Abstract
Class III peroxidases are members of a large plant-specific sequence-heterogeneous protein family. Several sequence-conserved homologs have been associated with lignin polymerization in Arabidopsis thaliana, Oryza sativa, Nicotiana tabacum, Zinnia elegans, Picea abies, and Pinus sylvestris. In Populus trichocarpa, a model species for studies of wood formation, the peroxidases involved in lignin biosynthesis have not yet been identified. To do this, we retrieved sequences of all PtrPOs from Peroxibase and conducted RNA-seq to identify candidates. Transcripts from 42 PtrPOs were detected in stem differentiating xylem (SDX) and four of them are the most xylem-abundant (PtrPO12, PtrPO21, PtrPO42, and PtrPO64). PtrPO21 shows xylem-specific expression similar to that of genes encoding the monolignol biosynthetic enzymes. Using protein cleavage-isotope dilution mass spectrometry, PtrPO21 is detected only in the cell wall fraction and not in the soluble fraction. Downregulated transgenics of PtrPO21 have a lignin reduction of ~20 % with subunit composition (S/G ratio) similar to wild type. The transgenics show a growth reduction and reddish color of stem wood. The modulus of elasticity (MOE) of the stems of the downregulated PtrPO21-line 8 can be reduced to ~60 % of wild type. Differentially expressed gene (DEG) analysis of PtrPO21 downregulated transgenics identified a significant overexpression of PtPrx35, suggesting a compensatory effect within the peroxidase family. No significant changes in the expression of the 49 P. trichocarpa laccases (PtrLACs) were observed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The evolution of lignin, a highly abundant plant phenolic polymer, enabled vascular plants to dominate terrestrial ecosystems (Lockhart 2013). Lignin provides mechanical strength, hydrophobicity, and pathogen resistance to secondary plant cell walls and limits the natural decay of plant tissue (Fry and White 1938; Sarkanen and Ludwig 1972; Vance et al. 1980; Koch et al. 2004). However, the deposition of lignin in plant cell walls limits the utilization of plant biomass, therefore requiring harsh chemical treatment for delignification (Novaes et al. 2010). Decreasing lignin content or modifying lignin structure can reduce the recalcitrance of biomass and increase the yield of extractable cellulose and fermentable sugars for pulp and paper or biofuel production (Hu et al. 1999; Eckardt 2002; Li et al. 2003a; Chen and Dixon 2007; Studer et al. 2011).
Lignin is principally polymerized from three monolignols (4-coumaryl, coniferyl, and sinapyl alcohols), which become monomeric subunits in lignin known as 4-hydroxyphenyl (H), guaiacyl (G), and syringyl (S) subunits, respectively (Higuchi 1997). In angiosperms, such as Populus trichocarpa, lignin is composed mainly of S and G subunits (S/G ratio is ~2) with a trace of H subunits. Combinatorial radical coupling of monolignols catalyzed by peroxidases and laccases is widely accepted as the current model for lignin polymerization (Erdtman 1933; Freudenberg 1959; Freudenberg 1965; Harkin and Obst 1973; Adler 1977; Higuchi 1985; Sterjiades et al. 1992; O’Malley et al. 1993; Dean and Eriksson 1994; Takahama 1995; Richardson et al. 1997; Ros Barcelo 1997; Hatfield and Vermerris 2001; Ralph et al. 2004; Weng et al. 2008; Zhao et al. 2013).
Plant peroxidases are heme-containing oxidases (Koua et al. 2009). Higher plants contain class I (EC 1.11.1.11) and class III (EC 1.11.1.7) peroxidases. Class I peroxidases are intracellular peroxidases, such as ascorbate peroxidase (AP), which removes hydrogen peroxides in the chloroplasts and the cytosol (Sharma and Dubey 2004). Class III peroxidases are plant-specific peroxidases, which can be secreted into the plant cell wall. They are not found in unicellular green algae, which do not produce lignin (Passardi et al. 2004).
Class III peroxidases participate in many cellular processes, such as removal of hydrogen peroxides, oxidation of toxic compounds, suberization, plant hormone metabolism, salt tolerance, senescence, pathogen resistance, wound healing, as well as cell wall biosynthesis (Lovrekovich et al. 1968; Espelie and Kolattukudy 1985; Fry 1986; Abeles et al. 1988; Campa 1991; Gazaryan et al. 1996; Lagrimini et al. 1997; Whetten et al. 1998; Amaya et al. 1999; Bernards et al. 1999; Llorente et al. 2002; Allison and Schultz 2004; Bernards et al. 2004). In plants, class III peroxidases comprise a large multigenic family resulting from multiple duplication events, suggesting functional redundancy among peroxidase genes (Hiraga et al. 2001; Duroux and Welinder 2003; Passardi et al. 2004). The class III peroxidase gene families have been described in several plant species, such as Arabidopsis thaliana (73 genes) (Ostergaard et al. 1998; Tognolli et al. 2002; Cosio and Dunand 2009), Oryza sativa (138 genes) (Passardi et al. 2004), P. trichocarpa (95 genes) (Ren et al. 2014), Physcomitrella patens (43 genes) and Selaginella moellendorffii (79 genes) (Weng and Chapple 2010).
Since 1967, peroxidases have been implicated in lignification (Nakamura 1967; Harkin and Obst 1973; Whetten et al. 1998). Lignin-associated peroxidases have been identified in Lycopersicon esculentum (tomato) (Botella et al. 1994; Quiroga et al. 2000), Phaseolus vulgaris (French bean) (Smith et al. 1994), Nicotiana tabacum (tobacco) (Lagrimini et al. 1987; Blee et al. 2003), A. thaliana (Ostergaard et al. 2000; Nielsen et al. 2001; Herrero et al. 2013b; Shigeto et al. 2013), Zinnia elegans (Masuda et al. 1983; Sato et al. 1995; Gabaldon et al. 2005; Gabaldón et al. 2006; Sato et al. 2006), poplars (Osakabe et al. 1995; Christensen et al. 1998; Tsutsumi et al. 1998; Christensen et al. 2001b; Aoyama et al. 2002; Sasaki et al. 2004; Sasaki et al. 2006; Sasaki et al. 2008), and other tree species, such as Picea abies (Norway spruce) and Pinus sylvestris (Scots pine) (Polle et al. 1994; Fagerstedt et al. 1998; McDougall 2001; Kärkönen et al. 2002; Koutaniemi et al. 2005; Marjamaa et al. 2006; Koutaniemi et al. 2007; Fagerstedt et al. 2010).
Direct functional evidence for involvement of peroxidases in lignin biosynthesis has come from studies of mutants and transgenic plants (Supplemental Table S1). Overexpression of a tomato peroxidase (TPX1) leads to a 40–220 % increase in lignin content in tomato (El Mansouri et al. 1999). Tobacco with downregulation of peroxidase TP60 had a 50 % reduction of lignin affecting both S and G subunits (Blee et al. 2003). Another TP60 downregulated tobacco with lignin content reduction of 23 % showed a reduction in the number of vessels and a striking enlargement in diameter of surrounding fibers (Kavousi et al. 2010). Downregulation of prxA3a in Populus sieboldii x Populus grandidentata reduced lignin content by 10–20 % (Li et al. 2003b). Arabidopsis mutations in peroxidases (AtPrx53, AtPrx2, AtPrx25, AtPrx71, and AtPrx72) all show significant reduction in lignin content (Ostergaard et al. 2000; Herrero et al. 2013b; Shigeto et al. 2013). However, different results from transgenic plants of other species make the role of peroxidases in lignin biosynthesis more problematic. Downregulation of the TP02 in tobacco to 40–80 % of wild type resulted in no significant change in lignin levels (McIntyre et al. 1996). Suppression of Pox25, Pox29 and Pox36 in P. sieboldii x P. grandidentata showed no significant reduction of lignin level (Tamura et al. 2001). These results are confounded by gene redundancy. More work is needed to learn the nature and extent of gene specific effects. Since 2004, a cationic cell-wall-peroxidase (CWPO-C) from Populus alba has been proposed for lignification (Sasaki et al. 2004; Sasaki et al. 2006; Sasaki et al. 2008). However, transgenics with perturbation of CWPO-C have not yet been produced and its role remains to be determined.
P. trichocarpa has been a model for studying wood formation because of its rapid growth and because its genome sequence is known (Tuskan et al. 2006). All the monolignol biosynthetic enzymes have been identified in P. trichocarpa based on xylem-specific expression (Shi et al. 2010) and most of these enzymes have been characterized in detail (Wang et al. 2014). However, the P. trichocarpa class III peroxidases (PtrPOs) for lignin biosynthesis have not yet been identified. The only peroxidase proposed for lignin biosynthesis in P. trichocarpa is PXP3-4, which showed no effect on lignin content and no phenotype when it was overexpressed 800-fold (Christensen et al. 2001a).
In this study, we aimed to identify class III peroxidases in P. trichocarpa involved in lignin biosynthesis using a more systematic search and to validate the candidates using transgenesis. Based on transcriptome analysis of different tissues, four PtrPOs (PtrPO12, PtrPO21, PtrPO42, and PtrPO64) were identified as the most xylem-abundant peroxidases. PtrPO21 showed xylem-specific expression similar to that of the genes encoding the monolignol biosynthetic enzymes. Using protein cleavage-isotope dilution mass spectrometry (PC-IDMS), RNA interference (RNAi) transgenesis, modulus of elasticity (MOE) measurements, and cell wall component analysis, PtrPO21 was identified as a cell wall-bound anionic peroxidase that may play an essential role in lignin biosynthesis.
