Abstract
Bacterium Pseudomonas aeruginosa BCH was able to degrade naphthylaminesulfonic azo dye Amaranth in plain distilled water within 6 h at 50 mg l−1 dye concentration. Studies were carried out to find the optimum physical conditions and which came out to be pH 7 and temperature 30 °C. Amaranth could also be decolorized at concentration 500 mg l−1. Presence of Zn and Hg ions could strongly slow down the decolorization process, whereas decolorization progressed rapidly in presence of Mn. Decolorization rate was increased with increasing cell mass. Induction in intracellular and extracellular activities of tyrosinase and NADH-DCIP reductase along with intracellular laccase and veratryl alcohol oxidase indicated their co-ordinate action during dye biodegradation. Up-flow bioreactor studies with alginate immobilized cells proved the capability of strain to degrade Amaranth in continuous process at 20 ml h−1 flow rate. Various analytical studies viz.—HPLC, HPTLC, and FTIR gave the confirmation that decolorization was due to biodegradation. From GC-MS analysis, various metabolites were detected, and possible degradation pathway was predicted. Toxicity studies carried out with Allium cepa L. through the assessment of various antioxidant enzymes viz. sulphur oxide dismutase, guaiacol peroxidase, and catalase along with estimation of lipid peroxidation and protein oxidation levels conclusively demonstrated that oxidative stress was generated by Amaranth.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Azo dyes are most widely and diversely used dyes. Among the 12 classes of chromogenic groups of dyes, they make about 70 % of the textile dyestuff produced and considered as very recalcitrant xenobiotic compounds to biodegradation processes (Lee and Pavlostathis 2004). In textile wastewater, concentration of reactive azo dyes ranges from 5 to 1,500 mg l−1 (Pierce 1994). About 3,000 azo dyes are used in various industries such as paper, food, cosmetics, and pharmaceuticals (Pan et al. 2012) and over 7 × 105 t are produced annually worldwide (Puvaneswari et al. 2006). Along with this, toxicity of these dyes is the main reason for prime intention of researchers towards their remediation. Sudan azo dyes and their metabolites have been shown to inhibit the growth of human intestinal bacteria (Pan et al. 2012). Azo dyes have been shown to be mutagenic to the human hepatoma cell line where frame shift mutation was observed (Ferraz et al. 2011a, b) and also found to induce the micronuclei formation in human lymphocytes and in HepG2 cells (Chequer et al. 2009). Mutagenicity of these dyes and their reduction products have been exposed by Jaochim et al. (1985) in Salmonella/microsome assay. Turkish government has banned the dyes azo dyes since 1 March 1995 due to the relevant aromatic amines. However, due to their efficiency of dyeing and cost, they have been still in use in textile dyeing processes in Turkey and in some countries (Isik and Sponza 2003). Conventional treatment approaches to the textile-dyeing wastewater consist of several combinations of chemical and physical methods which includes coagulation, precipitation and adsorption. However, large variability of the textile-dyeing wastewater is proving these methods as inadequate. Although many attempts have been made to apply electron-beam irradiation, photo-oxidation, supercritical water oxidation, and electrochemical methods for the treatment of wastewater, still no reliable textile-dyeing wastewater treatment process has been established so far (Mok and Jo 2007). Also, with time, potential hazards and disadvantages of these methods were noted as highly expensive coupled with formation of large amount of toxic sludge and the emission of even more toxic metabolites (Davis et al. 1994; Johnson et al. 1978).
Amaranth is one the representative of azo class of dyes. It is an anionic dye and which found its uses in natural and synthetic fibres, leather and paper industry, phenol-formaldehyde resins, and also as a food additive. However, there is toxicity concern for the use of this dye. Notably, exercise of this dye is not allowed in USA but not prohibited in other countries (Gomi et al. 2011). Hong et al. (2007) has reported the mutagenicity of Amaranth in Ames test but no cytotoxicity was observed. Dye was also found nontoxic in Microtox assay with Vibrio fischeri (Ramsay and Nguyen 2002).
Herein, we report the detailed investigation on biodegradation of Amaranth by Pseudomonas aeruginosa BCH in plain distilled water, including the enzymes involved and possible biodegradation pathway. This may be the first time report on bacterial decolorization in plain water. Efforts are also made to get insights into the toxicity of the dye with respect to generation of oxidative stress. And as an application part, we also examined the biodegradation of the dye in continuous decolorization study in up-flow bioreactor with alginate immobilized cells.
Materials and methods
Chemicals
Amaranth dye was purchased from Himedia Pvt. Ltd., India. Other chemicals such as o-tolidine, veratryl alcohol, l-ascorbic acid, Na-alginate, thiobarbituric acid, and DCIP were also obtained from Himedia Pvt. Ltd, India. Dichloromethane was purchased from Merck, India and ethyl acetate from sd-fine chem. limited, India. Nicotinamide adenine dinucleotide reduced salt (NADH) was purchased from SRL Chemicals, India.
Culture conditions and biodegradation assay
Bacterial strain P. aeruginosa BCH used in this study was previously isolated in our laboratory from dye-contaminated soil (Jadhav et al. 2010a). The pure culture was maintained at 4 °C on yeast extract agar medium having composition (grams per liter): yeast extract 5, NaCl 5, and agar 25.
