Abstract
The present work was focused on plant growth-promoting bacteria (PGPB) for the enhanced production of tomatoes. The effect of 14 indigenous PGPB isolated from the soil samples collected from Gir National forest areas (Dist. Junagadh, Gujarat, India) and the coastal region of Saurashtra (Dist. Gir-Somnath, Gujarat, India) on tomato seedlings was studied. Only 6 isolates showed positive results in in vitro biochemical and plant-growth-promoting assays such as phosphate and zinc solubilization, siderophores and HCN production and antibacterial and antifungal properties. The rest isolates were negative to one or more of the selected tests. Based on in vitro results, all PGPB were tested on tomato seedlings. Four stage screenings were used to select the best combinations. After the primary screening, 9 isolates were selected for further experiments based on the germination, height of plants and seedling vigor. Plant height, leaf count, seedling vigor and total chlorophyll were measured. Leaf anatomy was studied at the end of the quaternary trial to understand the changes at a cellular level and it revealed the anatomical changes such as increased chlorophyll at cellular level and enhanced starch production. Microbial consortia showed better results compated to single-inoculant treatment. Out of 503 different combinations tested at secondary trial, 129 were selected for tertiary trials and 24 from them were further qualified for quaternary trials. At the end of quaternary trials, total 04 combinations were selected for future experiments on field. The quaternary trial strategy helped to reduce the total 503 possible combinations to 04 combinations improving germination, height, seedling vigor, number of leaves and chlorophyll content compared to non-treated tomato plants. Further trials were carried out in field for long period of time to measure the profound effect of selected consortium in real life.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
For better growth and development, plants need the carbon which is obtained from the atmosphere, the hydrogen and oxygen from water, and the needed mineral ingredients from the soil. The rhizosphere is the root-surrounding soil helping the plant to get micro- and macro-nutrients available in soil. Bacteria found in this soil are known as rhizobacteria. Rhizobacteria are also known as plant-root bacteria. They are called “plant growth-promoting bacteria” (PGPB) because they help plants to develop. Numerous research works support the diverse benefits of such bacteria for plants by fixing atmospheric nitrogen, solubilizing phosphate, creating phytohormones, siderophores, controlling ethylene and HCN production and manufacturing 1-aminocyclopropane-1-carboxylic acid deaminase promoting systemic resistance, and producing antibiotic [1]. These bacteria nourish plants and protect them from dangerous microorganisms too [2].
The most crucial element of food is vegetables using a larger area for cultivation. Tomato is one of the most significant vegetables. Tomatoes belong to Solanaceae family. In 2020, tomato output tops 187 million tons. Over the previous decade, global tomato production and consumption rose 2.5% and the demand is increasing in coming years, thus, tomato production needs better growth promoters. Because of its high nutritional value and plenty of uses, tomato is grown throughout the world. Tomatoes are also a great source of vitamin C, potassium, folate, and vitamin K [3]. Tomato is not only an important vegetable, but it is also an important source of secondary metabolites like lycopene used in pharmaceutical, nutraceutical, healthcare and cosmetic industries [4].
The rising population and the demand for vegetables are directly enhancing the high need for tomato production. However, the amount of area, the kind of soil, and the amount of rainfall significantly affect the production of tomatoes [5]. Aside from the nutrients, the overall production of tomatoes is also affected by common fungal diseases called early blight and late blight.
Compared to chemical fertilizers, which have serious soil degradation, nitrogen leaching, soil compaction, reduction in soil organic matter, and loss of soil carbon, PGPB may minimise soil degradation via nitrogen fixation, reduction of soil compaction, increase of soil organic matter, solubilisation of phosphate, zinc, and siderophore activity [6]. Phosphorus helps in the growth and yield of tomatoes. PGPB makes phosphorus more plant-accessible by dissolving insoluble phosphate. PGPB’s antibiosis, cyanogenesis, and other biocontrol activities strengthen the immunity the of tomato plant and help in fighting against early and late blight diseases [7]. Rhizobacteria that get along nicely may colonize and combat illness. PGPBs not only increase plant growth but helped to reduce stress, and manage the disease. Research shows that PGPB co-inoculation boosts plant growth [8].
This study will evaluate whether native isolates PGPB isolated from the soil samples collected from different regions of India improve tomato development. Single and multiple-organism inoculations were performed. Additional laboratory tests include phosphate and zinc solubilization, indole-3-acetic acid (IAA) and siderophores production, and the biocontrol abilities of isolates.
