Abstract
Purpose
The pattern of eukaryotic macroorganisms varies with altitude is well-documented; by contrast, very little is known of how a bacterial pattern in soils varies with the elevation in a montane ecosystem. Mostly, previous studies on soil bacteria have either found a diversity decline, no trend, or a hump-back trend with increasing elevation. The aim of this study was to investigate the bacterial community composition and diversity patterns of Mount Nadu in Wolong Nature Reserve, Western Sichuan Plateau (3000–3945 masl).
Materials and methods
In total, 30 soil samples from the mountain at 10 sampling elevational zones (every 100 m from the baseband to the summit) were collected. High-throughput pyrosequencing approach was performed of soil bacterial 16S rRNA targeting V3 + V4 region by MiSeq PE300 and taxonomically classified based on Silva database. Bacterial community composition and diversity patterns were detected, and bacterial data were correlated with environmental factors to determine which factors influenced bacterial community composition.
Results and discussion
We obtained an average of 30,172 sequences per soil and found that the relative abundance of Acidobacteria and Proteobacteria count more than 70 % of the whole bacteria. Cooperative network analysis also revealed that Acidobacteria and Proteobacteria were important hubs in the community. Bacterial diversity pattern was found to be a significant hollow trend along altitudinal gradients and diversity of the dominant phyla (e.g., Acidobacteria, Proteobacteria) followed the results of the whole bacterial diversity. Moreover, distance-based linear model identified that soil pH and TN significantly provided 7.40 and 6.01 % of the total variation.
Conclusions
The hollow trend of bacterial diversity has rarely been observed in nature. It indicated that no unifying bacterial diversity pattern can be expected along elevational gradients among the mountain system, and our result suggested the importance of environmental factors in structuring bacterial communities in this montane ecosystem.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
The study of elevational gradients in montane ecosystem has long been a source of fascination for ecologists (Brown 2001; Korner 2007; Yang et al. 2014). Studies of distribution of species and communities have led to a hot research on vertical distribution patterns of macroorganisms (McCain 2005, 2009; Rahbek 2005). Despite a large number of researches on macroorganism, patterns of soil microorganisms in montane ecosystem remain largely unknown (Bryant et al. 2008; Singh et al. 2014). Microorganisms, especially bacteria, are abundant and diverse in mountain soil, which drives various important biogeochemical cycles in ecosystem (Swift et al. 1979; van der Heijden et al. 2008). Improved knowledge of vertical distribution patterns of bacteria is fundament to advancing basic ecology and comprehensive understanding and predicting the influences of climate change on montane ecosystem.
In the past few years, researches on diversity and composition of bacteria using molecular methods have yielded useful insights in bacterial distribution within montane ecosystem. A relative localized study in Mount Fuji suggested a significant hump-backed trend of total bacterial diversity (Singh et al. 2012) but a hollow trend of bacterial diversity was detected on Yeongsil (150–1700 masl) (Singh et al. 2014). And a pattern of decline in the bacterial diversity along the elevation was detected at all taxonomic levels on Mount Shegyla from 3106 to 4479 masl (Wang et al. 2015). Fierer et al. (2003) sampled different depths of soil (organic and mineral soil) across a montane elevational gradient, but did not find any bacterial diversity trend. Furthermore, a decline trend of Acidobacteria, mid-elevational bulge of Proteobacteria as well as Bacteroidetes with increasing elevation were reported (Singh et al. 2012). Bryant et al. (2008) focused only on Acidobacteria showed a monotonous decline towards higher elevations on the Rocky Mountains from 2460 to 3880 masl. Since the different methods of sampling and analysis, all studies above make it unclear how bacteria distributes with elevation, we could not get universal rules of microbes distribution. It is also suggesting that altitudinal distribution patterns of bacterial community should add many other studies of montane system.
Recent studies have shown that shifts in the structure of bacterial communities can be associated with changes in a number of soil physicochemistry including soil texture (Sessitsch et al. 2001; Kim et al. 2013), soil pH (Mannisto et al. 2007; Lauber et al. 2008; Xiong et al. 2012), soil moisture (Zhang et al. 2013), temperature (Lipson 2007), and soil nitrogen (Frey et al. 2004; Nilsson et al. 2007). From the baseband to the top of the mountain, environmental factors (including soil physicochemistry, plant and climate) change sharply (Korner 2007), indicating that bacterial distributions may exhibit differently owe to variations of environmental factors. It has been shown that the overall community composition along elevations, and in some cases bacterial richness and diversity of soil bacterial communities is often strongly correlated with soil pH (Shen et al. 2013). The finding from a pH controlled experiment indicated lower pH was stressful for most bacterial taxa such as Bacteriodetes, Nitrospira, Proteobacteria, etc. (Rousk et al. 2010). However, the strong correlation between soil pH and bacterial communities could be a result of soil pH integrating a number of other individual soil variables like nutrient, soil texture and soil moisture. It is not clear which of these factors has the major or minor influence on the biogeographical patterns exhibited by soil bacterial communities. To tease apart the explanation of these factors, our study focus on comparing their community structure along the elevation gradients and examining the effect of environmental factors on bacterial distribution.