Results
Identification of class III peroxidases in the genome of P. trichocarpa
A total of 101 P. trichocarpa class III peroxidases (PtrPOs) were retrieved from PeroxiBase (http://peroxibase.toulouse.inra.fr/) as candidates for lignin peroxidases. The list was shortened to 87 PtrPOs because 12 are pseudogenes and 2 are redundant gene records (See Supplemental Table S2). Then, we compared our list by sequence alignment to the 93 peroxidases studied by Ren et al. (2014). One extra peroxidase, PRX81, was included because it is a known functional peroxidase (Ren et al. 2014; See Supplemental Table S2). We also included PtPrx25, 28, 72 and 76 that were not included by Ren et al. (2014) (the PtPrx nomenclature for P. trichocarpa peroxidases follows Ren et al. 2014). After manual editing, a total of 88 PtrPOs were selected for this study.
PtrPO21 is a xylem-abundant and xylem-specific class III peroxidase in P. trichocarpa
To identify peroxidases functionally associated with lignin polymerization, we carried out transcriptome analysis to determine if transcripts of these 88 PtrPOs were detectable in the stem differentiating xylem (SDX) of P. trichocarpa (see “Materials and Methods”). Based on SDX RNA-seq data, 46 PtrPOs had no detectable transcripts in SDX and were excluded from further study. Of the remaining 42 PtrPOs, seven have abundant transcripts in P. trichocarpa SDX (Fig. 1). PtrPO42 accounts for 47.4 % of the total class III peroxidase transcripts, PtrPO12 for 35.1 %, PtrPO64 for 3.7 %, PtrPO21 for 3.1 %, PtPrx01 for 2.4 %, PtPrx26 for 2.2 %, and PtPrx18 for 2.0 % (Fig. 1). The remaining 35 PtrPOs represent 4.0 % of the total.
The quantification of transcript abundance of the 42 detectable class III peroxidases in the SDX of P. trichocarpa. Accession numbers of all PtrPOs are listed in Supplemental Table S2
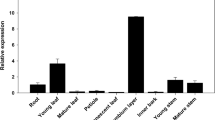
We further narrowed down the list of candidates of putative lignin peroxidases by tissue specificity. Transcript abundance of the 42 PtrPOs in SDX was compared to the transcript abundance in three other tissues, phloem (P), leaf (L), and young stem (S) (Fig. 2). Compared to the transcript pattern of the well-known xylem-specific monolignol biosynthetic genes, 4-coumaric acid: coenzyme A (CoA) ligases (Ptr4CL3 and Ptr4CL5) and hydroxycinnamoyl-CoA: shikimic acid hydroxycinnamoyl transferases (PtrHCT1 and PtrHCT6) (Fig. 2, bottom), five PtrPOs were identified as SDX-specific, which are PtrPO21, PtPrx15, PtPrx19, PtPrx37, and PtPrx82. Among these 5 xylem-specific PtrPOs, the transcript abundance of PtrPO21 is the highest and the transcript level is up to 255-fold more than the other 4 SDX-specific PtrPOs (PtPrx15, PtPrx19, PtPrx37, and PtPrx82). Therefore, we selected PtrPO21 as the best candidate of the xylem-abundant and xylem-specific peroxidases in P. trichocarpa (Figs. 1 and 2) for a role in lignin polymerization.
Transcript abundance of the 42 detectable class III peroxidases in different tissues in P. trichocarpa and SDX-specific expressed monolignol biosynthetic genes (Ptr4CL3, Ptr4CL5, Ptr4HCT1, and PtrHCT6). Transcript levels were counted from RNA-seq as RPM (reads per million) and the arrangement of 42 PtrPOs (left to right, top to bottom) is according to the transcript level (high to low) in SDX. X, SDX; P, phloem; L, leaf; S, young stem. Error bars represent one SE of three replicates
PtrPO21 shares conserved structural motifs with the coniferyl alcohol-specific peroxidase from A. thaliana and the sinapyl alcohol-specific peroxidase from Z. elegans
The amino acid sequence of PtrPO21 was compared to classic lignin peroxidases, the coniferyl alcohol (G)-specific peroxidase (ATP-A2) from A. thaliana (Ostergaard et al. 2000) and the sinapyl alcohol (S)-specific peroxidase (ZePrx) from Z. elegans (Gabaldon et al. 2005) to identify conserved structural motifs (Fig. 3a). Although the amino acid identity of PtrPO21 and ATP-A2 (32.4 %) or ZePrx (33.2 %) is low, the alignment still showed several conserved amino acids in active sites (black arrow head in Fig. 3b), substrate-binding sites (red dot in Fig. 3b), and a signature structural motif for G- or S-specific peroxidase (green bar in Fig. 3b) (Barceló et al. 2004).
Amino acid alignment of PtrPO21, the classic G peroxidase (ATP-A2) and the classic S peroxidase (ZePrx). a Identical amino acids are in yellow and conserved substitutions are in cyan. b Diagram of the primary structure of PtrPO21 protein. The structural motifs are indicated as active site (black box and arrow head), substrate-binding site (red box and dot) and S or G peroxidase structural motif (green box and bar)
PtrPO21 is an unusual anionic peroxidase in the stem differentiating xylem
Many plant peroxidases have been fractionated and characterized according to their isoelectric points (pI) (Zimmerlin et al. 1994; Christensen et al. 1998; Carpin et al. 1999; Quiroga et al. 2000; Gabaldon et al. 2005). Anionic peroxidases (pI < 7) were considered to contribute to the cell wall peroxidase activity and lignin polymerization, while cationic peroxidases (pI > 7) were implicated in auxin catabolism (Mader 1980; Campa 1991; Lagrimini et al. 1993; Aoyama et al. 2002). Plant peroxidases are also divided into cell wall or vacuolar types based on the presence or absence of a C-terminal extension peptide, which may function as a vacuolar sorting signal (VSS) (Matsui et al. 2011). Vacuolar peroxidases may function in defense against abiotic and biotic stresses (Ostergaard et al. 2000; Sasaki et al. 2004; Gabaldon et al. 2005; Bindschedler et al. 2006; Ren et al. 2014).
We compared the amino acid identity of PtrPO21 to lignin peroxidases from tomato, French bean, Z. elegans, tobacco, Arabidopsis, hybrid aspen (P. sieboldii x P. grandidentata), P. trichocarpa, and P. alba (Table 1). PtrPO21 showed about 30 % identity to most lignin peroxidases, except for higher identity of 53.9 % with TP60 in tobacco (underlined in Table 2). However, PtrPO21 is an anionic peroxidase with a pI of 5.87 and TP60 is a cationic peroxidase with a pI of 8.89 (Table 1). Two poplar peroxidases (prxA3a and PXP3-4) show high sequence identity (50–60 %) to FBP1 and ATP-A2 from Arabidopsis, but none of the known poplar lignin peroxidases (prxA3a from hybrid aspen, PXP3-4 from P. trichocarpa and CWPO-C from P. alba) is similar to PtrPO21.
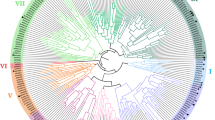
A phylogenetic analysis was performed with PtrPO21 and the 73 class III peroxidases in Arabidopsis (AtPrxs) (Fig. 4). Among the 73 class III peroxidases in Arabidopsis, all lignin peroxidases previously identified in Arabidopsis (AtPrx53, AtPrx2, AtPrx25, AtPrx71, and AtPrx72) show low amino acid identity to PtrPO21 (29.5–37.2 %). Of 73 AtPrxs, PtrPO21 shows highest identity to AtPrx21 (69.3 %), which is a peroxidase expressed in roots, leaves, and stems. AtPrx21 had been identified in a study of abiotic or biotic stresses in Arabidopsis, but a role in lignification needs to be investigated (Tognolli et al. 2002; Cosio and Dunand 2009).
Phylogenetic analysis of the PtrPOs, poplar lignin peroxidases (CWPO-C, prxA3a, and PXP3-4), Arabidopsis lignin peroxidases (ATP-A2, AtPrx2, AtPrx 25, AtPrx71, and AtPrx72) and all class III peroxidases in Arabidopsis. PtrPO21 belongs to a clade that is separate from prxA3a in P. sieboldii x P. grandidentata, PXP3-4 in P. trichocarpa and CWPO-C in P. alba
PtrPO21 is a cell wall-bound peroxidase
Similar to ATP-A2 and ZePrx, which are known cell wall-associated peroxidases (Ostergaard et al. 1998; Barcelo et al. 2007), PtrPO21 also lacks a C-terminal extension peptide (Fig. 3a). To confirm a cell wall localization for PtrPO21, absolute quantification by protein cleavage-isotope dilution mass spectrometry (PC-IDMS) was performed (Shuford et al. 2012) on the proteins in the cell wall and soluble fractions of SDX. PtrPO21 was only found in the cell wall fraction of SDX, while Ptr4CL3, a major soluble monolignol biosynthetic enzyme, was only found in the soluble SDX fraction (Fig. 5). The location of PtrPO21 in the cell wall is supporting evidence for a function in lignin polymerization. To obtain further support for a function of PtrPO21 in lignification, we next investigated the consequence of downregulation of PtrPO21 using RNA interference (RNAi) transgenesis.
Cellular localization of PtrPO21 and Ptr4CL3 using protein cleavage-isotope dilution mass spectrometry (PC-IDMS). a PtrPO21, b Ptr4CL3. Two wild-type trees were used for localization. Ptr4CL3 is a marker for the soluble fraction. The white bar is the cell wall fraction; the black bar is the soluble fraction. Error bars represent one SE obtained from three measurements of each sample. N.D. non-detectable
PtrPO21 downregulated transgenics in P. trichocarpa have reduced growth and reddish internodes in the stem wood
An RNAi construct was prepared containing an inverted repeat from a specific region of the PtrPO21 gene under the control of the Ptr4CL3 promoter as in our previous work (Wang et al. 2014). The construct was transformed into P. trichocarpa (Song et al. 2006) and six independent transgenic lines were obtained (Fig. 6). Three lines (PtrPO21-8, PtrPO21-9, and PtrPO21-10) were selected for further analysis because they show medium- and high-level downregulation of PtrPO21 transcript abundance. The specificity of the PtrPO21 RNAi downregulation was tested using real-time PCR. For the four most xylem-abundant class III peroxidases (PtrPO12, PtrPO21, PtrPO42, and PtrPO64) in SDX, only the transcript level of PtrPO21 was significantly downregulated (Supplemental Figure S1). Two lines of 6-month-old PtrPO21 downregulated transgenics (PtrPO21-8 and PtrPO21-9) with PtrPO21 transcript reduced to as little as ~5 % of wild type show significant height reduction (Fig. 7a).