MATERIALS AND METHODS
Rhizosphere soil sampling. Soil samples were collected from the Gir National Forest Area, Gujarat, India and Saurashtra Coastal Region of Gujarat, India. Collected samples were given by accession numbers for labelling and easy processing. Coordinates of the sample collection site are given in Table 1.
Isolation of bacteria from collected rhizosphere. Rhizospheric soil (1 g) was suspended in sterilized, deionized water (90 mL). Serial dilution was made for isolation of pure culture. Ashby’s agar (HiMedia, India) was used for the isolation of the bacteria, which were plated in triplicate and incubated at 25°C for 24 h. Ashby’s agar plates were used to sub-culture and purify the bacteria colonies.
In vitro screening of isolates for different plant growth-promoting activities. Screening of HCN-producing bacteria. Isolated bacteria’s capacity to produce HCN was tested using the technique outlined by Lorck [9]. Glycerine-containing agar was used to grow bacteria on the Luria Bertani agar (HiMedia, India). Whatman filter paper no. 1 (UK) soaked with 1% picric acid and wet with 10% sodium bicarbonate was applied to the inside of the Petri plate lead. The plates were incubated for 24–48 h at 37 ± 1°C with the paraffin film sealed on top. Light brown to dark brown coloration of tested paper is an indication of high levels of HCN production.
Screening of phosphate-solubilizing bacteria. The capacity of isolates to dissolve phosphate was tested using the Verma and associates' technique [10]. Pikovskaya’s agar (HiMedia, India) plates containing tri-calcium phosphate were inoculated with the isolates to test their ability to dissolve the phosphate. Phosphate solubilization activity was judged to be successful after 7 days of incubation at 28°C.
Screening of IAA-producing bacteria. The Bric et al. [11] approach was used to look for IAA-producing bacteria. The streak plate technique was used to inoculate bacterial isolates onto LB agar plate supplemented with 1% Trp. Incubation took place at 37°C for 24–48 h. The Salkowski reagent was used to treat the membrane after a satisfactory incubation period. A red halo around the colony indicates the presence of IAA produced by the bacteria.
Screening of siderophore-producing bacteria. We followed the Schwyn and Neilands’ technique to screen siderophore-producing bacteria [12]. Briefly, Chrome Azurol S (CAS) agar medium was used to confirm the siderophore formation. LB agar medium supplemented with CAS was inoculated with 24-h-old bacteria and cultured for 72 h at 30°C. Siderophore production is indicated by a shift in the colour of the medium from blue to orange or yellow to light orange surrounding the colony.
Screening of zinc-solubilizing bacteria. All bacterial strains were screened for their zinc-solubilizing ability as per the method described by Kamran and co-workers [1]. Briefly, 5 insoluble zinc compounds, viz., ZnSO4, ZnO, ZnCl2, Zn3(PO4)2 and ZnCO3 were used. A one-day-old bacterial culture was inoculated on LB medium supplemented with insoluble ZnCO3. Aluminium foil was used to cover the plates and they were kept in the dark at 37°C for 14 days. Clear zones were created around colonies by strains that dissolved zinc compound.
Activity of isolates against fungi (Fusarium oxysporum). A phytopathogenic fungus may be inhibited by bacteria. F. oxysporum NCIM (accession no. 1008) was investigated in accordance with Slama et al. recommended approach [13]. Potato dextrose agar (PDA; HiMedia, India) plates were used to test the antagonism of bacterial isolates to phytopathogenic fungi in vitro. Fungal agar discs were placed on the Petri plate 3 cm apart from each bacterial growth site. A negative control was also performed, which consisted of fungal agar discs without any bacterial culture spots. After that, the Petri plates were incubated for 7 days at 30°C. Recording the growth patterns of fungi and bacteria and the antifungal properties of isolated bacteria were documented.
Chitinase activity of isolates. The isolates were examined for chitinase activity in accordance with the procedure described by Cappuccino and Sherman [14]. Briefly, chitin agar plates were used to cultivate all of the bacteria that were found in the samples. After 5 days of incubation at 30°C, a clear zone formed around the colonies showed that bacteria were chitinase positive.
Molecular characterization of bacterial strains. Isolates that had the highest levels of growth-promoting characteristics in plants were selected for further characterisation and identification.
