Abstract
The identification of proteins involved in pollen germination and tube growth is important for fundamental studies of fertility and reproduction in flowering plants. We used 2-DE and MALDI-TOF-MS to identify differentially expressed proteins in mature (P0) and 1-h germinated (P1) maize pollen. Among about 470 proteins separated in 2D gels, the abundances of 26 protein spots changed (up- or down-regulation) between P0 and P1. The 13 up-regulated protein spots were mainly involved in tube wall modification, actin cytoskeleton organization, energy metabolism, signaling, protein folding and degradation. In particular, pectin methylesterase, inorganic pyrophosphatase, glucose-1-phosphate uridylyltransferase and rab GDP dissociation inhibitor α are highly up-regulated, suggesting their potential role in pollen tube growth. The down-regulated 13 protein spots mainly include defense-related proteins, pollen allergens and some metabolic enzymes. This study would contribute to the understanding of the changes in protein expression associated with pollen tube development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pollen germination and tube growth play a crucial role in sexual reproduction of flowering plants as pollen tubes deliver sperms into the sac, a necessary process for fertilization and seed setting (Feijo et al. 2004; Cai and Cresti 2009). The pollen tube grows exclusively at the tip and is very fast in some cases. It is well known that a steep calcium gradient within the tube tip and actin microfilaments are major biochemical factors driving the pollen tube elongation (Krichevsky et al. 2007; Cai and Cresti 2009). In spite of the great progress made in the elucidation of signaling pathway in pollen tube growth (Malho et al. 2006), only a small number of proteins have so far been reported to be differentially expressed during pollen germination and associated with pollen tube growth (e.g., Wittink et al. 2000; Yang 2002; Golovkin and Reddy 2003; Dai et al. 2007).
The proteome is the entire set of proteins expressed by a genome, cell, tissue or organism. (Wilkins et al. 1996). It is highly dynamic and depends on cell cycle, environmental influences and tissue/cell type. Currently, proteomic analysis based on two-dimensional electrophoresis (2-DE) and mass spectrometry (MS) is most effective in identifying differentially expressed proteins involving in pollen development and germination in Arabidopsis (Holmes-Davis et al. 2005; Noir et al. 2005; Sheoran et al. 2006; Grobei et al. 2009; Zou et al. 2009), rice (Dai et al. 2006, 2007), tomato (Sheoran et al. 2007), pine (Fernando 2005; Chen et al. 2006) and Lilium longiflorum (Pertl et al. 2009). Recently, dynamic proteomic analysis revealed that 344 were differentially expressed in mature and germinated pollen in canola (Brassica napus) (Sheoran et al. 2009). Obviously, proteomic identification of proteins differentially expressed in germinated pollen will provide new insights into the roles of protein composition and quantity in pollen germination and tube growth.
Maize is one of the most important crops throughout the world and the largest food crop in China. Its tassels produce a large quantity of pollen. Importantly, fresh maize pollen has high ability to germinate in vitro, and its tube can grow as fast as 1 cm/h and extend to about 30 cm in length within 24 h (Barnabas and Fridvalszky 1984). Compared to other plant species, it is easy to obtain sufficient germinated maize pollen for proteomic analysis.
A comparative proteomic analysis of mature maize pollen and germinated pollen is needed to reveal the complex molecular mechanisms of pollen germination and tube growth. To our knowledge, there has been no proteomic study of pollen germination of maize. Thus, we aimed to identify differentially expressed proteins during maize pollen germination. We have used 2-DE and MS/MS to compare the changes in protein profiles between mature and germinated maize pollen. The differentially expressed proteins were identified by MALDI-TOF-MS and homologous sequence comparison. The potential roles of the differentially expressed proteins in pollen germination and tube growth were discussed.
Materials and methods
Pollen collection and pollen germination in vitro
Zhengdan 958, one of the widely grown high-yield maize hybrids in China, was grown in a greenhouse. At anthesis, fresh pollen was collected in the morning by shaking the tassel in a plastic bag, while old pollen and anthers were removed from tassels by vigorous shaking the evening of the day before.