The medium for bacterial growth was composed of (grams per liter): yeast extract 2, NaCl 5. A single colony of bacterium was inoculated in 250 ml Erlenmeyer flasks containing 100 ml medium and grown for 8 h at 30 °C at shaking condition (120 rpm). From this inoculum, 1 ml of culture was then inoculated in the 100 ml fresh medium and grown for 24 h as above (idea was to grow the cells of the log phase for 24 h). Cells were harvested by centrifugation at 6,000 rpm for 10 min at 4 °C (dry cells weight was 30 ± 3 mg). Centrifuged cells were washed twice with sterile distilled water and resuspended in distilled water at a concentration of 1 g l−1 dry weight. Dye was added at a concentration of 50 mg l−l from the stock solution prepared in sterile distilled water. Aliquots of 1 ml from this decolorization assay sample were withdrawn immediately following dye addition and after dye decolorization; centrifuged at 5,500 rpm for 15 min at room temperature, and supernatants were analyzed at 525 nm on spectrophotometer. All experiments were performed in triplicates in aseptic conditions. Decolorization activity was calculated by using the formula,
Effect of physico-chemical parameters and increasing dye concentration
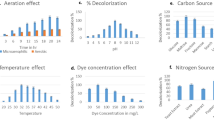
Effect of different pH was studied by adjusting the pH of distilled water (viz.—5, 6, 7, 8, and 9) with 0.1 M HCl and 0.1 M NaOH prior to cells and dye addition. Similarly, to study the effect of different temperatures (from 10 °C to 50 °C) on decolorization performance, flasks containing cells suspended in distilled water were kept at respective temperatures for 15 min before dye addition to attain the temperature. After dye addition, incubation was continued at respective temperature. Decolorization was monitored after 6 h of incubation.
In order to examine the effect of initial dye concentrations on the decolorization performance, different concentrations of dye—100, 200, 300, 400, and 500 mg l−1 were added into the decolorization flask containing 1 g 1−1 bacterial cells in distilled water. The percent decolorization was measured at different time intervals.
Effect of cell mass concentrations
We also scrutinized the impact of cell mass concentration on decolorization rate. Experiments were carried out at cell concentration viz.—0.5, 1.0, 1.5, 2.0, and 2.5 g l−1. Flasks were incubated at 30 °C, and decolorization was monitored at various time intervals.
Effect of heavy metal ions
To examine the impact of various metals such as Mg (MgCl2), Co (CoCl2), Zn (ZnSO4), Mn (MnCl2), and Hg (HgCl2) on decolorization activity, decolorization experiments were carried out in the presence of these metals at concentration 1 and 5 mM for all metals. Metals from stock solutions were added to the cell suspension (in distilled water) separately and incubated for 15 min, and then flasks were added with the dye aliquot. Decolorization was monitored at 30 °C for different time intervals.
Enzymatic analysis
Preparation of cell free extract
Bacterial cells grown for 24 h, as mentioned above, were centrifuged at 6,000 rpm for 10 min at 4 °C (control). Centrifuged cells (from 100 ml broth) were washed and resuspended in 20 ml potassium phosphate buffer (50 mM, pH 7.2). Further, these cells were homogenized in a glass homogenizer and sonicated (Sonics-vibracell ultrasonic processor) keeping sonicator output at 60 amplitudes and giving seven strokes each of 30 s. with 2 min interval at 4 °C. The homogenate was centrifuged at 6,000 rpm for 10 min at 4 °C, and supernatant was used as a source of crude enzyme. To quantify the enzyme activities after dye decolorization, cells were harvested from decolorization flask by centrifugation, and similar procedure was followed to get the cell-free extract as an enzyme source.
Enzyme assays
Activities of laccase, veratryl alcohol oxidase, NADH-DCIP reductase, and tyrosinase were assayed spectrophotometrically by using Shimadzu UV–vis spectrophotometer (UV 1800). Enzyme assays for laccase, veratryl alcohol oxidase, and NADH-DCIP reductase were carried out as mentioned in our previous report (Jadhav et al. 2011). For tyrosinase assay, protocol mentioned by Surwase and Jadhav (2012) was followed. All enzyme assays were carried out at room temperature; reference blanks contained all the components except enzyme.
Continuous dye decolorization in up-flow reactor
In this study, bacterial ability to carry out dye decolorization in continuous process was evaluated with Amaranth. For this, cells of P. aeruginosa BCH were immobilized in Na-alginate. Cells were centrifuged after growing for 24 h as mentioned above, and 180 mg cells were mixed with 4 % Na-alginate (100 ml). Beads of immobilized cells were formed in 2 % CaCl2 solution. These beads (3–4 mm diameter) were packed in a glass column of 2.8 cm inner diameter and 35 cm length. The packed volume of reactor was 170 ml in 30 cm packed section. Reactor was operated at 20 ml h−1 flow rate with the help of peristaltic pump, and decolorization was monitored at various time intervals. Nutrients were supplied after 6 days of incubation.
Analytical analysis
Extraction of the metabolites
Decolorized sample was centrifuged at 9,000 rpm for 20 min at room temperature, and the supernatant obtained was used to extract metabolites; firstly with equal volume of ethyl acetate and then with equal volume dichloromethane. Both extracts were combined, dried over anhydrous Na2SO4, and evaporated to dryness in rotary evaporator. This dried residue was redissolved in small volume of HPLC grade methanol and used for analytical studies like HPTLC, HPLC, and GC-MS.