Isolation of DNA and 16s rDNA gene amplification. Genomic study including DNA isolation, amplification of 16s rDNA gene and sequencing was performed at Gujarat biotechnology research center (GBRC) Gandhinagar, Gujarat, India as per the standard protocol of the laboratory. The amplification of selected gene was made using the universal bacterial primers 27F 5'-AGAGTTTGATCCTGGCTCAG-3' and 1492R 5'-GGTTACCTTGTTACGACTT-3'. The PCR was performed on thermal cycler Veriti, Applied Biosystems (USA) using EmeraldAmp® GT PCR Master Mix kit (Takara, Japan). Following conditions were used: initial denaturation for 5 min at 95°C, then 35 cycles of denaturation for 30 s at 94°C, annealing for 30 s at 62°C, extension for 1 min at 72°C, and finally another extension for 5 min at 72°C. Cycle sequencing was done using 24 capillary electrophoresis machine (3500 Genetic Analyzer, Applied Biosystems, USA) at GBRC.
Sequence alignment and phylogenetic analysis. In order to build a phylogenetic tree, MEGA software version 11 was used. The maximum parsimony approach was used to analyse the 16S rDNA sequences [15].
Preparation of inoculum and seed treatment. Bacterial in vitro activity was used as a selection guide. Selected bacteria were grown in LB broth for 24 h at 37°C to produce bacterial inoculum. Surface sterilization of tomato seeds took 2 min using 0.02% sodium hypochlorite. After sterilization, the seeds were thoroughly washed with sterile distilled water. Seeds were soaked in bacterial isolate suspensions (1 × 108 CFU/mL) for 30 min. In each of the three treatments, 3 replicates of inoculated seeds were used: 1% water agar, sterile soil, and non-sterile soil. For germination, the pots were kept at 30°C for 5 days in the dark. As a control, the seedlings that had been treated with sterile distilled water were used [5].
Field study. Field study was carried out during the period of May 2021 to April 2022 using the selected isolates. Initially single bacterial plant growth-promoting activity was assessed. After 7 days, out of 14 bacteria only 9 were selected for further studies. Combinations of bacteria were done as per equation 1. A total of 502 different combinations were calculated from the equation where 36 combinations were possible without repletion when any 2 or 7 bacteria were combined. When 3 or 6 bacteria were combined, 84 combinations were possible, 126 combinations exist for 4 and 5 bacterial combinations, 9 combinations were possible when 8 bacteria were combined, and 1 combination was possible when all 9 bacteria were mixed together at a time.
where nCr = number of combinations, n = number of objects, r = sample size, ni = n factorial, ri = r factorial
Microscopic analysis of plants. Control and treated plants were collected, fixed in FAA (70% ethanol—90 mL, 40% formalin—5 mL, glacial acetic acid—5 mL), separated into glass vials, dehydrated with an ethanol series, and embedded in paraffin [16]. Transverse serial sections, 12–14 µm thick, were cut with a rotary microtome. The histological sections were contrasted with 1% Toluidine blue (dissolved in 1% aqueous borax) and safranin O/fast green [17]. Slides were observed and photographed under different magnifications with help of DME research microscope (Leica, Germany).
Statistical analysis. All the study results were statistically evaluated by two-way ANOVA followed by Bonferroni post-tests for primary and secondary screening. One-way ANOVA followed by Dunnett’s Multiple Comparison Test was used to compare the effectiveness of the control v/s treatment plant at final screening. All the experiments were performed in triplicate and results were expressed as mean ± SEM. For statistical analysis and plotting graphs, GraphPad Prism 5 software was used.
RESULTS
In vitro screening of isolates with plant growth-promoting activities. Based on the plant growth promoting activity, total 14 bacteria were isolated and characterized. Code of microorganism, name of bacteria and GenBank accession number are depicted in Table 2. Bacteria having plant growth promoting activity were selected using in vitro screening (Table 2). Bacteria having phosphate and zinc solubilization properties and ability to produce siderophore and HCN were revealed. Apart from above mentioned properties, bacteria having antifungal activity, bacterial compatibility with other bacteria and chitinase production activity were suitable for primary screening. All isolated bacteria showed phosphate solubilization, siderophore and bacterial compatibility. Most of the bacteria had HCN production except GFS03C1, GFS16C1, SCS12C3 and SCS06C1. However, SCS06C1, SCS12C5, SCS12C2 and GFS16C1 did not showed antifungal activity. GFS03C1 and SCS06C1 did not have chitinase and zinc solubilization activities whereas GFS16C1 and SCS12C3 did not show only chitinase activity.
