Abstract
Metabolite profiles of green beans of Coffea arabica var. sigararutang processed with three different treatments, including dry, semi-dry, and wet postharvest methods, were evaluated with 1H NMR spectroscopy analysis coupled with multivariate data analysis. Partial least square discriminant analysis (PLSDA) models successfully classified the green coffee beans based on the postharvest methods. S-plots of two-classes orthogonal projection to latent structure-discriminant analysis (OPLSDA) revealed that the dry-processed coffee was characterized with fructose and GABA, while the wet-processed coffee was represented by chlorogenic acids (caffeoyl quinic acids, CQAs), malic acid, and sucrose. Meanwhile, the green coffee beans processed with semi-dry postharvest method possessed the intermediary metabolite profile between the dry and the wet coffees. Generally, the semi-dry coffee exhibited the highest antioxidant activity compared to the other coffee samples. In this report, 1H NMR-based metabolic profiling successfully evaluated metabolite profiles of green coffee beans treated with three different postharvest methods and also revealed the characteristic compounds for each coffee samples.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Coffee is the second most consumed beverage in the world after the plain water. Over 3 billion cups of coffee are consumed in the world every day [1]. Coffea arabica (arabica) and Coffea canephora (robusta) are the widely cultivated coffee species. Around 70% of available coffee products in the world market is arabica coffee, while the rest 30% mainly is robusta coffee [1]. Compared to the other coffee species, arabica is considered to have a better quality due to its diminished bitterness, lower caffeine content, pronounced aroma, heightened acidity, sweeter profile, milder taste, and richer flavor. These attributes contribute to its heightened desirability and higher market value [2, 3].
Coffee quality is determined by some factors, including the cultivar genetic properties, the geographic condition, the agricultural practices, and the postharvest treatment [4]. Among these factors, the postharvest treatment provides instant impacts to the coffee quality by altering the chemical constituent, the flavor, the aroma, and the physical properties [5]. At least, there are three common methods in the coffee postharvest, including the wet process called as the full-washed, the dry process commonly known as the natural, and semi-dry referred to the honey method. The wet process is considered able to produce a more acceptance coffee quality when compared to the dry method [6]. Meanwhile, the drying process is the most important step in the dry method since impacts the final coffee quality [7]. In the semi-dry method, the sorted coffee cherries are peeled, dried under the sun, and then hulled mechanically [8].
Some metabolomics approaches had been employed for discriminating the metabolite profiles of the green coffee beans processed with the different postharvest treatments. The analytical techniques applied in these studies were NIR spectroscopy [9], UV spectroscopy [10], ESI–MS [11], DESI-MS [12], ESI FT-ICR MS [13], GC [14], GCMS [14, 15] and LC–MS/MS [16]. Recently, LC–MS/MS was employed to tentatively identify 136 bioactive metabolites in coffee samples obtained from different continents [17]. Beside the analytical techniques mentioned before, 1H NMR spectroscopy is one of the most used methods in metabolomics. This method had been applied to study the coffee metabolite profiles based on the geographic origins [18], species [19], varieties [20], roasting levels [21], coffee quality [2], sensory [22], and the authenticity study [23]. Recently, 1H NMR-based metabolomics was also employed to investigate metabolite profiles of the wild and the caged civet (Luwak) coffees [24].
Indonesia is the fourth largest coffee producers in the world after Brazil, Vietnam, and Columbia [25]. One of the most cultivated arabica coffee varieties in Indonesia is sigararutang. The green beans of sigararutang coffee are also exported widely oversea. Sigararutang is a superior coffee variety having high productivity, interesting flavor, dry resistance, fast growth, and rapid fruiting [26]. This arabica coffee variety was initially found in Lintong, Humbang Hasundutan, North Sumatera, and is considered as a natural hybrid between blawan pasumah (derived from typica) and catimor [27].
In this report, the metabolite profiles of green coffee beans processed with dry, semi-dry and wet methods were evaluated with 1H NMR spectroscopy combined with chemometric analysis. The sample used in this work was the certified C. arabica var. sigararutang, widely cultivated in Indonesia coffee plantations. The antioxidant property of the green coffee beans was examined as well. To the best of our knowledge, it is the first report concerning the application of 1H NMR-based metabolomics in investigating the influence of the different postharvest methods on the metabolite profiles of the green coffee beans.
Materials and methods
Materials
Coffee samples used in this work were C. arabica var. sigararutang cultivated in Malabar Mountain (around 1800 m above sea level), Pangalengan, Bandung Residency, and processed with 3 different postharvest methods (dry, semi-dry, and wet) by Rahayu Farmer Group, West Java, Indonesia. The variety of this coffee sample was certified by the Directorate General of Plantation, Ministry of Agriculture, Republic of Indonesia, under license No. 65/Kpts/Sr.120/2/2014. All samples were deposited in Organic Chemistry Laboratory, Bandung Institute of Technology, with voucher numbers: the wet-processed coffee, BPM-1FW; the dry-processed coffee, BPM-1N; the semi-dry-processed coffee, BPM-1H. Deuterium oxide (D2O), methanol, ferric chloride (FeCl3), ethanol, sodium acetate trihydrate (CH3COONa.3H2O), acetic acid, potassium dihydrogen phosphate (KH2PO4), dipotassium hydrogen phosphate (K2HPO4), and 3-(trimethylsilyl)- 2,2,3,3-tetradeuteropropionic acid sodium salt (TSP) were bought from Merck (Darmstadt, Germany). Ascorbic acid, 1,1-diphenyl-2-picryl hydrazyl (DPPH,), 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) diammonium salts (ABTS), potassium persulfate (K2O8S2), and 2,4,6 tripyridyl-s-triazine (TPTZ), were purchased from Sigma-Aldrich (St. Louis, USA).