UV–vis spectral analysis, HPTLC, HPLC, FTIR, and GC-MS
UV–visible spectral analysis of cell free sample, before and after dye decolorization, was carried by using Shimadzu UV spectrophotometer (UV 1800) and changes in its absorption spectrum in the visible range (400–800 nm) were recorded. High performance thin layer chromatography (HPTLC) analysis was carried out by using HPTLC system (CAMAG, Switzerland). Samples of dye and its biodegradation metabolites (dissolved in HPLC grade methanol) were loaded on pre-coated HPTLC plates (Silica gel 60F 254, Merck, Germany), by using nitrogen as a spraying gas and TLC sample loading instrument (CAMAG LINOMAT 5). TLC plate was developed in solvent system toluene: ethyl acetate: methanol (2:4:13). After development, the plate was observed in UV chamber (CAMAG) and scanned at 254 nm with slit dimension 5 × 0.45 mm by using TLC scanner (CAMAG). Results were generated by using HPTLC software WinCATS 1.4.4.6337. HPLC analysis was performed in an isocratic Waters 2690 system equipped with dual absorbance detector, using C18 column (symmetry 4.6 × 250 mm) and HPLC grade methanol as mobile phase with flow rate 1 ml min−1. For FTIR analysis, residue obtained after evaporation of solvent extracts was mixed with stereoscopically pure KBr, and analysis was carried out using Simadzu 8400S spectrophotometer in the mid-infrared region of 400–4000 cm−1 with 16-scan speed. Identification of metabolites formed after biodegradation of Amaranth was carried out using a QP2010 gas chromatography coupled with mass spectroscopy (Shimadzu). The ionization voltage was 70 eV and helium was used as carrier gas with a flow rate 1 ml min−1 and 33 min run time. Gas chromatography was conducted in temperature programming mode with a Resteck column (0.25 × 30 mm; XTI-5). Column oven temperature was 280 °C, and injection temperature was 200 °C. Temperature was held at 50 °C for 1 min then rose to 280 °C at 10 °C rise per minute.
Toxicity evaluation of dye and its metabolites
Toxicity of Amaranth and its metabolites obtained after dye biodegradation was investigated with Allium cepa L. Generation of oxidative stress was studied which included the antioxidant enzyme status, lipid peroxidation, and protein oxidation.
Treatment
Small bulbs of A. cepa of equal size and shape were initially exposed to water to develop the roots. Bulbs were then grouped into three sets as I, II, and III for the control (distilled water treatment), treatment with dye (1,000 ppm), and metabolites obtained after dye biodegradation (1,000 ppm), respectively (Jadhav et al. 2010b). Bulbs in each case were exposed to respective treatment for 72 h.
Analysis of antioxidant enzymes, protein oxidation, and lipid peroxidation
For investigating the antioxidant enzyme status and levels of protein oxidation and lipid peroxidation, protocol of Achary et al. (2008) was followed. Antioxidant enzyme status was assessed by both spectrophotometric as well as in-gel enzyme assays. Antioxidant enzymes that were analyzed include (CAT, E.C.1.11.1.6), SOD (E.C.1.15.1.1), and GPX (E.C.1.11.1.7). Spectrophotometric assays were initiated by adding aliquots of enzyme extracts containing 50 μg of protein to 3 ml of reaction mixture. In case of in-gel enzyme assays, protein extracts containing equal amounts of soluble protein (50 μg) were subjected to native discontinuous polyacrylamide gel electrophoresis under non-reducing and non-denaturing conditions and stained by activity staining for respective enzymes.
Protein oxidation assay was based on the reaction of carbonyls resulting from free radical-mediated modification of proteins and 2, 4-dinitrophenyl hydrazine (DNPH, Qualigens, India). Difference in absorbance of test and blank was drawn on to calculate the carbonyl concentrations using extinction coefficient of DNPH (ε = 22 mM−1 cm−1) and expressed in nanomoles of DNPH incorporated per milligram of protein. In lipid peroxidation assay, amount of malondialdehyde (MDA) was measured which is produced from thiobarbituric acid (TBA) reaction. The concentration of MDA was calculated using an extinction coefficient (ε = 155 mM−1 cm−1) and expressed in nanomole gram FW. For all enzymatic studies, protein concentrations were estimated by the Lowry method (1951).
Statistical analysis
Data were analyzed by one-way analysis of variance (ANOVA) with Tukey–Kramer multiple comparisons test.
Results
Dye decolorization at various physicochemical conditions
Bacterium P. aeruginosa BCH had shown the ability to decolorize Amaranth in plain distilled water. Dye decolorization was confirmed through UV–visible spectral analysis which showed vanishing of the peak responsible for λmax of Amaranth after decolorization (Fig. not shown). Optimum decolorization was achieved after examining the effects of various pH and temperature values. Highest decolorization of 97 % was recorded at pH 7 and temperature at 30 °C (Fig. 1). However, we observe that there was no difference in optimum decolorization activity without adjusting the pH to 7 of distilled water (which was 6.7 ± 0.1). Therefore, further studies were carried out in plain distilled water. Dye decolorization was not occurred at shaking condition (120 rpm). It was noteworthy that heat-killed cell suspension did not show decolorization activity and also no biosorption of the dye was occurred by dead cell mass. However, adsorption of dye was observed only initially (0 time) and not after decolorization. Cell viability was checked in decolorized flask by streaking loop full culture on agar plate, and normal growth pattern was observed which signified that cells remain viable.
