Abstract
The prevalence and persistence of antibiotic resistance genes in wastewater treatment plants (WWTPs) is of growing interest, and residual sludge is among the main sources for the release of antibiotic resistance genes (ARGs). Moreover, heavy metals concentrated in dense microbial communities of sludge could potentially favor co-selection of ARGs and metal resistance genes (MRGs). Residual sludge treatment is needed to limit the spread of resistance from WWTPs into the environment. This study aimed to explore the fate of ARGs and MRGs during thermophilic two-phase (acidogenic/methanogenic phase) anaerobic digestion by metagenomic analysis. The occurrence and abundance of mobile genetic elements were also determined based on the SEED database. Among the 27 major ARG subtypes detected in feed sludge, large reductions (> 50%) in 6 ARG subtypes were achieved by acidogenic phase (AP), while 63.0% of the ARG subtypes proliferated in the following methanogenic phase (MP). In contrast, a 2.8-fold increase in total MRG abundance was found in AP, while the total abundance during MP decreased to the same order of magnitude as in feed sludge. The distinct dynamics of ARGs and MRGs during the two-phase anaerobic digestion are noteworthy, and more specific treatments are required to limit their proliferation in the environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Antibiotic resistance genes (ARGs) and their abundance in the natural environment are becoming a big health concern, and wastewater treatment plants (WWTPs) are demonstrated to be a hub and a significant source for their spread to the environment (Berendonk et al. 2015; LaPara et al. 2011; Yang et al. 2012). Sewage containing residual antibiotics from a variety of sources, including hospitals and households, veterinary clinics, and agriculture all end up in WWTPs (Rizzo et al. 2013). Meanwhile, heavy metals, another significant part of anthropogenic emissions in WWTPs, can hardly be removed by activated sludge processes and concentrate in the sludge. They could enhance the selection of multi-resistant bacteria and the spread of resistances into the environment (Di Cesare et al. 2016; Gao et al. 2015). Moreover, metal resistance genes (MRGs) could be linked to ARGs, particularly on plasmids, and their co-selection could be highly favored if they are located on the same mobile genetic element such as integrons, transposons, and plasmids (Baker-Austin et al. 2006), which could greatly facilitate their spread and proliferation via horizontal gene transfer (HGT) in return. Thus, residual solids of WWTPs have undoubtedly become an important node for controlling the dissemination of antibiotic and metal resistance genes into the natural environment.
In recent years, various existing technologies have been used to treat excess sludge from WWTPs in order to achieve safe disposal or land application, such as anaerobic and aerobic digestion, composting, air-drying, chemical stabilization, etc. Ma et al. (2011) have already investigated the effect of various anaerobic digestion conditions on the fate of ARGs and suggested that temperature is the crucial factor, and studies also showed that bacterial community composition drives the distribution of ARGs during composting (Cui et al. 2016; Su et al. 2015). However, owing to the specificity and limitation of quantitative polymerase chain reaction (q-PCR), these prior studies could only identify a handful of ARGs and did not uncover the full spectrum of functional resistance genes such as metal resistance genes and mobile genetic elements during these processes.
In the present study, a metagenomic analysis based on high-throughput DNA sequencing was used to simultaneously explore the wide-ranging profile of ARGs, MRGs, and mobile genetic elements during two-phase anaerobic digestion process. It is an optimization of the single-phase anaerobic digestion thereby achieving more efficient sludge stabilization with separation of acidogenic and methanogenic processes to two reactors in series (Pohland and Ghosh 1971). Apart from this, two-phase anaerobic digestion processes are designed to consider the different ecophysiological optima of the microbes involved in these processes (Demirel and Yenigun 2002; Pohland and Ghosh 1971). The first phase (acidogenesis) is maintained at low pH and short hydraulic residence time (HRT) resulting in a consortium of acid-consuming organisms, while the second phase (methanogenesis) is operated at longer HRT and neutral pH in order to facilitate proliferation of slow-growing methanogenic archaea. Thus, two distinctly different microbial communities are selected and enriched in the acidogenic and methanogenic phases, which is believed to harbor different communities of resistance genes as previously mentioned.