Western Sichuan Plateau is the main department of Southeast of Tibetan Plateau, which lies between the Tibetan Plateau and the Sichuan Basin. The climate is affected by the southwest monsoons of the Indian Ocean and plateau monsoon (Zhang 2014). The special conditions make the Western Sichuan Plateau a weak landscape that is highly sensitive to the global climate changes (Chen et al. 2002), resulting in a great effect on biodiversity and ecosystem function (Araujo et al. 2005). Meanwhile, a subalpine coniferous forest of the Western Sichuan Plateau is the majority of southwest forestry area in China which is an important ecological barrier of the upper Yangtze River. Wolong National Nature Reserve is the largest nature reserve in Western Sichuan Plateau with many endangered creatures (e.g., Ailuropoda melanoleuca, Davidia involucrata etc.), and the large biodiversity of Wolong National Nature Reserve play an important role in balancing subalpine ecosystem. In addition, there is little disturbance (e.g., graze, fire, road etc.) in this primeval forest on Mount Nadu of Wolong Nature Reserve. All the favorable conditions provide Mount Nadu a natural laboratory to investigate bacterial montane distribution patterns. Soil bacteria is the majority of the soil microbes, and a better understanding of bacteria distribution pattern along the elevation could provide detail materials for biodiversity study and predict the impacts of global climate changes (Prosser et al. 2007).
Here, in an effort to comparing soil bacterial distribution in various elevational gradients in Mount Nadu, we aimed to determine whether soil bacteria show clear pattern in this subalpine montane ecosystem. The main objectives of this study were (1) to directly compare variability of bacterial distribution pattern across elevational gradients and (2) to identify environmental factors controlling the distribution of bacterial communities in our study.
2 Materials and methods
2.1 Site selection
Wolong Nature Reserve (102° 52′–103° 24′ E, 30° 45′–31° 25′ N) is located in Western Sichuan Plateau, Southwest of China. The climate is a subtropical inland mountain climate. Along the increasing altitudinal gradients from 1218 to 6250 masl, mean annual temperature is 8.5 °C (ranging from −8.5 to 29.2 °C), and mean annual precipitation is about 800 mm. Sampling was carried out along the Mount Nadu (102° 57′ E, 30° 50′ N) which lies in the southwest of Wolong Nature Reserve starting from the base of the mountain at 3000 masl, to the summit area at around 3950 masl.
Climatic variations result in a vertical zonation of major soil and plant types that is especially distinct. Along elevation increasing, soil varies in the order: brown coniferous forest soil and subalpine meadow soil. The parent rock of brown coniferous forest soil and subalpine meadow soil is weathered killas, while soil eluviation is stronger in brown coniferous forest soil than that in subalpine meadow soil (Zhang 1983). Mount Nadu has the primitive and completed subalpine coniferous forest. At the base of the study mountain forest is theropencedrymion which is dominated by Betula, Fargesia nitida and Abies faxoniana. As is usually the case on mountains, there is a marked decline in tree and shrub species richness with increasing elevation. Above 3300 masl is a coniferous forest dominated by Rhododendron and Abies faxoniana, and the herbaceous cover becomes sparser, while the tree gives way to shrubs and herbs above 3800 masl such as Rhododendron, Rosa, Lonicera, and Rhodiola. With elevation increasing, the species richness of shrub and tree decreased with elevation, while the herbs exhibited no significant pattern in richness.
2.2 Soil sampling, chemical characterization, and DNA extraction
We sampled along a transect on the north face of the mountain at 10 sampling elevational zones every 100 m from the baseband to the summit. At each elevational level, soil samples were collected from three independent replicate plots (10 × 10 m) at least 20 m distant from each other to minimize the effect of spatial autocorrelation. In each plot, 0–10-cm soil samples were collected by sampling in five locations and composited together. Visible roots and residues were removed prior. The fresh soil samples were sieved through a 2-mm sieve and divided into two subsamples. 500 g of soil was stored at 4 °C to determine the physicochemical properties, and 50 g of soil was stored at −80 °C prior to DNA extraction.
Volumetric soil water content (moisture) was determined by TF-SC (DeltaT Devices Cambridge, DT3-ML2x, UK). Soil temperature (T) was detected by iButton DS1922L (Maxim, San Jose, CA, USA). Soil texture was evaluated using a Mastersizer 2000 Laser Diffraction Particle Analyzer (Malvern Instruments Ltd, Malvern, UK) according to the universal criteria of soil particle size (sand > 0.02 mm, 0.02 mm > silt > 0.002 mm, clay < 0.002 mm). Soil pH was measured after shaking a soil water (1:5 mass/volume) suspension for 30 min. Soil total carbon (TC) and total nitrogen (TN) were determined using a Vario EL III analyzer (Elementar Analysensysteme GmbH, Hanau, Germany). Soil DNA was extracted from the 0.25 g wet soil using the MOBIO Powersoil DNA extraction kit (MOBIO Laboratories, Carlsbad, CA, USA) as directed by the manufacturer. Isolater DNA was stored at -80 °C before sequencing.
