 0.429, and 0.538 in control, KC-5, KC-6, VLC-4, KC-5 + KC-6, KC-5 + VLC-4, and biofertilizer control respectively.
Discussion
The positive relationship of plants with the soil microbiome is studied extensively through the years to be having a potential future in the sustainability of the agricultural sector (Stamenković et al. 2018). Moreover, any PGPBs related scientific reports have suggested that tomato is a suitable crop for studying plant-microbial interactions (Ahirwar et al. 2015; Katsenios et al. 2021).
The results obtained in this study also demonstrated that the application of both the formulated PGPB consortia enhanced the growth and development of Solanum lycopersicum. In the present study, we isolated bacteria from nutrient-rich soil samples collected from Kanyakumari and Vellore. A total of 15 distinctive bacterial isolates were selected from the spread plates. These isolates were screened for PGPB characteristics such as the production of IAA, Siderophore, ACC deaminase, ammonia, HCN, and protease; solubilization of Phosphate, potassium, and zinc.
IAA (3-indoleacetic acid) is a plant hormone that is commonly known as auxin. Auxin has an influential role in plant growth and development; majorly in vascular differentiation, root architecture, and disease signaling. Plants generally produce this phytohormone in its various parts, while the soil microflora also contributes its share (Duca et al. 2014; Saxena 2014). In this context, the application of our bacterial combinations may have provided an additional supply of IAA which may have supported the growth and development of the host Solanum lycopersicum. Similarly, the production of siderophore -an iron-chelating compound that aids the plant in iron absorption, was evaluated. Siderophore is also a known anti-fungal agent and can also help bacteria to colonize the rhizosphere (Saha et al. 2016). 1-aminocyclopropane-1-carboxylic acid (ACC) is a precursor molecule of ethylene (plant stress-responsive hormone- that participates in stress-responsive inhibition of plant growth). ACC deaminase enzyme catalyzes the conversion of ACC to ammonia and α-ketobutyrate. Bacterial ACC deaminase promotes plant growth by reducing plant ACC level and eventually ethylene level thereby ameliorating plant growth inhibition encountered due to ethylene production (Moon and Ali 2022). Ammonia production by bacteria is associated with nitrogen supplementation to the host plants (Bhattacharyya et al. 2020). At times certain nutrients present in the soil may be in an unavailable (insoluble) form to plants. Some genera of bacteria can mineralize this unavailable form of such nutrients to the available form (Granada et al. 2018). Here in this study, we evaluated the ability of our isolates in solubilizing nutrients like phosphate, zinc, and potassium. The production of protease, HCN (Hydrogen Cyanide), and siderophore; and the formation of biofilm are associated with the bacterial aiding of plants for ISR. Protease is an enzyme that hydrolyses proteins and peptides which compose the cell wall of plant pathogens. Likewise, HCN is also a biocontrol agent as it is ascribed to be toxic against plant pathogens (Yasmin et al. 2020). Bacterial biofilm take part in biocontrol of phytopathogens. In addition, they could also enhance nutrient availability to plants and mitigate abiotic stress (Haque et al. 2020). Therefore, the application of bacterial cultures that produce the above-mentioned compounds is expected to benefit the growth and development of plants.
The isolates KC-5, KC-6, VLC-4, and VLC-6 were short-listed by the screening and they were subsequently found to be closely related to NR_042073 Lysnibacillus sphaericus (94.99% similarity), NR_117931.1 Acinetobacter nosocomialis (95.13% similarity), NR_044978.1 Enterobacter cloacae subsp. dissolvens (95.66% similarity), and NR_113599.1 Pseudomonas aeruginosa (96.81% similarity) respectively (Fig. S1). Species level identification of KC-5, KC-6 and VLC-4 was not possible, since the blast similarity index was below the threshold (97%)(Aw et al. 2016; Nguyen et al. 2016). However, the similarity index of VLC-6 was nearly the threshold value. Moreover, it showed the biochemical characteristics of Pseudomonas aeruginosa. Thus, confirming the molecular identification.
The compatibility test for each of these four bacterial isolates was carried out in different combinations. VLC-6 was found to inhibit the growth of KC-5 and KC-6. Henceforth, KC-5 + KC-6 and KC-5 + VLC-4 combinations were selected for the further plant application study.
The positive impact of our consortia on the germination and vigor of tomato seeds was evaluated by a seed germination study. All bacterial treatment formulations at least doubled the seed vigor index when compared to the control group. However, isolate KC-6 was found to promote seed vigor when treated individually rather than as a consortium. This phenomenon may have happened as in some instances PGPB mixtures do not play synergistic roles in growth promotion (Kang et al. 2014; Lobo et al. 2022).