The objective of present study, therefore, was to determine the effect of two-phase anaerobic digestion on the fate of ARGs and MRGs under thermophilic condition, and investigate their associations during the process. The occurrence and abundance of mobile genetic elements were also investigated. Moreover, metagenomics can overcome the limitations of PCR detection methods, such as applicability of primer/probe systems in a difficult matrix and possible biases in the amplification process (Volkmann et al. 2007). We believe this study provided a comprehensive profile and monitored the dynamics of these functional genes in the two-phase anaerobic digestion, and increased our knowledge on the role of two-phase anaerobic digestion in the spread and dissemination of ARGs and MRGs into the natural environment.
Materials and methods
Reactor start-up and operation
Two-phase anaerobic digesters were set up in the lab. An acidogenic phase reactor (AP) was 1 L continuously stirred tank reactors (CSTR) with flask working volume of 0.3 L preceding a methanogenic-phase reactor (MP), 1.5 L CSTR with flask working volume of 1 L. The reactors were maintained at 55 °C through circulating hot water around the flask. Bench-scale two-phase anaerobic digesters were initiated with seeding sludge obtained from an anaerobic bioreactor of Linan municipal wastewater treatment plant (Hangzhou, China). Dewatered sewage sludge was also collected from the Linan municipal wastewater treatment plant and stored at − 20 °C as the source of feed sludge through the duration of the experiment. The characteristic of feed sludge was described in our previous work (Wu et al. 2016).
To maintain the sludge retention time (SRT) of AP and MR at 3 and 10 days, 0.1 L effluent from AP was replaced by same sludge feedstock once a day, and 0.l L of effluent from MR was replaced by the AP effluent (0.1 L) once a day. The feeding was started after the effluent was withdrawn. pH of AP was controlled around 6.0 by adding 8 g/L NaHCO3, which could promote the growth of hydrolytic and acidogenic bacteria in the acidogenic phase (Luo et al. 2011). Biogas production in AP and MR was monitored by water replacement devices. Total solid content and volatile solid (VS) were analyzed according to standard methods (Zhan 2009). Volatile fatty acids (VFA) concentration was measured by using gas chromatograph (GC) (Agilent, 6890 N) equipped with a flame ionization detector and DB-624 column (30 m × 0.53 mm × 3.0 μm). Samples analyzed for volatile fatty acids were centrifuged at 10,000 rpm for 10 min and then the supernatant was filtrated though a 0.45-μm membrane. Formic acid was added to adjust the pH to approximately 2.0 before loading onto the GC. Both reactors had achieved stabilized performance with respect to biogas production, VS reduction, and VFA production at the time samples collection commenced.
Sample collection, DNA extraction, and high-throughput sequencing
Raw feed sludge, effluent of AP, and MR were collected over three sampling events with a time interval of 6 days. These three composite samples for each point were immediately centrifuged at 10,000 rpm for 10 min, and after freeze-drying and sieving through a 2-mm mesh, the pellet was stored at − 80 °C before DNA extraction. DNA extraction was performed using a FastDNA Spin Kit for Soil (MP, Biomedicals) from approximately 0.1 g of lyophilized samples. The purified DNA concentration and quality were determined by 1.5% agar gel electrophoresis and spectrophotometer analysis (NanoDrop ND-2000c, Thermo).
High-throughput sequencing was performed in Analysis Center of Agrobiology and Environmental Sciences of Zhejiang University using Ion Proton™ system. About 100 ng of DNA of the feed sludge, AP, and MR was used for library construction and DNA sequencing. Approximately 22.8 Gb of data was generated for all the samples. Metagenomic data were deposited in the NCBI Short Read Archive.