2.3 PCR amplification and barcoded pyrosequencing of bacterial communities
Primers 338F (5-CCTACGGGAGGCAGCAG-3) and 806R (5-GGACTACHVGGGTWTCTAAT-3) targeting V3 + V4 region were used to amplify an 16S rRNA gene fragment for the ABI GeneAmp® 9700. PCR using the TransGen Ap221-02: TransStart Fastpfu DNA Polymerase, 20 μl PCR reaction mixture. 4 μl 5 × FastPfu Buffer, 2 μl 2.5 mM dNTPs, 0.8 μl Forward Primer (5 μM), 0.8 μl Reverse Primer (5 μM), 0.4 μl FastPfu Polymerase, 10 ng Template DNA and add sterilized H2O to 20 μl. The resultant PCR amplicons were purified with a AxyPrepDNA Gel (AXYGEN, USA) and Tris-Hcl. Then, the PCR amplicons were detected by 2 % agarose gel electrophoresis method. The PCR products were detected on QuantiFluor™ -ST Blue fluorescence quantitative system (Promega, USA), the amplicons were mixed and sequenced on MiSeq PE300 (Illumina, USA).
2.4 Pyrosequencing data processing
The raw sequences obtained through MiSeq barcoded pyrosequencing were optimized using the Trimmomatic and FLASH (Magoc and Salzberg 2011; Lohse et al. 2012). Reads were quality screened using a moving window that was 50-bases long, then we discarded the downstream bases from the window which average quality score less than 20 and removed any trimmed reads were smaller than 50 base in length. After the trimming, reads in pair were merged into one sequence with a overlap length threshold of 10 base (mismatch ratio < 0.2). Qualified sequences were selected to be assigned to soil samples based on barcodes and adjusted the direction of the sequence before clustering (barcode: mismatch number = 0; primer: mismatch number < 2). Then dereplication and discard of singletons were performed by USEARCH (Edgar 2010), and the final sequences with no chimeras were then clustered into operational taxonomic units (OTUs) at 97 % similarity. Taxonomical classification of representative sequences to different classification levels used Quantitative Insights into Microbial Ecology (QIIME) pipeline (Caporaso et al. 2010). Ribosomal Database Project (RDP) Bayesian classifier (confidence threshold, 0.7) was chosen to assign the OTU at 97 % similarity level and count the community composition of each sample at different classification levels (Wang et al. 2007). The RDP classifier was trained using the Silva 16S rRNA database (Pruesse et al. 2007).
A network analysis was performed via an online tool, named the Molecular Ecological Network Analysis Pipeline (MENAP) (Zhou et al. 2011; Deng et al. 2012) to indentify key populations at OUT level in the subalpine coniferous forestry soil (Montoya et al. 2006). The first phase is network construction: Pearson correlation coefficient was used in pair-wise similarity matrix calculation and a threshold of 0.78 used for defining adjacency matrices by RMT-based approach. The second phase is network analyses: the topological role of each OUT was determined according to the within-module connectivity (Zi) and among-module connectivity (Pi) (Olesen et al. 2006). The network graph was visualized with Cytoscape 3.2.1 (Shannon et al. 2003).
A resampling procedure (single_rarefaction.py) was employed at a depth of 20,000 sequences per sample to eliminate the bias on diversity comparison as caused by unevenly sequencing (Gotelli and Colwell 2011). The subsampled OTU table was used for downstream beta and alpha diversity analyses. The unweighted Unifrac distance matrix was calculated to determine the amount of dissimilarity (beta diversity) between any pair of bacterial communities along altitude (Lozupone and Knight 2005). Shannon diversity, phylogenetic diversity (Faith’s PD (Faith 1992)), observed species, and Chao1 index were calculated to compare the alpha diversity. The filter_taxa_otu_table.py provided by QIIME was used to separate the OUT table of individual microbial taxa, according to which the diversity of specific taxa was analyzed.
2.5 Statistical analysis
Spearman’s correlation analysis was performed to examine the relationships among altitude and environmental factors in SPSS 22.0 (IBM Co, Armonk, NY, USA). Non-metric multidimensional scaling (NMDS) analysis based on unweighted Unifrac distance was performed to illustrate the beta diversity along elevation gradients using nmds.py script in QIIME. Mantel test between the unweighted Unifrac distance and altitude was performed in R 3.0 (permutation = 999) (R Core Team 2013). Regressions analysis against elevation, phylodiversity, richness and other parameters were drawn with SPSS 22.0 (IBM Co, Armonk, NY, USA). A distance-based multivariate linear model (DistLM) (Anderson 2001) was performed to evaluate the relative contribution of each environmental factor on bacterial distribution pattern. The contribution of each environmental variables was assessed using “marginal tests” to evaluate the statistical significance and percentage contribution of each variables taken alone, and then “sequential tests” to evaluate the cumulative effect of the environmental variables explaining microbial bacterial community similarity. Tests were performed using the computer program DISTLM_forward 3 (Anderson 2003).