Decolorization activity at higher dye concentrations
Ability of bacterium to tolerate higher dye concentration was assessed in this study, and it was found that strain could show the activity even at 500 mg l-1 dye concentration. Exposure of bacterial cells to concentrations above 100 mg l−1 resulted in a decrease in color removal efficiency. More than 95 % color removal at dye concentrations below 100 mg l−1 was obtained within or inside 6 h, whereas increase in dye concentration up to 500 mg l−1 showed a significant decrease in decolorization level to 87 % within 39 h (Fig. 2).
Decolorization using various cell mass concentrations
Assessment of cell mass effect was done by carrying out decolorization experiments under different cell mass concentrations. Linear type of relationship was obtained where time required for decolorization went on decreasing with increasing cell mass, that is, decolorization rate was increased with increasing cell mass concentration (Fig. 3). From this study, it was evident that cell mass concentration was the strong process affecting parameter.
Effect of heavy metals on decolorization performance
Dye containing effluents frequently contain heavy metals. Therefore, Amaranth decolorization was studied in presence of some metals at concentrations 1 and 5 mM. Figure 4 reflects the effects of Mn, Zn, Co, Mg, and Hg. It can be seen that Zn strongly inhibited the decolorization efficacy at 5 mM concentration resulting in only 30 % decolorization by P. aeruginosa BCH. It also inhibited the decolorization performance at 1 mM concentration but to a lesser extent (decolorization 81 %). Similarly, Hg too showed strong inhibition activity. It was notable that Mn had strong enhancing effect on decolorization performance where 98 % decolorization was attained with both tested concentrations. But with 5 mM concentration, same activity was observed at very high decolorization rate (within 105 min of incubation). Mg exhibited negligible effect while the presence Co was inhibitory to Amaranth decolorization.
Enzymatic status
Data presented in Table 1 shows the intracellular and extracellular enzyme activities before and after decolorization of Amaranth by P. aeruginosa BCH. It is evident that activity of intracellular laccase was induced significantly from 0.665 to 0.812 U. Activity of intracellular veratryl alcohol oxidase was remarkably induced from 1.086 to 1.256 U after dye decolorization. Activities of tyrosinase and NADH-DCIP reductase were induced significantly in both extracellular and intracellular samples.
Decolorization in up-flow bioreactor
Decolorization was monitored for 18 days at a flow rate of 20 ml h−1 with nutrient supply after 6 days of incubation in a continuous process. Figure 5a gives schematic representation of reactor and shows the decolorization pattern observed after every 12 h of run (Fig. 5b). It can be seen that immobilized cells decolorized the dye for the first 2 days without change in efficiency where 97 % decolorization was observed. Bioreactor was then slightly dropping down in its efficiency from the third day, and at sixth day, 71 % decolorization was recorded. Further on supply of nutrients decolorization activity was regained by the reactor with slightly low efficacy. Decolorization pattern observed in this second round/cycle was from 90 % to 61 % on the sixth day after nutrient supply. This decline in reduction of decolorization competence continued in the third cycle. Here, only 51 % decolorization occurred at the end of sixth day. Cell viability may be the reason for this decreasing efficiency. Abiotic decolorization was calculated at the end of the experiment with cell-free beads, and it was 5 % which served as the control.
Analytical confirmation of dye biodegradation
HPLC, HPTLC, and FTIR analyses
Figure 6 indicates the results of HPLC analysis. It showed single peak for Amaranth at retention time of 1.784 min. (Fig. 6a), whereas sample obtained after decolorization showed four peaks with altered retention times viz.—1.971, 2.483, 2.718, and 3.274 min (Fig. 6b).
Figure 7a shows three dimensional graph of Rf values versus absorbance units. Control sample (dye) ran on fluorescent HPTLC plate showed peak at Rf value of 0.76, whereas metabolite sample showed two different spots with altered Rf values viz.—0.72 and 0.90 (Fig. 7b).
When the dye was analyzed in FTIR analysis (Fig. 8a), several peaks responsible for different functional groups were obtained as follows. Peak at 638.70 cm−1 corresponds to stretching in terminal monosubstituted C–H deformation. Peak at 667.04 cm−1 corresponded to C–S stretching in sulphonyl group. C–H deformation in benzene ring was indicated by peak at 836.59 cm−1. S=O asymmetric stretching was evidenced by the presence of peak at 1,035.52 and 1,194.80 cm−1. While the peak at 1,374.94 cm−1 was in agreement with presence of phenolic −OH group indicating O–H deformation. Aromatic homocyclic compound was matched up by C=C stretching. Peak at 1,628.67 cm−1 confirms the azo nature of the dye indicating N=N stretching. 3,430.18 cm−1 was indicating the O–H stretching in free phenolic OH group.
Analysis of metabolites showed completely different spectrum (Fig. 8b). It showed peak at 698.36 cm−1 which corresponded to the C–S stretching, suggesting that one of the metabolites or intermediates may contain the sulphonyl group. Peak at 1,411.07 cm−1 was indicative of –C–O stretching plus O–H deformation in –COOH group. Peak value of 1,475.82 cm−1 denoted the aromatic homocyclic C=C stretching. Aryl carboxylic acid was suggested by peak at 1,699.85 cm−1. The values 2,854.20 and 2,925.00 cm−1 corresponded to O–H stretching in bonded OH.