Pink or red internodes of stem wood were observed when PtrPO21 was downregulated compared to the pale yellow color of the wild-type stems (Fig. 8d). The PtrPO21-10 line with PtrPO21 transcript reduced to ~40 % of wild type showed pink color in the stem wood (Fig. 8c) and both PtrPO21-8 and PtrPO21-9 (~5 % of wild-type transcript abundance) showed reddish color (Fig. 8a, b). Both PtrPO21-8 and PtrPO21-9 have shorter internodes than wild type (distance between arrows in Fig. 8) and gross morphology of the stem sections did not show any other obvious differences compared to wild type (data not shown). The extractive free wood powder of the PtrPO21 downregulated transgenics also showed brownish color compared to the pale yellow color of wild type (Supplemental Figure S2). Moreover, transgenic downregulation of the other three most xylem-abundant peroxidases (PtrPO12, PtrPO42, or PtrPO64) had neither significant lignin reduction nor pink or red color of the stems (Supplemental Figure S3A and S3B). Therefore, the growth, lignin content, and wood phenotype are likely to be effect of the downregulation of PtrPO21, and not the result of off-target effects of the RNAi construct on the other SDX-specific peroxidase transcripts, or to cryptic insertions affecting non-peroxidase genes.
Cell wall component analysis of PtrPO21 downregulated transgenics
To further investigate whether downregulation of PtrPO21 affects lignification and wood composition in P. trichocarpa, the stems of the transgenic and wild-type trees were analyzed for lignin and carbohydrate content. Compared to the wild type, the lignin content of the three lines, PtrPO21-8, PtrPO21-9, and PtrPO21-10, showed reduced Klason (acid insoluble) lignin content ranging from 17.8 to 23.2 % reduction (p < 0.001; Table 3). Total lignin content was also significantly reduced (p < 0.001; Table 3) in all PtrPO21 downregulated transgenics. No significant change was observed in the minor fraction of acid soluble lignin. This result adds support to the evidence that PtrPO21 has a role in lignification in P. trichocarpa. Xylan content was significantly increased (p < 0.005; Table 3), and the content of mannan was significantly decreased (p < 0.05; Table 3). Cellulose, arabinan, and galactan content had no significant changes. Similar changes in cell wall components were also observed when laccases in P. trichocarpa were downregulated (Lu et al. 2013), suggesting that both types of oxidases have similar functions in plant cell wall biosynthesis.
Mechanical properties of the wood of PtrPO21 downregulated transgenics
To examine the effect of PtrPO21 downregulation on mechanical properties of wood, we determined the modulus of elasticity (MOE) using three point bending (Horvath et al. 2010). MOE reveals the resistance of the stem to bending, which estimates the elasticity. All stem wood samples from PtrPO21 downregulated transgenics showed significantly lower MOE (37–60 %) than the wild type (Fig. 9), which means that the strength of the wood in PtrPO21 downregulated transgenics had significantly decreased.
Lignin composition of PtrPO21 downregulated transgenics
To determine whether the decrease in MOE results from altered lignin composition in the PtrPO21 downregulated transgenics, the lignin composition of wild type and PtrPO21 downregulated transgenics was examined by nitrobenzene oxidation. In wild type, the ratio of syringyl (S) to guaiacyl (G) subunits is 2.1, and with trace amounts (0.5 %) of 4-hydroxyphenyl (H) subunits (Table 4). The S subunits are represented by the sum of syringaldehyde and syringic acid, and the sum of vanillin and vanillic acid represents the G subunits. In PtrPO21-10 (transcript abundance reduced to ~40 % of wild type), the S subunits were reduced by 10.1 %, while the G subunit content was similar to wild type. In the severe PtrPO21 downregulated transgenic lines (PtrPO21-8 and PtrPO21-9), S subunits were reduced by 20 % and G subunits were decreased by around 10 % (Table 4). In angiosperm wood, such as P. trichocarpa, fiber cell lignin mainly contains S subunits. The reduction of S subunits may account for the reduction in the elasticity of the plant cell walls, as we observe in the reduction of the MOE (Fig. 9). Moreover, H subunits increased ~3-fold in PtrPO21 downregulated transgenics (Table 4). The increase in H subunits is known to result in lower molecular weight lignin polymers and more easily deconstructed biomass, which may also contribute to the lower MOE (Sangha et al. 2014); Fig. 9).
Differentially expressed genes (DEGs) in the PtrPO21 downregulated transgenics
Functional redundancy is expected among the class III peroxidase and laccase families in P. trichocarpa; therefore, we performed transcriptome analysis to identify any differentially expressed oxidases (peroxidases or laccases) in PtrPO21 downregulated transgenics, which may compensate for the reduction of PtrPO21. One of the 42 xylem-abundant class III peroxidases, PtPrx35, had significant overexpression (Table 5). PtPrx35 has 73.9 % amino acid sequence identity to AtPrx17 in Arabidopsis, and AtPrx17 had been suggested to have a role in lignification for pod shattering (Cosio and Dunand 2009; Cosio and Dunand 2010). None of the 49 gene models of P. trichocarpa laccases (PtrLACs) had significant transcript changes in any of the PtrPO21 downregulated transgenics (Supplemental Table S3). Of the 21 monolignol biosynthetic enzymes, P. trichocarpa phenylalanine ammonia-lyase 3 (PtrPAL3) showed significant overexpression in all PtrPO21 downregulated transgenic lines (Supplemental Table S4), suggesting specific and novel regulation involving these genes in lignin biosynthesis.
Discussion
Class III peroxidases have been considered to play important roles in lignification (Nakamura 1967; Harkin and Obst 1973; Whetten et al. 1998; Cosio and Dunand 2009). Because of the large gene family and the functional redundancy of the class III peroxidases, only a few specific cationic or anionic peroxidases involved in lignification have been identified and isolated from plant cell walls or plant suspension cells (Imberty et al. 1985; Lagrimini et al. 1987; Polle et al. 1994; Sato et al. 1995; Christensen et al. 1998; Barceló et al. 2004; Gabaldon et al. 2005; Sato et al. 2006). Using computational and structural simulation, several Arabidopsis class III peroxidases have been identified and validated using Arabidopsis mutants (Ostergaard et al. 1998; Nielsen et al. 2001; Tokunaga et al. 2009; Herrero et al. 2013a, b; Shigeto et al. 2013). In woody plants such as poplar, the identity of lignin peroxidases was still inconclusive (Christensen et al. 2001a; Tamura et al. 2001; Li et al. 2003b). Identification of lignin peroxidases in P. trichocarpa is important to understand the complete lignin biosynthetic pathway in a model woody plant and to design better strategies to reduce recalcitrance of biomass for pulp/paper and biofuel production.
Class III peroxidases involved in lignification
Identification of peroxidases with high amino acid similarity to other lignin peroxidases among plant species has led to the discovery of new lignin peroxidases (Tokunaga et al. 2009; Herrero et al. 2013a; Shigeto et al. 2013). For example, AtPrx2, AtPrx25, and AtPrx71 are closely related to CWPO-C. AtPrx66, AtPrx47, and AtPrx64 are closely related to ZPO-C, and AtPrx72 is very similar to ZePrx. However, PtrPO21 is unusual because of the low amino acid identity (30 %) compared to other lignin peroxidases (Table 2) and it is also distinct from TP60 (Table 1).
Several studies of recombinant peroxidases revealed that both cationic and anionic peroxidases are able to oxidize coniferyl alcohol or sinapyl alcohol, but with different preferences. The anionic peroxidase, ATP-A2, prefers coniferyl alcohol (Ostergaard et al. 2000; Nielsen et al. 2001) and the cationic peroxidases, ZePrx and CWPO-C, prefer sinapyl alcohol (Tsutsumi et al. 1998; Aoyama et al. 2002; Sasaki et al. 2004; Gabaldon et al. 2005; Sasaki et al. 2006; Sasaki et al. 2008). ZPO-C, a cationic peroxidase, shows activity for sinapyl alcohol as well as coniferyl alcohol (Sato et al. 2006) and the anionic peroxidases, PXP3-4 and TPX1, show syringaldazine-oxidizing activity (Christensen et al. 1998; Quiroga et al. 2000; Christensen et al. 2001b). In fact, the closest homolog to TP60 in P. trichocarpa is PtrPO12 (see note in Supplemental Table S2), which has 80.7 % amino acid identity and is also a cationic peroxidase with a pI of 7.73. However, the PtrPO12 downregulated P. trichocarpa did not show a reduction in lignin content (Supplemental Figure S3A).
PtrPO21 is distinct from previously identified poplar peroxidases, the PXP3-4, prxA3a, and CWPO-C (Table 2). Among the 88 class III peroxidases, PXP3-4 shows most identity to PtPrx03. PrxA3a has most sequence identity to PtPrx98, and PtPrx75 is the closest homolog to CWPO-C. PtPrx03, PtPrx98, and PtPrx75 were not studied here because they are not xylem-abundant or xylem-specific in P. trichocarpa.
