Abstract
The practice of utilizing animal manures on land is widespread in agriculture, but it has raised concerns about the possible spread of antibiotic resistance genes (ARGs) and the potential risk it poses to public health through food production. Fermentation bed culture is an effective circular agricultural practice commonly utilized in pig farming that minimizes the environmental impact of livestock farming. However, this method generates a significant amount of fermentation bed waste (FBW), which can be turned into organic fertilizer for land application. The objective of this research was to examine the impacts of amending agricultural soil samples with swine manure–derived FBW on microbial communities, mobile genetic elements (MGEs), and ARG profiles over different periods. The study findings indicated that the amendment of swine manure–derived FBW significantly increased the diversity and abundance of ARGs and MGEs during the early stages of amendment, but this effect diminished over time, and after 12 months of FBW amendments, the levels returned to those comparable to control samples. The shift in the bacterial communities played a significant role in shaping the patterns of ARGs. Actinobacteriota, Proteobacteria, and Bacteroidetes were identified as the primary potential hosts of ARGs through metagenomic binning analysis. Furthermore, the pH of soil samples was identified as the most important property in driving the composition of the bacterial community and soil resistome. These findings provided valuable insights into the temporal patterns and dissemination risks of ARGs in FBW-amended agriculture soil, which could contribute to the development of effective strategies to manage the dissemination risks of FBW-derived ARGs.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The prevalence of antimicrobial usage in intensive animal production systems is primarily for disease prevention, for treatment, and to meet the global demand for animal protein (Van Boeckel et al. 2015, 2019). Approximately 73% of global antimicrobial usage is attributed to animals, leading to the emergence of antibiotic resistance (Mulchandani et al. 2023; Van Boeckel et al. 2017). After administration, antibiotics are poorly absorbed by animals, with a significant proportion of 70–90% of antibiotics being excreted (Kumar et al. 2005). This excretion can lead to the emergence of antibiotic-resistant genes (ARGs) and antibiotic-resistant bacteria (ARB) in manure-affected environments, including those containing antibiotic residues and metabolites (Chen et al. 2019; Van den Meersche et al. 2020; Zhou et al. 2020). The application of manure to land, in particular, has facilitated the spread of ARGs from agricultural sources to surrounding ecosystems (Christou et al. 2017; Williams-Nguyen et al. 2016; Zhou et al. 2019).
Fermentation bed culture is a circular agriculture method commonly used in pig farming, referred to as breeding pig on litter, deep-litter-system, or in situ decomposition of pig manure (Morrison et al. 2007). Pigs are raised on a fermentation bed, and the manure is decomposed in situ. While this method reduces the environmental impact of livestock farming, it generates a large amount of fermentation bed waste (FBW), which comprises both manure and fermentation bed materials, such as agricultural straw, saw powder and rice husk (Pérez et al. 2022). FBW can be utilized as organic fertilizer in agriculture, enhancing microbial diversity and activity in soil, and improving soil nutrient availability and soil organic matter content for plant health and growth (Chen et al. 2017; Xu et al. 2022). However, the use of FBW can also contribute to the spread of ARB and ARGs. ARB can transfer genetic information via horizontal gene transfer mechanisms facilitated by mobile genetic elements (MGEs), including integrative conjugative elements, integrons, plasmids and insertion sequences (Ezugworie et al. 2021; Mbanga et al. 2021). Although numerous studies have examined the effects of manure application on ARG profiles and ARB, there is a scarcity of research on the impact of FBW, which is applied after the in situ decomposition of manure (Checcucci et al. 2020; Chen et al. 2016; He et al. 2020; Yu et al. 2023; Zou et al. 2020).
To address this research gap, we sampled three farmland plots with varying periods of FBW amendments near the swine feedlots equipped with fermentation beds. Our objective was to assess the influence of FBW on the microbial communities and ARG profiles in amended soil and identify the potential bacterial hosts of ARGs through metagenomic binning analysis.
Materials and methods
Sampling site
A total of 12 soil samples were collected in April 2021 from the main provinces in China for fermentation bed culture. Specifically, three soil samples were collected from Jilin (Jilin farmland), three from Shaanxi (Shaanxi farmland), three from Guizhou (Guizhou farmland), and three soil samples for control. The experimental stations were selected based on their continuous amendments of swine manure–derived FBW from nearby small-scale fattening pig production feedlots. These feedlots were equipped with fermentation beds managed in a similar manner, and the fermentation bed materials had been used for more than two years. The FBWs were collected from the surface layer of the fermentation bed, which comprised both swine manure and fermented bed materials, such as agricultural straw, saw powder and rice husk.
Soil amendments with FBW were applied to the farmland at a rate of approximately 15 tons per hectare once every two years. Soil samples were collected from specific locations with different amendment periods at each experimental station. Specifically, the soil samples from Jilin farmland were collected 6 months after the FBW amendment (designated JLF6); The farmland had not been cropped during this period. The soil samples from Guizhou farmland were collected 21 months after the FBW amendment (designated GZF21), and the farmland had been cropped with cabbage and green bean under rotation during this period. The soil samples from Shaanxi farmland were collected 12 months after the FBW amendment (designated SXF12), and the farmland had been cropped with apple trees during this period. The sampling site followed typical local practices, which usually involved the application of chemical fertilizer.
Control samples were collected from a pristine forest that had not been subjected to any amendment of FBW, swine manure, or anthropogenic antibiotics. These control samples enable a more comprehensive assessment of the potential risks associated with the application of FBW in agricultural settings, particularly the antibiotic resistome risk. Thus, we designated these soil samples as the control samples (CK).