For germination in vitro, 5 mg of pollen (on three replicas) was placed in a 100 × 15 mm Petri dish containing 1.5 ml of germination medium consisting of 100 ppm Ca(NO3)2, 10 ppm H3BO3, 37.5 ppm lysine, 5 ppm cysteine, 0.05 ppm glutamic acid and 15% (w/v) sucrose (Suen and Huang 2007). The dish was covered to retain moisture and the pollen was allowed to germinate at 25°C. Pollen germination was observed and counted with a Leica DMLB microscope. Pollen was considered germinated when the length of the tube was equal to or greater than the diameter (approximately 70 μm) of the pollen grain. The pollen tubes were collected and centrifuged at 1,500g at 4°C for 3 min. The pelleted tubes were pooled and subjected to protein extraction.
Protein extraction and 2-DE
Mature pollen and germinated pollen (equal to 20 mg of pollen) were respectively homogenized in 4 ml of the buffer consisting of 0. 1 M Tris–HCl, pH 7.8, 20 mM DTT and 1 mM PMSF in a mortar (4°C). After that, the homogenate was centrifuged at 20,000g for 15 min at 4°C. The supernatant was mixed with equal volume of Tris-buffered phenol (pH 8.0, Sigma) by shaking for 10 min. The mixture was centrifuged at 20,000g for 5 min and the phenol phase was recovered (Wang et al. 2006). Protein in the phenol phase was precipitated with five volumes of 0.1 M ammonium acetate in methanol for 1 h at −20°C, and centrifuged at 20,000g for 10 min at 4°C. The pellet was washed with cold acetone twice, air dried and solubilized in rehydration buffer (7 M urea, 2 M thiourea, 4% CHAPS, 20 mM DTT and 1% IPG buffer). Protein concentrations were determined using the BioRad protein assay dye reagent with BSA as a standard.
For 2-DE, protein (700 in 250 μl rehydration buffer) was loaded into 11-cm linear pH 4–7 strip (GE Healthcare, OK, USA) via passive rehydration overnight (Wang et al. 2009). IEF was performed with Ettan III system (GE Healthcare) at 300 V for 1 h, 3,000 V for 1 h and 6,000 V for 10 h (20°C). Focused strips were equilibrated for 15 min in a buffer containing 0.1 M Tris–HCl (pH 8.8), 2% SDS, 6 M urea, 30% glycerol and 0.1 M DTT, and for another 15 min in the same buffer but plus 0.25 M iodoacetamide. Afterward, SDS-PAGE was run in 12.5% polyacrylamide gel (20 × 15 × 0.1 cm). Gels were stained with 0.1% (w/v) colloidal CBB G for 24 h and destained in 10% (v/v) acetic acid until a clear background was obtained. Digital images of the gels were processed and analyzed using PDQUEST software (Bio-Rad) and proteins differentially expressed with twofold variations were selected for MALDI-TOF analysis.
MALDI-TOF-MS and protein identification
Differential protein spots were manually excised, reduced with 10 mM DTT, alkylated with 50 mM iodoacetamide and digested with 10 μg/μl trypsin (Promega, Madison, WI, USA) for 16 h at 37°C in 50 mM ammonium bicarbonate. The supernatants were vacuum dried and dissolved in 10 μL 0.1% trifluoroacetic acid and 0.5 μl added onto a matrix consisting of 0.5 μl of 5 mg/ml 2-5-dihydroxybenzoic acid in water:acetonitrile (2:1) (Wang et al. 2009).
The digested fragments were analyzed on Ettan MALDI-TOF Pro mass spectrometer (GE Healthcare, USA). The ion acceleration voltage was 20 kV. Each spectrum was internally calibrated with the masses of two trypsin autolysis products. The peptide masses and sequences obtained were automatically matched to proteins in a nonredundant database (NCBI) with the Mascot algorithm (http://www.matrixscience.com). The following parameters were adopted for database search: complete carbamidomethylation of cysteines and partial oxidation of methionines, peptide mass tolerance ±1.2 Da, fragment mass tolerance ±0.9 Da and missed cleavages 2. Searches were performed in the full range of Mr and pI. No species restriction was applied. All of the positive protein identification scores were significant (P < 0.05, score >60). Functional categorization and subcellular localization of identified proteins was performed using annotation in NCBI database and Swiss-prot database.