GC-MS and possible pathway
Three compounds having m/z values 258, 349, and 174 were detected in GC-MS and are depicted in Table 2 with their retention times obtained in GC spectrum. Based on these results, biodegradation pathway for Amaranth was proposed. First step in this dye biodegradation pathway was predicted as asymmetric cleavage by oxidoreductive enzymes (Fig. 9). This asymmetric cleavage resulted in two compounds which showed the m/z ratios as 258 and 349 and were identified as sodium-4-diazenyl-5, 8-dihydronaphthalene-1-sulhponate [1] (MW 260) and other one as disodium 3-hydroxynaphthalene-2, 7-disulphonate [2] (MW 348), respectively. Compound [1] further undergoes the oxidation reaction by laccase to yield an unidentified product. Further on, desulphonation of the 5-nitro-1, 4-dihydronaphthalene (MW 174) was produced which was detected in GC-MS analysis having m/z ratio of 176.
Toxicological appraisal before and after dye biodegradation
We herein analyzed various antioxidant enzymes—SOD, CAT, and GPX by spectrophotometric as well as in-gel enzyme assays in A. cepa root cells. Roots exposed to dye showed significant induction in the activity of SOD from 03.22 (control) to 09.75 U (Table 3). Likewise, GPX, the enzyme which represents the activity of overall peroxidase, was induced from 27.73 to 39.82 U. In contrast, CAT activity was reduced in dye-treated sample from 48.62 to 32.31 U. Activities of these enzymes obtained in A. cepa roots treated with metabolites formed after bacterial treatment of dye, were found to be 5.42, 35.09 and 49.98 U, respectively, for SOD, GPX, and CAT. As it is evident from Table 3, these values were very close to the control values. Figure 10 indicates the zymographs (activity staining with respective substrates) of these enzymes which are in agreement with results obtained in spectrophotometric assays. Subsequently, we evaluated the levels of lipid peroxidation and protein oxidation in same treatments (Table 3). Formation of MDA was 0.39 nmol g−1 FW in control treatment and 0.71 nmol g−1 FW in dye-treated sample, suggesting that there was increased lipid peroxidation due to dye. In the same way, carbonyl content in control sample was found to be 4.01 nmol mg−1 protein which singnificantly raised to 9.63 nmol mg−1 protein after dye treatment which demonstrated the damage to the proteins by Amaranth. It is important to note that values of lipid peroxidation and protein oxidation in metabolite-treated samples were 0.43 (MDA formed nmol g−1 FW) and 5.05 (nmol mg−1 protein carbonyl content), respectively, which are very close to that of control values.
Discussion
P. aeruginosa BCH showed the ability to decolorize xenobiotic azo dye Amaranth in the absence of organic or inorganic nutrients. These are significant findings with respect to that there is no report on biodegradation of such xenobiotic compounds by a bacterial species. Up till now, all studies regarding industrial dyes have been carried out in the presence of organic and inorganic growth factors with exception of reports on yeast species Saccharomyces cerevisiae MTCC 463 (Jadhav and Govindwar 2006; Jadhav et al. 2007). In that study, dye decolorization occurred through biosorption by yeast cells subsidiary to biotransformation. Results of the present study suggest that decolorization was mediated by bacterial biodegradation activity only; which was confirmed by experiments with heat-killed cell suspension, cell viability and enzymatic studies. Therefore, it can be said that different mode of dye decolorization was observed in present study from the earlier report of dye biodegradation in plain distilled water by yeast species (Jadhav et al. 2007). Our studies suggested that decolorization of Amaranth by P. aeruginosa BCH was a pH and temperature-dependent phenomenon like that of several reports in broth studies. However, bacterium showed decolorization activity in short pH and temperature ranges of 5–8 and 20–40 °C, respectively. Maximum initial dye concentration tolerated was 500 mg l−1 but as concentration goes to higher levels, the time required for decolorization was also increased. Besides this, reduction in decolorization rate also occurred. Decreased color removal at higher dye concentrations may be attributed to the inhibition of bioremediation enzymes and cellular metabolic activities due to increased toxic effect of dye at such high concentrations (Jadhav et al. 2011). Another possible reason might be the sticking or adhesion of dye molecules on microbial cell wall at higher dye concentrations, rendering the mass transfer difficult or slow. Sani and Banerjee (1999) have demonstrated that at higher dye concentrations, color removal efficiency of Kurthia sp. was not enough to degrade all the dye transferred through bacterial cell membrane.
In the next study, it was observed that higher cell mass concentration accelerated the decolorization rate of bacterium. It suggests that when cells were present in higher numbers, less time was required to utilize the available dye molecules as compared to the time required when cells were present in lower concentration. Although numbers of metals are essential for growth, some can be harmful for living cells. This is mainly due to the fact that heavy metals form complexes with protein molecules which render them inactive, for example, inactivation of enzymes. Many heavy metals are detrimental to microorganisms even at low concentrations present in natural waters. When decolorization performance was checked in the presence of different heavy metals, it was observed that their presence can alter the decolorization activity for Amaranth. Only Mg did not show the significant influence. However, MgCl2 has been used as a coagulant to decolorize the dye solutions (Tan et al. 2000; Gao et al. 2007). Therefore, we tested decolorization of Amaranth with MgCl2 in the absence of bacterial cells. We found that the dye was not decolorized in the pH range 5–9 (used in our study), suggesting that there was no influence of this coagulant in Amaranth decolorization, but we got decolorization at pH 12. These findings support the previous studies (Tan et al. 2000; Gao et al. 2007) and also point out the fact that higher basic pH values are required for MgCl2 to act as a coagulant for azo dye removal purpose.