Extraction
The green coffee beans were crushed into powder with a 600N coffee grinder (Yang Chia Machine Work, Taiwan). Around 400 mg of coffee powder was mixed with 2 mL of D2O containing sodium phosphate buffer (pH 6.00) and TSP in a plastic tube. The mixture was sonicated for 20 min, incubated at 95 °C in a Memmert wnb 22 waterbath (Memmert, Schwabach, Germany) for 30 min, and then cooled on water for 10 min. Afterward, the solution was centrifuged at 12,000 rpm with a MC-12 microcentrifuge (Benchmark, New Jersey, United States) for 5 min. Finally, 500 μL of the supernatant was placed into a 5 mm NMR tube.
1H NMR measurement
1H NMR spectra were recorded with a Variant Unity INOVA-500 Spectrometer (Agilent Technologies, Santa Clara, United States). The water signal was suppressed with the presauration method. The parameters of the 1H NMR measurement were number of scans, 128; number of data points, 64 K; spectral width, 8012.8 Hz; acquisition time, 2.720 s; and delay time, 2.0 s. The Free Induction Decay (FID) 1H NMR data were further processed with ACD/Labs 12.0 software (ACD/Labs, Toronto, Canada). The spectrum baseline was corrected, and the chemical shift was calibrated with the TSP signal.
Data extraction
1H NMR spectra of green coffee beans were further processed with alignment and bucketing techniques using the ACD/Labs 12.0 software (ACD/Labs, Toronto, Canada). Bucketing with the intelligent option was applied to produce integrated bins with the equal width of 0.04 ppm within δ 0.50–10.00 ppm. The buckets containing residual water signals (δ 4.75–5.20 ppm) were removed from the analysis. Caffeine buckets at δ 3.22–3.49 and 3.82–3.88 ppm were also excluded since their chemical shifts were changeable. The buckets of sucrose at δ 3.46–3.52, 3.54–3.60, 3.66–3.72, and 3.74–3.96 were ignored in the analysis since overlapped with the signals of other sugar compounds in the dry and semi-dry coffee samples. The extracted data were then transferred into Microsoft Excel software for normalization with the sum method to avoid the bias effects.
Multivariate data analysis
After normalization, the data were exported into SIMCA-P version 12.0 (Umetrics, Umeå, Sweden). The data were treated with Pareto scaling to reduce the mask effect in the data analysis. Principal component analysis (PCA), unsupervised approach, was first created to evaluate the intrinsic variation in the data. Hotelling’s T2 technique (an ellipse in the score plots) explained the 95% confidence interval of the model variation. In this analysis, partial least square discriminant analysis (PLSDA) was applied as the main model for classifying the coffee metabolomes. The green coffee bean data were grouped into three classes based on their postharvest methods, and then examined with PLSDA models. PLSDA model quality was explained by R2X, R2Y and Q2 values. R2X and R2Y described the data variation and indicated the fit goodness. Q2 explained the variation predicted by the models according to the cross validation. PLSDA models were validated by a permutation test with 200 iterations.
Relative quantification
Concentrations of some identified metabolites were determined relatively with 1H NMR quantitative analysis. TSP (1 mM) signal was used as the reference compound since having a singlet peak and does not overlap with other signals. The metabolite concentration was calculated by comparing the proton signal integration of the targeted metabolites with the singlet signal integration of TSP. One-way ANOVA and Tukey’s range test for the statistical calculation of the quantitative analysis were performed with Minitab 19 software (Minitab, Sydney, Australia).
Bioassays
In this work, the green coffee bean samples were examined with antioxidant tests, including the tests of DPPH radical (DPPH·) and ABTS radical cation (ABTS·+) scavenging activities, and ferric-reducing antioxidant power (FRAP) assay. These antioxidant tests were performed based on the procedures in our previous report [24].
Results and discussions
Metabolite identification
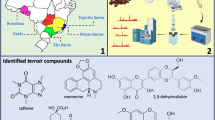
In this research, the metabolites were identified in the 1H NMR spectra of the green coffee beans (Fig. 1) by detecting their fingerprint peaks and comparing with the reference spectrum obtained from HMDB database (https://hmdb.ca/). The detected peaks were validated with 2D NMR spectra analysis, including COSY, TOCSY, and J-resolved. Moreover, the results were further clarified by comparing with the data acquired from literature [2, 28,29,30]. In total, 26 metabolites were successfully identified in the 1H NMR spectra of green coffee beans processed with the dry and the semi-dry postharvest methods. Meanwhile, 24 metabolites were detected in the 1H NMR spectra of the wet-processed green coffee beans. Figure 2 depicted molecular structures of some identified metabolites in the green coffee bean samples.