PtrPO21 possesses structural motifs similar to G- and S-specific peroxidases
Anionic peroxidase ATP-A2 is a well-studied G-specific peroxidase in Arabidopsis (Ostergaard et al. 2000; Nielsen et al. 2001; Barceló et al. 2004). The amino acids P69, I138, P139, S140, and R175 are key determinants of the conformation and hydrophobicity of the ATP-A2 substrate-binding site (Ostergaard et al. 2000). The oxidation of sinapyl alcohol is thought to be sterically hindered due to unfavorable hydrophobic interactions between the sinapyl alcohol methoxy side chain and the conserved I138 and P139 residues at the substrate-binding site (Ostergaard et al. 2000). PtrPO21 has conserved the I138 and P139 residues, suggesting that PtrPO21 may prefer coniferyl alcohol, as a G-specific peroxidase (2nd red box in Fig. 3). However, PtrPO21 also has the structural motif VSCAD, which is characteristic of an S peroxidase, where G peroxidases have a structural motif of VSCSD (green box in Fig. 3a) (Gomez Ros et al. 2007). Therefore, PtrPO21 has characteristics of both G and S peroxidases, which may explain why both G and S subunits are decreased in PtrPO21 downregulated transgenics (Table 4).
Reddish internodes of stems in PtrPO21 downregulated transgenics
Reddish stem wood has been observed previously in mutant or transgenic plants when the monolignol biosynthetic pathway is downregulated. The most prominent reddish stem phenotype is observed when cinnamyl alcohol dehydrogenase (CAD) is downregulated or silenced in maize brown midrib (bm) or sorghum (bmr) mutants (Grand et al. 1985; Pillonel et al. 1991; Halpin et al. 1998; Zhang et al. 2006), tobacco (Higuchi et al. 1994; Ralph et al. 1998), poplar (Baucher et al. 1996), loblolly pine (MacKay et al. 1997; Ralph et al. 1997; MacKay et al. 1999; Lapierre et al. 2000) and Arabidopsis (Sibout et al. 2005). In addition to CAD, the reddish color of stems is also observed in mutant or transgenics with reduced activity of 4CL (Voelker et al. 2010; Xu et al. 2011), caffeic acid O-methyltransferase (COMT) (Vignols et al. 1995), cinnamoyl-CoA reductase (CCR) (Van Acker et al. 2014), or caffeoyl-coenzyme A-O-methyltransferase (CCoAOMT) (Meyermans et al. 2000). The reddish color is considered to result from the polymerization of the hydroxycinnamaldehydes in the transgenics or incorporation of novel monolignol monomers (Saathoff et al. 2011; Kaur et al. 2012). Hydroxycinnamaldehydes can form a wine-red dehydrogenation polymer in vitro (Higuchi et al. 1994).
In this study, the reddish color of the stem internode is observed when PtrPO21 is downregulated (Fig. 8). This reddish phenotype in the PtrPO21 downregulated transgenics has not been reported for a downregulated lignin peroxidase. The reddish coloration of the stem may also be a consequence of lignin polymerization in P. trichocarpa (Voelker et al. 2010). The color is retained in extracted wood powder and therefore is not due to a soluble component.
The role of peroxidases and laccases in lignin polymerization
In early studies, the role of peroxidase was considered to be more likely than laccase in the lignification because of the lack of detection of laccase activity in some plants (Nakamura 1967; Harkin and Obst 1973). Lignification requires the presence of hydrogen peroxide (Kärkönen et al. 2002) and the first downregulation of laccase in poplar showed no change in lignin content (Ranocha et al. 2002). Moreover, peroxidase has far higher specific activities with phenolic substrates compared to laccase (Wallace and Fry 1999).
However, a role for laccase in lignification was re-examined in 1983 because of improved procedures for identification of laccase in Acer pseudoplatanus (Bligny and Douce 1983; Sterjiades et al. 1992), but laccase was still suggested to be involved only in the early stages of lignification. Laccase activity was then detected and correlated with lignification in several plants (Driouich et al. 1992; Bao et al. 1993; O’Malley et al. 1993; Dean and Eriksson 1994; Liu et al. 1994; Richardson and McDougall 1997; Richardson et al. 2000; Sato et al. 2001). In Arabidopsis, a triple mutant of LAC4, LAC11 and LAC17 dramatically reduced lignin deposition and arrested plant growth (Zhao et al. 2013). In P. trichocarpa, overexpression of a microRNA (Ptr-mirRNA397) downregulated 17 PtrLACs and reduced Klason lignin content as much as 22 % below the wild-type level (Lu et al. 2013). Therefore, both peroxidases and laccases should be considered important for lignification in P. trichocarpa.
In this study, we identified a new peroxidase PtrPO21 involved in lignin biosynthesis and validated its involvement by downregulation in transgenic P. trichocarpa affecting lignin content and composition. Because of the high level of gene redundancy in the PtrPO family, we cannot exclude the possibility that other PtrPOs expressed in xylem may also play roles in lignin biosynthesis. We suggest the involvement of other oxidases for lignin biosynthesis because lignin content can be reduced up to ~22 % by downregulation of Ptr-mirRNA397 and ~20 % by downregulation of PtrPO21. To identify more lignin peroxidases in P. trichocarpa, downregulation transgenics could be generated for multiple peroxidases or combinations of peroxidase and laccase genes.
Materials and methods
Plant materials
As in our previous work, stem differentiating xylem (SDX) was collected from 6-month-old P. trichocarpa (Nisqually-1). Our wild type and PtrPO transgenic plants maintained in a greenhouse following Li et al. (2011).
Identification of class III peroxidases in P. trichocarpa and phylogenetic analysis
The class III peroxidases of A. thaliana and P. trichocarpa were retrieved from PeroxiBase (http://peroxibase.toulouse.inra.fr/). Class III peroxidases of P. trichocarpa from the latest publication were also used for comparison (Ren et al. 2014). Amino acid alignment was performed using Vector NTI software (Invitrogen, Grand Island, NY) (Lu and Moriyama 2004). Phylogenetic analysis was carried out using MEGA 5.1 (Tamura et al. 2011) with a bootstrap resampling of 1000 replicates and probabilities >50 %.
RNA extractions
Fifty to 100 mg of xylem, phloem, leaf, or young stem frozen powder was used to purify total RNA using the RNeasy Plant RNA Isolation Kit (Qiagen, Limburg, Netherlands) following the manufacturer’s protocol with a DNase on-column digestion. A260/A280 ratios of the RNA ranged from 1.9 to 2.1, and RNA integrity was estimated using an Agilent 2100 Bioanalyzer. RNA integrity numbers (RIN) ranged from 8.6 to 10.0.
Transcriptome (RNA-seq) and differentially expressed gene analysis
RNA-seq and differentially expressed gene (DEG) analysis follow our previous methods (Lu et al. 2013). We use the term “differentially expressed gene (DEG)” following common usage, although we are aware that what is measured are changes in transcript abundance, which may also be due to changes in processing or degradation. For stem differentiating xylem (SDX) and tissue-specific P. trichocarpa peroxidase analysis, 1 μg RNA from SDX, leaf, phloem, and young stem of three wild-type plants were used. For transcriptome analysis of PtrPO21 transgenics, 1 μg of RNA from 2 trees from each transgenic line or wild-type P. trichocarpa was used, except for PtrPO21-8 where only one plant was available. Following the RNA-seq library preparation (Illumina, San Diego, CA), mRNA was purified using poly-T oligo-attached magnetic beads. The mRNA was then fragmented, reverse transcribed into double-strand cDNA, followed by end repair and 3′ end adenylation. The cDNA was ligated with multiplex adapters and PCR amplified to produce the RNA-seq libraries. The RNA-seq libraries were adjusted to 1 nM and pooled for multiplex sequencing on a Hiseq 2000 (Illumina, San Diego, CA). DEG analysis also followed Li et al. (2011). The resulting sequences were mapped to the P. trichocarpa genome v2.0, gene annotation v2.2 (www.phytozome.org) using TOPHAT (Trapnell et al. 2009). The frequency of raw counts was determined by BEDtools and normalized using the trimmed mean of M value (TMM), a scaling normalization method for analysis of differential gene expression (Quinlan and Hall 2010; Robinson et al. 2010). The genes with counts lower than 15 per million per library were filtered out. DEGs were obtained by pairwise comparisons of transgenic and wild-type libraries using edgeR/Bioconductor (Robinson et al. 2010). The statistical significance of DEGs is based on a false discovery rate (FDR) of 0.05.
Absolute quantification of PtrPO21 from cell fractionation using protein cleavage-isotope dilution mass spectrometry (PC-IDMS)
The extraction of xylem crude protein followed Shuford et al. (2012). Six grams of xylem tissue was ground in liquid nitrogen followed by 2 min of homogenization in 30 mL of extraction buffer on ice. Cellular debris was pelleted by centrifugation for 15 min at 3000×g at 4 °C, and the supernatant was collected as a soluble fraction. The pellet was prepared as in Aoyama et al. (2002) with little modification. The pellet was washed four times with 50 mM Tris-HC1 buffer (pH 7.5) to ensure the removal of the soluble protein. The resulting cell wall residue was incubated in the same buffer plus 1 M NaC1 to extract ionic-bound cell wall peroxidases. After centrifugation at 10,000×g at 4 °C, the supernatant was collected as the cell wall fraction. Absolute abundances of PtrPO21 and Ptr4CL3 in the soluble and cell wall fraction were obtained by PC-IDMS using labelled surrogate peptides following Shuford et al. (2012) and implementing our recently optimized digestion conditions for absolute quantification of proteins (Loziuk et al. 2013). To enhance sensitivity towards these target proteins, only PtrPO21 and Ptr4CL3 were targeted in the selected reaction monitoring assay. Peptides were confirmed to be quantifiable based on co-elution of labelled and native peptides as well as co-elution of specific fragments. Purity of these fragments was further confirmed based on the expected relative abundance of these fragments to one another above a threshold value.