3 Results
3.1 Environmental and soil condition
On Mount Nadu, variables were fitted with linear and quadratic regressions to reveal the relationships between elevation and environmental factors. With the increasing of altitude, the species richness of herbs showed no significant pattern, while trees (R 2 = 0.584, P < 0.01) and shrubs (R 2 = 0.154, P < 0.05) were found to decline linearly with altitude. Meanwhile, soil temperature (R 2 = 0.752, P < 0.01) and TN (R 2 = 0.443, P < 0.01) exhibited significantly hollow pattern along elevation. Soil pH ranging from 3.71 to 6.67, also behaved a significant hollow pattern (R 2 = 0.409, P < 0.01) with increasing altitude, while Clay (R 2 = 0.216, P < 0.05) was found to be a significant convex trend with altitude increasing. No significant relationship with altitude has been detected on TC, C/N, sand, silt, and soil moisture.
3.2 Bacterial community composition and diversity along elevation
Samples were classified by elevation; all the rarefaction curves were asymptotic lines, suggesting the sequencing effort is sufficient (Fig. 1). It interesting that the species accumulation in mid-altitudinal sites were slower than that in higher (3900 masl) or lower (3000 masl) sampling sites, indicating that the species distribution may vary with elevation.
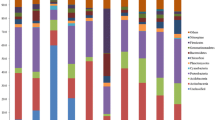
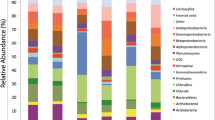
In total, we get 905,168 MiSeq 16S rRNA gene sequences after optimization from the 30 soil samples. On average, 30,172 sequences were found in each sample. When grouped at the 97 % similarity level, there were 4,108 phylotypes in the complete data set, with an average of 1,169 phylotypes per sample. Of the classifiable sequences, 33 phyla were identified across the sample set. Acidobacteria (41.58 %) and Proteobacteria (30.12 %) were the dominant phyla of the bacteria across the study site soils (Fig. 2). Phyla that were less abundances but still were found in most of the soils examined included the Actinobacteria (8.27 %), Chloroflexi (6.52 %), Verrucomicrobia (4.56 %), Bacteroidetes (1.97 %), Planctomycetes (1.73 %), WD272 (1.73 %) and Gemmatimonadetes (1.02 %). We identified sequences from an additional 24 rare phyla, which we define here as those phyla represented by <1 % of the sequences.
The interactions among different bacterial taxa in the forestry ecosystem were explored through construction of an OUT network. The final network included 214 nodes and 491 edges (Fig. 3). The overall topology indices (Table 1) revealed that network connectivity distribution was fitted well with the power-law model (R 2 = 0.872). Also, the average path length (GD) was 2.996, which were close to logarithms of the total number of network nodes. The topological roles of nodes were illustrated in ZP-plot (Fig. 3b). According the values of Zi (≥2.5) or Pi (≥0.62), a total of 9 nodes were module hubs and 2 nodes were connectors, and those 11 bacterial species were derived from Acidobacteria (OUT872, OUT4206, OUT3262, OUT2633, OUT2883, OUT4055, OUT3370, OUT3774, OUT879), Proteobacteria (OUT219) and Verrucomicrobia (OUT3995), which indicated that these species may represent potential keystone species.
The network of the forest soil bacteria. a The network graph structure by the fast greedy modularity optimization method. Each node signifies an OUT, which could correspond to a microbial population. Colors of the nodes indicate different major phyla. A blue edge indicates a positive interaction between two individual nodes, while a red edge indicates a negative interaction. b ZP-plot showing distribution of OTUs based on their module-based topological roles
Unweighted UniFrac distance on bacterial OTUs and a non-metric multidimensional scaling (NMDS) analysis was used to illustrate the dissimilarities in bacterial OTU community (beta diversity) between elevations (Fig. 4). Community from the middle altitudinal sites tend to cluster together, while higher and lower sites were more similar to each other. This finding was further collaborated by Mantel test through which a significant relationship was detected between altitude and the unweighted Unifrac distance of bacterial communities (Spearman r = 0.21, P < 0.01). Alpha diversity of bacterial community was found to be a hollow pattern along elevation gradients too (Fig. 5). Shannon diversity (R 2 = 0.420, P < 0.01), phylodiversity (R 2 = 0.526, P < 0.01), observed species (R 2 = 0.533, P < 0.01), and Chao1 index (R 2 = 0.531, P < 0.01) decreased from baseband to the mid-altitudinal sites, then increased up toward summit. Furthermore, the relative abundance and the alpha diversity (including Shannon diversity, phylodiversity, observed species, and Chao1 index) patterns of the dominant phyla along the altitude were characterized (Table 2). Alpha diversity of most phyla showed significant hollow pattern along elevation, such as Acidobacteria, Proteobacteria, Actinobacteria, Verrucomicrobia, etc.