Results
Pollen germination in vitro
Mature pollen of maize (cultivar Zhengdan 958) germinates in vitro fast (Fig. 1). About 80% of the pollen germinated within 30 min, and more than 90% germinated after 1-h incubation. On average, the pollen tube length reached 200 μm (1 h germination), about three times the diameter (ca. 70 μm) of the pollen. Thus, mature pollen (P0) and pollen germinated for 1 h (P1) were selected for further proteomic analysis.
Protein profile comparison
We aimed to characterize the difference in proteome maps between mature and germinated pollen. Technically speaking, it is impossible to resolve the proteome profile of a single pollen grain or pollen tube using the common 2-DE. Thus, we used pooled P0 and P1 for each batch 2-DE analysis. Thus, protein samples from pollen grains and tube here should be considered as “average samples” (Holmes-Davis et al. 2005) and not biological replicates.
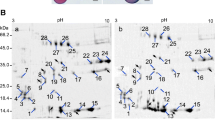
Proteins from P0 and P1 (700 μg of each extract) were compared by 2-DE, using linear pH 4–7 IPG strips (11 cm) in first dimension and 12% SDS-PAGE in the second dimension. After electrophoresis, proteins were visualized by CBB staining. The global protein pattern of P1 was largely unaltered compared to P0, as analyzed with the PDQUEST software. On average, 468 ± 6 and 473 ± 7 reproducible spots (n = 3) were detected in P0 and P1, respectively (Fig. 2). Spots of poor quality and very low raw volumes were discarded using criteria set by the PDQUEST software. These proteins had pI of 4.5–7 and Mr of 10–110 kDa. A total of 28 spots (5.9% of the total matched spots across all the 2D maps) in P1 were identified as differentially expressed (Fig. 2, up- or down-regulated, indicated by arrows). In particular, 13 spots were found to be up-regulated in P1 compared to P0, of which 2 were newly expressed (spot 22 and 23), and 13 down-regulated (Table 1). Therefore, in germinated maize pollen, there are both qualitative (newly induced) and quantitative (increased or decreased) changes in protein spots.
Protein profile comparison between P0 (a mature pollen) and P1 (b germinated for 1 h). The image displays two representative CBB-stained gels out of a total of six maps. Proteins (700 μg) were resolved in 11-cm linear immobilized pH gradient strips (4–7) and then in 12.5% SDS/acrylamide gels. Spot numbers indicated on the gel were subjected to MALDI-TOF analysis. pI and Mr (in kDa) are noted
Protein identification
All differentially expressed protein spots, indicated by arrows in Fig. 2, were excised from the gels and subjected to in-gel digestion and MALDI-TOF analysis. Automated Mascot software was used to search NCBI protein database. Of the 28 spots analyzed, 26 were identified, representing 21 distinct proteins in NCBI or SWISS-PROT protein databases (Table 1), while two spots were not identified (not shown).
Each identified protein was listed by its accession number, molecular weight (Mr) and isoelectric point (pI), expression level, cellular compartment and possible function. These proteins are mainly involved in actin cytoskeleton organization, tube wall expansion, carbohydrate metabolism, signaling, protein folding and degradation and oxidative stress tolerance. For most identified proteins, the calculated Mr and pI were close to the theoretical values, but exceptions were Zea m 1 and both isoforms of Zea m 13, which had much smaller pI in 2D gels than expected. Besides, five identified proteins, including pectin methylesterases (PMEs), actin, inorganic pyrophosphatase, phosphoglucomutase, and Zea m 13, each exist as two isoforms.
Discussion
Differential expression of proteins
The comparison of protein profiles between P0 and P1 shows that a large fraction of the proteome was similar, and about 28 spots changed greatly (increase or decrease in spot intensities) in both samples. This agrees with previous findings that the majority of the proteins in the pollen tube are already present in the pollen (Fernando 2005; Chen et al. 2006; Dai et al. 2007; Zou et al. 2009). For example, the pollen and pollen tubes in Pinus strobus share about 94% similarity based on quantitative variations (Fernando 2005); only 23 differentially expressed spots were detected in germinated Arabidopsis pollen compared to mature pollen (Zou et al. 2009).