RNA-interference (RNAi) plasmid constructions and plant transformation
An RNA-silencing construct about 300 base pairs, with an inverted repeat specific to PtrPO21, was prepared as in our previous study (Wang et al. 2014). A 680-bp GUS linker (GL) was amplified with a specific pair of primers (Li et al. 2011) and cloned into the pCR2.1 vector, resulting in pCR2.1-GL. The inverted repeats consisted of chimeric sequences from specific target gene PtrPOs obtained using specific primer sets (Supplemental Table S5). The sense and antisense fragments were inserted into pCR2.1-GL to produce pCR2.1-sense-GL-antisense. The sense-GL-antisense fragment was then cloned into pBI121-Ptr4CLp behind the P. trichocarpa 4-coumaric acid: coenzyme A (CoA) ligase (4CL) promoter (Ptr4CLp) replacing the GUS gene. The construct was introduced into Agrobacterium tumefaciens (C58) by the freeze thaw method (Holsters et al. 1978) and transferred into P. trichocarpa (Nisqually 1) following our established method for Agrobacterium-based transformation (Song et al. 2006).
Real-time PCR (RT-PCR)
Gene-specific primer sets were designed for PtrPO12, PtrPO21, PtrPO42, and PtrPO64 (Supplemental Table S5). RT-PCR was performed following Li et al. (2011). Total RNA (150 ng) was reverse-transcribed using TaqMan reverse transcription reagents (Life Technologies, Grand Island, NY). Real-Time PCR (RT-PCR) reactions were carried out in a 25-μL mixture of first strand cDNA (equivalent to 5 ng of total RNA), 5 pmol of specific primer sets, and 12.5 μL of 2X SYBR green PCR master mix (Roche, Basel, Switzerland). The products of the RT-PCR were detected using the 7900 HT Sequence Detection System (Life Technologies, Grand Island, NY). The program of RT-PCR is: 95 °C for 10 min, then 45 cycles of 95 °C for 15 s and 60 °C for 1 min, after which a thermal denaturing cycle was added to determine the dissociation curve of the PCR products to check the amplification specificity.
Lignin content and carbohydrate determination
The procedure followed our lab protocol (Lu et al. 2013). Debarked stems were placed in 100 % acetone and held for 2 days at room temperature to remove extractives. The acetone was replaced three times by 90 % acetone at 48-h intervals. After drying, wood was ground in a Wiley mill with a 40-mesh screen and the wood powder was further screened between 40 mesh and 60 mesh. The resulting wood meal (40–60 mesh) was used for lignin content determination to estimate acid-soluble lignin (ASL), acid-insoluble lignin (Klason lignin), and the total lignin (ASL plus Klason lignin) (Yeh et al. 2005). Oven-dried wood powder (100 mg) was mixed with 1.5 mL of 72 % sulfuric acid for 90 min and then diluted to 57.5 mL with distilled water. The suspension was heated at 121 °C for 150 min, then filtered through a crucible. The supernatant was collected for ASL and carbohydrate analysis. The residual lignin in the crucible was used to measure the acid-insoluble lignin by weight. For ASL, the filtrate solution was 10-fold diluted with distilled water and absorbance measured at 205 nm. The extinction coefficient for ASL is 110 g−1·cm−1 at 205 nm (Dence 1992). For carbohydrate analysis, the filtrate without dilution was neutralized with calcium carbonate overnight at room temperature. Then, the neutralized solution was further filtered and the filtrate was analyzed using analytical HPLC (Shodex SUGAR SPO 810, 8 × 30 mm, Pb2+ cation exchange column, Showa Denko America, Inc., NY) with distilled water as eluent, at a flow rate of 0.5 mL/min, at 80 °C. Sugars were identified by refractive index based on authentic compounds.
Nitrobenzene oxidation for lignin composition
Following Chen (1992), 200 mg of oven-dried extractive free wood powder was used to determine the lignin composition using nitrobenzene oxidation. Following oxidation, 5 μL of sample was analyzed by analytical HPLC using a Zorbax SB-C3 5 μm, 4.6 × 150-mm column (Agilent, Santa Clara, CA). Analysis of reactions was carried out using an HPLC gradient (solvent A, 10 mM formic acid in water; solvent B, 10 mM formic acid in acetonitrile; 10 % B for 3 min, 10 to 20 % B for 5 min, 20 to 30 % B for 6 min, 30 to 100 % B for 2 min, 100 % B for 2 min; flow rate: 1.5 mL/min, at 40 °C). A standard curve was established using a Diode-Array Detector SL (Agilent, Santa Clara, CA) based on authentic compounds of 4-hydroxybenzoic acid, 4-hydroxybenzaldehyde, vanillic acid, syringic acid, vanillin, and syringaldehyde (Liu et al. 2012).
Mechanical Properties of Wood
About 20-cm-long stem sections were cut from the bottom of the stems of the transgenic trees. The stems were kept in plastic bags to prevent drying. One segment with a length-to-width ratio of 22 was cut from each stem section and debarked before measuring the mechanical properties. A three-point bending test was conducted to measure the modulus of elasticity (MOE) by an MTS Alliance RF/300 universal mechanical tester (Li et al. 2011).
References
Abeles FB, Dunn LJ, Morgens P, Callahan A, Dinterman RE, Schmidt J (1988) Induction of 33-kD and 60-kD peroxidases during ethylene-induced senescence of cucumber cotyledons. Plant Physiol 87:609–615
Adler E (1977) Lignin chemistry—past, present and future. Wood Sci Technol 11:169–218
Allison SD, Schultz JC (2004) Differential activity of peroxidase isozymes in response to wounding, gypsy moth, and plant hormones in northern red oak (Quercus rubra L.). J Chem Ecol 30:1363–1379
Amaya I, Botella MA, de la Calle M, Medina MI, Heredia A, Bressan RA, Hasegawa PM, Quesada MA, Valpuesta V (1999) Improved germination under osmotic stress of tobacco plants overexpressing a cell wall peroxidase. FEBS Lett 457:80–84
Aoyama W, Sasaki S, Matsumura S, Mitsunaga T, Hirai H, Tsutsumi Y, Nishida T (2002) Sinapyl alcohol-specific peroxidase isoenzyme catalyzes the formation of the dehydrogenative polymer from sinapyl alcohol. J Wood Sci 48:497–504
Bao W, O’Malley DM, Whetten R, Sederoff RR (1993) A laccase associated with lignification in loblolly pine xylem. Science 260:672–672
Barceló AR, Gómez Ros LV, Gabaldón C, López-Serrano M, Pomar F, Carrión JS, Pedreño MA (2004) Basic peroxidases: the gateway for lignin evolution? Phytochem Rev 3:61–78
Barcelo AR, Ros LV, Carrasco AE (2007) Looking for syringyl peroxidases. Trends Plant Sci 12:486–491
Baucher M, Chabbert B, Pilate G, Van Doorsselaere J, Tollier MT, Petit-Conil M, Cornu D, Monties B, Van Montagu M, Inze D, Jouanin L, Boerjan W (1996) Red xylem and higher lignin extractability by down-regulating a cinnamyl alcohol dehydrogenase in poplar. Plant Physiol 112:1479–1490
Bernards MA, Fleming WD, Llewellyn DB, Priefer R, Yang X, Sabatino A, Plourde GL (1999) Biochemical characterization of the suberization-associated anionic peroxidase of potato. Plant Physiol 121:135–146
Bernards MK, Summerhurst DK, Razem FA (2004) Oxidases, peroxidases and hydrogen peroxide: the suberin connection. Phytochem Rev 3:113–126
Bindschedler LV, Dewdney J, Blee KA, Stone JM, Asai T, Plotnikov J, Denoux C, Hayes T, Gerrish C, Davies DR, Ausubel FM, Bolwell GP (2006) Peroxidase-dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. Plant J 47:851–863
Blee KA, Choi JW, O’Connell AP, Schuch W, Lewis NG, Bolwell GP (2003) A lignin-specific peroxidase in tobacco whose antisense suppression leads to vascular tissue modification. Phytochemistry 64:163–176
Bligny R, Douce R (1983) Excretion of laccase by sycamore (Acer pseudoplatanus L.) cells. Purification and properties of the enzyme. Biochem J 209:489–496
Botella MA, Quesada MA, Medina MI, Pliego F, Valpuesta V (1994) Induction of a tomato peroxidase gene in vascular tissue. FEBS Lett 347:195–198
Campa A (1991) Biological roles of plant peroxidases: known and potential function. Peroxidases Chem Biol 2:25–50
Carpin S, Crèvecoeur M, Greppin H, Penel C (1999) Molecular cloning and tissue-specific expression of an anionic peroxidase in zucchini. Plant Physiol 120:799–810
Chen CL (1992) Nitrobenzene and cupric oxide oxidations. In: Lin S, Dence C (eds) Methods in lignin chemistry. Springer, Berlin Heidelberg, pp 301–321
Chen F, Dixon RA (2007) Lignin modification improves fermentable sugar yields for biofuel production. Nat Biotechnol 25:759–761
Christensen JH, Bauw G, Welinder KG, Van Montagu M, Boerjan W (1998) Purification and characterization of peroxidases correlated with lignification in poplar xylem. Plant Physiol 118:125–135
Christensen JH, Montagu MV, Bauw G, Boerjan W (2001a) Xylem peroxidases: purification and altered expression. Prog Biotechnol 18:171–176
Christensen JH, Overney S, Rohde A, Diaz WA, Bauw G, Simon P, Van Montagu M, Boerjan W (2001b) The syringaldazine-oxidizing peroxidase PXP 3-4 from poplar xylem: cDNA isolation, characterization and expression. Plant Mol Biol 47:581–593
Cosio C, Dunand C (2009) Specific functions of individual class III peroxidase genes. J Exp Bot 60:391–408