Bioinformatic analysis
The metagenomic data sets were filtered using a self-written script to remove the reads containing five or more ambiguous nucleotides and those with lengths of less than 100 bp. The detailed information about the sequencing reads for each sample was described in Table S1. In order to compare the abundance and diversity of functional genes in different sludge samples, sequencing data was normalized to 10,000,000 reads for each dataset. ARG-like sequences were identified by aligning the sequences against the Antibiotic Resistance Database (ARDB) using BLAST with the e-value at 1 × 10−5. The sequence was annotated as an ARG-like sequence if its best hit in the ARDB had ≥ 90% amino acid identity and an alignment length longer than 25 amino acids (Kristiansson et al. 2011; Zhang et al. 2011). MRGs annotation was conducted similarly by searching against the BacMet database (Pal et al. 2014). Mobile genetic elements were obtained by aligning the sequences against the SEED subsystems database (Overbeek et al. 2005) using a maximum e-value of 10−5.
Statistical analysis
The portion of types or subtypes of sequences with different functions in “total metagenome sequences” was defined as “abundance” (using the unit of “ppm”, one read in a million reads), and the composition of functional gene sequences was described using the unit of “%” to represent the portion of a type or a subtype of functional gene-like sequences in total sequences with the same function. Previous studies about the functional genes in the environmental samples also applied the units of “ppm” and “%” (Li et al. 2015; Zhang et al. 2015). Correlations between the prevalence of ARGs, MRGs, and mobile genetic elements were analyzed using MATLAB R2016a (The MathWorks, Natick, MA). In addition, the correlation analyses between antibiotic or metal resistance genes and microbial community were performed, respectively. A p value < 0.05 was considered statistically significant.
Results
Occurrence and abundance of ARGs in two-phase anaerobic digesters
ARG-like reads corresponding to 6 ARG types were identified in feed sludge with a total abundance of 229.8 ppm. Sulfonamide resistance genes (121.6 ppm, 53%), aminoglycoside resistance genes (51.8 ppm, 23%), and tetracycline resistance genes (27.4 ppm, 12%) were the most abundant ARG types found in feed sludge (Fig. 1). A fluctuation of total ARG abundance was found during two-phase anaerobic digestion. The total ARG abundance in AP (169.8 ppm) decreased by 26.1% on basis of feed sludge, while that of MP rebounded to 355.7 ppm (1.1 fold of AP), specifically regarding tetracycline resistance genes. The abundance of tetracycline resistance genes in MP was approximately eight times higher than that in AP.
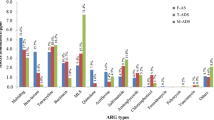
The abundance of 27 ARG subtypes from the top 3 abundant resistance genes of sulfonamide, aminoglycoside, and tetracycline is shown in Fig. 2. Sul1 (114.8 ppm), aph6Id (17.7 ppm), and tetC (17.2 ppm) were the most abundant ARG subtypes in feed sludge. Through two-phase anaerobic digestion, the major ARG subtypes changed to sul1, aph33Ib, and ant3Ia in the AP, and sul1, tetC, and tetM in the MP. After the AP digestion, 9 ARG subtypes decreased based on feed sludge, especially aph3Ib, sul2, tetX, aph6Id, tetG, and sul1, which all had a reduction of over 50% based on feed sludge, while another 13 ARGs increased within the same order of magnitude as feed sludge. In contrast to AP, 17 ARG subtypes proliferated in the methanogenic phase, specifically tetM and tetW which increased from 0 to 46.8 ppm and from 0.2 to 43.2 ppm, respectively, while 5 subtypes of aminoglycoside resistance genes and 4 subtypes of tetracycline resistance genes decreased.
Occurrence and abundance of MRGs in two-phase anaerobic digesters
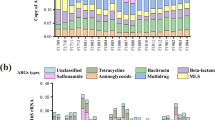
Nine MRG types were identified in the feed sludge with a total abundance of 145.3 ppm, and all of them were shared by AP and MP including MRGs corresponding to copper, arsenic, mercury, chromium, lead, zinc, iron, nickel, as well as multi-metal resistance (MRGs that encoded resistance to two or more metals) (Fig. 3). Copper, arsenic, and chromium resistance genes were the major MRG types in feed sludge, which occupied the largest fraction of 71%. During two-phase anaerobic digestion, the abundance of total MRG-like sequences changes. An obvious increase of total MRG abundance was found in AP, which added up to 399.7 ppm, a 2.8-fold increase compared to the feed sludge. In contrast, the total MRG abundance falling in MP was 194.7 ppm, which was well within the same order of magnitude as the feed sludge.