In order to check the ability of bacterial strain to carry out dye decolorization continuously and to assess the potential of strain for possible real-scale application, we constructed an up-flow reactor. Gel-entrapment method was used for immobilization of bacterial cells. This is the most widely used method of immobilization of whole cells, and a suitable matrix material is alginate because it is non-toxic (Enayatzamir et al. 2010). When decolorization was observed at different flow rates (10 to 50 ml h−1), higher flow rates resulted in lower dye decolorization efficiency. Hence, we carried out the further studies using 20 ml h−1. Experimental results suggested that P. aeruginosa BCH strain has the potential to work in a continuous process for longer time periods as well and can offer more practical approach for the removal of Amaranth.
Biodegradation of Amaranth by P. aeruginosa BCH was confirmed through various analytical studies before and after dye decolorization. Single peak or band was observed in HPLC and HPTLC analysis for Amaranth, whereas metabolite sample showed almost complete disappearance of this single peak, and there was appearance of multiple peaks and bands with altered retention times and retention factors. FTIR analysis also showed different patterns of spectrum for control dye and metabolites. One important thing to be noted is that the peak at 1,628.67 cm−1 which corresponded to the azo group in control dye was not detected in metabolites sample. It puts forward that azo group was removed after bacterial treatment. All these analytical analyses advocated that the decolorization was due to structural degradation of Amaranth. Therefore, GC-MS analysis was carried out to identify the metabolites formed after dye biodegradation.
As suggested in several reports, various oxidoreductive enzymes attack the dye molecule to cleave it symmetrically or asymmetrically, and subsequent cleavage of produced intermediates took place. Enzymatic analysis in our study indicated that laccase, veratryl alcohol oxidase, tyrosinase, and NADH-DCIP reductase were involved in Amaranth decolorization by P. aeruginosa BCH. Activity of azoreductase was found to be absent. In contrast, during biodegradation of methyl red in plain distilled water by yeast species S. cerevisiae, extracellular azoreductase played prominent role (Jadhav et al. 2007). These enzymatic studies showed that dye was degraded by the contribution of intracellular as well as extracellular oxidoreductive enzymes. Jadhav et al. (2009) have recently showed the ability of bacterial veratryl alcohol oxidase to degrade textile dyes. In addition, purified laccase have been shown to degrade the textile dyes (Zucca et al. 2011). Tyrosinases are also having shown to degrade the variety of aromatic pollutants (Girelli et al. 2006; Montiel et al. 2004). These enzymes are vastly studied and their role and mode of action are well established. On the contrary, action of NADH-DCIP reductase on a particular pollutant structure or metabolite is not well documented. Activity of this enzyme in various microorganisms has been shown to be induced during dye biodegradation for its reductase activity (Patil et al. 2008; Phugare et al. 2010). Generally, this enzyme is known to catalyze the reduction reaction using NADH as an electron donor. Based on enzymatic analysis, FTIR, and GC-MS analyses, we predicted herein the possible metabolic pathway for Amaranth biodegradation. Because of observed induction in the activities of oxidative as well as reductive enzymes, it was difficult to predict which enzyme is responsible for this first step cleavage of dye molecule. Here, oxidation of azo bond was predicted to be carried out by laccase, because this enzyme was previously shown to degrade the azo dyes (Chivukula and Renganathan 1995). Mechanistic breakdown of azo bond without the contribution of azoreductase was also disclosed by Pereira et al. (2009). Similar action of laccase could be seen in the present biodegradation pathway.
To detoxify pollutants is the main aim at which bioremediation processes are directed. Therefore, it becomes necessary for any bioremediation technique to verify it through toxicological analysis. We selected A. cepa plant species which is a part of human diet. Oxidative stress is the toxicological parameter that we have tested in A. cepa roots. Toxicity of Amaranth has been assessed previously with Ames test for mutagenicity (Hong et al. 2007). Hence, we choose to study the role of Amaranth in generation of oxidative stress and its release after bacterial treatment. Oxidative stress is generally produced when a plant experiences stress conditions. It is usually mediated by the production of highly toxic chemically reactive molecules called as free radicals or reactive oxygen species (ROS) which includes superoxide anion (O −2 ), hydrogen peroxide (H2O2), and hydroxyl radical (OH). These are formed from the partial reduction metabolism of oxygen as byproducts. However, plants do possess the mechanisms to overcome such stress conditions. Protection of the cells from such free radicals mediated damage is done by scavenging these ROS by antioxidants or free radical scavenging enzymes. These antioxidants include lactoperoxidases, SOD, CAT, APX, GPX, glutathione reductase, and peroxiredoxins. Such oxidative stress can be monitored by analyzing up- or down-regulation of these antioxidant enzymes (Achary et al. 2008). Our study results showed induction in activities of SOD, GPX, and inhibition of CAT activity, which suggest that there was formation of free radicals in dye-treated samples. A similar pattern of up/down-regulation of these enzymes was observed as compared to previous reports on oxidative stress studies with textile dye Red HE3B (Phugare et al. 2010) and Al-mediated toxicity (Achary et al. 2008). Free radicals are capable of altering all major classes of biomolecules, such as lipids, nucleic acids, and proteins, with changes in their structure and function. And hence, the free radical-mediated oxidative stress is usually accompanied by lipid peroxidation and protein oxidation. In our study, it was observed that damage to these biomolecules was significantly higher in Amaranth-treated samples. However, these levels were significantly reduced and were close to the control values in the case of metabolites. Anbazhagan and Chellappan (2009) pointed out that prime targets of free radicals are the polyunsaturated fatty acids in cell membranes, and their interaction results in lipid peroxidation. It has been reported that free radical-mediated injury to proteins may be initiated by electron leakage, metal-ion-dependent reactions, and autoxidation of lipids and sugars (Dean et al. 1997). Overall, this toxicity analysis suggested that Amaranth was responsible for generating oxidative stress in A. cepa, and it was unimpeded after the biodegradation activity by P. aeruginosa BCH.
