Abstract
Purpose
Dynamic susceptibility contrast MRI (DSC) and dynamic contrast-enhanced MRI (DCE) are useful tools in the diagnosis and follow-up of brain gliomas; nevertheless, both techniques leave the open issue of data reproducibility. We evaluated the reproducibility of data obtained using two different commercial software for perfusion maps calculation and analysis, as one of the potential sources of variability can be the software itself.
Methods
DSC and DCE analyses from 20 patients with gliomas were tested for both the intrasoftware (as intraobserver and interobserver reproducibility) and the intersoftware reproducibility, as well as the impact of different postprocessing choices [vascular input function (VIF) selection and deconvolution algorithms] on the quantification of perfusion biomarkers plasma volume (Vp), volume transfer constant (K trans) and rCBV. Data reproducibility was evaluated with the intraclass correlation coefficient (ICC) and Bland–Altman analysis.
Results
For all the biomarkers, the intra- and interobserver reproducibility resulted in almost perfect agreement in each software, whereas for the intersoftware reproducibility the value ranged from 0.311 to 0.577, suggesting fair to moderate agreement; Bland–Altman analysis showed high dispersion of data, thus confirming these findings. Comparisons of different VIF estimation methods for DCE biomarkers resulted in ICC of 0.636 for K trans and 0.662 for Vp; comparison of two deconvolution algorithms in DSC resulted in an ICC of 0.999.
Conclusions
The use of single software ensures very good intraobserver and interobservers reproducibility. Caution should be taken when comparing data obtained using different software or different postprocessing within the same software, as reproducibility is not guaranteed anymore.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Dynamic susceptibility contrast MRI (DSC) and Dynamic contrast-enhanced MRI (DCE) can be useful tools in the diagnosis and follow-up of brain gliomas. Most studies published on this topic used perfusion-derived biomarkers for gliomas grading [1–5], patients’ outcome prediction [6–11] and differentiation of tumor recurrence from treatment-related effects like radionecrosis and pseudoprogression [12–16].
However, reproducibility is still an open issue for both techniques: no standardized protocol for acquisition and analysis of perfusion techniques is still available and, consequently, different research groups have used different protocols, generating a wide range of results [17–22]. Recently some steps towards standardization have been made, with the publication of an evidence- and consensus-based standards document for DCE [23] and a white paper for DSC [24].
One of the potential sources of variability is the software chosen for perfusion map calculation and analysis [25–32] since each software implements different analysis tools [i.e. leakage correction for DSC maps or automatic detection of vascular input function (VIF)] and sometimes different algorithms.
The aim of this study was to test the reproducibility of data obtained from studying the same cohort of patient with two commercial software, which were chosen based on the rationale that they could be used independently from the MR machine available for the acquisition, thus reducing data spread and improving standardization of perfusion analysis. We tested both intrasoftware (including the intraobserver and interobservers reproducibility) and intersoftware reproducibility and the impact of different postprocessing choices, such as VIF selection methods and deconvolution algorithms, on the quantification and reproducibility of perfusion biomarkers obtained from DSC and DCE analyses.
Methods
Patients
DCE and DSC data derived from 20 patients (mean age 51 years; range 16–76 years; 10 M/10 F) with histologically confirmed gliomas (8 WHO II, 4 WHO III and 8 WHO IV) were retrospectively evaluated. This study was approved by the ethical committee of our Institution.
MR imaging
MR imaging was performed on a 3T scanner (Achieva, Philips Healthcare, Best, The Netherlands) equipped with 80 mT/m gradients using a phased-array head 8 channel-coil. Online Resource 1 summarizes the imaging parameters of the MRI sequence protocol.
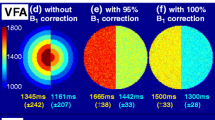
DCE–MRI was performed with a dynamic gradient-echo T1-weighted sequence using the following parameters: TR/TE 3.9/1.8 ms; flip angle 15°; matrix 96 × 84; FOV 230 × 201 mm; section thickness, 2.5 mm; in-plane acquisition voxel size, 2.4 × 2.4 mm. Seventy dynamic scans were performed with a temporal resolution of 5.1 s. The total acquisition time for DCE–MRI was 6 min and 10 s. DCE–MRI was preceded by a variable flip angle (VFA) axial sequence for T1 mapping.