Sucrose, the main sugar compound in the green coffee beans, was clearly detected in the 1H NMR spectra. The peaks at δ 3.48 (t), 3.57 (dd), 3.78 (t), 3.86 (m), and 5.42 (d) ppm were the proton signals belonging to the glucose moiety of sucrose, namely H-4, H-2, H-3, H-5, and H-1, respectively. Meanwhile, the proton signals of the fructose moiety of sucrose were detected at δ 3.69 (s), 3.84 (m), 3.90 (m), 4.07 (t), and 4.23 (d) ppm, assigned as H-1’, H-6’, H-5’, H-3’, and H-4’, respectively. The free monosaccharides forming sucrose, namely glucose and fructose, were detected in the 1H NMR spectra as well. The fingerprint signals of fructose protons were identified at δ 3.56 (H-3; H-4, m), 3.72 (H-1, H-6, m), and 4.04 (H-5, m). Meanwhile, the proton peaks of glucose were detected at δ 3.26 (H-2, m) and 4.66 (H-1, d). Another sugar compound discovered in the green coffee beans was myo-inositol. The fingerprint proton signals of myo-inositol were identified at δ 3.27 (H-5, t), 3.53 (H-2, m), and 3.62 ppm (H-4, H-6, m).
Alpha amino acids were successfully detected in the 1H NMR spectra, including alanine, asparagine, and valine. The fingerprint proton signal of alanine was clearly found at δ 1.49 ppm with doublet multiplicity and assigned as H-3, while the proton signals of asparagine were identified at δ 2.87 (H-3b, dd), 2.97 ppm (H-3a, dd) and 4.02 (H-2, m). The proton signals at δ 1.01 (d) and 1.05 ppm (d), only found in 1H NMR spectra of the dry and semi-dry coffees, were designated as H-3 and H-4 of valine, respectively. Gamma butyric acid (GABA) was another amino acid identified in all coffee samples. The peaks at δ 1.92 (m), 2.32 (t) and 3.03 ppm (t) were appointed as H-3, H-2, and H-4 of GABA, respectively.
Caffeine, chlorogenic acids (CQAs), and trigonelline were other metabolites detected obviously in the 1H NMR spectra indicating as major compounds in all coffee samples. The proton signals of 3 methyl groups attached to nitrogen atoms of caffeine were found at δ 3.22 (H-11, s), 3.39 (H-10, s), and 3.85 ppm (H-12, s), while its aromatic proton was identified at δ 7.77 ppm (H-8) as a singlet peak. Proton signals of trigonelline were discovered explicitly at δ 4.43 (H-8, s), 8.08 (H-5, t), 8.82 (H-6, m), 8.84 (H-4, m), and 9.12 (H-2, s). The detailed proton signals of CQAs, including 3-caffeoyl quinic acid (3-CQA), 4-caffeoyl quinic acid (4-CQA), and 5-caffeoyl quinic acid (5-CQA), were described in Table 1.
Quinic acid, a precursor compound of chlorogenic acids was successfully recognized in the 1H NMR spectra. The signals at δ 1.89 (dd), 1.98 (m), 2.08 (m), 4.03 (m), 4.16 (m) were assigned as H-2a, H-6a, H-2e and H-6e, H-3, and H-5 of quinic acid, respectively. Acidic short chain compounds were also detected in the coffee samples, including acetic acid (δ 1.95 ppm), citric acid (δ 2.60 and 2.72 ppm), formic acid (δ 8.47 ppm), fumaric acid (δ 6.54 ppm), lactic acid (δ 1.34 and 4.13 ppm), malic acid (δ 2.44, 2.2.72, and 4.34 ppm), and succinic acid (δ 2.46 ppm).
Fatty acid lipids were successfully identified in the green coffee bean samples, including palmitic acid, stearic acid, and linoleic acid. The methyl group of palmitic acid was noted at δ 0.91 ppm (H-16), while its methylene signal close to carboxylic acid functional group was recorded at δ 1.66 (H-3). Other methylene signals of palmitic acid were detected at δ 1.31 ppm (H-4 up to H-15). The proton signals at δ 1.02, 1.42, 1.73, and 2.47 ppm corresponded to H-18, H-4 up to H-17, H-3, and H-2 of stearic acid, respectively. Meanwhile, the proton signals assigned to linoleic acid were explained completely in Table 1. The identification of linoleic acid was in accordance with the recent report identifying this compound in the coffee with LC–MS/MS [17]. Other metabolites successfully detected in the coffee samples were choline (δ 3.21 and 3.53 ppm) and nicotinic acid (δ 8.62 and 8.94 ppm).
The identification of caffeine, trigonelline, chlorogenic acids, sucrose, fructose, myo-inositol, quinic acid, citric acid, malic acid, fumaric acid, succinic acid, lactic acid, and acetic acid, was in accordance with the previous report that employed LC–MS/MS for detecting these compounds in the green coffee beans treated with different postharvest methods as well [16]. It indicated that, despite its low sensitivity, 1H NMR-based metabolomics has the capability to identify the same compounds as detected by LC–MS/MS.