Soil sampling and physicochemical properties analysis
For soil sampling, five surface soil cores (0-20 cm) were randomly collected from each site and combined into one plot sample. Three replicates were taken from each farmland. After sieving through a 2 mm mesh and thoroughly mixed, one portion was air-dried for analysis of soil physicochemical properties, while the other portion was stored at -80 ℃ for further DNA extraction.
Soil physicochemical properties, including electrical conductivity (EC), soil pH, total organic carbon (TOC), total nitrogen (TN) and C/N ratio were analyzed using the methods outlined by Cheng et al. (Cheng et al. 2019). Soil pH was measured in suspensions with a 1:5 soil/distilled water ratio (w/v) using a pH meter. Soil EC was measured in the water extracts with a 1:5 soil/distilled water ratio (w/v). TN content was measured by the Kjeldahl method, while soil TOC was measured through dichromate oxidation and titration with a ferrous sulfate solution.
DNA extraction
The genomic DNA of each soil sample was extracted by utilizing the PowerSoil DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA), following the manufacturer’s instructions, from 0.25 g of freeze-dried soil samples. The purity and quality of the extracted DNA were assessed on 1% agarose gels, and then, the DNA was stored at -20 ℃ for further experiments.
Library preparation, sequencing, and assembly
The extracted DNA was transported to Allwegene Company (Beijing, China) on dry-ice for library preparation and shotgun metagenomics sequencing using the Illumina Novaseq platform. Trimmomatic (v0.36) was used to remove adapter sequences and low-quality reads from the raw metagenomic reads (Chen et al. 2018). MEGAHIT (v1.1.2) was then used to assemble quality-controlled reads into contigs, with contigs over 500 bp as the final assembling result (Li et al. 2015). The raw metagenomic sequence data were publicly available in the National Center for Biotechnology Information (NCBI) BioProject database with the accession number of PRJNA953804.
Gene prediction, taxonomy, and ARG and MGE analysis
Open reading frames (ORFs) were identified by Prodigal (Hyatt et al. 2010). The predicted ORFs with a length of 100 bp or greater were selected and translated into amino acid sequences. CD-HIT was then used to construct the non-redundant gene with 95% sequence identity (Fu et al. 2012). Bowtie (Langmead and Salzberg 2012) was employed to align the sequences with the non-redundant gene set, and gene abundance information was counted. The high-quality sequences were analyzed using Diamond (v 0.8.35) to be blasted against the NCBI NR database for taxonomic annotations.
ARG-like sequences were annotated from the metagenomic sequencing reads using an online analysis pipeline, ARGs-OAP, with default settings (Yang et al. 2016). The plasmids, integrons, insertion sequences and integrative conjugative elements were identified based on NCBI RefSeq, INTEGRALL, ISfinder and ICEberg database by Diamond (v 0.8.35) with the following parameters: under a cutoff of E value of < 10−5, identify ≥ 90%, and over an alignment of at least 50 bp or 50 amino acids (Zhu et al. 2023). The abundance of MGEs was conducted using gene abundance information.
Metagenomic binning and function annotation
Metagenome-assembled genome (MAG) was recovered from assembly contigs of metagenomic data in soil samples by MetaBAT2 (v2.12.1) with default parameters (Kang et al. 2019). The completeness and contamination of MAGs were evaluated with CheckM (v1.1.3) (Parks et al. 2015). MAGs with completeness ≥ 50% and contamination ≤ 10% were selected for further refinement by secondary assembly. The relative abundance of each MAG was calculated by Salmon v0.9.1 (Patro et al. 2017), and the taxonomy of MAG was assigned by the GTDB-Tk (v1.3.0) with default options (Chaumeil et al. 2020). ARGs carried by MAGs were identified using Diamond (V0.8.35) against the SARG database (v2.2) with an e value cutoff of 10−5, 70% query coverage, and 70% similarity (Yin et al. 2018). MGEs carried by MAGs were identified by matching the results of NR annotation to keywords: plasmid, relaxase, resolvase, transposase, transposon, conjugal, recombinase, mobilization, conjugative, integrase, integron, recombination, invertase, and translocase (Li et al. 2017).
Results
Physicochemical properties of soil samples
The physicochemical properties of soil samples with varying periods of FBW amendments were assessed through one-way ANOVA and least significant difference (LSD) analysis. The results revealed that there were significant variations in the physicochemical properties among the different soil samples (P < 0.05, Table 1). The JLF6 soil samples showed the lowest pH value, while GZF21 soil samples displayed the highest levels of TN, TOC, and C/N ratio. As for CK soil samples, they showed the highest soil pH and EC but the lowest TN content.
Diversity of the microbial community
Metagenomic taxonomic annotation was performed to investigate the diversity of the microbial community. Bacteria were found to be the dominant domain across all soil samples, with a mean value of 98.44% of total sequences (Table S1). Principal component analysis (PCA) indicated significant separation of microbiota composition across soil samples with varying periods of FBW amendments (Fig. 1a). A total of 86 bacterial phyla were detected. The most common bacterial phyla present in all soil samples were Proteobacteria, Actinobacteria, Bacteroidetes, Acidobacteria, Planctomycetes, Chloroflexi, Gemmatimonadetes, Firmicutes, and Verrucomicrobia (Fig. 1b). Among these, Proteobacteria, Actinobacteria, and Bacteroidetes were the dominant phyla in all soil samples including CK samples. In JLF6, the relative abundances of Proteobacteria and Bacteroidetes were much higher than in other soil samples, accounting for 50.3% and 14.3% of the community, respectively. Conversely, the relative abundances of Planctomycetes, Acidobacteria, and Chloroflexi were lower in JLF6 than in other soil samples.