3.3 Effect of environmental factors in structuring bacterial distribution patterns along elevation
(DistLM) was used to distinguish the effects of each environmental factors in shaping the bacterial distribution pattern, and Mantel tests (permutations = 999) showing the significance (Table 3). Of all of the environmental variable examined, soil pH, TN and soil moisture were mostly closely correlated with total bacterial community when considered individually (Table 3). The sequential model indicated three significant variables (pH, TN, TC, P < 0.05 in all cases) that explained 19.12 % of the total variation of the microbial community composition, with pH providing the greatest explanatory power (7.4 % of the total variation; Table 3).
4 Discussion
4.1 Composition of soil bacterial community
Although an average of 1169 phylotypes per sample were detected in our study, we still have not survey the full extent of bacterial diversity. A numbers of other studies had demonstrated that soil bacterial communities harbor an enormous number of unique taxa (Tringe et al. 2005). We could not document the all bacteria species in a given soil, because most bacterial taxa can be considered rare and high degree of endemism at finer levels of taxonomic resolution (Elshahed et al. 2008; Fulthorpe et al. 2008). In this study, we sampling from Mount Nadu, the key phyla are Acidobacteria and Proteobacteria from the result of relative abundance (Fig. 2) and ecological network analyses (Fig. 3). Our study suggested that Acidobacteria and Proteobacteria play important roles in Wolong Natural Reserve forestry soil, and it is in good agreement with previous study on bacterial relative abundance in montane ecosystems of other sites (Fierer et al. 2011; Singh et al. 2012). The network analyses indicated hubs derived from Acidobacteria, Proteobacteria and Verrucomicrobia are the key populations in subalpine coniferous forestry soil. These species were also detected in the broadleaved evergeen and the broadleaved deciduous forest soil of Shennongjia National Nature Reserve (Ding et al. 2015). However, broadleaved forestry soil contained more Actinobacteria hubs, known as copiotrophic bacteria preferentially living in abundant resources (Ramirez et al. 2012), suggesting that the key species of soil can be disparate in different forestry ecosystem.
4.2 Diversity of soil bacterial community
The whole bacterial community showed significant hollow patterns in diversity, in direct contrast to the decline patterns observed for tree and shrub richness. Our finding here parallels the studies of microbes that bacterial diversity do not follow the elevational diversity patterns of macroorganisms (Fierer et al. 2011; Shen et al. 2014). Hillebrand and Azovsky (2001) pointed out that small size strongly promotes high dispersal ability, and large population size and short generation times may result in a better chance of long-distance dispersal (Finlay 2002; McCain 2006). In addition, we observed that Acidobacteria and Proteobacteria respond similarly to the whole bacterial communities with a hollow pattern along elevations (Table 2). The result that the diversity patterns of dominant phyla were similar to that of the total community is not surprising because taxonomic and phylogenetic diversity are typically related (Morlon et al. 2010). The characteristic of Acidobacteria and Proteobacteria across the subalpine were similar to that of the whole bacteria coupled with their high abundance and connection in ecological network, suggesting that these phyla are likely driving the overall community. Similar result was found along the deglaciated chronosequence that Acidobacteria responded similarly to the whole bacteria in phylogenetic structure (Brown and Jumpponen 2015). Our study further illustrated that the restriction mechanism which can affect the diversity pattern may different between macroorganisms and microorganisms, and the rules of dominant phyla may play a decisive role in total bacterial community distribution.