Metabolome comparison of green coffee beans
Multivariate data analysis was performed to investigate the metabolome profiles of green coffee beans treated with different postharvest methods. In the initial approach, the data extracted from the 1H NMR spectra of all green coffee bean samples, were analyzed with PCA model, unsupervised approach. After pareto scaling applied, the PCA model had 3 components explaining 83.6% of cumulative variances (R2X) and 69.7% of cumulative cross-validated variance (Q2) of spectral data. PCA score plot combining PC1 (59.1%) and PC2 (16 0.2%), successfully described 75.3% of variation in the data and almost provided 3 well-separated clusters based on the differences in postharvest methods. In this score plot, one sample of the dry-processed was situated within the cluster corresponding to the semi-dry coffee, as illustrated in Fig. 3a. Nonetheless, this score plot successfully distinguished the wet coffee from the rest of the samples, indicating its unique metabolite profile.
Multivariate data analysis calculated for the green beans of C. arabica var. sigararutang processed with dry, semi-dry, and wet methods. a PCA score plot separated the wet coffee from the others. b PLSDA score plot successfully classified the coffee samples based on their postharvest methods. c PLSDA loading plot unveiled the significant buckets influencing to the discrimination
To obtain better classification, the data were further evaluated with PLSDA model, a supervised approach. In this model, the samples were labeled based on their postharvest methods. This PLSDA model consisted of 4 components with 85.0% and 97.5% cumulative variances (R2X and R2Y, respectively), in which the first 2 components explained 71.5% of total variation. Meanwhile, this model possessed 86.3% of the variance based on the cross validation (Q2), indicating a good predictive ability. The best sample classification in the score plot was obtained by combining the first (57.9%) and the second (13.6%) components as depicted in Fig. 3b. This score plot separated the coffee samples based on their postharvest methods. The first component discriminated the wet coffee from the others, while the second component distinguished the dry coffee from the semi-dry coffee samples. As seen in the PLSDA score plot (Fig. 3b), the position of dry coffee cluster was closer to the semi-dry coffee cluster compared to the wet coffee cluster, suggesting both had more similarity in the metabolite profile term.
To identify the responsible buckets for the sample classification in the PLSDA score plot, the corresponding loading plot was examined. As described in Fig. 3c, at least there were 20 buckets contributing to the classification. These buckets corresponded to some major metabolites, including sucrose (δ 4.04–4.10, 4.20–4.25, and 5.39–5.45 ppm) and CQAs (δ 2.03–2.09, 2.12–2.18, 2.18–2.14, 4.25–4.31, and 5.31–5.39 ppm). Buckets belong to acetic acid (δ 2.41–2.44 and 1.93–1.98 ppm), malic acid (δ 4.31–4.37 ppm), fructose (δ 3.63–3.66 and 4.00–4.04 ppm), glucose (δ 3.72–3.74 ppm), and GABA (δ 2.29–2.35 and 3.00–3.06 ppm) were also detected influencing the discrimination. Other buckets found contributing to the classification, were bucket at δ 1.31–1.37 ppm corresponded to lactic acid, buckets at δ 0.88–0.94 and 1.28–131 ppm assigned to fatty acids, including linoleic acid and palmitic acid, and bucket at δ 1.40–1.46 ppm designed to stearic acid, another major fatty acid found in the green coffee beans. Among those, the buckets belonging to sucrose, CQAs, lactic acid, acetic acid, fructose, and GABA were identified as the more influence buckets to the classification of green coffee bean samples based on their postharvest methods.
Three models of two-classes OPLSDA were produced for comparing the metabolite profiles of green coffee bean samples one on one. Furthermore, the corresponding S-plots of the two-classes OPLSDA models were evaluated to reveal the most discriminant metabolites for each green coffee bean sample. The first two-classes OPLSDA model was created to discriminate wet-processed coffee from the dry-processed coffee. This OPLSDA model had 3 components describing 84.9% and 99.8% of total variances (R2X and R2Y, respectively), and 97.8% of total variance according to the cross validation (Q2). The S-plot of this two-classes OPLSDA model (Fig. 4a) revealed that the wet coffee was characterized with sucrose, CQAs, and malic acid, while the dry coffee was represented with acetic acid, fructose, glucose, and GABA.
The second two-classes OPLSDA model was generated to compare the metabolite profiles between the wet coffee with the semi-dry coffee. This model possessed 3 components with 80.7% of R2X, 99,6% of R2Y, and 98.9% of Q2. Interestingly, the characteristic metabolites of the wet coffee in the first two-classes OPLSDA model were also identified as the discriminant compounds for this coffee type in the corresponding S-plot (Fig. 4b). Thus, it suggested that sucrose, CQAs, and malic acid are potential markers for the green coffee bean processed with the wet method. Meanwhile, acetic acid, fructose, glucose, and GABA were detected as the discriminant compounds of the semi-dry coffee. These metabolites were also the characteristic compounds of the dry coffee when compared to the wet coffee (Fig. 4a). Therefore, it indicated that the wet coffee exhibited the lowest levels of acetic acid, fructose, glucose, and GABA among the samples.