Cosio C, Dunand C (2010) Transcriptome analysis of various flower and silique development stages indicates a set of class III peroxidase genes potentially involved in pod shattering in Arabidopsis thaliana. BMC Genomics 11:528
Dean JFD, Eriksson KEL (1994) Laccase and the deposition of lignin in vascular plants. In Holzforschung, p 21
Dence CW (1992) The determination of lignin. In: Lin S, Dence C (eds) Methods in lignin chemistry. Springer, Berlin Heidelberg, pp 33–61
Driouich A, Lainé A-C, Vian B, Faye L (1992) Characterization and localization of laccase forms in stem and cell cultures of sycamore. Plant J 2:13–24
Duroux L, Welinder KG (2003) The peroxidase gene family in plants: a phylogenetic overview. J Mol Evol 57:397–407
Eckardt NA (2002) Probing the mysteries of lignin biosynthesis: the crystal structure of caffeic acid/5-hydroxyferulic acid 3/5-O-methyltransferase provides new insights. Plant Cell 14:1185–1189
El Mansouri I, Mercado JA, Santiago-Dómenech N, Pliego-Alfaro F, Valpuesta V, Quesada MA (1999) Biochemical and phenotypical characterization of transgenic tomato plants overexpressing a basic peroxidase. Physiol Plant 106:355–362
Erdtman H (1933) Dehydrierungen in der Coniferylreihe. II. Dehydrodi‐isoeugenol. Justus Liebigs Annalen der Chemie 503:283–294
Espelie KE, Kolattukudy PE (1985) Purification and characterization of an abscisic acid-inducible anionic peroxidase associated with suberization in potato (Solanum tuberosum). Arch Biochem Biophys 240:539–545
Fagerstedt K, Saranpää P, Piispanen R (1998) Peroxidase activity, isoenzymes and histological localisation in sapwood and heartwood of Scots pine (Pinus sylvestris L.). J For Res 3:43–47
Fagerstedt KV, Kukkola EM, Koistinen VV, Takahashi J, Marjamaa K (2010) Cell wall lignin is polymerised by class III secretable plant peroxidases in Norway spruce. J Integr Plant Biol 52:186–194
Freudenberg K (1959) Biosynthesis and constitution of lignin. Nature 183:1152–1155
Freudenberg K (1965) Lignin: its constitution and formation from p-hydroxycinnamyl alcohols: lignin is duplicated by dehydrogenation of these alcohols; intermediates explain formation and structure. Science 148:595–600
Fry SC (1986) Cross-linking of matrix polymers in the growing cell-walls of angiosperms. Annu Rev Plant Physiol Plant Mol Biol 37:165–186
Fry W, White J (1938) Big trees. Palo Alto. Stanford Univ. Press, CA
Gabaldon C, Lopez-Serrano M, Pedreno MA, Barcelo AR (2005) Cloning and molecular characterization of the basic peroxidase isoenzyme from Zinnia elegans, an enzyme involved in lignin biosynthesis. Plant Physiol 139:1138–1154
Gabaldón C, López-Serrano M, Pomar F, Merino F, Cuello J, Pedreño MA, Barceló AR (2006) Characterization of the last step of lignin biosynthesis in Zinnia elegans suspension cell cultures. FEBS Lett 580:4311–4316
Gazaryan IG, Lagrimini LM, Ashby GA, Thorneley RN (1996) Mechanism of indole-3-acetic acid oxidation by plant peroxidases: anaerobic stopped-flow spectrophotometric studies on horseradish and tobacco peroxidases. Biochem J 313(Pt 3):841–847
Gomez Ros LV, Aznar-Asensio GJ, Hernandez JA, Bernal MA, Nunez-Flores MJ, Cuello J, Ros Barcelo A (2007) Structural motifs of syringyl peroxidases are conserved during angiosperm evolution. J Agric Food Chem 55:4131–4138
Grand C, Parmentier P, Boudet A, Boudet A (1985) Comparison of lignins and of enzymes involved in lignification in normal and brown midrib (bm3) mutant corn seedlings. Physiol Veg 23:905–911
Halpin C, Holt K, Chojecki J, Oliver D, Chabbert B, Monties B, Edwards K, Barakate A, Foxon GA (1998) Brown-midrib maize (bm1)—a mutation affecting the cinnamyl alcohol dehydrogenase gene. Plant J 14:545–553
Harkin JM, Obst JR (1973) Lignification in trees: indication of exclusive peroxidase participation. Science 180:296–298
Hatfield R, Vermerris W (2001) Lignin formation in plants. The dilemma of linkage specificity. Plant Physiol 126:1351–1357
Herrero J, Esteban-Carrasco A, Zapata JM (2013a) Looking for Arabidopsis thaliana peroxidases involved in lignin biosynthesis. Plant Physiol Biochem 67:77–86
Herrero J, Fernandez-Perez F, Yebra T, Novo-Uzal E, Pomar F, Pedreno MA, Cuello J, Guera A, Esteban-Carrasco A, Zapata JM (2013b) Bioinformatic and functional characterization of the basic peroxidase 72 from Arabidopsis thaliana involved in lignin biosynthesis. Planta 237:1599–1612
Higuchi T (1985) Biosynthesis of lignin. Biosynthesis and biodegradation of wood components:141–160
Higuchi T (1997) Biochemistry and molecular biology of wood. Springer, Berlin
Higuchi T, Ito T, Umezawa T, Hibino T, Shibata D (1994) Red-brown color of lignified tissues of transgenic plants with antisense CAD gene: wine-red lignin from coniferyl aldehyde. J Biotechnol 37:151–158
Hiraga S, Sasaki K, Ito H, Ohashi Y, Matsui H (2001) A large family of class III plant peroxidases. Plant Cell Physiol 42:462–468
Holsters M, de Waele D, Depicker A, Messens E, van Montagu M, Schell J (1978) Transfection and transformation of Agrobacterium tumefaciens. Mol Gen Genet 163:181–187
Horvath L, Peszlen I, Peralta P, Kasal B, Li L (2010) Mechanical properties of genetically engineered young aspen with modified lignin content and/or structure. Wood Fiber Sci 42:310–317
Hu WJ, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD, Tsai CJ, Chiang VL (1999) Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nat Biotechnol 17:808–812
Imberty A, Goldberg R, Catesson A-M (1985) Isolation and characterization of Populus isoperoxidases involved in the last step of lignin formation. Planta 164:221–226
Kärkönen A, Koutaniemi S, Mustonen M, Syrjänen K, Brunow G, Kilpeläinen I, Teeri TH, Simola LK (2002) Lignification related enzymes in Picea abies suspension cultures. Physiol Plant 114:343–353
Kaur H, Shaker K, Heinzel N, Ralph J, Gális I, Baldwin IT (2012) Environmental stresses of field growth allow cinnamyl alcohol dehydrogenase-deficient Nicotiana attenuata plants to compensate for their structural deficiencies. Plant Physiol 159:1545–1570
Kavousi B, Daudi A, Cook CM, Joseleau JP, Ruel K, Devoto A, Bolwell GP, Blee KA (2010) Consequences of antisense down-regulation of a lignification-specific peroxidase on leaf and vascular tissue in tobacco lines demonstrating enhanced enzymic saccharification. Phytochemistry 71:531–542
Koch GW, Sillett SC, Jennings GM, Davis SD (2004) The limits to tree height. Nature 428:851–854
Koua D, Cerutti L, Falquet L, Sigrist CJ, Theiler G, Hulo N, Dunand C (2009) PeroxiBase: a database with new tools for peroxidase family classification. Nucleic Acids Res 37:D261–D266
Koutaniemi S, Toikka MM, Karkonen A, Mustonen M, Lundell T, Simola LK, Kilpelainen IA, Teeri TH (2005) Characterization of basic p-coumaryl and coniferyl alcohol oxidizing peroxidases from a lignin-forming Picea abies suspension culture. Plant Mol Biol 58:141–157
Koutaniemi S, Warinowski T, Karkonen A, Alatalo E, Fossdal CG, Saranpaa P, Laakso T, Fagerstedt KV, Simola LK, Paulin L, Rudd S, Teeri TH (2007) Expression profiling of the lignin biosynthetic pathway in Norway spruce using EST sequencing and real-time RT-PCR. Plant Mol Biol 65:311–328
Lagrimini LM, Burkhart W, Moyer M, Rothstein S (1987) Molecular cloning of complementary DNA encoding the lignin-forming peroxidase from tobacco: Molecular analysis and tissue-specific expression. Proc Natl Acad Sci U S A 84:7542–7546
Lagrimini LM, Vaughn J, Erb WA, Miller SA (1993) Peroxidase overproduction in tomato: wound-induced polyphenol deposition and disease resistance. HortSci 28:218–221
Lagrimini LM, Joly RJ, Dunlap JR, Liu T-TY (1997) The consequence of peroxidase overexpression in transgenic plants on root growth and development. Plant Mol Biol 33:887–895
Lapierre C, Pollet B, MacKay JJ, Sederoff RR (2000) Lignin structure in a mutant pine deficient in cinnamyl alcohol dehydrogenase. J Agric Food Chem 48:2326–2331
Li L, Zhou Y, Cheng X, Sun J, Marita JM, Ralph J, Chiang VL (2003a) Combinatorial modification of multiple lignin traits in trees through multigene cotransformation. Proc Natl Acad Sci U S A 100:4939–4944
Li Y, Kajita S, Kawai S, Katayama Y, Morohoshi N (2003b) Down-regulation of an anionic peroxidase in transgenic aspen and its effect on lignin characteristics. J Plant Res 116:175–182
Li Q, Min D, Wang JP-Y, Peszlen I, Horvath L, Horvath B, Nishimura Y, Jameel H, Chang H-M, Chiang VL (2011) Down-regulation of glycosyltransferase 8D genes in Populus trichocarpa caused reduced mechanical strength and xylan content in wood. Tree Physiol 31:226–236
Liu L, Dean JFD, Friedman WE, Eriksson K-EL (1994) A laccase-like phenoloxidase is correlated with lignin biosynthesis in Zinnia elegans stem tissues. Plant J 6:213–224
Liu J, Shi R, Li Q, Sederoff RR, Chiang VL (2012) A standard reaction condition and a single HPLC separation system are sufficient for estimation of monolignol biosynthetic pathway enzyme activities. Planta 236:879–885