Plants inoculated with both of our combinations showed better physiological characteristics compared to the control group. However, the application with the consortium KC-5 + KC-6 was found to significantly influence the plant height, number of branches, and number of leaves. This consortium was equally influential in plant physiology like the commercially available biofertilizer control used in this study.
Plant biochemical tests namely, estimation of carbohydrates, protein, phenol, and pigments were performed in 15 days intervals to know the influence of the treated bacterial isolates at different times of growth and development. The concentration of carbohydrates was clearly improved by the application of the formulated consortia. However, KC-5 + KC-6 were found to be stimulating more production of carbohydrates when compared with KC-5 + VLC-4. The 30th-day carbohydrate content suggests that isolate VLC-4 is comparatively weaker improving the carbohydrate content. The applied culture treatments were found not to influence the plants till the 30th day as there was no significant change in any of the treatments including the biofertilizer control when compared to the protein content in control. However, this scenario seems to be changed on 45th and 60th-day protein estimations; as protein content in the control plant was found to be reduced to below 0.5 mg/mL. However, plants applied with bacterial formulations were found to conserve the protein content. Among the culture treatments, KC-6 when applied alone showed the highest influence on protein production. As a consortium KC-5 + VLC-4 was found to be better than its competitor. A clear influence of the culture treatments on the phenol content level was seen across each time interval quantification. However, the influence rate varied from time to time. In the case of pigments, in both the consortia, the formulation was found to be improving all the pigment contents significantly better than the biofertilizer control.
Conclusion
In conclusion, the formulated consortia KC-5 + KC-6 and KC-5 + VLC-4 considerably enhanced the growth and development of Solanum lycopersicum. When compared to an already marketized bacterial bio fertilizer, our consortia were found to be showing equally good performance in various levels of plant growth promotion. Still, the influence of these consortia on the plants is required to be further explored on a molecular level for a complete understanding of their positive effects on plants for sustainable biofertilizer development.
Data availability
The genome sequences of KC-5, KC-6, VLC-4 and VLC-6 strains analysed during the current study are available in the GenBank repository.
[https://www.ncbi.nlm.nih.gov/nuccore/2277478657, https://www.ncbi.nlm.nih.gov/nuccore/2277723799, https://www.ncbi.nlm.nih.gov/nuccore/2276735244, https://www.ncbi.nlm.nih.gov/nuccore/2277100149].
Abbreviations
- PGPB:
-
Plant Growth Promoting Bacteria
- BSA:
-
Bovine Serum Albumin
- NCBI:
-
National Center for Biotechnology Information
- IAA:
-
Indole-3-Acetic Acid
- KC-5:
-
Kanyakumari Culture-5
- KC-6:
-
Kanyakumari Culture -6
- VLC-4:
-
Vellore Lake Culture -4
- BFC:
-
Biofertilizer Control
- HD:
-
Halo Diameter
- CD:
-
Colony Diameter
- HCN:
-
Hydrogen Cyanide
- ACC:
-
1-Aminocyclopropane-1-Carboxylate
References
Abdelwahed S, Trabelsi E, Saadouli I, Kouidhi S, Masmoudi AS, Cherif A, Mnif W, Mosbah A (2022) A new pioneer colorimetric micro-plate method for the estimation of ammonia production by plant growth promoting rhizobacteria (PGPR). Main Group Chem 21:55–68. https://doi.org/10.3233/MGC-210077
Ahirwar NK, Gupta G, Singh V, Rawlley RK, Ramana S (2015) Influence on growth and fruit yield of tomato (Lycopersicon esculentum Mill.) plants by inoculation with Pseudomonas fluorescence (SS5): Possible role of plant growth promotion. Int J Curr Microbiol Appl Sci 4:720–730