Besides, 5 out of 21 proteins identified here exist as isoforms. Isoforms are generally considered to diversify the function of a protein (Lockhart and Winzeler 2000; Wang et al. 2004a). The presence of isoforms may result from sequence-related proteins encoded by distinct genes and/or polypeptide variants encoded by the same gene (splice variants and/or PTM) (Holmes-Davis et al. 2005; Dai et al. 2007; Sheoran et al. 2007).
Functional significance of differentially expressed proteins
In the present study, the identified differentially expressed proteins were classified into seven major categories according to their subcellular localization and molecular function (Table 1). These proteins are involved in a wide variety of cellular processes, as found in previous studies (Fernando 2005; Chen et al. 2006; Dai et al. 2006, 2007; Wang et al. 2008; Zou et al. 2009).
Cell wall-modifying proteins
PMEs can regulate the cell wall dynamics of growing pollen tubes (Bosch and Hepler 2005; Bosch et al. 2005). During the pollen tube elongation process, pectins are secreted as methylesters and subsequently deesterified by PMEs. Here, PMEs were found to exist in undetectably low abundance in P0, but their two isoforms (spots 22 and 23) synthesized de novo were present in higher abundance in P1, indicating their important role in pollen tube growth.
Se-wap41 is a salt-extractable and wall-associated protein (41 kDa) in etiolated maize seedlings (Sagi et al. 2005). Its presence is necessary for the growth of the pollen tube of Picea meyeri (Chen et al. 2006). However, we found that cell wall-modifying protein Golgi-associated se-wap41 was detected in down-regulation in germinated maize pollen.
Energy-related proteins and metabolic enzymes
Pollen germination and tube growth require a high rate of carbohydrate metabolism to meet energetic and biosynthetic demands (Krichevsky et al. 2007). It is not surprising that a high percentage (19%) of proteins related to energy metabolism was identified in mature tomato pollen (Sheoran et al. 2007). Other proteomic studies also show the presence of a large quantity of energy-related proteins and metabolic enzymes in pollen (Holmes-Davis et al. 2005; Noir et al. 2005; Dai et al. 2007).
In the present study, six enzymes involved in energy pathway were found up-regulated in P1, such as ATP production-related protein ATPase subunit 1 and basic energy supply-related soluble inorganic pyrophosphatase (Table 1). Triosephosphate isomerase, phosphoglucomutase and enolase 2 were down-regulated (Table 1).
Cytoskeletal proteins
Pollen tube growth is a typical tip growth process, which is highly dependent on the actin cytoskeleton (Geitmann et al. 2000; Vidali et al. 2001; Cai and Cresti 2009). Actin cytoskeleton supports cytoplasmic streaming and vesicular transport critical for pollen tube growth, and appears to be a target for calcium-mediated tube growth (Cardenas et al. 2005). Here, two proteins (spots 13 and 14) with increased intensity in germinated maize pollen were identified as actin.
Signaling proteins
In the present study, two protein spots were identified as signaling proteins, including rab GDP dissociation inhibitor α (rab-GDI) and adenosine kinase (ADK). Rab-GDI binds all rab GTPases and plays a key role in the rab GTPase cycle, which plays a critical role in the vesicular trafficking in pollen tube growth (Krichevsky et al. 2007). The increased level of Rab-GDI found here was in agreement with the recent finding (Szumlanski and Nielsen 2009).
ADK is an enzyme involved in the adenylate metabolic network, by which adenosine (Ado) is converted into adenosine 5′-monophosphate (AMP) using one molecule of adenosine 5′-triphosphate (ATP) (Moffatt et al. 2000). In this study, ADK was observed to be up-regulated in P1, indicating a possible involvement of ADK in pollen germination. However, Zou et al. (2009) found that ADK1 was only expressed in mature pollen, but not in pollen tubes.
Proteins involved in protein metabolism
Mitochondrial-processing peptidase (MPP) cleaves the vast majority of mitochondrial proteins, which is structurally and functionally conserved across species (Gakh et al. 2002). MPP (-like) was identified in the pollen proteome of rice (Dai et al. 2006) and tomato (Sheoran et al. 2007). We found here that MPP α subunit (spot 25) was up-regulated in germinated maize pollen, as in rice (Dai et al. 2006), suggesting a possible role of MPP in pollen tube growth.