DSC–MRI was performed with an axial gradient-echo T2*-weighted EPI sequence using the following parameters: TR/TE 1500/40 ms; flip angle 75°; matrix 96 × 77 mm; FOV 230 × 230 mm; section thickness, 5 mm; in-plane acquisition voxel size, 2.4 × 2.9. Eighty dynamic scans were performed with a temporal resolution of 1.5 s. The total acquisition time for DSC–MRI was 2 min and 4 s.
A cumulative fixed dose of 10 ml of gadobutrol (Gadovist, 1 mmol/ml; Bayer Schering Pharma, 6 Berlin, Germany) was administered, splitted in two boluses of 5 ml. The first bolus of 5 ml was injected 50 s after the start of the DCE sequence using a power injector (Spectris Solaris MR injector; MedRad, Indianola, Pennsylvania) at a rate of 2 ml/s, immediately followed by a 20 ml continuous saline flush at the same injection rate. The second bolus of 5 ml was injected 16 s after the start of the DSC sequence using the same power injector at a rate of 5 ml/s, followed by a 20 ml saline flush at the same injection rate. Thus, the contrast administration during DCE sequence pre-saturated the tissue for the following DSC–MR imaging.
Preprocessing
Perfusion MR analysis was performed using Olea Sphere (v. 2.3, Olea Medical Solutions, La Ciotat, France) and NordicIce (v. 2.3.12, NordicNeuroLab, Bergen, Norway). In both software, preprocessing steps included automatic motion correction by a rigid-body registration, automatic spatial smoothing and background segmentation.
DCE analysis
For DCE analysis, patient-specific baseline T1 maps were derived from VFA axial sequences (see Online Resource 1 for acquisition details).
In both software DCE analysis was based on the extended Tofts model [33]. In NordicIce, a venous VIF was obtained by manually drawing a ROI in the superior sagittal sinus (SSS). In Olea Sphere both a manual venous VIF and an automatic one were obtained; the venous one was obtained from the SSS, while the automatic one was obtained using a tool implemented in the software [34], which selected both arteries and veins as sources of input. Parametric maps of volume transfer constant (K trans) and plasma volume (Vp) were then obtained.
DSC analysis
In both software, DSC analysis was performed using the singular value decomposition (SVD) algorithm for deconvolution [35]. However, while Olea Sphere offers different methods for SVD, and both the standard truncated SVD (sSVD) and oscillation-index cSVD (oSVD) were used for analysis, in NordicIce only sSVD was available. An arterial VIF was obtained in Olea Sphere using an automatic method [34] while in NordicIce it was obtained semi-automatically: the observer selected the axial slice of interest and then the software searched for a valid vascular signal. Leakage correction was not applied during the postprocessing since it was already obtained with the contrast administration during the DCE acquisition, which served as a pre-bolus. Parametric maps of cerebral blood volume (CBV) were then obtained.
A fixed hematocrit value of 0.45 was set in both software.
Image analysis
All the parametric maps were independently analyzed by two observers (GMC, with 3 years experience in perfusion analysis, NA with 7 years experience in perfusion analysis).
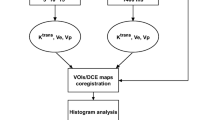
In both software, perfusion maps were automatically co-registrated with FLAIR and/or post-contrast T1 images by performing a rigid transformation of the datasets.
For each parametric map, multiple circular ROIs, 25–30 mm2 in area, were drawn in the tumor area that showed the highest value of perfusion biomarkers, then the one showing the highest mean value for each perfusion biomarker was selected. To normalize CBV values, a single elliptic ROI, 250–300 mm2 in area, was drawn in the contralateral normal appearing white matter (NAWM) in the centrum semiovale (nCBV). The ratio (rCBV) between the tumor-CBV and nCBV was then considered. The analysis was repeated after one month to assess intraobserver reproducibility.
From DCE analysis, for each parametric map and each observer three datasets were obtained: one with the automatic VIF selection (Olea automatic) and two with the manual VIF selection (Olea Manual and NordicIce); from DSC two datasets (Olea automatic and NordicIce) were obtained.