Compared with previous studies which reported a monotonous decline, hump-backed, and no pattern in diversity on bacteria with increasing elevation (Bryant et al. 2008; Zhang et al. 2009; Singh et al. 2012; Wang et al. 2015), our results illustrated that the diversity pattern found in soil bacterial community is for a significant mid-elevation hollow. A hollow diversity pattern is rarely detected in nature (Rahbek 2005), and those various distribution patterns suggest that there may be some different factors could be responsible for diversity patterns. For example, a hump-backed pattern of bacterial diversity on Mt. Fuji may owe to disturbance gradients with its volcanic ash of surface soil from mid-elevation upwards (Singh et al. 2012); it is not surprising that our result is different because Mount Nadu is mainly vegetated and there is little disturbance. The comparable study of bacteria with elevation has been that on Laojun Mountain (1820 to 4050 masl), the range of altitude contained the actually elevational variation in our study (3000–3950 masl). That study found that richness and phylodiversity on biofilm bacterial communities was also a hollow pattern (Wang et al. 2012). But from 3000 to 3950 masl on Laojun Mountain, bacterial diversity increased with the altitude. It is worth mentioning that the sampling scheme here is very different: they sampled from surface of stones along a stream and with no replicates. This diversity pattern owes a lot to the declining temperature and frequent disturbances towards high elevation; although, a hollow bacterial pattern was detected along elevational gradients from 150 to 1700 masl on vegetated Yeongsil (Singh et al. 2014). It not clear what exactly had caused the elevational diversity trend, and the two possibilities maybe that some other undetected factors and disturbance. Those studies all found no exact factor related with diversity pattern and the natural disturbance rates in soil (e.g., the high day-night temperature variation, UV exposure) is no obvious quantifiable disturbance factor available for Mount Nadu. Nevertheless, Wang et al. (2015) found that the decline pattern of bacterial diversity on Mt. Shegyla was affected by soil pH and clay content definitely. Therefore, those quantitative evaluated factors (e.g., soil temperature, soil moisture, vegetation etc.) were used to explore the cause for the hollow pattern in our study.
Among the factors examined, our study found that the pattern of bacterial biodiversity along elevation is strongly associated with soil pH and TN. When the environmental parameters were considered individually, TN was found significantly correlated with bacterial community, which is agree with the result observed on Mt. Shegyla along elevations (Wang et al. 2015), suggesting nitrogen could affect the elevational distribution patterns of soil bacteria via determining the metabolism of soil bacteria (Yang et al. 2014). As the most important feature in determining bacterial distribution in our study, soil pH generally found to be the most important identifiable factor controlling soil bacterial diversity. The important role of pH has recently been found to influence bacterial diversity on horizontal soils (Griffiths et al. 2011; Philippot et al. 2013), and vertical soils across an elevational gradient (Xiong et al. 2012; Shen et al. 2014). The finding that diversity of bacteria was positively related to pH was consistent with that from a pH controlled experiment indicated lower pH was stressful for most taxa (Rousk et al. 2010). Most bacterial taxa exhibit relatively narrow pH range permissive, usually 3–4 pH units between minimum and maximum (Rosso et al. 1995). With the pH ranging from 3.71 to 6.67, some endurable taxa may become abundant and some new species may appear, resulting in the increase of bacterial diversity. In our study, the parent rock related with soil pH mainly derives from the collapsed killas, and humus accumulation drives the pedogenesis (Shi et al. 2000). The significant hollow pattern of soil pH with elevation in our study is related with the strong eluviation at mid-elevations. Under the Rhododendron-Abies faxoniana forest, water retention in top soil is strong since the well-developed living moss and ground litter, and it creates the favorable conditions for eluviation. It can be further demonstrated by the pattern of soil clay content. From the baseband to the summit, clay content also exhibited a maximum value in the Rhododendron-Abies faxoniana forest, indicating that mid-elevations hold stronger eluviation and clayization than that in lower or higher elevational sites. Taken together, this finding highlighted the role of soil environmental factors, and further suggested that pH is a universal predictor of bacterial diversity.
5 Conclusions
Overall, diversity of the soil bacterial community along the elevation gradients on Mount Nadu changed vertically with a clear hollow distribution pattern, and the dominant phylotypes corresponded to the distribution of the whole community along elevation. In addition, our results suggest that a range of environmental factors, including total nitrogen, soil moisture, and pH, operated together to explain variation in soil microbial community distribution along elevation gradients. Especially, the important role of pH in structuring bacterial diversity was also demonstrated. Comparing with other studies, given different patterns seen, suggesting that no simple rule of diversity pattern can be applied to mountain system. This finding also confirmed the important role of environmental factors in bacterial distribution on mountain system and would better help develop more insights into bacterial distribution pattern.