Molecular characterization of bacterial strains. Molecular characterization of isolated bacteria helps to understand the phenotypes and heterogenicity. The sequences of the 16S rDNA gene of the isolates were compared with the NCBI nucleotide database using the BLAST to determine the identity of the isolated bacteria (Bethesda, USA). Phylogenetic analysis revealed the relationship between the isolates (Fig. 1). Out of 14 isolates, 7 belong to Bacillus species.
All the solitary bacterium’s evolutionary connections. The maxim parsimony approach was used to infer the evolutionary history. The evolutionary history of the species under study is assumed to be represented by the bootstrap consensus tree constructed from 1000 repetitions of the data. MEGA-11 was used for the evolutionary analyses.
Plant growth-promotion assay. Primary screening. Primary screening of PGPB were done by testing all bacteria. Figure 2 explains the effect of bacteria on plant growth. For primary screening, only seedling vigor of plant was considered. Primary screening revealed that out of 14 only 9 bacteria have potential plant growth-promoting activity. Results obtained after 7 days of tomato seed germination were considered for primary screening of bacteria for 15 days trial. Figure 3 describes the seedling vigor of isolated bacteria after 15 days germination.
Seedling vigor data after 7 days of tomato seeds germinated in water agar, sterile soil and non-sterile soil and treated with 14 bacterial isolates. Results are expressed as mean ± SEM. Results are analysed by two way ANOVA followed by Bonferroni posttests. 1—Control; 2—GFS01C1; 3—GFS03C1; 4—GFS11C15—GFS15C2; 6—GFS16C1; 7—GFS16C2; 8—GFS19C2; 9— SCS03C1; 10—SCS07C3; 11—SCS12C1; 12—SCS12C2; 13—SCS12C3; 14—SCS12C5; 15—SCS06C1.
Seedling vigor data after 15 days tomato seeds germinated in water agar, sterile soil and non-sterile soil and treated with selected 9 bacterial strains. Results are expressed as mean ± SEM. Results are analysed by two way ANOVA followed by Bonferroni posttests. 1—Control; 2—GFS01C1; 3—GFS11C1; 4—GFS15C2; 5—GFS16C2; 6—GFS19C2; 7—SCS03C1; 8—SCS07C3; 9—SCS12C2; 10—SCS12C5.
Secondary screening. Combination of bacteria and its effect on plant growth after 7-days germination is described in Table S1. Seedling vigor higher than 270 index were taken further for the tertiary screening. Out of 36 combinations of any two bacteria, only 9 showed seedling vigor > 270 index, whereas, out of 84 possible combinations of 3 bacteria only 23 combinations showed promising results. Total 36 combinations of 4 bacteria had high seedling vigor, 30 combinations of 5 bacteria, 22 combinations of 6 bacteria, 6 combinations of 7 bacteria, 2 combinations of 8 bacteria showed seedling vigor more than 270 index. Interestingly, consortium of all bacteria showed < 270 seedling vigor. Out of 502 combinations 128 had seedling vigor higher than 270 index whereas, only 43 showed seedling vigor more than 290 index. However, only 16 combinations had seedling vigor more than 300. Control showed seedling vigor in between 170–185.
Tertiary screening. Those combinations showing seedling vigor more than 270 index in secondary screening were taken for tertiary screening. Total 128 combinations passed secondary screening and considered for tertiary screening. From tertiary screening, top 50% of the plants having average 5 leaves or more leaves were taken for quaternary trials. Detailed results of all 128 trials are described in Table S2. Tertiary screening helped to reduce the total no trials from 128 to 24.
Quaternary screening. Quaternary screening was done for the 24 bacterial consortia selected from the results of tertiary screening. Results are described in Table 3. Study results revealed that bacterial consortium is highly effective in terms of total chlorophyll (p < 0.0001). Out of 24 combinations, only 4 were selected for future experimentation based on criterion that plant germination was more than 70%. Top 50% bacteria were selected based on seedling vigor height and total chlorophyll. Details of the selected consortia compared to control group are given in Table 3. Compared to control group, all consortium groups showed reliable results (p > 0.05).
Microscopic analysis of plant. Leaf. Tomato leaf showed dorsiventral type of structure anatomically (Figs. 4a, 4b). It possesses upper and lower epidermis. Both the epidermis are covered with cuticle. Simple and glandular trichomes are present towards both layers of epidermis. In the center of the leaf, single midrib is present which shows collenhymatous tissues after upper epidermis while presence of parenchyma tissue above the lower epidermis, and vascular bundle is present between them. In the vascular bundle, phloem is facing towards the lower epidermis. In the lamina region pallisade tissue is present towards the upper epidermis with 2–3 layers, while spongy chlorenchyma tissue is present towards the 3–4 layers of lower epidermis (Fig. 4c). These anatomical characteristics are found in untreated/control plant leaves while treated plant leaves had spongy chlorenchyma in the lower epidermis, a more compact arrangement of cells, and an increase in cell number (Fig. 4d).