The last OPLSDA model, built to distinguish the dry coffee from the semi-dry coffee, possessed 3 components with 74.5% of R2X, 98.5% of R2Y, and 82.3% of Q2. In the S-plot of this model (Fig. 4c), sucrose and CQAs were identified as the characteristic compounds of the semi-dry coffee. As seen in Fig. 4b, oppositely these metabolites did not correlate to the semi-dry coffee but represented the wet coffee. Thus, it suggested that the metabolite profile of the semi-dry coffee was the intermediary between the dry and the wet coffees. Meanwhile, lactic acid, fructose, GABA, alpha-linoleic acid, linoleic acid, palmitic acid, stearic acid, and lipid were observed as the discriminant metabolites of the dry coffee. Among these metabolites, fructose and GABA were always found as the characteristic compound of the dry coffee in the corresponding S-plot (Fig. 4a and c), indicating as the potential marker candidate for the green coffee beans processed with the dry postharvest method.
Relative quantification
In this work, concentrations of 16 identified metabolites in the green beans of the coffee samples were determined relatively with quantitative 1H NMR analysis as depicted in Table 2. The two hexoses glucose and fructose exhibited the highest levels in the coffees processed with the dry method, and the lowest in the wet coffee. Meanwhile, both compounds in the semi-dry coffee were in the intermediate amounts. The lower contents of glucose and fructose in the green coffee beans processed with the wet method were suggested as a consequence of sugar metabolism in anaerobic fermentation on the coffee endosperm during the soaking process [31]. These results indicated that the concentrations of glucose and fructose were significantly affected by the postharvest processing methods. It also confirmed the results of multivariate data analysis explained previously and was in agreement with the previous reports [16, 31].
Previous works reported that the sucrose content, the major sugar compound in the green coffee beans, was not influenced by the postharvest methods but affected by the preharvest factors, including the year of production [31, 32]. In contrast, other reports documented that the green coffee beans processed with the wet method contained lower sucrose level compared to the dry and semi-dry coffee samples [16, 33]. Thus, the previous reports indicated the impact of postharvest methods on sucrose is still unclear. Interestingly, as depicted in Table 2, the highest sucrose concentration (43.49 ± 1.89 mM) was detected in the green coffee beans processed with the wet method, while the lowest level of this compound (31.90 ± 1.16 mM) was found in the dry coffee samples. Moreover, sucrose was also identified as the characteristic compound of the wet coffee sample in the discrimination of the coffee samples described in the previous section. It had been reported that during the storage, the sucrose concentration decreased as a consequence of the hydrolyzation yielding fructose and glucose [34]. The authors mentioned the sucrose hydrolyzation was accelerated with the increase of the humidity and the storage time [34]. The green coffee beans processed with the dry method require the longest drying time compared to the other postharvest methods since it is dried in the form of intact fruits. In addition, Indonesia is well known as a tropical country with the high humidity. Hence, it is plausible to propose that the reduced concentration of sucrose detected in the dry coffee could be attributed to the hydrolysis of sucrose, resulting in the formation of glucose and fructose. This transformation might have occurred during the prolonged drying process under sunlight with elevated humidity levels. It possibly correlated with the high concentrations of glucose and fructose observed in the dry coffee (Table 2).
Other metabolites found in the highest concentration in the wet coffee were 4-CQA, 5-CQA, and malic acid. This result verified those compounds as the discriminant compounds of the wet coffee described in the metabolome comparison section. The highest contents of CQAs in the wet coffee confirmed the data in the previous reports [1, 2, 11, 33]. A different case was found when comparing the concentrations of acidic compounds, including acetic, formic, and lactic acids, yielded from the microbial fermentation. These compounds were observed with the highest concentration on the dry coffee samples, while the lowest was on the wet coffee. This result confirmed the data in the previous report [16]. In the dry and semi-dry postharvest methods, the green beans were still covered by the mucilage during the drying under the sun. It facilitated the longer microbial fermentation on the mucilage producing more acidic compounds which can diffuse and accumulate into the beans [35, 36] as we observed in the dry and the semi-dry coffees. Quinic acid was the other acidic compound detected with the highest concentration on the dry coffee. Hydrolyzation of CQAs yielding quinic acid possibly led to the accumulation of this compound in the dry coffee. It was supported by the lower content of CQAs in this coffee.
The dry and the semi-dry coffees possessed the higher concentrations of GABA, while the wet coffee contained GABA in a trace level. This compound is associated as a metabolite involved in the response to various stress conditions and formed through α-decarboxylation of glutamic acid [37]. The higher content of GABA in the dry and the semi-dry coffees was suggested as a consequence of the longer drying period triggering the intense stress response [38]. The contents of other quantified compounds, including alanine, asparagine, and caffeine, in all coffee samples were not much differ. For instance, concentrations of caffeine in the dry, the semi-dry, and the wet coffees were 7.21 ± 0.13, 6.78 ± 0.02, and 7.02 ± 0.07 mM, respectively. These results suggested that the compounds were not influenced significantly by the postharvest methods, and it was in agreement with the previous reports [16, 33, 38].