Sixty-five subtypes of MRGs were found in feed sludge and two-phase anaerobic digesters (Fig. 4). DnaK (16 ppm), arsB (14.5 ppm), ruvB (11.7 ppm), and copB (10.1 ppm) had the highest abundances in the feed sludge, while the abundance of other MRG subtypes was lower than 10 ppm. Through two-phase anaerobic digestion, the major MRG subtypes changed to ziaA, dnaK, and mgtA in AP, and arsB, dnaK, and ziaA in MP. In addition, obvious fluctuations were found in the abundance of a few MRG subtypes, especially the zinc resistance gene ziaA. After the acidogenic phase of anaerobic digestion, ziaA significantly increased from 0.5 to 220.6 ppm, while most MRG subtypes fluctuated at a much smaller scale and the abundances of cinA, cueR, mmco, and arsR did not change. In the following MP, 56.9% subtypes of MRGs decreased within a small range of 0.1–5.9 ppm based on AP, while ziaA decreased by 88.1% of the value at AP. In contrast, 29.2% MRGs slightly increased within a small range of 0.1–3.2 ppm while arsB increased by 2-fold based on AP.
Mobile genetic elements in thermophilic two-phase anaerobic digester
The occurrence and abundance of genes encoding mobile genetic elements were obtained from the SEED database (Table S2). Fifty-two types of mobile genetic elements were found in feed sludge and two-phase anaerobic digesters. The total abundance of mobile genetic elements was 253.2 ppm in feed sludge, and traC (32.6 ppm, 12.9%), traG (23.6 ppm, 9.3%) as well as integron integrase gene (22.4 ppm, 8.8%) were the major types of mobile genetic elements. After the two-phase anaerobic digestion, obvious reductions of the total relative abundance of mobile genetic elements were found in both AP and MP. The total abundance of mobile genetic elements in AP and MP was 43.7 and 76.5 ppm, respectively, which decreased by 209.5 ppm (82.7%) and 176.7 ppm (69.8%) based on feed sludge. In addition, sulfonamide resistance genes were positively correlated with IntIPac and TraI (p < 0.05), and tetracycline resistance genes were found to be correlated with nine types of conjugative transposon significantly (data not shown). In contrast, Pb and Fe resistance genes were both correlated with mobile genetic elements of IncF plasmid. No positive correlations were found between ARGs and MRGs during thermophilic two-phase anaerobic digestion (p > 0.05).
Microbial community structures during two-phase digestion
The taxonomic structure including both bacteria at the phylum level and two archaea classes is shown in Fig. S1. It revealed that microbial communities in two-phase digestors are dynamic, capable of changes at acidogenic and methanogenic phase relative to feed sludge. Proteobacteria occupied the largest part (53.3%) in the feed sludge, followed by Nitrospirae (28.8%) and Actinobacteria (9.2%). Large differences in microbial community structures were observed after two-phase digestion. The AP reactor was dominated by Firmicutes with a high percentage up to 95.3%, which is a unique pattern greatly affected by acidogenic condition. Although Firmicutes was still the most abundant organism (54.5%) in the MP reactor, a more diverse microbial community was found, especially the methane-generating archaea such as Methanomicrobia and Methanobacteria increased to a larger percentage, 19.3 and 11.5%, respectively. Thus, very distinct microbial communities have been formed in the AP and MP reactor, separately. In the correlation analysis between microbial community and antibiotic resistance genes (Table S9), Methanomicrobia was negatively correlated with β-lactam resistance genes while Firmicutes and Synergistetes both were correlated with macrolide resistance genes positively (p < 0.05). In the aspect of metal resistance genes, both Thermotogae and Aquificae were positively associated with zinc resistance genes, and Nitrospirae was associated with arsenic resistance genes negatively (p < 0.05).