At the genus level, heatmap analysis displayed significant differences in the top 30 abundant genera among different soil samples (Fig. 1c). For instance, the relative abundances of Mycobacterium, Pseudomonas, Rhizobium, Devosia, Mesorhizobium, Rhodanobacter, Flavobacterium, and Pedobacter were more abundant in JLF6 compared to the GZF21, SXF12, and CK samples. In contrast, the relative abundances of Pyrinomonas, Streptomyces, Solirubrobacter, and Conexibacter were highest in CK samples, followed by SXF12 and GZF21, while JLF6 had the lowest abundance. An interesting observation was that the relative abundance of Arthrobacter, Sphingomonas, and Nocardioides in JLF6, SXF12, and GZF21 gradually decreased with the increasing amendment period of FBW (Fig. 1c).
Variations in the abundance and diversity of ARGs
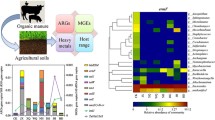
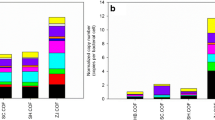
In terms of ARGs, a total of 129 unique ARGs were detected across all soil samples (Table S2). The FBW-amended soils (JLF6, SXF12, and GZF21) exhibited a significantly higher number of identified ARGs compared to the control samples (CK), showing a seven-fold increase (Fig. 2a). Kruskal–Wallis ANOVA confirmed a significant difference in the number of detected ARGs among soil samples with different durations of FBW amendments. The range of detected ARGs in FBW-amended soil samples was 12 to 87, with JLF6 having the highest number, accounting for 69% of the total number of detected ARGs. SXF12 had the second-highest number, which was significantly greater than that of CK (P < 0.05). However, GZF21 exhibited the lowest number of detected ARGs, similar to that of CK (Fig. 2a). An inverse linear relationship was observed between the number of detected ARGs in the soil samples and the duration of FBW amendments (Fig. 2a). The most frequently encountered ARGs were those conferring resistance to multidrug (21.11%), tetracycline (20.58%), and aminoglycoside (15.04%), followed by macrolide-lincosamide-streptogramin (MLS) (13.98%) and trimethoprim (6.07%) (Fig. 2b).
The number of detected ARG types in different soil samples (Kruskal–Wallis ANOVA test: *p < 0.05, **p < 0.01) (a). The percentage in the number of detected ARG types in all soil samples (b). The relative abundance (normalized to 16S rRNA) of ARG types in different soil samples (c). The heatmap analysis of top 20 ARG subtypes in different soil samples (d)
The relative abundance of ARGs (normalized to 16S rRNA) followed a similar trend as the number of detected ARGs, decreasing as the duration of FBW amendments increased. The highest abundance of ARG was observed in JLF6, 6 months after FBW amendments without any cropped, which was 20-fold higher than that in CK (Fig. 2c). However, over the course of 12 months of FBM amendments (SXF12 and GZF21), the relative abundance of ARGs decreased and returned to a level comparable to that of CK, with no significant difference observed (P > 0.05) (Fig. S1). Additionally, the principal coordinates analysis (PCoA) based on the Barry Curtis distances of ARGs also indicated that SXF12, GZF21, and CK exhibited greater similarity to each other than to JLF6 (Fig. S2). In JLF6, the most prevalent types of ARGs were those conferring resistance to sulfonamide (37.8%, 0.04 copies per 16S rRNA gene), aminoglycoside (16.9%, 0.018 copies per 16S rRNA gene), tetracycline (16.32%, 0.017 copies per 16S rRNA gene), and chloramphenicol (13.3%, 0.014 copies per 16S rRNA gene). In contrast, these ARGs were present at very low abundance (below 0.00025 copies per 16S rRNA gene) in SXF12, GZF21, and CK. Instead, the most abundant types of ARG in SXF12, GZF21, and CK were vancomycin and multidrug (Fig. 2c and Fig. S3). Furthermore, the heatmap analysis of top 20 ARGs revealed significant enrichment of sulfonamide resistance genes (sul1 and sul2), aminoglycoside-resistant genes (aadA, aac(6')-I, aac(6')-II, and aadE), and tetracycline resistance genes (tetG, tetX2, tetA, and tetZ) in JLF6, which were decreased significantly in SXF12, GZF21, and CK. Conversely, the vancomycin-resistant gene (vanR) showed a high level of enrichment in SXF12, GZF21, and CK (Fig. 2d).