Antioxidant activity
The antioxidant activity of the green coffee bean samples was evaluated using DPPH·, ABTS·+, and FRAP tests. For DPPH· and ABTS·+ tests, the IC50 values, indicating the concentration of the sample required to effectively inhibit 50% of DPPH· and ABTS·+, were calculated. Meanwhile, the EC50 value for the FRAP test, representing the effective concentration of the sample in exhibiting 50% of the FRAP capacity, was determined. As depicted in Table 3, the semi-dry coffee possessed the highest antioxidant activity in all tests compared to the others. This result confirmed the previous report [39]. The semi-dry coffee showed the IC50 values of 18.07 ± 0.06 mg/L (p value < 0.05) and 21.88 ± 0.68 mg/L (p value < 0.05) in the DPPH· and ABTS·+ tests, respectively. Meanwhile, the EC50 value of this coffee in FRAP capacity assay was 19.05 ± 0.04 mg/L (p value < 0.05). The dry coffee sample had slightly greater antioxidant activity than the wet coffee. In the ABTS·+ assay, the IC50 value for dry coffee was recorded at 29.77 ± 0.01 mg/L (p value < 0.05), indicating slightly higher activity compared to wet coffee (31.68 ± 0.09 mg/L, p value < 0.05). This trend was also observed in the FRAP capacity test, where the dry coffee had an EC50 value of 26.96 ± 0.01 mg/L (p value < 0.05), while the wet coffee had an EC50 value of 28.88 ± 0.04 mg/L (p value < 0.05). However, both the dry and wet coffees demonstrated comparable IC50 values in the DPPH assay, specifically 24.01 ± 0.06 mg/L (p value < 0.05) for the dry coffee, and 23.01 ± 1.68 mg/L (p value < 0.05) for the wet coffee.
Conclusions
Green beans of Coffea arabica var. sigararutang processed with three different postharvest, including dry, semi-dry, and wet methods, were successfully evaluated with 1H NMR-based metabolic profiling and antioxidant assays. Multivariate data analysis classified the coffee samples based on the postharvest methods and revealed the important discriminant compounds. The green coffee bean treated with the wet method was characterized with CQAs, sucrose, and malic acid. The coffee sample processed with the dry technique was discriminated with acetic acid, fructose, glucose, and GABA. Meanwhile, the green coffee beans subjected to the semi-dry postharvest method exhibited an intermediary metabolite profile, positioned between that of the dry and wet coffee samples. Relative quantification with 1H NMR spectroscopy method, confirmed that the contents of CQAs, acetic acid, formic acid, fructose, GABA, glucose, lactic acid, malic acid, quinic acid, and sucrose in the green coffee beans were influenced significantly by the postharvest methods. Bioactivity assays suggested that the semi-dry postharvest method led to the better antioxidant activity in the green coffee beans.
References
International Coffee Organization (ICO), Coffee development report 2019, International Coffee Organization (2019)
D.-J. Kwon, H.-J. Jeong, H. Moon, H.-N. Kim, J.-H. Cho, J.-E. Lee, K.S. Hong, Y.-S. Hong, Assessment of green coffee bean metabolites dependent on coffee quality using a 1H NMR-based metabolomics approach. Food Res. Int. 67, 175–182 (2015). https://doi.org/10.1016/j.foodres.2014.11.010
P. Lashermes, F. Anthony, Coffee, in Technical crops: genome mapping and molecular breeding in plants. ed. by B.C. Kole (Springer, Berlin, 2007), pp.109–118
F. Echeverria-Beirute, S.C. Murray, P. Klein, C. Kerth, R. Miller, B. Bertrand, Rust and thinning management effect on cup quality and plant performance for two cultivars of Coffea arabica L. J. Agric. Food Chem. 66, 5281–5292 (2018). https://doi.org/10.1021/acs.jafc.7b03180
D. Selmar, M. Kleinwachter, G. Bytof, Metabolic responses of coffee beans during processing and their impact on coffee flavor, in Cocoa and coffee fermentations. ed. by R.F. Schwan, G.H. Fleet (CRC Press, Boca Raton, Florida, USA, 2014), pp.431–476. https://doi.org/10.1201/b17536
K.B. do Carmo, J.C.B. do Carmo, Marcelo Rodrigo Krause, Aldemar Polonini Moreli, Paola Alfonsa Vieira Lo Monaco, Quality of Arabic coffee under different processing systems, drying methods, and altitudes. Biosci. J. 36, 1116–1125 (2020). https://doi.org/10.14393/BJ-v36n4a2020-47890
Y. Hamdouche, J.C. Meile, D.N. Nganou, N. Durand, C. Teyssier, D. Montet, Discrimination of post-harvest coffee processing methods by microbial ecology analyses. Food Control 65, 112–120 (2016). https://doi.org/10.1016/j.foodcont.2016.01.022
G.V.M. Pereira, D.P.C. Neto, A.I.M. Júnior, Z.S. Vásquez, A.B.P. Medeiros, L.P.S. Vandenberghe, C.R. Soccol, Exploring the impact of postharvest processing on the aroma formation of coffee beans—a review. Food Chem. 272, 441–452 (2019). https://doi.org/10.1016/j.foodchem.2018.08.061