Llorente F, Lopez-Cobollo RM, Catala R, Martinez-Zapater JM, Salinas J (2002) A novel cold-inducible gene from Arabidopsis, RCI3, encodes a peroxidase that constitutes a component for stress tolerance. Plant J 32:13–24
Lockhart J (2013) Breaking down the complex regulatory web underlying lignin biosynthesis. Plant Cell 25:4282
Lovrekovich L, Lovrekovich H, Stahmann MA (1968) Tobacco mosaic virus-induced resistance to Pseudomonas tabaci in tobacco. Phytopathology 58:1034–1035
Loziuk PL, Wang J, Li Q, Sederoff RR, Chiang VL, Muddiman DC (2013) Understanding the role of proteolytic digestion on discovery and targeted proteomic measurements using liquid chromatography tandem mass spectrometry and design of experiments. J Proteome Res 12:5820–5829
Lu G, Moriyama EN (2004) Vector NTI, a balanced all-in-one sequence analysis suite. Brief Bioinform 5:378–388
Lu S, Li Q, Wei H, Chang M-J, Tunlaya-Anukit S, Kim H, Liu J, Song J, Sun Y-H, Yuan L, Yeh T-F, Peszlen I, Ralph J, Sederoff RR, Chiang VL (2013) Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc Natl Acad Sci U S A 110:10848–10853
MacKay JJ, O’Malley DM, Presnell T, Booker FL, Campbell MM, Whetten RW, Sederoff RR (1997) Inheritance, gene expression, and lignin characterization in a mutant pine deficient in cinnamyl alcohol dehydrogenase. Proc Natl Acad Sci U S A 94:8255–8260
MacKay J, Presnell T, Jameel H, Taneda H, O’Malley D, Sederoff R (1999) Modified lignin and delignification with a CAD-deficient loblolly pine. In Holzforschung, p 403
Mader M (1980) Origin of the heterogeneity of peroxidase isoenzyme group Gr from Nicotiana tabacum. I. Conformation. Z Pflanzenphysiol 96:283–296
Marjamaa K, Kukkola E, Lundell T, Karhunen P, Saranpää P, Fagerstedt KV (2006) Monolignol oxidation by xylem peroxidase isoforms of Norway spruce (Picea abies) and silver birch (Betula pendula). Tree Physiol 26:605–611
Masuda H, Fukuda H, Komamine A (1983) Changes in peroxidase isoenzyme patterns during tracheary element differentiation in a culture of single cells isolated from the mesophyll of Zinnia elegans. Z Pflanzenphysiol 112:417–426
Matsui T, Tabayashi A, Iwano M, Shinmyo A, Kato K, Nakayama H (2011) Activity of the C-terminal-dependent vacuolar sorting signal of horseradish peroxidase C1a is enhanced by its secondary structure. Plant Cell Physiol 52:413–420
McDougall GJ (2001) Cell-wall-associated peroxidases from the lignifying xylem of angiosperms and gymnosperms: monolignol oxidation. Holzforschung 55:246–249
McIntyre CL, Bettenay HM, Manners JM (1996) Strategies for the suppression of peroxidase gene expression in tobacco. II. In vivo suppression of peroxidase activity in transgenic tobacco using ribozyme and antisense constructs. Transgenic Res 5:263–270
Meyermans H, Morreel K, Lapierre C, Pollet B, De Bruyn A, Busson R, Herdewijn P, Devreese B, Van Beeumen J, Marita JM, Ralph J, Chen C, Burggraeve B, Van Montagu M, Messens E, Boerjan W (2000) Modifications in lignin and accumulation of phenolic glucosides in poplar xylem upon down-regulation of caffeoyl-coenzyme a o-methyltransferase, an enzyme involved in lignin biosynthesis. J Biol Chem 275:36899–36909
Nakamura W (1967) Studies on the biosynthesis of lignin. I. Disproof against the catalytic activity of laccase in the oxidation of coniferyl alcohol. J Biochem 62:54–61
Nielsen KL, Indiani C, Henriksen A, Feis A, Becucci M, Gajhede M, Smulevich G, Welinder KG (2001) Differential activity and structure of highly similar peroxidases. Spectroscopic, crystallographic, and enzymatic analyses of lignifying Arabidopsis thaliana peroxidase A2 and horseradish peroxidase A2. Biochemistry 40:11013–11021
Novaes E, Kirst M, Chiang V, Winter-Sederoff H, Sederoff R (2010) Lignin and biomass: a negative correlation for wood formation and lignin content in trees. Plant Physiol 154:555–561
O’Malley DM, Whetten R, Bao W, Chen C-L, Sederoff RR (1993) The role of laccase in lignification. Plant J 4:751–757
Osakabe K, Koyama H, Kawai S, Katayama Y, Morohoshi N (1994) Molecular cloning and the nucleotide sequences of two novel cDNAs that encode anionic peroxidases of Populus kitakamiensis. J Plant Science 103:167–175
Osakabe K, Koyama H, Kawai S, Katayama Y, Morohoshi N (1995) Molecular cloning of two tandemly arranged peroxidase genes from Populus kitakamiensis and their differential regulation in the stem. Plant Mol Biol 28:677–689
Ostergaard L, Pedersen AG, Jespersen HM, Brunak S, Welinder KG (1998) Computational analyses and annotations of the Arabidopsis peroxidase gene family. FEBS Lett 433:98–102
Ostergaard L, Teilum K, Mirza O, Mattsson O, Petersen M, Welinder KG, Mundy J, Gajhede M, Henriksen A (2000) Arabidopsis ATP A2 peroxidase. Expression and high-resolution structure of a plant peroxidase with implications for lignification. Plant Mol Biol 44:231–243
Passardi F, Longet D, Penel C, Dunand C (2004) The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochemistry 65:1879–1893
Pillonel C, Mulder MM, Boon JJ, Forster B, Binder A (1991) Involvement of cinnamyl-alcohol dehydrogenase in the control of lignin formation in Sorghum bicolor L. Moench Planta 185:538–544
Polle A, Otter T, Seifert F (1994) Apoplastic peroxidases and lignification in needles of Norway Spruce (Picea abies L.). Plant Physiol 106:53–60
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842
Quiroga M, Guerrero C, Botella MA, Barcelo A, Amaya I, Medina MI, Alonso FJ, de Forchetti SM, Tigier H, Valpuesta V (2000) A tomato peroxidase involved in the synthesis of lignin and suberin. Plant Physiol 122:1119–1127
Ralph J, MacKay JJ, Hatfield RD, O’Malley DM, Whetten RW, Sederoff RR (1997) Abnormal lignin in a loblolly pine mutant. Science 277:235–239
Ralph J, Hatfield RD, Piquemal J, Yahiaoui N, Pean M, Lapierre C, Boudet AM (1998) NMR characterization of altered lignins extracted from tobacco plants down-regulated for lignification enzymes cinnamylalcohol dehydrogenase and cinnamoyl-CoA reductase. Proc Natl Acad Sci U S A 95:12803–12808
Ralph J, Lundquist K, Brunow G, Lu F, Kim H, Schatz PF, Marita JM, Hatfield RD, Ralph SA, Christensen JH (2004) Lignins: natural polymers from oxidative coupling of 4-hydroxyphenyl-propanoids. Phytochem Rev 3:29–60
Ranocha P, Chabannes M, Chamayou S, Danoun S, Jauneau A, Boudet AM, Goffner D (2002) Laccase down-regulation causes alterations in phenolic metabolism and cell wall structure in poplar. Plant Physiol 129:145–155
Ren L-L, Liu Y-J, Liu H-J, Qian T-T, Qi L-W, Wang X-R, Zeng Q-Y (2014) Subcellular relocalization and positive selection play key roles in the retention of duplicate genes of Populus class III peroxidase family. Plant Cell 26:2404–2419
Richardson A, McDougall GJ (1997) A laccase-type polyphenol oxidase from lignifying xylem of tobacco. Phytochemistry 44:229–235
Richardson A, Stewart D, McDougall GJ (1997) Identification and partial characterization of a coniferyl alcohol oxidase from lignifying xylem of Sitka spruce (Picea sitchensis). Planta 203:35–43
Richardson A, Duncan J, McDougall GJ (2000) Oxidase activity in lignifying xylem of a taxonomically diverse range of trees: identification of a conifer laccase. Tree Physiol 20:1039–1047
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Ros Barcelo A (1997) Lignification in plant cell walls. Int Rev Cytol 176:87–132
Saathoff AJ, Sarath G, Chow EK, Dien BS, Tobias CM (2011) Downregulation of cinnamyl-alcohol dehydrogenase in switchgrass by RNA silencing results in enhanced glucose release after cellulase treatment. PLoS One 6, e16416
Sangha AK, Davison BH, Standaert RF, Davis MF, Smith JC, Parks JM (2014) Chemical factors that control lignin polymerization. J Phys Chem B 118:164–170
Sarkanen KV, Ludwig CH (1972) Lignins: occurrence, formation, structure and reactions. Wiley-Interscience, New York
Sasaki S, Nishida T, Tsutsumi Y, Kondo R (2004) Lignin dehydrogenative polymerization mechanism: a poplar cell wall peroxidase directly oxidizes polymer lignin and produces in vitro dehydrogenative polymer rich in beta-O-4 linkage. FEBS Lett 562:197–201
Sasaki S, Baba K, Nishida T, Tsutsumi Y, Kondo R (2006) The cationic cell-wall-peroxidase having oxidation ability for polymeric substrate participates in the late stage of lignification of Populus alba L. Plant Mol Biol 62:797–807
Sasaki S, Nonaka D, Wariishi H, Tsutsumi Y, Kondo R (2008) Role of Tyr residues on the protein surface of cationic cell-wall-peroxidase (CWPO-C) from poplar: potential oxidation sites for oxidative polymerization of lignin. Phytochemistry 69:348–355
Sato Y, Sugiyama M, Komamine A, Fukuda H (1995) Separation and characterization of the isoenzymes of wall-bound peroxidase from cultured Zinnia cells during tracheary element differentiation. Planta 196:141–147
Sato Y, Bao W, Sederoff R, Whetten R (2001) Molecular cloning and expression of eight laccase cDNAs in loblolly pine (Pinus taeda). J Plant Res 114:147–155