Discussion
Municipal wastewater from different sources such as households, hospitals, farms, and factories are mixed in the WWTP, often containing antibiotics, heavy metals, and other organic matters. Accordingly, the dense microbial communities of residue sludge could be a hotspot for the potential co-selection of ARGs and MRGs (Di Cesare et al. 2016; Li et al. 2015). The technical literature, however, contains relatively little information about the influence of two-phase anaerobic digestion on the changing pattern of genes that encode for antibiotic and metal resistance determinants. In the present work, we thus used a metagenomic approach to investigate the effect of two-phase anaerobic digestion on the abundances of ARGs and MRGs and mobile genetic elements, and explore the link between these resistance genes and the microbial community.
Sulfonamide, aminoglycoside, and tetracycline resistance genes were the major ARG types found in feed sludge and the two-phase anaerobic digesters. The prevalence of sulfonamide and tetracycline resistance genes in sludge samples may result from the frequent use of sulfonamides and tetracycline for livestock (Cheng et al. 2013; Sui et al. 2016). Moreover, high concentrations of sulfamethoxazole and oxytetracycline were also detected in this WWTP according to our previous study (Li et al. 2016). In addition, aminoglycoside antibiotics play an important role in the therapy of serious staphylococcal infections, and prevalence of aminoglycoside resistance genes were widely discovered in clinical isolates (Schmitz et al. 1999). It indicated that a hospital might be one of the sources that altered the composition of ARGs in sewage, and further contributed to the widespread occurrence of aminoglycoside resistance genes in the WWTP.
A decrease in abundances of most ARGs within AP would suggest either that the organisms harboring genes encoding antibiotic resistance determinants are inactivated within the acidogenic phase or that the rate of HGT is low. The relatively low pH and acidogenic conditions within AP lead to a very different microbial structure, which enriched Clostridia and disinfect others from the feed sludge. On the other hand, the abundance of mobile genetic elements in AP was significantly lower than in feed sludge and the two abundant types of ARGs (sulfonamide and tetracycline resistance genes) were both positively correlated with mobile genetic elements; thus, it might influence the spread of these antibiotic resistance genes by HGT (Broaders et al. 2013). This finding may be responsible for the decrease of ARGs in the AP. On the contrary, the abundance of macrolide resistance genes was increased in AP reactor, and it was positively associated with Firmicutes in the correlation analysis. Soge et al. reported that macrolide resistance gene (mefA) was found in environmental Clostridium perfringens, one species belonging to the Firmicutes phylum (Soge et al. 2009). This indicates that microbes in the phylum of Firmicutes may be the host microorganism of macrolide resistance genes in our study.
In feed sludge and two-phase anaerobic digesters, the genes encoding resistance determinants for copper, arsenic, zinc, and chromium were the most common in all the detected MRGs, because these metal ions are essential for most microorganisms, yet they can be toxic at high concentrations (Ji and Silver 1995). Meanwhile, Copper and arsenic compounds have been widely used as antimicrobial, pesticidal, and antifungal agents, and as animal feed additives (Hobman and Crossman 2015). Zinc was the most primary heavy metal in sewage sludge as galvanized pipes were quite widespread in China, while chromium is commonly found in effluents from tanneries and relevant industries (Aravind et al. 2016). The extensive use of those metals could contribute to the high abundance of copper, arsenic, zinc, and chromium resistance genes in sewage sludge.
Compared to ARGs, more subtypes of MRGs were discovered in feed sludge and two-phase anaerobic digesters, but their abundance was much lower. This may indicate that various types of metals were present in sludge, but the bioavailability and mobility of metals were different due to their speciation, both physical and chemical. Moreover, previous studies had found that sewage sludge exhibited higher concentrations of heavy metals bound to the oxidizable and residue fractions, compared to the mobile ones (Jamali et al. 2007; Tytla et al. 2016). Thus, large amount of metals may not be freely available to bacteria, and this may be one explanation for the lower abundance of MRGs in present study.