Moreover, to avoid any possible variability due to different ROIs positioning between observers, intersoftware reproducibility was re-tested drawing the ROIs in the same tumor area in the two software. To do so, co-registered anatomical series were used as reference to guarantee the same ROIs positioning in the two software. Mean value of selected ROIs was then used for the comparison.
To test the impact of different postprocessing options on the quantitative data, additional analyses were performed using Olea Sphere: for DCE, the reproducibility of data obtained using the automatic and the manual VIF selection and for DSC the data obtained using the sSVD and oSVD methods. For all these analyses, the ROIs positions were kept constant.
Statistical analysis
Intraclass Correlation Coefficient analysis (ICC) selecting a two-way random effect model was performed to assess intra- and intersoftware reproducibility using SPSS (SPSS 20.0 for MacOSX, IBM, Chicago, IL, USA) and interpreted as follows: 0–0.2 indicates poor agreement, 0.3–0.4 indicates fair agreement, 0.5–0.6 indicates moderate agreement, 0.7–0.8 indicates strong agreement and >0.8 indicates almost perfect agreement.
Additionally, to compare data obtained with the two software and with the different postprocessing options, Bland–Altman analysis was performed using Graph Pad Prism 6 (GraphPad Software, La Jolla, California, USA, http://www.graphpad.com).
Results
Intrasoftware reproducibility
Intraobserver reproducibility resulted in an ICC always superior than 0.8 except for two comparisons (Table 1). Similar results were obtained for interobservers agreement (Table 1).
Intersoftware reproducibility
Intersoftware reproducibility resulted fair to moderate for all perfusion biomarkers (Table 2): ICC ranged from 0.354 to 0.577; similar result was found drawing the ROIs in the same tumor area in the two software (Table 3). Selecting the same SVD algorithm in the two software only slightly improved reproducibility of rCBV (Table 3).
Graphs derived from Bland–Altman analysis are summarized in Fig. 1; when comparing Olea Automatic and NordicIce, bias for K trans, Vp and rCBV were 0.0035, −0.8030 and −4.604 while 95% limits of agreement range were (−0.16, 0.17), (−8.6, 6.99), (−17.29, 8.07), respectively; comparing Olea Manual and NordicIce, bias for K trans and Vp were 0.0210 and −0.4285 while 95% limits of agreement range were (−0.09, 0.13) and (−6.62, 5.76), respectively.
Impact of postprocessing options on reproducibility
Reproducibility of different VIF selection methods for DCE-derived biomarkers resulted in an ICC of 0.636 for K trans and 0.662 for Vp (Table 4). Regarding Bland–Altman analysis (Fig. 1), bias for K trans and Vp were −0.0175 and −0.3745 while 95% limits of agreement range were (−0.16, 0.12) and (−6.50, 5.75), respectively.
Comparison of the two SVD methods for DSC resulted in an ICC of 0.999 for rCBV in the same software (Table 4).
Discussion
One of the major limitations to the use of PWI in the diagnostic workflow of brain tumors is the lack of standardization of acquisition and analysis protocols [17–20]. This leads to highly heterogeneous findings among studies, thus decreasing reproducibility. Some of the known PWI analysis variables are operator-dependent, such as the VIF selection [17, 18, 25, 26, 28] and ROI positioning [20].
Another source of variability is the choice of the PWI analysis software itself [25–32]: different FDA and CE-approved software are available, such as the software platforms offered by all the major MR manufacturers, and some commercial software packages for advanced MR imaging analyses. Moreover, especially in neuroimaging, some groups analyze PWI datasets using non CE and FDA approved in-house software.
The issue of the reproducibility of data derived from different software seems to be independent from imaging technique and anatomical district: previous studies focused on data obtained using different software or different versions of the same software in both CT-PWI [27, 29, 30] and MR-PWI [25, 26, 28, 31, 32]. To our best knowledge, this study is the first reporting data on DCE reproducibility in brain imaging.
Intrasoftware reproducibility
Regarding the intrasoftware analysis, we found good reproducibility for all comparisons (Table 1). One of the two software, Olea Sphere, showed narrower and higher range of ICC values (Fig. 2). Better reproducibility was especially observed when the automatic selection of VIF was used, probably because this method avoids one of the possible observer-dependent variables. Nevertheless, our data shows that the manual selection of VIF also ensures a good intrasoftware reproducibility when selected in a fixed position (in our study, the superior sagittal sinus).