S. Buratti, N. Sinelli, E. Bertone, A. Venturello, E. Casiraghi, F. Geobaldo, Discrimination between washed Arabica, natural Arabica and Robusta coffees by using near infrared spectroscopy, electronic nose and electronic tongue analysis. J. Sci. Food Agric. 95, 2192–2200 (2015). https://doi.org/10.1002/jsfa.6933
D. Suhandy, M. Yulia, Classification of Lampung robusta specialty coffee according to differences in cherry processing methods using UV spectroscopy and chemometrics. Agriculture 11, 109 (2021). https://doi.org/10.3390/agriculture11020109
A.C.L. Amorim, A.M.C. Hovell, A.C. Pinto, M.N. Eberlin, N.P. Arruda, E.J. Pereira, H.R. Bizzo, R.R. Catharino, Z.B.M. Filho, C.M. Rezende, Green and roasted arabica coffees differentiated by ripeness, process and cup quality via electrospray ionization mass spectrometry fingerprinting. J. Braz. Chem. Soc. 20, 313–321 (2009). https://doi.org/10.1590/S0103-50532009000200017
R. Garrett, N.V. Schwab, E.C. Cabral, B.V.M. Henrique, D.R. Ifa, M.N. Eberlin, C.M. Rezende, Ambient mass spectrometry employed for direct analysis of intact arabica coffee beans. J. Braz. Chem. Soc. 25, 1172–1177 (2014). https://doi.org/10.5935/0103-5053.20140094
A. Tsukui, P.H. Vendramini, R. Garrett, M.B.S. Scholz, M.N. Eberlin, H.R. Bizzo, C.M. Rezende, Direct-infusion electrospray ionization-mass spectrometry analysis reveals atractyligenin derivatives as potential markers for green coffee postharvest discrimination. LWT 103, 205–211 (2019). https://doi.org/10.1016/j.lwt.2018.12.078
A.P.L.R. de Oliveira, P.C. Correa, E.L. Reis, G.H.H. de Oliveira, Comparative study of the physical and chemical characteristics of coffee and sensorial analysis by principal components. Food Anal. Methods 8, 1303–1315 (2014). https://doi.org/10.1007/s12161-014-0007-4
F. Amalia, P. Aditiawati, Yusianto, S.P. Putri, E. Fukusaki, Gas chromatography/mass spectrometr-based metabolite profiling of coffee beans obtained from different altitudes and origins with various postharvest processing. Metabolomics 17, 69 (2021). https://doi.org/10.1007/s11306-021-01817-z
F. De Bruyn, S.J. Zhang, V. Pothakos, J. Torres, C. Lambot, A.V. Moroni, M. Callanan, W. Sybesma, S. Weckx, L. De Vuys, Exploring the impacts of postharvest processing on the microbiota and metabolite profiles during green coffee bean production. Appl. Environ. Microbiol. 83, e02398-e12316 (2017). https://doi.org/10.1128/aem.02398-16
A. Ali, H.F. Zahid, J.J. Cottrell, F.R. Dunshea, A comparative study for nutritional and phytochemical profiling of Coffea arabica (C. arabica) from different origins and their antioxidant potential and molecular docking. Molecules 27, 5126 (2022). https://doi.org/10.3390/molecules27165126
R. Consonni, L.R. Cagliani, C. Cogliati, NMR based geographical characterization of roasted coffee. Talanta 88, 420–426 (2012). https://doi.org/10.1016/j.talanta.2011.11.010
V.A. Arana, J. Medina, R. Alarcon, E. Moreno, H. Schaefer, J. Wist, Coffee’s country of origin determined by NMR: the Colombian case. Food Chem. 175, 500–506 (2015). https://doi.org/10.1016/j.foodchem.2014.11.160
N. Happyana, A. Pratiwi, H.H. Hakim, Metabolite profiles of the green beans of Indonesian Arabica coffee varieties. Int. J. Food Sci. 2021, 782578 (2021). https://doi.org/10.1155/2021/5782578
F. Wei, K. Furihata, M. Koda, F. Hu, T. Miyakawa, M. Tanokura, Roasting process of coffee beans as studied by nuclear magnetic resonance: time course of changes in composition. J. Agric. Food Chem. 60, 1005–1012 (2012). https://doi.org/10.1021/jf205315r
F. Wei, K. Furihata, T. Miyakawa, M. Tanokura, A pilot study of NMR-based sensory prediction of roasted coffee bean extracts. Food Chem. 152, 363–369 (2014). https://doi.org/10.1016/j.foodchem.2013.11.161
A.T. Toci, M.V.M. Ribeiro, P.R.A.B. Toledo, N. Boralle, H.R. Pezza, L. Pezza, Fingerprint and authenticity roasted coffees by 1H-NMR: the Brazilian coffee case. Food Sci. Biotechnol. 27, 19–26 (2017). https://doi.org/10.1007/s10068-017-0243-7
L. Febrina, N. Happyana, Y.M. Syah, Metabolite profiles and antidiabetic activity of the green beans of Luwak (civet) coffees. Food Chem. 355, 129496 (2021). https://doi.org/10.1016/j.foodchem.2021.129496
International Coffee Organization (ICO), Coffee development report 2020, International Coffee Organization (2020)