The presence, abundance, and diversity of MGEs
We aimed to investigate the abundance, diversity, and presence of mobile genetic elements (MGEs) among all soil samples, including integrative conjugative elements (ICEs), insertion sequences, plasmids, and integrons. MGEs played a pivotal role in facilitating the horizontal transfer of ARGs among microorganisms. Similar to the patterns of ARG distribution in different soil samples, we found that MGEs including plasmids, ICEs, insertion sequences, and integrons were most abundant in JLF6 compared to SXF12, GZF21, and CK (Fig. 3). The number of gene counts of JLF6 matched to plasmids was 3780, while SXF12, GZF21, and CK exhibited lower gene counts with 1883, 2753, and 2541 gene counts, respectively (Table S3). Integrons were also analyzed by aligning them against the INTEGRALL database, and JLF6 had the highest number of gene counts, with intI1 being the most abundant. In terms of insertion sequences, JLF6 had a higher count of 105 compared to CK, SXF12, and GZF21 (Table S3). The dominant insertion sequences in JLF6 were IS1247, IS1245, and ISMlu5, while ISBj17, ISSsp2, and ISBj5 were dominant insertion sequences in CK, SXF12, and GZF21, respectively (Fig. S4). Furthermore, we evaluated the presence of ICEs in soil samples with different durations of FBW amendments. JLF6 had a total of 212 gene counts matched to ICEs, with the dominant ICEs being ICE6440, ICEPaePA14-1, and ICEEa1. In contrast, gene counts matched to ICEs in SXF12, GZF21, and CK were below 23, and the dominant ICEs were ICEMloMAF-1, ICEMaSym (WSM2073)-alpha, and ICEMcSym (1284)-alpha, respectively (Fig. S5). These results indicated that JLF6 exhibited greater MGE abundance and more diverse MGE profiles than SXF12, GZF21, and CK.
Relationships among soil physicochemical properties, ARG profiles, and bacterial communities
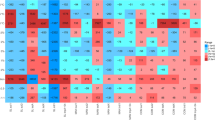
We further assessed the impact of soil physicochemical properties including EC, pH, TOC, TN, and C/N ratio on the composition of bacterial communities and ARGs through redundancy analysis (RDA). The result indicated that RDA1 and RDA2 elucidated 95.84% and 98.2% of the total variation in bacterial communities and ARG profiles, respectively (Fig. 4a and b). The RDA vectors indicated that pH played a more significant role in driving the composition of bacterial communities and ARGs, which aligned with the results of the Mantel test (Table S4). Additionally, Spearman’s heatmap analysis showed that the relative abundance of Actinobacteria was positively correlated with pH and EC but negatively correlated with TOC, TN, and C/N ratio. In contrast, Proteobacteria and Bacteroidetes exhibited a negative correlation with pH, but a positive correlation with TN, TOC, and C/N ratio (Fig. 4c). Moreover, the relative abundance of vancomycin-resistant genes was positively correlated with pH, but negatively correlated with TOC and TN (Fig. 4d). These results suggested that soil physicochemical properties influence the distribution of soil bacterial communities and ARGs.
Redundancy analysis (RDA) of quantitative correlation between bacterial communities (a) or ARG types (b) and soil chemical properties in different soil samples. Spearman heatmap of correlation between bacterial communities (c) or ARG types (d) and soil chemical properties (*p < 0.05, **p < 0.01, and ***p < 0.001)
The recovered metagenome-assembled genomes (MAGs) binning and their distribution to ARGs and MGEs
Based on metagenomic binning analysis, a total of 281 MAGs were assembled from soil samples. After quality control, 55 high-quality MAGs were retained for downstream analysis, comprising 10 near-complete MAGs (≥ 90% completeness and ≤ 5% contamination), 20 medium-quality MAGs (≥ 70% completeness and ≤ 10% contamination), and 25 partial MAGs (completeness ≥ 50% and contamination ≤ 10%) (Fig. 5a). Among these 55 recovered MAGs, 12 MAGs were found to carry ARGs, with their relative abundance ranging from 1.0 to 35.2 cpm (copies per million) in each soil sample. Notably, the relative abundance of ARG-carrying MAGs in JLF6 was more than tenfold higher than that in CK, SXF12, and GZF21 (Fig. 5b and Table S5).
Quality assessment of the recovered MAGs carrying ARGs (a). Genome quality was defined as completeness − 5 × contamination, and only genomes with quality of ≥ 50% were retained. Near-complete genomes (completeness ≥ 90%; contamination ≤ 5%) are shown in red, medium-quality genomes (completeness ≥ 70%; contamination ≤ 10%) in blue, and partial genomes (completeness ≥ 50%; contamination ≤ 10%) in black. The relative abundance of ARG-carrying MAGs in different soil samples (Kruskal–Wallis test: p = 0.02) (b)
As shown in Fig. 6, these ARG-carrying MAGs predominantly harbored eight types of ARGs, with tetracycline, multidrug, and vancomycin being the most frequently detected types. Most ARG-carrying MAGs belonged to Actinobacteriota, Proteobacteria, and Bacteroidota, followed by Chloroflexota and Verrucomicrobiota. Many of these bacterial hosts exhibited potential for multi-antibiotic resistance. For example, one MAG (bins.30) belonging to genus Mycobacterium of the phylum Actinobacteriota could confer multiple resistance to tetracycline (tetX2 and tetV), rifamycin (ADP-ribosylating transferase_arr) and quinolone (mfpA). Additionally, the genus Herbaspirillum affiliated to the phylum Proteobacteria and family CSP1-4 were also found to harbor two or more ARG subtypes (Fig. 6 and Table S6). These bacterial hosts could potentially play a significant role in the dissemination and acquisition of ARGs in the environment. According to mobilome profile analysis, Acidobacteriota harbored the highest number of MGEs (accounting for 49.86%), followed by Patescibacteria (accounting for 23.04%), Bacteroidota (accounting for 22.49%), and Gemmatimonadota (accounting for 2.98%) (Fig. S6). These findings suggested that these bacterial hosts have a greater potential to facilitate the transfer of ARGs in the environment.