Slight differences were found in DCE and DSC reproducibility: Vp and K trans showed slightly better results compared to rCBV for both intraobserver and interobserver reproducibility (Table 1; Fig. 2); this may be due to the need of drawing two ROIs for rCBV estimation, one in the tumor area, and one in the NAWM, thus adding an additional variable compared to K trans and Vp.
Intersoftware reproducibility
Regarding the intersoftware analysis, the ICC values obtained suggest only fair to moderate reproducibility for both DSC and DCE parameters; moreover, the 95% confidence interval for ICC values showed a wide range of results for all comparisons (Table 2). This ICC value dispersion suggests that even if the two software were used with similar parameters, some differences do exist in how the two software generate the perfusion maps, and are probably due to options inaccessible to the users.
Similar results were also obtained drawing the ROIs in the same tumor area in the two software, with only a slight improvement of ICC values, showing that ROIs positioning might not be a major issue for PWI reproducibility if the analysis is performed by trained readers (Table 3).
The Bland–Altman analysis confirmed this data, since the limits of agreement and the dispersion of data were too wide to define the results reproducible (Fig. 1); the automatic VIF selection seems to slightly underestimate parameters if compared to the manual method; this may be due to the different composition of the input of the automatic method (see Sect. “VIF selection in DCE” paragraph).
Slight differences in DSC and DCE reproducibility were found when different methods of VIF selection were applied in the two software for DCE analysis, obtaining better results for rCBV (Tables 2, 3); these differences disappeared when the manual selection of VIF was applied in both software for DCE analysis (Tables 2, 3), suggesting that is important to use the same VIF selection method when performing perfusion analysis (see Sect. “VIF selection in DCE” paragraph).
To better clarify why these ICC values were obtained in intersoftware comparison, further analysis on the impact of different postprocessing options on data reproducibility were performed considering hypotheses and results of published studies. In particular, the impact of deconvolution algorithms on DSC and VIF selection on DCE data reproducibility were investigated.
Deconvolution algorithms in DSC
For DSC, one of the proposed factors that may affect reproducibility is the choice of the deconvolution algorithm [27, 30]. In our study DSC analyses were performed applying both different and same deconvolution algorithms in the two software, finding low intersoftware reproducibility, that only slightly increased from an ICC of 0.455–0.582 when selecting the same algorithm (sSVD) (Table 3). It is possible that the choice of a different algorithm per se does not introduce significant variability; this was proved comparing the data obtained using the two different SVD methods in the same software, resulting in an almost perfect agreement with an ICC of 0.999 (Table 4). Bland–Altman analysis confirmed this finding since the dispersion of data is very low (Fig. 1). This could mean that the two software apply the same algorithm in a different way or that the main source of variability may be due to other postprocessing options. Lastly, it must be noticed that we compared rCBV value, while some differences are expected in CBF when comparing oSVD and sSVD [36].
VIF selection in DCE
A possible source of variability for DCE analysis is the definition of VIF, as proposed by Heye [25] and Beuzit [28]. Our data confirms their hypothesis, since different quantitative values and thus different reproducibility of DCE data according to different VIF estimation methods were obtained. In the intersoftware comparison, a better reproducibility was obtained for both K trans and Vp when the VIF was selected manually in both software (Tables 2, 3). These differences may arise due to a different vessel selection in the two methods: the automatic selection of VIF selects both arteries and veins as a source of input, and this may determine a different peak characterization of the VIF curve (Figs. 3, 4); on the other hand, when performing the manual VIF selection, only venous signal is chosen, making the peaks of the curves probably more similar between the two software. Nevertheless, even considering this slight improvement, reproducibility still remains low. To precisely define the impact of different VIF on reproducibility, the two VIF estimation methods were compared in the same software (Figs. 3, 4). Our results show that this single change in postprocessing step has a significant impact on data as only moderate agreement was obtained, even if the ROIs were positioned in the same area and no other differences exist between the datasets compared (Table 4). Probably the VIF selection itself cannot entirely explain the poor reproducibility but it seems to play a major role in it.