It should be noted that zinc resistance gene subtype ziaA largely increased during the acidogenic phase of anaerobic digestion. Zn was found to be significantly present in the acid-extractable fraction but had a very low proportion in the oxidizable fraction in sewage sludge, which suggests that zinc is more likely released into the surrounding environment and taken up by organisms (Fadiran et al. 2014; Tou et al. 2017). Accordingly, the low pH in acidogenic phase would result in a high concentration of zinc in the AP reactor. In order to deal with excess zinc, microorganisms have zinc-specific efflux pumps, encoded by ziaA, that transport Zn2+ from the cytoplasm to the periplasmic space. This efflux system expression is induced by zinc and is regulated by a zinc-specific repressor protein, ZiaR (Barnett et al. 2012; Thelwell et al. 1998). Thus, the surge of ziaA is activated by significantly high bioavailable zinc concentrations in the acidogenic phase. On the other hand, although zinc resistance genes were found to significantly correlated with Thermotogae and Aquificae positively (p < 0.05), no direct evidence in literature shows that microorganisms in these two phyla harbored zinc resistance genes. In the following MP, however, more than half of MRG subtypes decreased especially ziaA, indicating that the abundance of bioavailable metal ions in environmental niches of MP may decrease, thus declining the selection pressure. On the other hand, no significant correlations were found between MRGs and ARGs, suggesting that the changing mechanism of MRGs and ARGs is different. And the mobile genetic elements such as IncF plasmid may also play a role in the spread of Pb and Fe resistance genes, which is comparable with the early recognition of Pb resistance locating on pMOL30 plasmid (Diels et al. 1989).
The potential of two-phase anaerobic digestion process to mitigate the spread of ARGs and MRGs requires further researches. In this study, ARG patterns may be affected by bacterial community structures in the two-phase anaerobic digesters as previous study suggested (Wu et al. 2016). In contrast, the abundance of MRGs seems to be driven by the metal concentrations, more so than the different composition of bacterial community in the two phases. Thus, more specific measures should be taken to control the spread of ARGs and MRGs in the environment. The results of this study could also be used to guide selection for q-PCR assays to target representative ARGs, MRGs, and mobile genetic elements of importance, and provide support for advanced biological risk assessment evaluations, which are needed to determine how polluted environments affect the proliferation of antibiotic resistance.
Conclusion
This study has revealed the impact of two-phase anaerobic digestion on the profile of ARGs and MRGs. ARGs showed higher prevalence in feed sludge and two-phase anaerobic digesters in comparison to MRGs. AP was an effective phase to eliminate most ARGs. The elevated abundance of zinc resistance gene and no significant correlations between ARGs and MRGs indicates their distinct fate during two-phase anaerobic digestion. Mobile genetic elements possibly mediated the spread and proliferation of ARGs and MRGs. Further research is needed to explore the potential transfer mechanisms of specific ARGs and MRGs in the digesters to develop advanced sludge treatment.