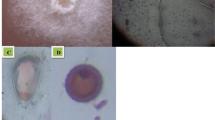
Microscopic anatomical features comparisoin of control vs treated tomato plants. (a) Transverse section of control tomato leaf; (b) transverse section of treated with PGPB tomato leaf; (c) lamina region of control tomato leaf (arrow shows loosely arranged spongy parenchyma); (d) lamina region of treated with PGPB tomato leaf (arrow shows compactly arranged spongy parenchyma); (e) transverse section of control tomato plant root; (f) transverse section of treated with PGPB tomato root; (g) cortex of control root shows starch grains (arrow); (h) cortex of treated with PGPB root shows more amount of starch grains (arrow).
Root. Tomato’s root reveals an exterior layer of the epidermis (periderm) and a well-developed core cylinder called a stele that includes xylem toward the inner side and phloem near the periderm. Parenchymal cortical cell and well-developed core cylinder called a stele includes xylem toward the inner side and phloem near the periphery. There was procambium in between them, and it was responsible for the subsequent development of vascular tissues (Figs. 4e, 4f). After the periderm, cortical region is present which is made up of by parenchymal cells compactly arranged. After those cells, vascular tissues are arranged. In the cortical region starch grains are present (Fig. 4g). These tissues are present in untreated plant root. After PGPB-treatment, the increase in the number of starch grains was observed (Fig. 4h), which indicates the increase in chlorophyll amount in leaf.
DISCUSSION
Photosynthesis is the key process through which light energy is transformed into chemical energy. One of the most important molecules in photosynthesis is phosphorus. Due to its insoluble nature, phosphorus compounds hampered plant development in general, as well as the efficiency with which photosynthesis and respiration were carried out [18]. Phosphate has a critical function in plant development, yet only plants can use it. Phosphorus is an essential component of the photosynthetic center of any high-yield crop production system, where light energy is converted into sugar and then into all the unique plant components [19]. Thus, the ability to dissolve phosphate would be a crucial criterion in selecting PGPB. As depicted above we have selected phosphate solubilization in vitro qualitative assay for selecting PGPB for the tomato’s growth and development in terms of increased height, number of leaves and chlorophyll content.
Iron is important element in electron chains and cofactor in many essential enzymes. Due to its extremely reactive nature, iron must be chelated before it can undergo intracellular trafficking [20]. Siderophores are a class of secondary metabolites that are produced by a broad range of organisms to remove iron from their surrounding environments and make it available to cells. Literature suggests that many different types of microorganisms create siderophores, which chelate iron, reduce its toxicity, and enhance its utilisation in cellular biochemistry Multiple research highlight siderophores potential role in fostering plant development [21]. Positive siderophore secretion was detected in all of the isolated bacteria, according to the results of the in vitro study. These microorganisms were suited for the growth of tomato plants.
The HCN is well recognized as biocontrol agent against several plant pathogens including bacteria and fungi [22]. The research implies that the covalent attachment of HCN to cytochrome oxidase is responsible for its toxicity. Since the PGPB use a cytochrome oxidase that is invulnerable to cyanide, they are unaffected by HCN production [23]. Out of 14 isolated bacteria, 10 bacteria showed in vitro HCN production. Based on the previous finding, we can suggest that consortia of isolates would not only help the plant growth but also protect the plant from variety of pathogens. Nine isolates out of 14 had antifungal activity. Surprisingly, 2 of the 5 isolates that lacked antifungal activity also failed to secrete HCN in vitro. Indications from this study suggest that tomato resistance to fungal infections may be due in part to the bacterial capacity to produce HCN.
Zinc is an essential trace element for the metabolism, enzyme processes, and oxidation-reduction reactions of plant, animal, and bacterial cells. Zinc deficiency has been linked to malfunctions in nitrogen metabolism, protein synthesis, and energy transfer in plant cells. Zinc aids in plant growth and development and also boosts the plant nutritional value [24]. It was discovered that 12 out of 14 bacteria tested were effective zinc solubilizes. The current study found that PGPB resulted in superior development of tomato plants compared to normal plants.