Aw YK, Ong KS, Lee LH, Cheow YL, Yule CM, Lee SM (2016) Newly isolated Paenibacillus tyrfis sp. nov., from Malaysian tropical peat swamp soil with broad spectrum antimicrobial activity. Front Microbiol 7:219. https://doi.org/10.3389/FMICB.2016.00219/BIBTEX
Ayyaz K, Zaheer A, Rasul G, Mirza MS (2016) Isolation and identification by 16S rRNA sequence analysis of plant growth-promoting azospirilla from the rhizosphere of wheat. Braz J Microbiol 47:542–550. https://doi.org/10.1016/j.bjm.2015.11.035
Bhattacharyya C, Banerjee S, Acharya U, Mitra A, Mallick I, Haldar A, Haldar S, Ghosh A, Ghosh A (2020) Evaluation of plant growth promotion properties and induction of antioxidative defense mechanism by tea rhizobacteria of Darjeeling, India. Sci Rep 10:1–19. https://doi.org/10.1038/s41598-020-72439-z
Bihon W, Ognakossan KE, Tignegre JB, Hanson P, Ndiaye K, Srinivasan R (2022) Evaluation of Different Tomato (Solanum lycopersicum L.) Entries and Varieties for Performance and Adaptation in Mali, West Africa. Horticulturae 8:579. https://doi.org/10.3390/horticulturae8070579
Chauhan A, Jindal T (2020) Biochemical and molecular methods for bacterial identification. Microbiol Methods Environ Food and Pharm Anal 425–468. https://doi.org/10.1007/978-3-030-52024-3_10
Chow PS, Landhäusser SM (2004) A method for routine measurements of total sugar and starch content in woody plant tissues. Tree Physiol 24:1129–1136. https://doi.org/10.1093/treephys/24.10.1129
Duca D, Lorv J, Patten CL, Rose D, Glick BR (2014) Indole-3-acetic acid in plant-microbe interactions. Antonie Van Leeuwenhoek 106:85–125. https://doi.org/10.1007/s10482-013-0095-y
Gaete A, Mandakovic D, González M (2020) Isolation and identification of soil bacteria from extreme environments of Chile and their plant beneficial characteristics. Microorganisms 8:1–13. https://doi.org/10.3390/microorganisms8081213
García-Fraile P, Menéndez E, Rivas R (2015) Role of bacterial biofertilizers in agriculture and forestry. AIMS Bioengineering 2:183–205. https://doi.org/10.3934/bioeng.2015.3.183
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica 2012:1–15. https://doi.org/10.6064/2012/963401
Gomiero T, Pimentel D, Paoletti MG (2011) Environmental impact of different agricultural management practices: Conventional vs. Organic agriculture. Crit Rev Plant Sci 30:95–124. https://doi.org/10.1080/07352689.2011.554355
Granada CE, Passaglia LMP, de Souza EM, Sperotto RA (2018) Is Phosphate Solubilization the Forgotten Child of Plant Growth-Promoting Rhizobacteria? Front Microbiol 9:2054. https://doi.org/10.3389/FMICB.2018.02054
Gupta SK, Kumar A, Hussain SA (2013) Extraction of PCR-amplifiable DNA from a variety of biological samples with uniform success rate. Conserv Genet Resour 5:215–217. https://doi.org/10.1007/s12686-012-9772-9
Haque MM, Mosharaf MK, Khatun M, Haque MA, Biswas MS, Islam MS, Islam MM, Shozib HB, Miah MMU, Molla AH, Siddiquee MA (2020) Biofilm producing rhizobacteria with multiple plant growth-promoting traits promote growth of tomato under water-deficit stress. Front Microbiol 11:542053. https://doi.org/10.3389/fmicb.2020.542053
Helal DS, El-khawas H, Elsayed TR (2022) Molecular characterization of endophytic and ectophytic plant growth promoting bacteria isolated from tomato plants (Solanum lycopersicum L.) grown in different soil types. J Genet Eng Biotechnol 20:79. https://doi.org/10.1186/s43141-022-00361-0
Kamran S, Shahid I, Baig DN, Rizwan M, Malik KA, Mehnaz S (2017) Contribution of zinc solubilizing bacteria in growth promotion and zinc content of wheat. Front Microbiol 8:2593. https://doi.org/10.3389/fmicb.2017.02593
Kang Y, Shen M, Yang X, Cheng D, Zhao Q (2014) A plant growth-promoting rhizobacteria (PGPR) mixture does not display synergistic effects, likely by biofilm but not growth inhibition. Microbiology (russian Federation) 83:666–673. https://doi.org/10.1134/S0026261714050166
Karthik T, Sarkar G, Babu S, Amalraj LD, Jayasri MA (2020) Preparation and evaluation of liquid fertilizer from Turbinaria ornata and Ulva reticulata. Biocatal Agric Biotechnol 28:101712. https://doi.org/10.1016/j.bcab.2020.101712