Besides, protein disulfide isomerase, belonging to the thioredoxin family, was found to be up-regulated in P1. The protein is localized in ER and known as molecular chaperone involved in proper protein folding, a necessary process for protein transport (Sheoran et al. 2006).
Defense-related proteins
In the present study, three spots differentially expressed in P1 were identified as defense-related proteins: glutathione S-transferase (spot 10), ascorbate peroxidase (spot 7) and peroxidase 1 (spot 9). They were all down-regulated, as in germinated rice pollen (Dai et al. 2007). This may be explained by the fact that pollen germinated in vitro is under less oxidative stress compared to germination in vivo in field conditions. On the contrary, the presence of defense-related proteins in pollen (Holmes-Davis et al. 2005; Noir et al. 2005; Sheoran et al. 2006; Dai et al. 2006) may provide protection against oxidative stress. For example, three isoforms of ascorbate peroxidase were up-regulated in developing rice pollen (Imin et al. 2004).
Pollen allergens
The structures and biological functions of pollen allergens were recently reviewed (Pomés 2008). Zea m 1 (group 1) and Zea m 13 (group 13) were the most prominent allergens in maize pollen (Petersen et al. 2006). Zea m 1 represents an expansin, involved in the stretching of cell walls and in the growth of pollen tubes for fertilization (Cosgrove et al. 1997; Li et al. 2003). Its reduced abundance in pollen was associated with a male-sterile phenotype of maize (Wang et al. 2004b). At present, the function of Zea m 13 is not yet clear. In the present study, two down-regulated proteins were identified as Zea m 13 (NCBI accession no. ×57627), with a pI similar to previously reported value (6.8–7.6) (Petersen et al. 2006).
Additionally, two differentially expressed spots in P1 were not identified from the protein database (not shown). A large number of proteins reported in Arabidopsis and rice pollen (Holmes-Davis et al. 2005; Noir et al. 2005; Dai et al. 2006) also belong to the group with unknown function. Besides, our study identified 26 protein spots, which can be explained by the limitations of 2-DE in detecting low-abundance and integral membrane proteins. Further study is required to identify additional proteins, especially low-abundance and integral membrane proteins.
In conclusion, we found that the abundances of 28 protein spots, representing 21 distinct proteins, changed between mature and germinated maize pollen. The up-regulated proteins were mainly involved in tube wall modification, actin cytoskeleton organization, energy metabolism, signaling, protein folding and degradation. In particular, pectin methylesterase (spots 22 and 23), inorganic pyrophosphatase (spot 3), glucose-1-phosphate uridylyltransferase (spot 21) and rab GDP dissociation inhibitor α (spot 20) are highly up-regulated, suggesting their potential role in pollen tube growth. The down-regulated proteins mainly include defense-related proteins, pollen allergens and some metabolic enzymes. In addition, many proteins identified are found in different subcellular locations. This study would contribute to the understanding of the changes in protein expression associated with pollen germination.