Discussion
Impact of swine manure–derived fermentation bed waste amendment on ARGs
The application of swine manure–derived fermentation bed waste (FBW) as an amendment in agricultural fields was found to have a significant impact on the diversity and detection frequency of ARGs. The number of detected ARGs in FBW-amended soil samples was significantly higher compared to control samples. However, as the duration of FBW amendments increased, the number of detected ARGs decreased, suggesting a reduction in the potential ARG risk associated with FBM amendments over time. The relative abundance of ARGs followed a similar trend, with an increase in their abundance following FBW amendments. Even after 6 months of FBW amendments without cropping, the relative abundance of ARGs remained higher in JLF6. The higher detected ARGs in JLF6 may be attributed to FBW from swine feedlot, with sulfonamide, aminoglycoside, and tetracycline resistance genes found to be the dominant types of ARGs (Fig. 2c). These three types of ARGs were universally distributed in the conventional concrete-floor swine feedlot and in agriculture soils that received swine manure (Ben et al. 2017; Wang et al. 2016). To further support this, initial FBW samples were collected from the same swine feedlots in Jilin province as in our previous study (referred to as JLW, data not yet published). The dominant types of ARGs in JLW were sulfonamide (32.3%)-, aminoglycoside (23.2%)-, and tetracycline (19.9%)-resistant genes, accounting for 75.4% of total abundance of ARGs (Fig. S7a). The ARG profiles in JLW were similar to those in JLF6, and the relative abundance of ARG increased significantly during the first 6 months after FBW amendments in agricultural fields (Fig. S7b). These results suggested that the amendment of FBW in agricultural soils could pose a risk of transferring ARGs from swine feedlots to the surrounding environment within the first 6 months. However, after more than 12 months of FBW amendments, the relative abundance of these three types of ARGs decreased significantly and the profiles of ARGs returned to a pristine level similar to that of control samples, indicating a decrease in the soil resistome risk over time. This finding was supported by the results of clustering analysis on heatmap and PCoA (Fig. 2d and Fig. S2). The most abundant types of ARGs were those conferring resistance to vancomycin and multidrug, after more than 12 months of FBW amendments (SXF12 and GZF21), which were similar to the control samples. These types of ARG were ancient and commonly found in soils worldwide. Extensive research had been conducted on these ARGs, supporting their widespread presence (D’Costa et al. 2011; Zheng et al. 2022). Specifically, vanR was consistently the most frequently detected ARG subtype in SXF12, GZF21, and CK. It was important to note that vanR was highly prevalent in “non-human-associated” environments such as soil and was not human-associated and the least likely to endanger human health, which was classified as low risk to humans (Rank IV) (Zhang et al. 2021).
The underlying mechanisms in the dissemination of ARGs
The addition of manure in agricultural soil could cause significant changes to the soil microbial community, potentially impacting the prevalence of ARGs (Guan et al. 2022; Li et al. 2022). In this research, we investigated the impact of swine manure–derived FBW amendment on the soil resistome. A mantel test revealed a strong correlation between bacterial communities and ARG profiles (Fig. S8). The significant differences in bacterial community structure were observed between early stage of FBW-amended soil samples (JLF6) and control samples (CK) or long-term FBW-amended soil samples (SXF12 and GZF21) (Fig. 1). These findings suggested that the shift in bacterial community might lead to the changes in soil resistome, which aligned with the impact of manure amendment on soil resistome (Gu et al. 2021). However, reports on the bacterial communities and ARG profiles of the early stage and long-term FBW-amended soil were scarce.
In this study, we found that early stage of FBW-amended soil samples (JLF6) had a higher proportion of Proteobacteria, Actinobacteriota, and Bacteroidetes, which were known to be the most abundant phyla in manures and could possibly carry ARGs (Han et al. 2018) (Fig. 1a). As these bacteria were derived from swine manure in FBW, they may potentially contribute to the dissemination of ARGs into agricultural soil during the early stage of the amendment. Additionally, metagenome-assembled genome binning revealed that Actinobacteriota, Proteobacteria, and Bacteroidota were the dominant bacterial hosts of ARGs in this study (Fig. 6). This finding was consistent with previous research which reported that Actinobacteriota, Proteobacteria, and Bacteroidetes were the most common hosts of ARGs in wastewater (Zhu et al. 2023). We also found that many bacterial hosts of ARGs showed possible multi-antibiotic resistance characteristics, indicating higher risks for the spread of ARGs. For instance, Mycobacterium elephantis was detected as the carrier of multiple antibiotic resistance, including tetX2, tetV, and ADP-ribosylating transferase_arr and mfpA, which had previously been identified as a most important multi-drug resistant bacteria in wastewater treatment plants (Zhao et al. 2020). Mycobacterium were commonly found in soil and played a significant role in the spread and transfer of ARGs in soil. Some Mycobacterium species, including pathogenic species such as M. tuberculosis, would pose the risk of ARG from environment to human health (Peterson and Kaur 2018). Additionally, genus Herbaspirillum and family CSP1-4 harbored two or more ARGs, which were also enriched in JLF6 (Fig. S9).
In addition to changes in the bacteria community, MGEs were also prominently correlated with soil resistome, potentially facilitating the dissemination of ARGs (Zheng et al. 2022). Swine manure–derived FBW-borne bacteria can transfer into indigenous bacteria in agriculture soil via MGEs, providing an alternative mechanism for the soil resistome (Du et al. 2020). The largest number and abundance of MGEs were detected during the early stages of FBW amendment (JLF6) (Table S3, Fig. 3), indicating that MGEs originating from swine manure–derived FBW bacteria could establish themselves in soils and promote soil resistome more extensively (Heuer et al. 2011). The MAGs binning results further revealed that Acidobacteriota, Patescibacteria, and Bacteroidota were the main hosts of MGEs in soil samples, suggesting that these bacterial hosts possess better abilities to facilitate the transfer of ARGs in the environment. However, after more than 12 months of FBW amendments, the relative abundance of ARGs and MGEs decreased substantially, and profiles of bacteria community and ARGs were brought back to a level similar to those of CK. These results suggested that the risk of soil resistome decreases over time. The structure of the bacteria community may have a significant impact on the dissemination and acquisition of antibiotic resistance in the environment.