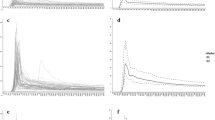
Comparison of automatic and manual VIF selection in the same patient; in this case automatic VIF selection on DCE maps shows both arterial (a) and venous (b) input; manual VIF selection on DCE maps consists in venous input only (d); c, e Concentration time curves show different peaks, with the one derived from automatic VIF being lower. The curve derived from the manual VIF is the mean of four different input located around the pixel selected (e)
Comparison of automatic and manual VIF selection in the same patient; in this case automatic VIF selection on DCE maps shows only venous input (a, b); manual VIF selection on DCE maps consists in venous input only (d); c, e Concentration time curves shows similar peaks. The curve derived from the manual VIF is the mean of four different input located around the pixel selected (e)
Our study confirms the recent findings by Orsingher [26], Milchenko [31] and Kelm [32], who also found low intersoftware reproducibility of DSC parameter CBV. Moreover, we confirmed its good interobserver reproducibility found by Kelm [32]. Very similar findings were found for DCE parameters. This has several consequences on the application of perfusion analysis.
The excellent intrasoftware reproducibility suggests that each software can be used in a reliable way even by multiple readers. It must be said, however, that a thorough training is necessary to obtain this result and the acquisition and analysis protocol must be followed very strictly.
The poor intersoftware reproducibility and the impact of postprocessing options on data reproducibility have consequences on the design of follow-up studies, on the application of data published in literature and on the daily clinical routine use of perfusion analysis. Follow-up studies should be designed and performed using the same software and the same postprocessing options (i.e. VIF selection), otherwise it could be difficult to define if any observed variation in pharmacokinetic parameters is a true biological phenomenon or it is simply related to the chosen analysis method. Moreover, it could be risky to apply published findings to other datasets, unless the same software, analysis protocol and acquisition protocol have been used. This is particularly relevant for those studies that define cut-off values for PWI biomarkers (i.e. in tumors grading). It must be said, however, that, as shown in the work by Kelm [32], different diagnostic accuracy is unlikely to be found among different FDA-cleared software.
Conclusions
In conclusion, we found a very good intrasoftware reproducibility and poor intersoftware reproducibility for DCE and DSC analyses, suggesting that while the use of a single software ensures reliable results, caution should be taken when comparing data obtained using different software and different postprocessing options, since the reproducibility in this case is not guaranteed anymore.
Abbreviations
- DSC:
-
Dynamic susceptibility contrast
- DCE:
-
Dynamic contrast-enhanced
- Vp:
-
Plasma volume
- K trans :
-
Volume transfer constant
- VIF:
-
Vascular input function
- SVD:
-
Singular value decomposition
- ICC:
-
Intraclass correlation coefficient
References
Nguyen TB, Cron GO, Perdrizet K et al (2015) Comparison of the diagnostic accuracy of DSC- and dynamic contrast-enhanced MRI in the preoperative grading of astrocytomas. Am J Neuroradiol 36:2017–2022. doi:10.3174/ajnr.a4398
Emblem K, Nedregaard B, Nome T et al (2008) Glioma grading by using histogram analysis of blood volume heterogeneity from MR-derived cerebral blood volume maps. Radiology 247:808–817. doi:10.1148/radiol.2473070571
Jung SC, Yeom JA, Kim JH et al (2014) Glioma: application of histogram analysis of pharmacokinetic parameters from T1-weighted dynamic contrast-enhanced MR imaging to tumor grading. Am J Neuroradiol 35:1103–1110. doi:10.3174/ajnr.A3825
Law M, Yang S, Babb J et al (2004) Comparison of cerebral blood volume and vascular permeability from dynamic susceptibility contrast-enhanced perfusion MR imaging with glioma grade. AJNR Am J Neuroradiol 25:746–755