Concluding remarks
Bacterium P. aeruginosa BCH possesses the remarkable potential for biodegradation of azo dye Amaranth in the absence of any kinds of nutrients where decolorization was achieved through biodegradation activity. Strain also bears the capacity to degrade higher dye concentrations. Presence of Mn ions can accelerate the decolorization process so as the higher biomass. Laccase, veratryl alcohol oxidase, tyrosinase, and NADH–DCIP reductase found to have the role in Amaranth biodegradation. Up-flow reactor studies signified that the potential of this strain could be explored for real scale application. Overall toxicity studies with A. cepa roots pointed out that Amaranth was responsible for generating the oxidative stress. However, after bacterial treatment, this stress was found to be released which demonstrated the capability of strain to reduce toxic effects of dye. This superior aptitude of P. aeruginosa BCH can offer feasible approach for bioremediation of xenobiotic azo dye.
References
Achary VM, Jena S, Panda KK, Panda BB (2008) Aluminium induced oxidative stress and DNA damage in root cells of Allium cepa L. Ecotoxicol Environ Saf 70:300–310. doi:10.1016/j.ecoenv.2007.10.022
Anbazhagan M, Chellappan PR (2009) Activities of antioxidant enzyme and lipid peroxidation in ovarian cancer patients. Acad J Cancer Res 2:68–72
Chequer FMD, Angeli F, Ferraz ERA, Tsuboy MS, Marcarini JC, Mantovani MS, Oliveira DP (2009) The azo dyes Disperse Red 1 and Disperse Orange 1 increase the micronuclei frequencies in human lymphocytes and in HepG2 cells. Mutat Res 676:83–86. doi:10.1016/j.mrgentox.2009.04.004
Chivukula M, Renganathan V (1995) Phenolic azo dye oxidation by laccase from Pyricularia oryzae. Appl Environ Microbiol 61:4374–4377
Davis RJ, Gainer JL, O’Neal G, Wu IW (1994) Photocatalytic decolorization of waste water dyes. Water Environ Res 66:50–53, http://dx.doi.org/10.2175/WER.66.1.8
Dean RT, Shanlin FU, Stocker R, Davies MJ (1997) Biochemistry and pathology of radical-mediated protein oxidation. Biochem J 324:1–18
Enayatzamir K, Alikhani HA, Yakhchali B, Tabandehi F, Rodríguez-Couto S (2010) Decolouration of azo dyes by Phanerochaete chrysosporium immobilised into alginate beads. Environ Sci Pollut Res 17:145–153. doi:10.1007/s11356-009-0109-5
Ferraz ERA, Grando MD, Oliveira DP (2011a) The azo dye Disperse Orange 1 induces DNA damage and cytotoxic effects but does not cause ecotoxic effects in Daphnia similis and Vibrio fischeri. J Hazard Mater 192:628–633. doi:10.1016/j.jhazmat.2011.05.063
Ferraz ERA, Umbuzeiro GAG, Caloto-Oliveira A, Chequer FMD, Zanoni MVB, Dorta DJ, Oliveira DP (2011b) Differential toxicity of Disperse Red 1 and Disperse Red 13 in the Ames test, HepG2 cytotoxicity assay, and Daphnia acute toxicity test. Environ Toxicol 26:489–497. doi:10.1002/tox.20576
Gao BY, Yue QY, Wang Y, Zhou WZ (2007) Color removal from dye-containing wastewater by magnesium chloride. J Environ Manage 82:167–172. doi:10.1016/j.jenvman.2005.12.019
Girelli AM, Mattei E, Messina A (2006) Phenols removal by immobilized tyrosinase reactor in on-line high performance liquid chromatography. Anal Chim Acta 580:271–277. doi:10.1016/j.aca.2006.07.088
Gomi N, Yoshida S, Matsumoto K, Okudomi M, Konno H, Hisabori T, Sugano Y (2011) Degradation of the synthetic dye Amaranth by the fungus Bjerkandera adusta Dec 1: inference of the degradation pathway from an analysis of decolorized products. Biodegradation 22:1239–1245. doi:10.1007/s10532-011-9478-9
Hong Y, Guo J, Xu Z, Mo C, Xu M, Sun G (2007) Reduction and partial degradation mechanisms of naphthylaminesulfonic azo dye Amaranth by Shewanella decolorationis S12. Appl Microbiol Biotechnol 75:647–654. doi:10.1007/s00253-007-0838-7
Isik M, Sponza DT (2003) Effect of oxygen on decolorization of azo dyes by Escherichia coli and Pseudomonas sp. and fate of aromatic amines. Process Biochem 38:1183–1192
Jadhav JP, Govindwar SP (2006) Biotransformation of malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323. doi:10.1002/yea.1356
Jadhav JP, Parshetti GK, Kalme SD, Govindwar SP (2007) Decolourization of azo dye methyl red by Saccharomyces cerevisiae MTCC 463. Chemosphere 68:394–400. doi:10.1016/j.chemosphere.2006.12.087
Jadhav UU, Dawkar VV, Tamboli DP, Govindwar SP (2009) Purification and characterization of veratryl alcohol oxidase from Comamonas sp. UVS and its role in decolorization of textile dyes. Biotechnol Bioprocess Eng 14:369–376. doi:10.1007/s12257-008-0300-4