Abbreviations
- 2D:
-
Two dimensional
- 2-DE:
-
Two-dimensional electrophoresis
- ADK:
-
Adenosine kinase
- MPP:
-
Mitochondrial-processing peptidase
- MALDI-TOF:
-
Matrix-assisted laser desorption/ionization time-of-flight
- Mr:
-
Molecular weight
- MS:
-
Mass spectrometry
- P0:
-
Mature pollen
- P1:
-
Pollen germinated for 1 h
- pI:
-
Isoelectric point
- PMEs:
-
Pectin methylesterases
- rab-GDI:
-
rab GDP dissociation inhibitor
References
Barnabas B, Fridvalszky L (1984) Adhesion and germination of differently treated maize pollen grains on the stigma. Acta Bot Hung 30:329–332
Bosch M, Hepler PK (2005) Pectin methylesterases and pectin dynamics in pollen tubes. Plant Cell 17:3219–3226
Bosch M, Cheung AY, Hepler PK (2005) Pectin methylesterase, a regulator of pollen tube growth. Plant Physiol 138:1334–1346
Cai G, Cresti M (2009) Organelle motility in the pollen tube: a tale of 20 years. J Exp Bot 60:495–508
Cardenas L, Lovy-Wheeler A, Wilsen KL, Hepler PK (2005) Actin polymerization promotes the reversal of streaming in the apex of pollen tubes. Cell Motil Cytoskel 61:112–127
Chen YM, Chen T, Shen SH, Zheng MZ, Guo Y, Lin J, Baluska F, Samaj J (2006) Differential display proteomic analysis of Picea meyeri pollen germination and pollen-tube growth after inhibition of actin polymerization by latrunculin B. Plant J 47:174–195
Cosgrove DJ, Bedinger P, Durachko DM (1997) Group I allergens of grass pollen as cell wall-loosening agents. Proc Natl Acad Sci USA 94:6559–6564
Dai S, Li L, Chen T, Chong K, Xue Y, Wang T (2006) Proteomic analysis of Oryza sativa mature pollen reveal novel proteins associated with pollen germination and tube growth. Proteomics 6:2504–2529
Dai S, Chen T, Chong K, Xue Y, Liu S, Wang T (2007) Proteomics identification of differentially expressed proteins associated with pollen germination and tube growth reveals characteristics of germinated Oryza sativa pollen. Mol Cell Proteomics 6:207–230
Feijo JA, Costa SS, Prado AM, Becker JD, Certal AC (2004) Signalling by tips. Curr Opin Plant Biol l7:589–598
Fernando DD (2005) Characterization of pollen tube development in eastern white pine (Pinus strobus) through proteomic analysis of differentially expressed proteins. Proteomics 5:4917–4926
Gakh O, Cavadini P, Isaya G (2002) Mitochondrial processing peptidases. Biochem Biophys Acta 1592:63–77
Geitmann A, Snowman BN, Emons AMC, Franklin-Tong VE (2000) Alterations in the actin cytoskeleton of pollen tubes are induced by the self-incompatibility reaction in Papaver rhoeas. Plant Cell 12:1239–1252
Golovkin M, Reddy ASN (2003) A calmodulin-binding protein from Arabidopsis has an essential role in pollen germination. Proc Natl Acad Sci USA 100:10563–10588
Grobei MN, Qeli E, Brunner E, Rehrauer H, Zhang RX, Roschitzki B, Basler K, Ahrens CH, Grossniklaus U (2009) Deterministic protein inference for shotgun proteomics data provides new insights into Arabidopsis pollen development and function. Genome Res 19:1786–1800
Holmes-Davis R, Tanaka CK, Vensel WH, Hurkman WJ, McCormick S (2005) Proteome mapping of mature pollen of Arabidopsis thaliana. Proteomics 5:4864–4884
Imin N, Kerim T, Rolfe BG, Weinman JJ (2004) Effect of early cold stress on the maturation of rice anthers. Proteomics 4:1873–1882
Krichevsky A, Kozlovsky SV, Tian GW, Chen MH, Zaltsman A, Citovsky V (2007) How pollen tubes grow. Dev Biol 303:405–420
Li LC, Bedinger PA, Volk C, Jones D, Cosgrove DJ (2003) Purification and characterization of four β-expansins (Zea m 1 isoforms) from maize pollen. Plant Physiol 132:2073–2085
Lockhart DJ, Winzeler EA (2000) Genomics, gene expression and DNA arrays. Nature 405:827–836
Malho R, Camacho L, Moutinho A (2006) Signaling pathways in pollen tube growth and reorientation. Ann Bot 85:59–68
Moffatt BA, Wang L, Allen MS, Stevens YY, Qin W, Snider J, von Schwartzenberg K (2000) Adenosine kinase of Arabidopsis. Kinetic properties and gene expression. Plant Physiol 124:1775–1785
Noir S, Brautigam A, Colby T, Schmidt J, Panstruga R (2005) A reference map of the Arabidopsis thaliana mature pollen proteome. Biochem Biophys Res Commun 337:1257–1266