Conclusions
As a result of this study, it has been demonstrated that the amendment of soil samples with swine manure–derived FBW can initially increase the abundance and diversity of ARGs and MGEs, particularly within the first 6 months of amendment. However, the abundance of ARGs and MGEs declined over time, and after 12 months of FBW amendments, their levels returned to similar to control samples. The structure of the bacterial community played a critical role in shaping the ARG patterns, with Actinobacteriota, Proteobacteria, and Bacteroidota identified as the main potential hosts of ARGs. Furthermore, Acidobacteriota, Bacteroidota, and Patescibacteria were found to harbor more MGEs, potentially facilitating the transfer of ARGs. These findings provided insights into the temporal patterns and dissemination risk of ARGs in swine manure–derived FBW-amended agricultural soil. Further research about the dissemination risk of ARGs in swine manure–derived FBW-amended agricultural soil with different amendment periods could be conducted in one experimental station with or without cropping, in order to better verify the risk of ARG dissemination from swine manure-derived FBW.
Data availability
The raw metagenomic sequence data were publicly available in the NCBI BioProject database with the accession number of PRJNA953804.
References
Ben W, Wang J, Pan X, Qiang Z (2017) Dissemination of antibiotic resistance genes and their potential removal by on-farm treatment processes in nine swine feedlots in shandong province, china. Chemosphere 167:262–268. https://doi.org/10.1016/j.chemosphere.2016.10.013
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2020) Gtdb-tk: A toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36:1925–1927. https://doi.org/10.1093/bioinformatics/btz848
Checcucci A, Trevisi P, Luise D, Modesto M, Blasioli S, Braschi I, Mattarelli P (2020) Exploring the animal waste resistome: The spread of antimicrobial resistance genes through the use of livestock manure. Front Microbiol 11:1416. https://doi.org/10.3389/fmicb.2020.01416
Chen C, Pankow CA, Oh M, Heath LS, Zhang L, Du P, Xia K, Pruden A (2019) Effect of antibiotic use and composting on antibiotic resistance gene abundance and resistome risks of soils receiving manure-derived amendments. Environ Int 128:233–243. https://doi.org/10.1016/j.envint.2019.04.043
Chen Q, An X, Li H, Su J, Ma Y, Zhu Y-G (2016) Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environ Int 92:1–10. https://doi.org/10.1016/j.envint.2016.03.026
Chen Q, Liu B, Wang J, Che J, Liu G, Guan X (2017) Diversity and dynamics of the bacterial community involved in pig manure biodegradation in a microbial fermentation bed system. Annals Microbiol 67:491–500. https://doi.org/10.1007/s13213-017-1278-y
Chen SF, Zhou YQ, Chen YR, Gu J (2018) Fastp: An ultra-fast all-in-one fastq preprocessor. Bioinformatics 34:884–890. https://doi.org/10.1093/bioinformatics/bty560
Cheng JH, Tang XY, Cui JF (2019) Effect of long-term manure slurry application on the occurrence of antibiotic resistance genes in arable purple soil (entisol). Sci Total Environ 647:853–861. https://doi.org/10.1016/j.scitotenv.2018.08.028
Christou A, Agüera A, Bayona JM, Cytryn E, Fotopoulos V, Lambropoulou D, Manaia CM, Michael C, Revitt M, Schröder P (2017) The potential implications of reclaimed wastewater reuse for irrigation on the agricultural environment: The knowns and unknowns of the fate of antibiotics and antibiotic resistant bacteria and resistance genes–a review. Water Res 123:448–467. https://doi.org/10.1016/j.watres.2017.07.004
D’Costa VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477:457–461. https://doi.org/10.1038/nature10388
Du S, Shen J-P, Hu H-W, Wang J-T, Han L-L, Sheng R, Wei W-X, Fang Y-T, Zhu Y-G, Zhang L-M (2020) Large-scale patterns of soil antibiotic resistome in chinese croplands. Sci Total Environ 712:136418. https://doi.org/10.1016/j.scitotenv.2019.136418
Ezugworie FN, Igbokwe VC, Onwosi CO (2021) Proliferation of antibiotic-resistant microorganisms and associated genes during composting: An overview of the potential impacts on public health, management and future. Sci Total Environ 784:147191. https://doi.org/10.1016/j.scitotenv.2021.147191
Fu LM, Niu BF, Zhu ZW, Wu ST, Li WZ (2012) Cd-hit: Accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Gu Q, Sun M, Lin T, Zhang Y, Wei X, Wu S, Zhang S, Pang R, Wang J, Ding Y, Liu Z, Chen L, Chen W, Lin X, Zhang J, Chen M, Xue L, Wu Q (2021) Characteristics of antibiotic resistance genes and antibiotic-resistant bacteria in full-scale drinking water treatment system using metagenomics and culturing. Front Microbiol 12:798442. https://doi.org/10.3389/fmicb.2021.798442
Guan Y, Xue X, Jia J, Li X, Xing H, Wang Z (2022) Metagenomic assembly and binning analyses the prevalence and spread of antibiotic resistome in water and fish gut microbiomes along an environmental gradient. J Environ Manage 318:115521. https://doi.org/10.1016/j.jenvman.2022.115521