Sato Y, Demura T, Yamawaki K, Inoue Y, Sato S, Sugiyama M, Fukuda H (2006) Isolation and characterization of a novel peroxidase gene ZPO-C whose expression and function are closely associated with lignification during tracheary element differentiation. Plant Cell Physiol 47:493–503
Sharma P, Dubey RS (2004) Ascorbate peroxidase from rice seedlings: properties of enzyme isoforms, effects of stresses and protective roles of osmolytes. Plant Sci 167:541–550
Shi R, Sun YH, Li Q, Heber S, Sederoff R, Chiang VL (2010) Towards a systems approach for lignin biosynthesis in Populus trichocarpa: transcript abundance and specificity of the monolignol biosynthetic genes. Plant Cell Physiol 51:144–163
Shigeto J, Kiyonaga Y, Fujita K, Kondo R, Tsutsumi Y (2013) Putative cationic cell-wall-bound peroxidase homologues in Arabidopsis, AtPrx2, AtPrx25, and AtPrx71, Are involved in lignification. J Agric Food Chem 61:3781–3788
Shuford CM, Sederoff RR, Chiang VL, Muddiman DC (2012) Peptide production and decay rates affect the quantitative accuracy of protein cleavage isotope dilution mass spectrometry (PC-IDMS). Mol Cell Proteomics 11:814–823
Sibout R, Eudes A, Mouille G, Pollet B, Lapierre C, Jouanin L, Seguin A (2005) Cinnamyl alcohol dehydrogenase-C and -D are the primary genes involved in lignin biosynthesis in the floral stem of Arabidopsis. Plant Cell 17:2059–2076
Smith CG, Rodgers MW, Zimmerlin A, Ferdinando D, Bolwell GP (1994) Tissue and subcellular immunolocalisation of enzymes of lignin synthesis in differentiating and wounded hypocotyl tissue of French bean (Phaseolus vulgaris L.). Planta 192:155–164
Song J, Lu S, Chen ZZ, Lourenco R, Chiang VL (2006) Genetic transformation of Populus trichocarpa genotype Nisqually-1: a functional genomic tool for woody plants. Plant Cell Physiol 47:1582–1589
Sterjiades R, Dean JF, Eriksson KE (1992) Laccase from sycamore maple (Acer pseudoplatanus) polymerizes monolignols. Plant Physiol 99:1162–1168
Studer MH, DeMartini JD, Davis MF, Sykes RW, Davison B, Keller M, Tuskan GA, Wyman CE (2011) Lignin content in natural Populus variants affects sugar release. Proc Natl Acad Sci U S A 108:6300–6305
Takahama U (1995) Oxidation of hydroxycinnamic acid and hydroxycinnamyl alcohol derivatives by laccase and peroxidase. Interactions among p-hydroxyphenyl, guaiacyl and syringyl groups during the oxidation reactions. Physiol Plant 93:61–68
Tamura T, Morohoshi N, Yasuda S (2001) Suitability of peroxidase-suppressed transgenic hybrid aspen (Populus sieboldii X Populus gradidentata) for Pulping. In Holzforschung, p 335
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tognolli M, Penel C, Greppin H, Simon P (2002) Analysis and expression of the class III peroxidase large gene family in Arabidopsis thaliana. Gene 288:129–138
Tokunaga N, Kaneta T, Sato S, Sato Y (2009) Analysis of expression profiles of three peroxidase genes associated with lignification in Arabidopsis thaliana. Physiol Plant 136:237–249
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25:1105–1111
Tsutsumi Y, Matsui K, Sakai K (1998) Substrate-specific peroxidases in woody angiosperms and gymnosperms participate in regulating the dehydrogenative polymerization of syringyl and guaiacyl type lignins. In Holzforschung, p 275
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Dejardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjarvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leple JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouze P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, Van de Peer Y, Rokhsar D (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Van Acker R, Leplé J-C, Aerts D, Storme V, Goeminne G, Ivens B, Légée F, Lapierre C, Piens K, Van Montagu MCE, Santoro N, Foster CE, Ralph J, Soetaert W, Pilate G, Boerjan W (2014) Improved saccharification and ethanol yield from field-grown transgenic poplar deficient in cinnamoyl-CoA reductase. Proc Natl Acad Sci U S A 111:845–850
Vance C, Kirk T, Sherwood R (1980) Lignification as a mechanism of disease resistance. Annu Rev Phytopathol 18:259–288
Vignols F, Rigau J, Torres MA, Capellades M, Puigdomènech P (1995) The brown midrib3 (bm3) mutation in maize occurs in the gene encoding caffeic acid O-methyltransferase. Plant Cell 7:407–416
Voelker SL, Lachenbruch B, Meinzer FC, Jourdes M, Ki C, Patten AM, Davin LB, Lewis NG, Tuskan GA, Gunter L, Decker SR, Selig MJ, Sykes R, Himmel ME, Kitin P, Shevchenko O, Strauss SH (2010) Antisense down-regulation of 4CL expression alters lignification, tree growth, and saccharification potential of field-grown poplar. Plant Physiol 154:874–886
Wallace G, Fry SC (1999) Action of diverse peroxidases and laccases on six cell wall-related phenolic compounds. Phytochemistry 52:769–773
Wang JP, Naik PP, Chen H-C, Shi R, Lin C-Y, Liu J, Shuford CM, Li Q, Sun Y-H, Tunlaya-Anukit S, Williams CM, Muddiman DC, Ducoste JJ, Sederoff RR, Chiang VL (2014) Complete proteomic-based enzyme reaction and inhibition kinetics reveal how monolignol biosynthetic enzyme families affect metabolic flux and lignin in Populus trichocarpa. Plant Cell 26:894–914
Weng JK, Chapple C (2010) The origin and evolution of lignin biosynthesis. New Phytol 187:273–285
Weng J-K, Li X, Bonawitz ND, Chapple C (2008) Emerging strategies of lignin engineering and degradation for cellulosic biofuel production. Curr Opin Biotechnol 19:166–172
Whetten RW, MacKay JJ, Sederoff RR (1998) Recent advances in understanding lignin biosynthesis. Annu Rev Plant Physiol Plant Mol Biol 49:585–609
Xu B, Escamilla-Treviño LL, Sathitsuksanoh N, Shen ZX, Shen H, Zhang PYH, Dixon RA, Zhao BY (2011) Silencing of 4-coumarate:coenzyme A ligase in switchgrass leads to reduced lignin content and improved fermentable sugar yields for biofuel production. New Phytol 192:611–625
Yeh TF, Yamada T, Capanema E, Chang HM, Chiang V, Kadla JF (2005) Rapid screening of wood chemical component variations using transmittance near-infrared spectroscopy. J Agric Food Chem 53:3328–3332
Zhang K, Qian Q, Huang Z, Wang Y, Li M, Hong L, Zeng D, Gu M, Chu C, Cheng Z (2006) Gold hull and internode2 encodes a primarily multifunctional cinnamyl-alcohol dehydrogenase in rice. Plant Physiol 140:972–983
Zhao Q, Nakashima J, Chen F, Yin Y, Fu C, Yun J, Shao H, Wang X, Wang Z-Y, Dixon RA (2013) Laccase is necessary and nonredundant with peroxidase for lignin polymerization during vascular development in Arabidopsis. Plant Cell 25:3976–3987
Zimmerlin A, Wojtaszek P, Bolwell GP (1994) Synthesis of dehydrogenation polymers of ferulic acid with high specificity by a purified cell-wall peroxidase from French bean (Phaseolus vulgaris L.). Biochem J 299:747–753
Acknowledgments
The National Science Foundation (USA), Plant Genome Research Program (Grant DBI-0922391) to V.L.C., the NC State University Jordan Family Endowment, NIH/NCSU Molecular Biotechnology Training Program (Grant 5T32GM00-8776-08), and the NC State University Forest Biotechnology Industrial Research Consortium supported this work.
Author contributions
C.Y.L. generated the PtrPO transgenic plants. Q.L. prepared the RNAi constructs and RNAi libraries. R.S. and Y.H.S. selected the peroxidase candidate. C.Y.L. and S.T.-A. performed the transcriptome analysis. C.Y.L. and J.L. performed the lignin and cell wall composition analysis. P.L. and D.C.M. performed the PC-IDMS protein quantification. C.Y.L., C.W.E. and Z.D.M. analyzed the modulus of elasticity of the wood samples. C.Y.L., J.P.W., I.P., R.R.S. and V.L.C. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We do not have any conflict of interest to report.
Data archiving statement
The sequence of class III peroxidases for Arabidopsis thaliana (AtPrx) and Populus trichocarpa (PtrPO) reported here are achieved and publicly available at the PeroxiBase database (http://peroxibase.toulouse.inra.fr/). The genome information of P. trichocarpa is available in Phytozome (http://www.phytozome.org). The accession numbers for plant peroxidases in National Center for Biotechnology Information (NCBI; https://www.ncbi.nlm.nih.gov/) are available as follows: TPX1 (L13654), FBP1 (AF149277), ZPO-C (AB023959), ZePrx (AJ880392), TP60 (AF149251), ATP-A2 (X99952), prxA3a (Q43049), PXP3-4 (X97350), and CPWPO-C (AB210901). The accession numbers for PtrPOs are provided in Supplementary Material as Table S2.
Additional information
Communicated by A. Brunner
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 1127 kb)
Rights and permissions
About this article
Cite this article
Lin, CY., Li, Q., Tunlaya-Anukit, S. et al. A cell wall-bound anionic peroxidase, PtrPO21, is involved in lignin polymerization in Populus trichocarpa . Tree Genetics & Genomes 12, 22 (2016). https://doi.org/10.1007/s11295-016-0978-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-016-0978-y