Katsenios N, Andreou V, Sparangis P, Djordjevic N, Giannoglou M, Chanioti S, Stergiou P, Xanthou MZ, Kakabouki I, Vlachakis D, Djordjevic S, Katsaros G, Efthimiadou A (2021) Evaluation of plant growth promoting bacteria strains on growth, yield and quality of industrial tomato. Microorganisms 9:1–17. https://doi.org/10.3390/microorganisms9102099
Li Z, Chang S, Lin L, Li Y, An Q (2011) A colorimetric assay of 1-aminocyclopropane-1-carboxylate (ACC) based on ninhydrin reaction for rapid screening of bacteria containing ACC deaminase. Lett Appl Microbiol 53:178–185. https://doi.org/10.1111/j.1472-765X.2011.03088.x
Li Y, Zhang J, Zhang J, Xu W, Mou Z (2019) Characteristics of inorganic phosphate-solubilizing bacteria from the sediments of a eutrophic lake. Int J Environ Res Public Health 16:2141. https://doi.org/10.3390/ijerph16122141
Lobo LLB, da Silva MSRDA, Carvalho RF, Rigobelo EC (2022) The negative effect of coinoculation of plant growth-promoting bacteria is not related to indole-3-acetic acid synthesis. J Plant Growth Regul 42:2317–2326. https://doi.org/10.1007/S00344-022-10706-1/METRICS
Lorck H (1948) Production of hydrocyanic acid by bacteria. Physiol Plant 1:142–146. https://doi.org/10.1111/J.1399-3054.1948.TB07118.X
Maheswari NU, Elakkiya T (2014) Effect of Liquid Biofertilizers on Growth and Yield of Vigna mungo L. Int J Pharm Sci Rev Res 29:42–45
Mathur C, Rai S, Sase N, Krish S, Jayasri MA (2015) Enteromorpha intestinalis Derived Seaweed Liquid Fertilizers as Prospective Biostimulant for Glycine max. Braz Arch Biol Technol 58:813–820. https://doi.org/10.1590/S1516-89132015060304
Moon YS, Ali S (2022) Possible mechanisms for the equilibrium of ACC and role of ACC deaminase-producing bacteria. Appl Microbiol Biotechnol 106:877–887. https://doi.org/10.1007/S00253-022-11772-X
Nguyen NP, Warnow T, Pop M, White B (2016) A perspective on 16S rRNA operational taxonomic unit clustering using sequence similarity. npj Biofilms Microbiomes 2(1):1–8. https://doi.org/10.1038/npjbiofilms.2016.4
O’Toole GA, Kolter R (1998) Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: a genetic analysis. Mol Microbiol 28:449–461. https://doi.org/10.1046/J.1365-2958.1998.00797.X
Prasad AA, Babu S (2017) Compatibility of Azospirillum Brasilense and Pseudomonas Fluorescens in Growth Promotion of Groundnut (Arachis Hypogea L.). Anais Da Academia Brasileira De Ciencias 89:1027–1040. https://doi.org/10.1590/0001-3765201720160617
Rajawat MVS, Singh S, Tyagi SP, Saxena AK (2016) A modified plate assay for rapid screening of potassium-solubilizing bacteria. Pedosphere 26:768–773. https://doi.org/10.1016/S1002-0160(15)60080-7
Sadashiva AT, Hanson P, Reddy KM, Ravishankar KV, Prasad M, Prasanna HC, Reddy KM, Singh TH, Saritha RK, Hussain Z, Mythili JB, Shivashankara KS, Bhatt RM, Laxman RH, Tiwari RB, Sridhar V, Sowmya V, Kumar NP, Kumar M, Kaushal A, Rai AK, Jatav V, Bhat L (2017) Breeding tomato ( Solanum lycopersicum L.) for resistance to biotic and abiotic stresses. J Hortl Sci 12:91–105
Saha M, Sarkar S, Sarkar B, Sharma BK, Bhattacharjee S, Tribedi P (2016) Microbial siderophores and their potential applications: a review. Environ Sci Pollut Res 23:3984–3999. https://doi.org/10.1007/s11356-015-4294-0
Santiago CD, Yagi S, Ijima M, Nashimoto T, Sawada M, Ikeda S, Asano K, Orikasa Y, Ohwada T (2017) Bacterial compatibility in combined inoculations enhances the growth of potato seedlings. Microbes Environ 32:14. https://doi.org/10.1264/JSME2.ME16127
Sarkar G, Jatar N, Goswami P, Cyriac R, Suthindhiran K, Jayasri MA (2018) Combination of different marine algal extracts as biostimulant and biofungicide. J Plant Nutr 41:1163–1171. https://doi.org/10.1080/01904167.2018.1434201
Sarker A, Al-Rashid J (2013) Analytical Protocol for determination of Indole 3 acetic acid (IAA) production by Plant Growth Promoting Bacteria (PGPB). Technical report of Quantification of IAA by microbes, pp 1–4