Pertl H, Schulze WX, Obermeyer G (2009) The pollen organelle membrane proteome reveals highly spatial–temporal dynamics during germination and tube growth of lily pollen. J Proteome Res 8:5142–5152
Petersen A, Dresselhaus T, Grobe K, Becker WM (2006) Proteome analysis of maize pollen for allergy-relevant components. Proteomics 6:6317–6325
Pomés A (2008) Allergen structures and biologic functions: the cutting edge of allergy research. Curr Allergy Asthm Rep 8:425–432
Sagi G, Katz A, Guenoune-Gelbart D, Epel BL (2005) Class 1 reversibly glycosylated polypeptides are plasmodesmal-associated proteins delivered to plasmodesmata via the Golgi apparatus. Plant Cell 17:1788–1800
Sheoran IS, Sproule KA, Olson DJH, Ross ARS, Sawhney VK (2006) Proteome profile and functional classification of proteins in Arabidopsis thaliana (Landsberg erecta) mature pollen. Sex Plant Reprod 19:185–196
Sheoran IS, Ross AR, Olson DJ, Sawhney VK (2007) Proteomic analysis of tomato (Lycopersicon esculentum) pollen. J Exp Bot 58:3525–3535
Sheoran IS, Pedersen EJ, Ross AR, Sawhney VK (2009) Dynamics of protein expression during pollen germination in canola (Brassica napus). Planta 230:779–793
Suen DF, Huang AHC (2007) Maize pollen coat xylanase facilitates pollen tube penetration into silk during sexual reproduction. J Biol Chem 282:625–636
Szumlanski AL, Nielsen E (2009) The rab GTPase rabA4D regulates pollen tube tip growth in Arabidopsis thaliana. Plant Cell 21:526–544
Vidali L, McKenna ST, Hepler PK (2001) Actin polymerization is essential for pollen tube growth. Mol Biol Cell 12:2534–2545
Wang W, Vignani R, Scali M, Cresti M (2004a) Post-translational modifications of α-tubulin in Zea mays are highly tissue specific. Planta 218:460–465
Wang W, Scali M, Vignani R, Milanesi C, Petersen A, Sari-Gorla M, Cresti M (2004b) Male-sterile mutation alters Zea m 1 (β-expansin 1) accumulation in a maize mutant. Sex Plant Reprod 17:41–47
Wang W, Vignani R, Scali M, Cresti M (2006) A universal and rapid protocol for protein extraction from recalcitrant plant tissues for proteomic analysis. Electrophoresis 27:2782–2786
Wang Y, Zhang WZ, Song LF, Zou JJ, Su Z, Wu WH (2008) Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis. Plant Physiol 148:1201–1211
Wang W, Bianchi L, Scali M, Liu L, Bini L, Cresti M (2009) Proteomic analysis of β-1,3-glucanase in grape berry tissues. Acta Physiol Plant 31:597–604
Wilkins MR, Pasquali C, Appel RD, Ou K, Golaz O, Sanchez JC, Yan JX, Gooley AA, Hughes G, Humphery-Smith I, Williams KL, Hochstrasser DF (1996) From proteins to proteomes: large scale protein identification by two-dimensional electrophoresis and amino acid analysis. Nature Biotech 14:61–65
Wittink FRA, Knuiman B, Derksen J, Čapková V, Twell D, Schrauwen JAM, Wullems GJ (2000) The pollen-specific gene Ntp303 encodes a 69-kDa glycoprotein associated with the vegetative membranes and the cell wall. Sex Plant Reprod 12:276–284
Yang Z (2002) Small GTPases: versatile signaling switches in plants. Plant Cell 14:S375–S388
Zou JJ, Song LF, Zhang WZ, Wang Y, Ruan S, Wu WH (2009) Comparative proteomic analysis of Arabidopsis mature pollen and germinated pollen. J Integr Plant Biol 51:438–455
Acknowledgments
This work was supported by the National Natural Science Foundation of China (project 30971705) by the Program for Science and Technology Innovation Talents in Universities of Henan Province (project 2008HASTIT005) and by Innovation Scientists and Technicians Troop Construction Projects of Henan Province (project 9410051003).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Z.-L. Zhang.
The authors Y. Zhu and P. Zhao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhu, Y., Zhao, P., Wu, X. et al. Proteomic identification of differentially expressed proteins in mature and germinated maize pollen. Acta Physiol Plant 33, 1467–1474 (2011). https://doi.org/10.1007/s11738-010-0683-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11738-010-0683-7