References
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Anderson MJ (2003) DISTLM forward: a FORTRAN computer program to calculate a distance-based multivariate analysis for a linear model using forward selection. University of Auckland, Auckland
Araujo MB, Whittaker RJ, Ladle RJ, Erhard M (2005) Reducing uncertainty in projections of extinction risk from climate change. Glob Ecol Biogeogr 14:529–538
Brown JH (2001) Mammals on mountainsides: elevational patterns of diversity. Glob Ecol Biogeogr 10:101–109
Brown SP, Jumpponen A (2015) Phylogenetic diversity analyses reveal disparity between fungal and bacterial communities during microbial primary succession. Soil Biol Biochem 89:52–60
Bryant JA, Lamanna C, Morlon H, Kerkhoff AJ, Enquist BJ, Green JL (2008) Microbes on mountainsides: contrasting elevational patterns of bacterial and plant diversity. Proc Natl Acad Sci U S A 105:11505–11511
Caporaso JG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Chen S, Fang X, Wang S (2002) Relation between the loess stratigraphy on the eastern Tibetan plateau and Indian monsoon. Mar Geol Quat Geol 22:41–46
Deng Y, Jiang YH, Yang YF, He ZL, Luo F, Zhou JZ (2012) Molecular ecological network analyses. BMC Bioinformatics 13:113
Ding JJ, Zhang YG, Wang MM, Sun X, Cong J, Deng Y, Lu H, Yuan T, Van Nostrand JD, Li DQ, Zhou JZ, Yang YF (2015) Soil organic matter quantity and quality shape microbial community compositions of subtropical broadleaved forests. Mol Ecol 24:5175–5185
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Elshahed MS, Youssef NH, Spain AM, Sheik C, Najar FZ, Sukharnikov LO, Roe BA, Davis JP, Schloss PD, Bailey VL, Krumholz LR (2008) Novelty and uniqueness patterns of rare members of the soil biosphere. Appl Environ Microbiol 74:5422–5428
Faith DP (1992) Conservation evaluation and phylogenetic diversity. Biol Conserv 61:1–10
Fierer N, Schimel JP, Holden PA (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35:167–176
Fierer N, McCain CM, Meir P, Zimmermann M, Rapp JM, Silman MR, Knight R (2011) Microbes do not follow the elevational diversity patterns of plants and animals. Ecology 92:797–804
Finlay BJ (2002) Global dispersal of free-living microbial eukaryote species. Science 296:1061–1063
Frey SD, Knorr M, Parrent JL, Simpson RT (2004) Chronic nitrogen enrichment affects the structure and function of the soil microbial community in temperate hardwood and pine forests. For Ecol Manag 196:159–171
Fulthorpe RR, Roesch LFW, Riva A, Triplett EW (2008) Distantly sampled soils carry few species in common. ISME J 2:901–910
Gotelli NJ, Colwell RK (2011) Estimating species richness. Frontiers in measuring biodiversity. Oxford University Press, New York, pp 39–54
Griffiths RI, Thomson BC, James P, Bell T, Bailey M, Whiteley AS (2011) The bacterial biogeography of British soils. Environ Microbiol 13:1642–1654
Hillebrand H, Azovsky AI (2001) Body size determines the strength of the latitudinal diversity gradient. Ecography 24:251–256
Kim M, Boldgiv B, Singh D, Chun J, Lkhagva A, Adams JM (2013) Structure of soil bacterial communities in relation to environmental variables in a semi-arid region of Mongolia. J Arid Environ 89:38–44
Korner C (2007) The use of “altitude” in ecological research. Trends Ecol Evol 22:569–574
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415
Lipson DA (2007) Relationships between temperature responses and bacterial community structure along seasonal and altitudinal gradients. FEMS Microbiol Ecol 59:418–427
Lohse M, Bolger AM, Nagel A, Fernie AR, Lunn JE, Stitt M, Usadel B (2012) RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res 40:W622–W627
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Mannisto MK, Tiirola M, Haggblom MM (2007) Bacterial communities in Arctic fields of Finnish Lapland are stable but highly pH-dependent. FEMS Microbiol Ecol 59:452–465
McCain CM (2005) Elevational gradients in diversity of small mammals. Ecology 86:366–372
McCain CM (2006) Do elevational range size, abundance, and body size patterns mirror those documented for geographic ranges? A case study using Costa Rican rodents. Evol Ecol Res 8:435–454
McCain CM (2009) Global analysis of bird elevational diversity. Glob Ecol Biogeogr 18:346–360
Montoya JM, Pimm SL, Sole RV (2006) Ecological networks and their fragility. Nature 442:259–264
Morlon H, Potts MD, Plotkin JB (2010) Inferring the dynamics of diversification: a coalescent approach. PLoS Biol 8, e1000493
Nilsson LO, Baath E, Falkengren-Grerup U, Wallander H (2007) Growth of ectomycorrhizal mycelia and composition of soil microbial communities in oak forest soils along a nitrogen deposition gradient. Oecologia 153:375–384
Olesen JM, Bascompte J, Dupont YL, Jordano P (2006) The smallest of all worlds: pollination networks. J Theor Biol 240:270–276
Philippot L, Spor A, Henault C, Bru D, Bizouard F, Jones CM, Sarr A, Maron PA (2013) Loss in microbial diversity affects nitrogen cycling in soil. ISME J 7:1609–1619