Falk A, Fahlström M, Rostrup E et al (2014) Discrimination between glioma grades II and III in suspected low-grade gliomas using dynamic contrast-enhanced and dynamic susceptibility contrast perfusion MR imaging: a histogram analysis approach. Neuroradiology 56:1031–1038. doi:10.1007/s00234-014-1426-z
Law M, Young R, Babb J et al (2008) Gliomas: predicting time to progression or survival with cerebral blood volume measurements at dynamic susceptibility-weighted contrast-enhanced perfusion MR imaging. Radiology 247:490–498. doi:10.1148/radiol.2472070898
Schmainda KM, Prah M, Connelly J et al (2014) Dynamic-susceptibility contrast agent MRI measures of relative cerebral blood volume predict response to bevacizumab in recurrent high-grade glioma. Neuro Oncol 16:880–888. doi:10.1093/neuonc/not216
Piludu F, Marzi S, Pace A et al (2015) Early biomarkers from dynamic contrast-enhanced magnetic resonance imaging to predict the response to antiangiogenic therapy in high-grade gliomas. Neuroradiology 57:1269–1280. doi:10.1007/s00234-015-1582-9
Kim R, Choi S, Yun T et al (2016) Prognosis prediction of non-enhancing T2 high signal intensity lesions in glioblastoma patients after standard treatment: application of dynamic contrast-enhanced MR imaging. Eur Radiol. doi:10.1007/s00330-016-4464-6
Khalifa J, Tensaouti F, Chaltiel L et al (2016) Identification of a candidate biomarker from perfusion MRI to anticipate glioblastoma progression after chemoradiation. Eur Radiol. doi:10.1007/s00330-016-4234-5
Sanz-Requena R, Revert-Ventura A, Martí-Bonmatí L et al (2013) Quantitative MR perfusion parameters related to survival time in high-grade gliomas. Eur Radiol 23:3456–3465. doi:10.1007/s00330-013-2967-y
Yoon R, Kim H, Paik W et al (2016) Different diagnostic values of imaging parameters to predict pseudoprogression in glioblastoma subgroups stratified by MGMT promoter methylation. Eur Radiol. doi:10.1007/s00330-016-4346-y
Larsen V, Simonsen H, Law I et al (2012) Evaluation of dynamic contrast-enhanced T1-weighted perfusion MRI in the differentiation of tumor recurrence from radiation necrosis. Neuroradiology 55:361–369. doi:10.1007/s00234-012-1127-4
Wang S, Martinez-Lage M, Sakai Y et al (2015) Differentiating tumor progression from pseudoprogression in patients with glioblastomas using diffusion tensor imaging and dynamic susceptibility contrast MRI. Am J Neuroradiol 37:28–36. doi:10.3174/ajnr.A4474
Thomas AA, Arevalo-Perez J, Kaley T et al (2015) Dynamic contrast enhanced T1 MRI perfusion differentiates pseudoprogression from recurrent glioblastoma. J Neurooncol 125:183–190. doi:10.1007/s11060-015-1893-z
Prager AJ, Martinez N, Beal K et al (2015) Diffusion and perfusion MRI to differentiate treatment-related changes including pseudoprogression from recurrent tumors in high-grade gliomas with histopathologic evidence. Am J Neuroradiol 36:877–885. doi:10.3174/ajnr.A4218
Heye AK, Culling RD, Hernández MDCV et al (2014) Assessment of blood-brain barrier disruption using dynamic contrast-enhanced MRI. a systematic review. Neuroimage Clin 6:262–274. doi:10.1016/j.nicl.2014.09.002
Ellingson B, Bendszus M, Sorensen G, Pope W (2014) Emerging techniques and technologies in brain tumor imaging. Neuro Oncol 16:vii12–vii23. doi:10.1093/neuonc/nou221
Shiroishi M, Boxerman J, Pope W (2015) Physiologic MRI for assessment of response to therapy and prognosis in glioblastoma. Neuro Oncol. doi:10.1093/neuonc/nov179
Usinskiene J, Ulyte A, Bjørnerud A et al (2016) Optimal differentiation of high- and low-grade glioma and metastasis: a meta-analysis of perfusion, diffusion, and spectroscopy metrics. Neuroradiology. doi:10.1007/s00234-016-1642-9
Goh V, Schaeffter T, Leach M (2013) Reproducibility of dynamic contrast-enhanced MR imaging: why we should care. Radiology. doi:10.1148/radiol.12122447
Viallon M, Cuvinciuc V, Delattre B et al (2015) State-of-the-art MRI techniques in neuroradiology: principles, pitfalls, and clinical applications. Neuroradiology 57:441–467. doi:10.1007/s00234-015-1500-1