References
Aravind J, Kanmani P, Sudha G, Balan R (2016) Optimization of chromium(VI) biosorption using gooseberry seeds by response surface methodology. Global J Environ Sci Manage 2:61–68
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182
Barnett JP, Millard A, Ksibe AZ, Scanlan DJ, Schmid R, Blindauer CA (2012) Mining genomes of marine cyanobacteria for elements of zinc homeostasis. Front Microbiol 3:142
Berendonk TU, Manaia CM, Merlin C, Fatta-Kassinos D, Cytryn E, Walsh F, Buergmann H, Sorum H, Norstrom M, Pons MN, Kreuzinger N, Huovinen P, Stefani S, Schwartz T, Kisand V, Baquero F, Luis Martinez J (2015) Tackling antibiotic resistance: the environmental framework. Nat Rev Microbiol 13:310–317
Broaders E, Gahan CGM, Marchesi JR (2013) Mobile genetic elements of the human gastrointestinal tract: potential for spread of antibiotic resistance genes. Gut Microbes 4:271–280
Cheng W, Chen H, Su C, Yan S (2013) Abundance and persistence of antibiotic resistance genes in livestock farms: a comprehensive investigation in eastern China. Environ Int 61:1–7
Cui E, Wu Y, Zuo Y, Chen H (2016) Effect of different biochars on antibiotic resistance genes and bacterial community during chicken manure composting. Bioresour Technol 203:11–17
Demirel B, Yenigun O (2002) Two-phase anaerobic digestion processes: a review. J Chem Technol Biotechnol 77:743–755
Di Cesare A, Eckert EM, D'Urso S, Bertoni R, Gillan DC, Wattiez R, Corno G (2016) Co-occurrence of integrase 1, antibiotic and heavy metal resistance genes in municipal wastewater treatment plants. Water Res 94:208–214
Diels L, Sadouk A, Mergeay M (1989) Large plasmids governing multiple resistance to heavy metals—a genetic approach. Toxicol Environ Chem 23:79–89
Fadiran AO, Tiruneh AT, Mtshali JS (2014) Assessment of mobility and bioavailability of heavy metals in sewage sludge from Swaziland through speciation analysis. Am J Environ Protect 3:198–208
Gao P, He S, Huang S, Li K, Liu Z, Xue G, Sun W (2015) Impacts of coexisting antibiotics, antibacterial residues, and heavy metals on the occurrence of erythromycin resistance genes in urban wastewater. Appl Microbiol Biotechnol 99:3971–3980
Hobman JL, Crossman LC (2015) Bacterial antimicrobial metal ion resistance. J Med Microbiol 64:471–497
Jamali MK, Kazi TG, Afridi HI, Arain MB, Jalbani N, Memon AR (2007) Speciation of heavy metals in untreated domestic wastewater sludge by time saving BCR sequential extraction method. J Environ Sci Health A Tox Hazard Subst Environ Eng 42:649–659
Ji GY, Silver S (1995) Bacterial-resistance mechanisms for heavy-metals of environmental concern. J Ind Microbiol 14:61–75
Kristiansson E, Fick J, Janzon A, Grabic R, Rutgersson C, Weijdegard B, Soderstrom H, Larsson DGJ (2011) Pyrosequencing of antibiotic-contaminated river sediments reveals high levels of resistance and gene transfer elements. PLoS One 6:e17038
LaPara TM, Burch TR, McNamara PJ, Tan DT, Yan M, Eichmiller JJ (2011) Tertiary-treated municipal wastewater is a significant point source of antibiotic resistance genes into Duluth-superior harbor. Environ Sci Technol 45:9543–9549
Li AD, Li LG, Zhang T (2015) Exploring antibiotic resistance genes and metal resistance genes in plasmid metagenomes from wastewater treatment plants. Front Microbiol 6
Li J, Cheng W, Xu L, Jiao Y, Baig SA, Chen H (2016) Occurrence and removal of antibiotics and the corresponding resistance genes in wastewater treatment plants: effluents’ influence to downstream water environment. Environ Sci Pollut Res 23:6826–6835
Luo G, Xie L, Zhou Q, Angelidaki I (2011) Enhancement of bioenergy production from organic wastes by two-stage anaerobic hydrogen and methane production process. Bioresour Technol 102:8700–8706
Ma Y, Wilson CA, Novak JT, Riffat R, Aynur S, Murthy S, Prudens A (2011) Effect of various sludge digestion conditions on sulfonamide, macrolide, and tetracycline resistance genes and class I integrons. Environ Sci Technol 45:7855–7861
Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M, de Crecy-Lagard V, Diaz N, Disz T, Edwards R, Fonstein M, Frank ED, Gerdes S, Glass EM, Goesmann A, Hanson A, Iwata-Reuyl D, Jensen R, Jamshidi N, Krause L, Kubal M, Larsen N, Linke B, McHardy AC, Meyer F, Neuweger H, Olsen G, Olson R, Osterman A, Portnoy V, Pusch GD, Rodionov DA, Ruckert C, Steiner J, Stevens R, Thiele I, Vassieva O, Ye Y, Zagnitko O, Vonstein V (2005) The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33:5691–5702
Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DGJ (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42:D737–D743
Pohland FG, Ghosh S (1971) Developments in anaerobic stabilization of organic wastes—the two-phase concept. Environ Lett 1:255–266
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassinos D (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447:345–360
Schmitz FJ, Fluit AC, Gondolf M, Beyrau R, Lindenlauf E, Verhoef J, Heinz HP, Jones ME (1999) The prevalence of aminoglycoside resistance and corresponding resistance genes in clinical isolates of staphylococci from 19 European hospitals. J Antimicrob Chemother 43:253–259
Soge OO, Tivoli LD, Meschke JS, Roberts MC (2009) A conjugative macrolide resistance gene, mef (A), in environmental Clostridium perfringens carrying multiple macrolide and/or tetracycline resistance genes. J Appl Microbiol 106:34–40
Su JQ, Wei B, Ou-Yang WY, Huang FY, Zhao Y, Xu HJ, Zhu YG (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol 49:7356–7363
Sui QW, Zhang JY, Chen MX, Tong J, Wang R, Wei YS (2016) Distribution of antibiotic resistance genes (ARGs) in anaerobic digestion and land application of swine wastewater. Environ Pollut 213:751–759
Thelwell C, Robinson NJ, Turner-Cavet JS (1998) An SmtB-like repressor from Synechocystis PCC 6803 regulates a zinc exporter. Proc Nati Acad Sci 95:10728–10733
Tou F, Yang Y, Feng J, Niu Z, Pan H, Qin Y, Guo X, Meng X, Liu M, Hochella MF (2017) Environmental risk implications of metals in sludges from waste water treatment plants: the discovery of vast stores of metal-containing nanoparticles. Environ Sci Technol 51:4831–4840
Tytla M, Widziewicz K, Zielewicz E (2016) Heavy metals and its chemical speciation in sewage sludge at different stages of processing. Environ Technol 37:899–908
Volkmann H, Schwartz T, Kirchen S, Stofer C, Obst U (2007) Evaluation of inhibition and cross-reaction effects on real-time PCR applied to the total DNA of wastewater samples for the quantification of bacterial antibiotic resistance genes and taxon-specific targets. Mol Cell Probes 21:125–133
Wu Y, Cui E, Zuo Y, Cheng W, Rensing C, Chen H (2016) Influence of two-phase anaerobic digestion on fate of selected antibiotic resistance genes and class I integrons in municipal wastewater sludge. Bioresour Technol 211:411–421
Yang Y, Zhang T, Zhang XX, Liang DW, Zhang M, Gao DW, Zhu HG, Huang QG, Fang HHP (2012) Quantification and characterization of beta-lactam resistance genes in 15 sewage treatment plants from East Asia and North America. Appl Microbiol Biotechnol 95:1351–1358
Zhan D (2009) Resource utilization of sludge. China Ocean University Press, Qingdao
Zhang T, Yang Y, Pruden A (2015) Effect of temperature on removal of antibiotic resistance genes by anaerobic digestion of activated sludge revealed by metagenomic approach. Appl Microbiol Biotechnol 99:7771–7779
Zhang T, Zhang XX, Ye L (2011) Plasmid metagenome reveals high levels of antibiotic resistance genes and mobile genetic elements in activated sludge. PLoS One 6
Acknowledgements
The authors would like to thank the managers of Linan municipal wastewater treatment plant for providing raw sludge, and the support of the IBM high performance computing cluster of Bio-macromolecules Analysis Lab, Zhejiang University.
Funding
This work was supported by the Natural Science Foundation of China (21677121, 41571130064).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Gerald Thouand
Electronic supplementary material
ESM 1
(DOC 879 kb)
Rights and permissions
About this article
Cite this article
Wu, Y., Cui, E., Zuo, Y. et al. Fate of antibiotic and metal resistance genes during two-phase anaerobic digestion of residue sludge revealed by metagenomic approach. Environ Sci Pollut Res 25, 13956–13963 (2018). https://doi.org/10.1007/s11356-018-1598-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-1598-x