Han XM, Hu HW, Chen QL, Yang LY, Li HL, Zhu YG, Li XZ, Ma YB (2018) Antibiotic resistance genes and associated bacterial communities in agricultural soils amended with different sources of animal manures. Soil Biol Biochem 126:91–102. https://doi.org/10.1016/j.soilbio.2018.08.018
He Y, Yuan Q, Mathieu J, Stadler L, Senehi N, Sun R, Alvarez PJJ (2020) Antibiotic resistance genes from livestock waste: Occurrence, dissemination, and treatment. NPJ Clean Water 3:4. https://doi.org/10.1038/s41545-020-0051-0
Heuer H, Schmitt H, Smalla K (2011) Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol 14:236–243. https://doi.org/10.1016/j.mib.2011.04.009
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: Prokaryotic gene recognition and translation initiation site identification. Bmc Bioinformatics 11. https://doi.org/10.1186/1471-2105-11-119
Kang DD, Li F, Kirton E, Thomas A, Egan R, An H, Wang Z (2019) Metabat 2: An adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7:e7359. https://doi.org/10.7717/peerj.7359
Kumar K, Gupta SC, Baidoo SK, Chander Y, Rosen CJ (2005) Antibiotic uptake by plants from soil fertilized with animal manure. J Environ Qual 34:2082–2085. https://doi.org/10.2134/jeq2005.0026
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9:357-U354. https://doi.org/10.1038/Nmeth.1923
Li DH, Liu CM, Luo RB, Sadakane K, Lam TW (2015) Megahit: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de bruijn graph. Bioinformatics 31:1674–1676. https://doi.org/10.1093/bioinformatics/btv033
Li LG, Xia Y, Zhang T (2017) Co-occurrence of antibiotic and metal resistance genes revealed in complete genome collection. Isme J 11:651–662. https://doi.org/10.1038/ismej.2016.155
Li T, Li R, Cao Y, Tao C, Deng X, Ou Y, Liu H, Shen Z, Li R, Shen Q (2022) Soil antibiotic abatement associates with the manipulation of soil microbiome via long-term fertilizer application. J Hazard Mater 439:129704. https://doi.org/10.1016/j.jhazmat.2022.129704
Mbanga J, Amoako DG, Abia ALK, Allam M, Ismail A, Essack SY (2021) Genomic analysis of enterococcus spp. Isolated from a wastewater treatment plant and its associated waters in umgungundlovu district, south africa. Front Microbiol 12:648454. https://doi.org/10.3389/fmicb.2021.648454
Morrison RS, Johnston LJ, Hilbrands AM (2007) The behaviour, welfare, growth performance and meat quality of pigs housed in a deep-litter, large group housing system compared to a conventional confinement system. Appl Anim Behav Sci 103:12–24. https://doi.org/10.1016/j.applanim.2006.04.002
Mulchandani R, Wang Y, Gilbert M, Van Boeckel TP (2023) Global trends in antimicrobial use in food-producing animals: 2020 to 2030. PLOS Global Public Health 3:e0001305. https://doi.org/10.1371/journal.pgph.0001305
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) Checkm: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055. https://doi.org/10.1101/gr.186072.114
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14:417-+. https://doi.org/10.1038/nmeth.4197
Pérez P, Solís-Tejeda MÁ, Lango-Reynoso F, Díaz-Rivera PD-R, Aguilar-Ávila J, Asiain-Hoyos A (2022) Deep litter pig production system as a sustainable alternative for small farmers. Agrociencia. https://doi.org/10.47163/agrociencia.v56i6.2755
Peterson E, Kaur P (2018) Antibiotic resistance mechanisms in bacteria: Relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Front Microbiol 9:2928. https://doi.org/10.3389/fmicb.2018.02928
Van Boeckel TP, Brower C, Gilbert M, Grenfell BT, Levin SA, Robinson TP, Teillant A, Laxminarayan R (2015) Global trends in antimicrobial use in food animals. P Natl Acad Sci USA 112:5649–5654. https://doi.org/10.1073/pnas.1503141112
Van Boeckel TP, Glennon EE, Chen D, Gilbert M, Robinson TP, Grenfell BT, Levin SA, Bonhoeffer S, Laxminarayan R (2017) Reducing antimicrobial use in food animals. Science 357:1350–1352. https://doi.org/10.1126/science.aao1495
Van Boeckel TP, Pires J, Silvester R, Zhao C, Song J, Criscuolo NG, Gilbert M, Bonhoeffer S, Laxminarayan R (2019) Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 365:1266-+. https://doi.org/10.1126/science.aaw1944
Van den Meersche T, Rasschaert G, Vanden Nest T, Haesebrouck F, Herman L, Van Coillie E, Van Weyenberg S, Daeseleire E, Heyndrickx M (2020) Longitudinal screening of antibiotic residues, antibiotic resistance genes and zoonotic bacteria in soils fertilized with pig manure. Environ Sci Pollut R 27:28016–28029. https://doi.org/10.1007/s11356-020-09119-y
Wang J, Ben W, Yang M, Zhang Y, Qiang Z (2016) Dissemination of veterinary antibiotics and corresponding resistance genes from a concentrated swine feedlot along the waste treatment paths. Environ Int 92–93:317–323. https://doi.org/10.1016/j.envint.2016.04.020