Duijff and co-workers [25] reported Pseudomonas fluorescens WCS417r showed an anatomical effect on tomato plants. Compant et al. [26] explained that endophytic colonization of bacteria caused notified anatomical changes in the plant in terms of characteristics like increased cortical cell layers, and increased strengthening of the exodermis cell wall. The outer peripheral and outermost part of the radial side of the first layer of the cortical cell walls showed an increase in thickening. Microscopic analysis revealed that the PGPB-treated tomato plants showed improved chlorophyll content and increased amount of starch accumulation in cells compared to normal plants (Fig. 4). This data lends credence to the theory that higher levels of chlorophyll and photosynthesis are associated with higher concentrations of soluble phosphate and zinc, as well as increased production of siderophores.
As discovered by Timmusk et al. [27], Paenibacillus polymyxa GSF01C1 is rhizobacterium that promotes plant growth and forms biofilms in root tip. Other researchers revealed nitrogen fixing and antifungal properties of this bacterium. The current research confirms the outcomes of similar studies. P. polymyxa is the member of 3 consortia out of 4 chosen from quaternary trials.
Researchers have looked into Bacillus toyonensis GFS03C1 extensively because of its potential to aid growth in a wide variety of plants, including tomato [7]. We observed that the isolated bacteria not only promoted tomato growth, but also acted as phosphate solubilizers, siderophore producers, and a fungal killer. However, this bacterium is not a part of finally chosen 4 consortia after quaternary trials.
Bacillus sonorensis GFS11C1 was well studied for its PGP action in tomato [3], etc. In the present study, in vitro tests for B. sonorensis were all positive. The presence of B. sonorensis is helping tomato overall growth, including plant height, number of leaves, and enhanced chlorophyll content. It was one of the member of 3 consortia out of 4 chosen from quaternary trials.
Peribacillus asahii GFS15C2 has received less attention in the scientific community. P. asahii’s potential in agriculture was highlighted for the first time in our paper. This bacterium performed well in all in vitro tests. It was member of one consortium out of 4 chosen from quaternary trials.
Phosphate, zinc, and siderophores are all made more soluble by Stenotrophomonas bentonitica GFS16C1. However, this bacterium cannot produce antifungal agents or chitinases.
Isolating Flavobacterium anhuiense GFS16C2 from cucumber roots, Jeong et al. [28] found that this strain exhibited potent anti-oomycete biocontrol activity. In the current study, promising PGP effects were revealed for F. anhusiense under all test conditions. It was the member of 2 consortia out of 4 chosen from quaternary trials.
Metabacillus endolithicus GFS19C2 has been isolated by Parag et al. [29]. Insufficient information is currently available on this bacterium in the literature. In the present study, this bacterium was part of 8 of 10 chosen consortia and performed well in vitro tests. The effect it has on tomato plant development is new information. It was member of one consortium out of 4 chosen from quaternary trials.
Isolation of Acinetobacter pitti (SCS03C1) from the maize rhizosphere and its use in boosting paddy plant development were reported recently [30]. However, its agricultural potential in the connection with PGP activity was not explored. In the present study, A. pitti had positive tests in vitro and in vivo in all trials. It was the member of 2 consortia out of 4 chosen from quaternary trials.
The most extensively researched PGPB is Bacillus pumilus SCS06C1. Several researches have shown its effectiveness. According to Gutiérrez-Mañero et al. [2], B. pumilus exhibited superior PGP properties. The gibberellins produced in large quantities by this bacterium, make it capable of promoting overall plant development. Antifungal activity activities were revealed in this bacteium. Joo et al. [31] analyzed this bacterium extensively for its ability to stimulate growth in red pepper plugs. Although it passed our phosphate solubilization assay and siderophore activity tests, this bacterium did not place in the top 4 final consortia for further study.
We know that Pseudomonas extremorientalis SCS07C3 is a promising example of PGPB due to its detailed characterization. The growth-promoting activity of P. extremorientalis was studied on many plants, such as wheat [32]. Our research shows that it has very positive effects on tomato development. P. extremorientalis is part of 2 out of 4 carefully put together groups, and all of them have shown that they can promote tomato growth.
Despite promising in vitro results, Bacillus licheniformis SCS12C1 failed to boost plant growth in the field. There were no reports of this bacterium’s PGP properties in the literature.
Sahu et al. [33] recently reported PGP activity in Bacillus haynesii SCS12C2 for rice. This bacterium was determined to be a part of 3 of 4 investigated consortia.
Recently, Park et al. [34] revealed the rice-targeting PGP activity in Bacillus vallismortis (SCS12C3). In the current study, we investigated whether B. vallismortis does improve the tomato growth or not. Results shown B. vallismortis is associated with negative outcomes.