S. Suswati, S. Hutapea, R.I. Barus, S. Setiawan, A.P. Hutapea, Integrated control of coffee bean borer (Hypothenemus Hampei) on sigararutang coffee, Motung Village, Ajibata SubDistrict, Toba Samosir District, Sumatera Utara. Bp. Int. Res. Exact Sci. 2, 52–61 (2020). https://doi.org/10.33258/birex.v2i1.700
The Ministry of Agriculture of the Republic of Indonesia, Decree of the Minister of Agriculture No. 205/Kpts/SR.120/4/ 2005: The Release of the Sigararutang Coffee Variety as the Superior Variety, Jakarta (2005)
A. Azizan, M.S.A. Bustamam, M. Maulidiani, K. Shaari, I.S. Ismail, N. Nagao, F. Abas, Metabolite profiling of the microalgal diatom Chaetoceros calcitrans and correlation with antioxidant and nitric oxide inhibitory activities via 1H NMR-based metabolomics. Mar. Drugs 16, 154 (2018). https://doi.org/10.3390/md16050154
B. de Falco, G. Incerti, R. Bochicchio, T.D. Phillips, M. Amato, V. Lanzotti, Metabolomic analysis of Salvia hispanica seeds using NMR spectroscopy and multivariate data analysis. Ind. Crops Prod. 99, 86–96 (2017). https://doi.org/10.1016/j.indcrop.2017.01.019
F. Wei, K. Furihata, F. Hu, T. Miyakawa, M. Tanokura, Complex mixture analysis of organic compounds in green coffee bean extract by two-dimensional NMR spectroscopy. Magn. Reson. Chem. 48, 857–865 (2010). https://doi.org/10.1002/mrc.2678
S.E. Knopp, G. Bytof, D. Selmar, Influence of processing on the content of sugars in green Arabica coffee beans. Eur. Food Res. Technol. 223, 195–201 (2006). https://doi.org/10.1007/s00217-005-0172-1
M.B. dos SantosScholz, S.H. Prudencio, C.S.G. Kitzberger, R.S. dos Santos Ferreira da Silva, Physico–chemical characteristics and sensory attributes of coffee beans submitted to two post-harvest processes. J. Food Meas. Charact. 13, 831–839 (2019). https://doi.org/10.1007/s11694-018-9995-x
G.S. Duarte, A.A. Pereira, A. Farah, Chlorogenic acids and other relevant compounds in Brazilian coffees processed by semi-dry and wet post-harvesting methods. Food Chem. 118, 851–855 (2010). https://doi.org/10.1016/j.foodchem.2009.05.042
P. Bucheli, I. Meyer, A. Pittet, G. Vuataz, R. Viani, Industrial storage of green robusta coffee under tropical conditions and its impact on raw material quality and ochratoxin A content. J. Agric. Food Chem. 46, 4507–4511 (1998). https://doi.org/10.1021/jf980468
S.R. Evangelista, C.F. Silva, M.G.P.C. Miguel, C.S. Cordeiro, A.C.M. Pinheiro, W.F. Duarte, R.F. Schwan, Improvement of coffee beverage quality by using selected yeasts strains during the fermentation in dry process. Food Res. Int. 61, 183–195 (2014). https://doi.org/10.1016/j.foodres.2013.11.033
G.V.M. Pereira, E. Neto, V.T. Soccol, A.B.P. Medeiros, A.L. Woiciechowski, C.R. Soccol, Conducting starter culture-controlled fermentations of coffee beans during on-farm wet processing: growth, metabolic analyses and sensorial effects. Food Res. Int. 75, 348–356 (2015). https://doi.org/10.1016/j.foodres.2015.06.027
B.J. Shelp, A.W. Bown, M.D. MacLean, Metabolism and functions of gamma-aminobutyric acid. Trends Plant Sci. 4, 446–452 (1999). https://doi.org/10.1016/s1360-1385(99)01486-7
G. Bytof, S.E. Knopp, P. Schieberle, I. Teutsch, D. Selmar, Influence of processing on the generation of g-aminobutyric acid in green coffee beans. Eur. Food Res. Technol. 220, 240–245 (2005). https://doi.org/10.1007/s00217-004-1033-z
C.T. Cortes-Macía, C.F. Lopez, P. Gentile, J. Giron-Hernandez, A.F. Lopez, Impact of post-harvest treatments on physicochemical and sensory characteristics of coffee beans in Huila, Colombia. Postharvest Biol. Technol. 187, 111852 (2022). https://doi.org/10.1016/j.postharvbio.2022.111852
Acknowledgements
The present study was financially supported by the BOPTN Penelitian 2022 Program (Grant No. 187/E5/PG.02.00.PT/2022) granted by the Directorate of Research, Technology, and Community Service under the Ministry of Education, Culture, Research, and Technology of the Republic of Indonesia. We are grateful to Supriatnadinuri from Rahayu Farmer Group for supplying the green coffee bean samples. The researchers express their gratitude to Prof. Yana Maolana Syah and Dr. Elvira Hermawati from the Organic Chemistry Division and the Laboratory of the Integrated Chemistry, Faculty of Mathematics and Natural Sciences, Bandung Institute of Technology, for their valuable assistance in facilitating the NMR measurements.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Happyana, N., Diniresna, A., Pratiwi, A. et al. 1H NMR-based metabolic profiling of green beans of Coffea arabica var. sigararutang with different postharvest treatments. Food Measure 18, 2587–2597 (2024). https://doi.org/10.1007/s11694-023-02338-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11694-023-02338-0