DCE-MRI Technical Committee (2012) DCE-MRI quantification profile, quantitative imaging biomarkers alliance. Version 1.0. Publicly Reviewed Version. QIBA. http://RSNA.org/QIBA
Welker K, Boxerman J, Kalnin A et al (2015) ASFNR recommendations for clinical performance of MR dynamic susceptibility contrast perfusion imaging of the brain. AJNR Am J Neuroradiol 36:E41–E51. doi:10.3174/ajnr.A4341
Heye T, Davenport M, Horvath J et al (2013) Reproducibility of dynamic contrast-enhanced MR imaging. Part I. Perfusion characteristics in the female pelvis by using multiple computer-aided diagnosis perfusion analysis solutions. Radiology 266:801–811. doi:10.1148/radiol.12120278
Orsingher L, Piccinini S, Crisi G (2014) Differences in dynamic susceptibility contrast MR perfusion maps generated by different methods implemented in commercial software. J Comput Assist Tomogr 38:647–654. doi:10.1097/RCT.0000000000000115
Kudo K, Sasaki M, Yamada K et al (2010) Differences in CT perfusion maps generated by different commercial software: quantitative analysis by using identical source data of acute stroke patients1. Radiology. doi:10.1148/radiol.254082000
Beuzit L, Eliat P-AA, Brun V et al (2015) Dynamic contrast-enhanced MRI: study of inter-software accuracy and reproducibility using simulated and clinical data. J Magn Reson Imaging. doi:10.1002/jmri.25101
Goh V, Shastry M, Engledow A et al (2010) Commercial software upgrades may significantly alter perfusion CT parameter values in colorectal cancer. Eur Radiol 21:744–749. doi:10.1007/s00330-010-1967-4
Kudo K, Christensen S, Sasaki M et al (2013) Accuracy and reliability assessment of CT and MR perfusion analysis software using a digital phantom. Radiology 267:201–211. doi:10.1148/radiol.12112618
Milchenko M, Rajderkar D, LaMontagne P et al (2014) Comparison of perfusion- and diffusion-weighted imaging parameters in brain tumor studies processed using different software platforms. Acad Radiol 21:1294–1303. doi:10.1016/j.acra.2014.05.016
Kelm ZS, Korfiatis PD, Lingineni RK, Daniels JR, Buckner JC, Lachance DH, Parney IF, Carter RE, Erickson BJ (2015) Variability and accuracy of different software packages for dynamic susceptibility contrast magnetic resonance imaging for distinguishing glioblastoma progression from pseudoprogression. J Med Imaging (Bellingham) 2(2):026001
Tofts PS, Brix G, Buckley DL et al (1999) Estimating kinetic parameters from dynamic contrast-enhanced T(1)-weighted MRI of a diffusable tracer: standardized quantities and symbols. J Magn Reson Imaging 10:223–232
Mouridsen K, Christensen S, Gyldensted L, Østergaard L (2006) Automatic selection of arterial input function using cluster analysis. Magn Reson Med 55:524–531. doi:10.1002/mrm.20759
Østergaard L, Sorensen A, Kwong K et al (1996) High resolution measurement of cerebral blood flow using intravascular tracer bolus passages. Part II: experimental comparison and preliminary results. Magnet Reson Med 36:726–736. doi:10.1002/mrm.1910360511
Wu O, Østergaard L, Weisskoff RM et al (2003) Tracer arrival timing-insensitive technique for estimating flow in MR perfusion-weighted imaging using singular value decomposition with a block-circulant deconvolution matrix. Magn Reson Med 50:164–174. doi:10.1002/mrm.10522
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Nicoletta Anzalone served as a consultant for Bayer HealthCare. Marcello Cadioli reports personal fees from Philips Spa and from San Raffaele Scientific Institute, outside the study. All the other authors declare that they have no potential conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Funding
No funding was received for this study.
Informed consent
For this type of study formal consent is not required.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Conte, G.M., Castellano, A., Altabella, L. et al. Reproducibility of dynamic contrast-enhanced MRI and dynamic susceptibility contrast MRI in the study of brain gliomas: a comparison of data obtained using different commercial software. Radiol med 122, 294–302 (2017). https://doi.org/10.1007/s11547-016-0720-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11547-016-0720-8