Williams-Nguyen J, Sallach JB, Bartelt-Hunt S, Boxall AB, Durso LM, McLain JE, Singer RS, Snow DD, Zilles JL (2016) Antibiotics and antibiotic resistance in agroecosystems: State of the science. J Environ Qual 45:394–406. https://doi.org/10.2134/jeq2015.07.0336
Xu Y, Li P, Gao Y, Li Y, Li S, Li Y, Sui P, Wang X, Gao W, Chen Y (2022) Sustainability evaluation for a circular maize-pig system driven by indigenous microbes: A case study in northeast china. Ecosyst Health Sustain 8:2080591. https://doi.org/10.1080/20964129.2022.2080591
Yang Y, Jiang X, Chai B, Ma L, Li B, Zhang A, Cole JR, Tiedje JM, Zhang T (2016) Args-oap: Online analysis pipeline for antibiotic resistance genes detection from metagenomic data using an integrated structured arg-database. Bioinformatics 32:2346–2351. https://doi.org/10.1093/bioinformatics/btw136
Yin XL, Jiang XT, Chai BL, Li LG, Yang Y, Cole JR, Tiedje JM, Zhang T (2018) Args-oap v2.0 with an expanded sarg database and hidden markov models for enhancement characterization and quantification of antibiotic resistance genes in environmental metagenomes. Bioinformatics 34:2263–2270. https://doi.org/10.1093/bioinformatics/bty053
Yu P, Dong P, Zou Y, Wang H (2023) Effect of ph on the mitigation of extracellular/intracellular antibiotic resistance genes and antibiotic resistance pathogenic bacteria during anaerobic fermentation of swine manure. Bioresour Technol 373:128706. https://doi.org/10.1016/j.biortech.2023.128706
Zhang AN, Gaston JM, Dai CL, Zhao S, Poyet M, Groussin M, Yin X, Li LG, van Loosdrecht MCM, Topp E, Gillings MR, Hanage WP, Tiedje JM, Moniz K, Alm EJ, Zhang T (2021) An omics-based framework for assessing the health risk of antimicrobial resistance genes. Nat Commun 12:4765. https://doi.org/10.1038/s41467-021-25096-3
Zhao RX, Yu K, Zhang JY, Zhang GJ, Huang J, Ma LP, Deng CF, Li XY, Li B (2020) Deciphering the mobility and bacterial hosts of antibiotic resistance genes under antibiotic selection pressure by metagenomic assembly and binning approaches. Water Research 186. https://doi.org/10.1016/j.watres.2020.116318
Zheng D, Yin G, Liu M, Hou L, Yang Y, Van Boeckel TP, Zheng Y, Li Y (2022) Global biogeography and projection of soil antibiotic resistance genes. Sci Adv 8:eabq8015. https://doi.org/10.1126/sciadv.abq8015
Zhou SY, Zhu D, Giles M, Yang XR, Daniell T, Neilson R, Zhu YG (2019) Phyllosphere of staple crops under pig manure fertilization, a reservoir of antibiotic resistance genes. Environ Pollut 252:227–235. https://doi.org/10.1016/j.envpol.2019.05.098. (Barking, Essex : 1987)
Zhou SY, Zhu D, Giles M, Daniell T, Neilson R, Yang XR (2020) Does reduced usage of antibiotics in livestock production mitigate the spread of antibiotic resistance in soil, earthworm guts, and the phyllosphere? Environ Int 136:105359. https://doi.org/10.1016/j.envint.2019.105359
Zhu L, Yuan L, Shuai XY, Lin ZJ, Sun YJ, Zhou ZC, Meng LX, Ju F, Chen H (2023) Deciphering basic and key traits of antibiotic resistome in influent and effluent of hospital wastewater treatment systems. Water Res 231:119614. https://doi.org/10.1016/j.watres.2023.119614
Zou Y, Xiao Y, Wang H, Fang T, Dong P (2020) New insight into fates of sulfonamide and tetracycline resistance genes and resistant bacteria during anaerobic digestion of manure at thermophilic and mesophilic temperatures. J Hazard Mater 384:121433. https://doi.org/10.1016/j.jhazmat.2019.121433
Acknowledgements
This work was supported by Fujian Special Fund for Scientific Research Institutes in the Public Interest (2020R1034002); Natural Science Foundation of Xiamen, China (3502Z202272400); and external cooperation project of Fujian Academy of Agricultural Sciences (DWHZ2021-15). The authors thank Yunfeng Agricultural and Animal Husbandry Cooperative, Maiping Li from the Baota District Animal Husbandry and Veterinary Service Center, and Xiyong Zhang for assisting with the sampling process.
Author information
Authors and Affiliations
Contributions
Zhizhen Pan, conceptualization, methodology, investigation, writing (original draft), visualization, data curation; Zheng Chen, methodology, investigation, resources; Liting Zhu, methodology, software, formal analysis; Ricardo David Avellán-Llaguno, methodology, software, formal analysis; Bo Liu, writing (review and editing), funding acquisition; Qiansheng Huang: conceptualization, methodology, writing (original draft), writing (review and editing), visualization, supervision, funding acquisition. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pan, Z., Chen, Z., Zhu, L. et al. Antibiotic resistome and associated bacterial communities in agricultural soil following the amendments of swine manure–derived fermentation bed waste. Environ Sci Pollut Res 30, 104520–104531 (2023). https://doi.org/10.1007/s11356-023-29691-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-29691-3