Jadhav JP, Kalyani DC, Phugare SS, Govindwar SP (2010a) Evaluation of the efficacy of a bacterial consortium for the removal of color, reduction of heavy metals, and toxicity from textile dye effluent. Bioresour Technol 101:165–173. doi:10.1016/j.biortech.2009.08.027
Jadhav JP, Phugare SS, Dhanve RS, Jadhav SB (2010b) Rapid biodegradation and decolorization of direct orange 39 (Orange TGLL) by an isolated bacterium Pseudomonas aeruginosa strain BCH. Biodegradation 21:453–463. doi:10.1007/s10532-009-9315-6
Jadhav SB, Phugare SS, Patil PS, Jadhav JP (2011) Biochemical degradation pathway of textile dye Remazol red and subsequent toxicological evaluation by cytotoxicity, genotoxicity and oxidative stress studies. Int Biodeterior Biodegrad 65:733–743. doi:10.1016/j.ibiod.2011.04.003
Jaochim F, Burrel A, Anderson J (1985) Mutagenicity of azo dyes in Salmonella/microsome assay using in-vitro and in-vivo activation. Mutat Res 156:131–138. doi:10.1016/j.bbr.2011.03.031
Johnson RF, Zenhausen A, Zollinger H (1978) Azo dyes. pp. 868–869. In: Mark HF, Mc ketta JJ, Othmer DF, Standen A (eds) Krik-othmer encyclopedia of chemical technol., vol. 2, 2nd edn. John Wiley, New York
Lee YH, Pavlostathis SG (2004) Decolorization and toxicity of reactive anthraquinone textile dyes under methanogenic conditions. Water Res 38:1838–1852. doi:10.1016/j.watres.2003.12
Lowry O, Rosbrough N, Farr A, Randall R (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275
Mok Y, Jo J (2007) Degradation of a textile azo dye by pulsed arc discharge to the surface of wastewater. Korean J Chem Eng 24:607–611. doi:10.1007/s11814-007-0011-0
Montiel AM, Fern'andez FJ, Marcial J, Soriano J, Barrios-Gonz’alez J, Tomasini A (2004) A fungal phenoloxidase (tyrosinase) involved in pentachlorophenol degradation. Biotechnol Lett 26:1353–1357. doi:10.1023/B:BILE.0000045632.36401.86
Pan H, Feng J, He G, Cerniglia C, Chen H (2012) Evaluation of impact of exposure of Sudan azo dyes and their metabolites on human intestinal bacteria. Anaerobe. doi:10.1016/j.anaerobe.2012.05.002
Patil PS, Shedbalkar UU, Kalyani DC, Jadhav JP (2008) Biodegradation of Reactive Blue 59 by isolated bacterial consortium PMB11. J Ind Microbiol Biotechnol 35:1181–1190. doi:10.1007/s10295-008-0398-6
Pereira L, Coelho AV, Viegas CA, Santos MM, Robalo MP, Martins LO (2009) Enzymatic biotransformation of the azo dye Sudan Orange G with bacterial CotA-laccase. J Biotechnol 139:68–77. doi:10.1016/j.jbiotec.2008.09.001
Phugare SS, Kalyani DC, Patil AV, Jadhav JP (2010) Textile dye degradation by bacterial consortium and subsequent toxicological analysis of dye and dye metabolites using cytotoxicity, genotoxicity and oxidative stress studies. J Hazard Mater 186:713–723. doi:10.1016/j.jhazmat.2010.11.049
Pierce J (1994) Colour in textile effluents—the origins of the problem. J Soc Dyers Colourists 110:131–133. doi:10.1111/j.1478-4408.1994.tb01624.x
Puvaneswari N, Muthukrishnan J, Gunasekaran P (2006) Toxicity assessment and microbial degradation of azo dyes. Indian J Exp Biol 44:618–626
Ramsay JA, Nguyen T (2002) Decoloration of textile dyes by Trametes versicolor and its effect on dye toxicity. Biotechnol Lett 24:1757–1761. doi:10.1023/A:1020644817514
Sani RK, Banerjee UC (1999) Decolorization of triphenylmethane dyes and textile and dye-stuff effluent by Kurthia sp. Enzyme Microb Technol 24:433–437
Surwase SN, Jadhav JP (2012) Efficient microbial conversion of l-tyrosine to l-DOPA by Brevundimonas sp. SGJ. Appl Biochem Biotechnol. doi:10.1007/s12010-012-9564-4
Tan BH, Ten TT, Omar AK (2000) Removal of dyes and industrial dye wastes by magnesium chloride. Water Res 34:597–601. doi:10.1016/S0043-1354(99)00151-7
Zucca P, Rescigno A, Olianas A, Maccioni S, Sollai FA, Sanjust E (2011) Induction, purification, and characterization of a laccase isozyme from Pleurotus sajor-caju and the potential in decolorization of textile dyes. J Mol Catal B Enzym 68:216–222. doi:10.1016/j.molcatb.2010.11.008
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Robert Duran
Rights and permissions
About this article
Cite this article
Jadhav, S.B., Patil, N.S., Watharkar, A.D. et al. Batch and continuous biodegradation of Amaranth in plain distilled water by P. aeruginosa BCH and toxicological scrutiny using oxidative stress studies. Environ Sci Pollut Res 20, 2854–2866 (2013). https://doi.org/10.1007/s11356-012-1155-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-012-1155-y