Sarwar M, Kremer RJ (1995) Determination of bacterially derived auxins using a microplate method. Lett Appl Microbiol 20:282–285. https://doi.org/10.1111/j.1472-765X.1995.tb00446.x
Saxena S (2014) Microbial metabolites for development of ecofriendly agrochemicals. Allelopath J 33:1–24
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
Senthilraj R, Prasad GS, Janakiraman K (2016) Sequence-based identification of microbial contaminants in non-parenteral products. Braz J Pharm Sci 52:329–336. https://doi.org/10.1590/S1984-82502016000200011
Sharma SK, Sharma MP, Ramesh A, Joshi OP (2011) Characterization of zinc-solubilizing Bacillus isolates and their potential to influence zinc assimilation in soybean seeds. J Microbiol Biotechnol 22:352–359. https://doi.org/10.4014/jmb.1106.05063
Singh RP, Jha PN (2015) Original research article plant growth promoting potential of ACC Deaminase Rhizospheric Bacteria Isolated from Aerva javanica: A Plant Adapted to Saline Environments. Int J Curr Microbiol App Sci 4(142):152
Singh VK, Singh AK, Singh PP, Kumar A (2018) Interaction of plant growth promoting bacteria with tomato under abiotic stress: A review. Agr Ecosyst Environ 267:129–140. https://doi.org/10.1016/J.AGEE.2018.08.020
Stamenković S, Beškoski V, Karabegović I, Lazić M, Nikolić N (2018) Microbial fertilizers: A comprehensive review of current findings and future perspectives. Span J Agric Res 16:1–18. https://doi.org/10.5424/sjar/2018161-12117
Sun HJ, Kataoka H, Yano M, Ezura H (2007) Genetically stable expression of functional miraculin, a new type of alternative sweetener, in transgenic tomato plants. Plant Biotechnol J 5:768–777. https://doi.org/10.1111/J.1467-7652.2007.00283.X
Vaikuntapu PR, Dutta S, Samudrala RB, Rao VRVN, Kalam S, Podile AR (2014) Preferential Promotion of Lycopersicon esculentum (Tomato) Growth by Plant Growth Promoting Bacteria Associated with Tomato. Indian Journal of Microbiology 54:403–412. https://doi.org/10.1007/S12088-014-0470-Z
Vasseur-Coronado M, du Boulois HD, Pertot I, Puopolo G (2021) Selection of plant growth promoting rhizobacteria sharing suitable features to be commercially developed as biostimulant products. Microbiol Res 245:126672. https://doi.org/10.1016/j.micres.2020.126672
Yasmin H, Naz R, Nosheen A, Hassan MN, Ilyas N, Sajjad M, Anjum S, Gao X, Geng Z (2020) Identification of new biocontrol agent against charcoal rot disease caused by Macrophomina phaseolina in Soybean (Glycine max L.). Sustainability 12:6856. https://doi.org/10.3390/SU12176856
Yousef-Coronado F, Travieso ML, Espinosa-Urgel M (2008) Different, overlapping mechanisms for colonization of abiotic and plant surfaces by Pseudomonas putida. FEMS Microbiol Lett 288:118–124. https://doi.org/10.1111/j.1574-6968.2008.01339.x
Acknowledgements
The authors thank management of the Vellore Institute of Technology for providing ’VIT SEED GRANT (RGEMS)-Sanctioned order NO: SG20230005 for carrying out this work.
Funding
The authors confirm that this research article was not written under the financial sponsorship of any funding agency.
Author information
Authors and Affiliations
Contributions
A T Naveen conceived, designed and performed the research experiments and wrote the manuscript under the direction of R Bharath, R Vidya and R Sudhakaran. Rajan, Kittichon U-taynapun, and Nion Chirapongsatonkul proofread and validated the results and analysis. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that there were no competing interests either financial or non-financial.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thaiparambil, N.A., Raja, B., Radhakrishnan, V. et al. Exploring the impact of formulated bacterial consortia on the growth of Solanum lycopersicum; towards sustainable biofertilizer development. Biologia 79, 1939–1955 (2024). https://doi.org/10.1007/s11756-024-01663-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-024-01663-z