Priestia aryabhattai SCS12C5 is well studied for its plant growth promotion for a variety of plants and crops [35]. In our investigation, we confirmed that P. aryabhattai is a promising plant growth promoter for tomatoes. It is the member of 2 of 4 consortia.
* *
Finally, we found that 9 isolates together increased tomato plant growth. These isolates include GFS01C1, GFS19C2, GFS11C1, GFS15C2, GFS16C2, SCS03C1, SCS073, SCS12C2, and SCS12C5. As such, they will aid in the development of tomato plants. More study of colonisation capacity and probable interactions between strains is needed to increase PGPB effectiveness.
REFERENCES
Kamran, S., Shahid, I., Baig, D.N., Rizwan, M., Malik, K.A., and Mehnaz, S., Front. Microbiol., 2017, vol. 8, p. 2593. https://doi.org/10.3389/fmicb.2017.02593
Gutiérrez-Mañero, F.J., Ramos-Solano, B., Probanza, A.N., Mehouachi, J.R., Tadeo, F., and Talon, M., Physiol. Plant., 2001, vol. 111, no. 2, pp. 206–211. https://doi.org/10.1034/j.1399-3054.2-001.1110211.x
Shakuntala, N., Nachu, N., Ashwin, R., and Bagyaraj, D., J. Soil Biol. Ecol., 2019, vol. 39, pp. 32–38.
Agarwal, S. and Rao, A.V., Can. Med. Assoc. J., 2000, vol. 163, no. 6, pp. 739–744.
Xu, M., Sheng, J., Chen, L., Men, Y., Gan, L., et al., World J. Microbiol. Biotechnol., 2014, vol. 30, no. 3, pp. 835–845. https://doi.org/10.1007/s11274-013-1486-y
Gupta, A. and Gopal, M., Ind. J. Agric. Res., 2008, vol. 42, no. 2, pp. 153–156.
Rojas-Solis, D., Vences-Guzmán, M.A., Sohlenkamp, C., and Santoyo, G., Curr. Microbiol., 2020, vol. 77, no. 10, pp. 2735–2744. https://doi.org/10.1007/s00284-020-02069-1
Egamberdieva, D., Li, L., Lindström, K., and Räsänen, L.A., Appl. Microbiol. Biotechnol., 2016, vol. 100, no. 6, pp. 2829–2841. https://doi.org/10.1007/s00253-015-7147-3
Lorck, H., Physiol. Plant., 1948, vol. 1, no. 2, pp. 142–146. https://doi.org/10.1111/j.1399-3054.1948.tb07118.x
Verma, S.C., Ladha, J.K., and Tripathi, A.K., J. Biotechnol., 2001, vol. 91, no. 2, pp. 127–141. https://doi.org/10.1016/S0168-1656-(01)00333-9
Bric, J.M., Bostock, R.M., and Silverstone, S.E., Appl. Environ. Microbiol., 1991, vol. 57, no. 2, pp. 535–538. https://doi.org/10.1128/aem.57.2.535-538.1991
Schwyn, B. and Neilands, J., Anal. Biochem., 1987, vol. 160, no. 1, pp. 47–56. https://doi.org/10.1016/0003-2697(87)-90612-9
Slama, H.B., Cherif-Silini, H., Chenari Bouket, A., Qader, M., Silini, A., et al., Front. Microbiol., 2019, vol. 9, pp. 1–24. https://doi.org/10.3389/fmicb.2018.03236
Cappuccino, J. and Sherman, N., Microbiology: A Laboratory Manual, New York, NY: Benjamin-Cummings Publishing Company, 1999, 3rd ed., pp. 125–179.
Tamura, K., Stecher, G., and Kumar, S., Mol. Biol. Evol., 2021, vol. 38, no. 7, pp. 3022–3027. https://doi.org/10.1093/molbev/msab120
Johansen, D.A., Plant Microtechnique, London: McGraw-Hill Book Company, Inc., 1940.
Sass, J.E., Botanical Microtechnique, Ames, Iowa, USA: The Iowa State College Press, 1958, 2nd ed.