Prosser JI, Bohannan BJM, Curtis TP, Ellis RJ, Firestone MK, Freckleton RP, Green JL, Green LE, Killham K, Lennon JJ, Osborn AM, Solan M, van der Gast CJ, Young JPW (2007) Essay—the role of ecological theory in microbial ecology. Nat Rev Microbiol 5:384–392
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J, Glockner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196
R Core Team (2013) R: a language and environment for statistical computing. R Foundatin for Statistical Computing, Vienna
Rahbek C (2005) The role of spatial scale and the perception of large-scale species-richness patterns. Ecol Lett 8:224–239
Ramirez KS, Craine JM, Fierer N (2012) Consistent effects of nitrogen amendments on soil microbial communities and processes across biomes. Glob Chang Biol 18:1918–1927
Rosso L, Lobry JR, Bajard S, Flandrois JP (1995) Convenient model to describe the combined effects of temperature and pH on microbial-growth. Appl Environ Microbiol 61:610–616
Rousk J, Baath E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351
Sessitsch A, Weilharter A, Gerzabek MH, Kirchmann H, Kandeler E (2001) Microbial population structures in soil particle size fractions of a long-term fertilizer field experiment. Appl Environ Microbiol 67:4215–4224
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Shen CC, Xiong JB, Zhang HY, Feng YZ, Lin XG, Li XY, Liang WJ, Chu HY (2013) Soil pH drives the spatial distribution of bacterial communities along elevation on Changbai Mountain. Soil Biol Biochem 57:204–211
Shen CC, Liang WJ, Shi Y, Lin XG, Zhang HY, Wu X, Xie G, Chain P, Grogan P, Chu HY (2014) Contrasting elevational diversity patterns between eukaryotic soil microbes and plants. Ecology 95:3190–3202
Shi P-L, Li W-H, Wang J-X, Liu X-L (2000) Species-abundance relation of herb communities in subalpine timberline ecotone of Wolong Natural Reserve, Sichuan Province, China (in Chinese). Acta Ecol Sin 20:384–390
Singh D, Takahashi K, Kim M, Chun J, Adams JM (2012) A hump-backed trend in bacterial diversity with elevation on Mount Fuji, Japan. Microb Ecol 63:429–437
Singh D, Lee-Cruz L, Kim WS, Kerfahi D, Chun JH, Adams JM (2014) Strong elevational trends in soil bacterial community composition on Mt. Halla, South Korea. Soil Biol Biochem 68:140–149
Swift MJ, Heal OW, Anderson JM (1979) Decomposition in terrestrial ecosystems. Blackwell Scientific Publications, Oxford
Tringe SG, von Mering C, Kobayashi A, Salamov AA, Chen K, Chang HW, Podar M, Short JM, Mathur EJ, Detter JC, Bork P, Hugenholtz P, Rubin EM (2005) Comparative metagenomics of microbial communities. Science 308:554–557
van der Heijden MGA, Bardgett RD, van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang JJ, Soininen J, He JZ, Shen J (2012) Phylogenetic clustering increases with elevation for microbes. Environ Microbiol Rep 4:217–226
Wang JT, Cao P, Hu HW, Li J, Han LL, Zhang LM, Zheng YM, He JZ (2015) Altitudinal distribution patterns of soil bacterial and archaeal communities along Mt. Shegyla on the Tibetan Plateau. Microb Ecol 69:135–145
Xiong JB, Liu YQ, Lin XG, Zhang HY, Zeng J, Hou JZ, Yang YP, Yao TD, Knight R, Chu HY (2012) Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ Microbiol 14:2457–2466
Yang YF, Gao Y, Wang SP, Xu DP, Yu H, Wu LW, Lin QY, Hu YG, Li XZ, He ZL, Deng Y, Zhou JZ (2014) The microbial gene diversity along an elevation gradient of the Tibetan grassland. ISME J 8:430–440
Zhang WR (1983) The forest soils of Wolong Nature Reserve and its vertical zonalities distribution (in Chinese). Scientia Silvae Sci 19:254–268
Zhang H (2014) A study on the characteristics of climate change on northwestern Sichuan plateau (in Chinese). J Southwest Univ (Nat Sci Ed) 36:148–156
Zhang LM, Wang M, Prosser JI, Zheng YM, He JZ (2009) Altitude ammonia-oxidizing bacteria and archaea in soils of Mount Everest. FEMS Microbiol Ecol 70:208–217
Zhang XF, Zhao L, Xu SJ, Liu YZ, Liu HY, Cheng GD (2013) Soil moisture effect on bacterial and fungal community in Beilu River (Tibetan Plateau) permafrost soils with different vegetation types. J Appl Microbiol 114:1054–1065
Zhou J, Deng Y, Luo F, He Z, Yang Y (2011) Phylogenetic molecular ecological network of soil microbial communities in response to elevated CO2. mBio 2:e00122-11
Acknowledgments
This study was supported by the Ministry of Science and Technology of China (2011BAC09B08, 2012CB417103), the Strategic Priority Research Program of the Chinese Academy of Science (XDA05060100) and National Natural Science Foundation of China (41071039).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Xilong Wang
Rights and permissions
About this article
Cite this article
Liu, D., Wu, X., Shi, S. et al. A hollow bacterial diversity pattern with elevation in Wolong Nature Reserve, Western Sichuan Plateau. J Soils Sediments 16, 2365–2374 (2016). https://doi.org/10.1007/s11368-016-1422-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-016-1422-5