Thuynsma, R., Kleinert, A., Kossmann, J., Valentine, A.J., and Hills, P.N., S. Afr. J. Bot., 2016, vol. 104, pp. 244–248. https://doi.org/10.1016/j.sajb.2016.03.001
Foyer, C. and Spencer, C., Planta, 1986, vol. 167, no. 3, pp. 369–375. https://doi.org/10.1007/BF00391341
Schmidt, W., Thomine, S., and Buckhout, T.J., Front. Plant Sci., 2020, vol. 10, p. 1670. https://doi.org/10.3389/fpls.2019.01670
Andleeb, S., Shafique, I., Naseer, A., Abbasi, W.A., Ejaz, S., et al., PLoS One, 2022, vol. 17, no. 6, p. e0269946. https://doi.org/10.1371/journal.pone.026-9946
Rijavec, T. and Lapanje, A., Front. Microbiol., 2016, vol. 7, p. 1785. https://doi.org/10.3389/fmicb.2016.01785
Sehrawat, A., Sindhu, S.S., and Glick, B.R., Pedosphere, 2022, vol. 32, no. 1, pp. 15–38. https://doi.org/10.1016/S1002-0160-(21)60058-9
Yuvaraj, M. and Subramanian, K., Biot. Res. Today, 2020, vol. 2, no. 8, pp. 823–825.
Duijff, B.J., Gianinazzi-Pearson, V., and Lemanceau, P., New Phytol., 1997, vol. 135, no. 2, pp. 325–334. https://doi.org/10.1046/j.1469-8137.-1997.00646.x
Compant, S., Reiter, B., Sessitsch, A., Nowak, J., Clément, C., and Ait Barka, E., Appl. Environ. Microbiol., 2005, vol. 71, no. 4, pp. 1685–1693. https://doi.org/10.1128/AEM.71.4.1685-1693.2005
Timmusk, S., Grantcharova, N., and Wagner, E.G.H., Appl. Environ. Microbiol., 2005, vol. 71, no. 11, pp. 7292–7300. https://doi.org/10.1128/AEM.71.11.7292-7300.2005
Jeong, J.-J., Sajidah, S., Oh, J.Y., Sang, M.K., Kim, K.-S., and Kim, K.D., Data Br., 2019, vol. 25, p. 104270. https://doi.org/10.1016/j.dib.2019.104270
Parag, B., Sasikala, C., and Ramana, C.V., Int. J. Syst. Evol. Microbiol., 2015, vol. 65, pt. 12, pp. 4568–4573. https://doi.org/10.1099/ijsem.0.000612
Ashfaq, M., Hassan, H.M., Ghazali, A.H.A., and Ahmad, M., Environ. Monit. Assess., 2020, vol. 192, no. 11, p. 697. https://doi.org/10.1007/s10661-020-08655-x
Joo, G.-J., Kim, Y.-M., Lee, I.-J., Song, K.-S., and Rhee, I.-K., Biotechnol. Lett., 2004, vol. 26, no. 6, pp. 487–491. https://doi.org/10.1023/B:BILE.0000019555.87121.34
Akimova, E.E., Tereshchenko, N.N., Zyubanova, T.I., and Minaeva, O.M., Bull. Tomsk Polytech. Univ., 2019, vol. 45, pp. 128–141. https://doi.org/10.17223/19988591/45/7
Sahu, P.K., Singh, S., Gupta, A.R., Gupta, A., Singh, U.B., Manzar, N., et al., Biol. Control, 2020, vol. 150, p. 104353. https://doi.org/10.1016/j.biocontrol.-2020.104353
Park, K.-S., Paul, D., and Yeh, W.-H., J. Plant Pathol., 2006, vol. 22, no. 3, pp. 278–282. https://doi.org/10.5423/PPJ.2006.22.3.278
Cochard, B., Giroud, B., Crovadore, J., Chablais, R., Arminjon, L., and Lefort, F., Microorganisms, 2022, vol. 10, no. 4, p. 765. https://doi.org/10.3390/microorganisms10040765
ACKNOWLEDGMENTS
The authors are very grateful to Dr. D. Vaishnav, an assistant professor in the Department of Pharmaceutical Sciences at Saurashtra University, for helping with statistical analysis. Authors would also like to thank Mr. V. Joshi, Mr. P. Jyani and Mr. S. Sonara from Shree H. N. Shukla College of Science for their assistance with this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest.
This article does not contain any studies involving animals or human participants performed by any of the authors.
Supplementary Information
Rights and permissions
About this article
Cite this article
Pattani, V., Kaneriya, J., Joshi, K. et al. Effect of Growth-promoting Bacterial Consortia on Overall Growth of Tomato Plants. Appl Biochem Microbiol 59, 511–521 (2023). https://doi.org/10.1134/S0003683823040105
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683823040105