Abstract
Electron Tomography (ET) is a powerful three-dimensional (3D) imaging technique used in structural biology and biomedicine to allow the visualization of the interior of cells at close-to-molecular resolution. Interpretation of the 3D volumes in ET is usually challenging due to the complexity of the cellular environment, noise conditions and other factors. Automated segmentation methods focused on membranes of the cells and organelles greatly facilitate visualization and interpretation of the 3D volumes. However, they are typically computationally expensive and spend significant processing time on standard computers. In this work, we introduce efficient implementations of one of the methods most commonly used in the ET field for membrane segmentation. They were developed by using High Performance Computing (HPC) techniques to make the most of modern CPU-based and GPU-based computing platforms. A thorough evaluation of the performance on state-of-the-art machines was carried out. The HPC implementations succeed in achieving remarkable speedups, which are around \(100\times\) on GPUs, and making it possible to process large 3D volumes in the order of seconds or a few minutes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Electron Tomography (ET) has emerged as a powerful technique in structural biology and biomedicine for three-dimensional (3D) visualization of the subcellular architecture at the nanometer scale [1]; furthermore, Cryo-ET faithfully preserves structures beyond nanometer scale by rapid freezing the sample. This technique relies on the same principles as Computed Tomography (CT) usually employed in Medicine [2]. In ET, a series of electron microscopy images is acquired from a specimen at different orientations around a single axis. These images are then combined by means of tomographic reconstruction methods to yield the 3D volume, which is then visualized and analyzed [3].
One essential stage for the interpretation of the reconstructed 3D volume is the segmentation into its constitutive structural components. However, such segmentation proves to be challenging because of a number of factors such as the molecular crowding often found in the cellular environment, artefacts inherent to the ET technique and the low signal-to-noise ratio (SNR) [3]. Thus, segmentation is still a major bottleneck in ET.
Therefore, there is compelling need for automated segmentation methods that facilitate the interpretation of the overwhelming structural information available in the 3D volumes in ET [3,4,5]. There have been numerous attempts to develop segmentation methods in the ET field (e.g. based on template matching, watershed) [3, 6, 7]. However, none has produced results of general applicability, and thus, manual segmentation is still a common choice. Recently, deep-learning techniques have emerged with promising prospects [8,9,10,11]. Nevertheless, they are characterized by enormous computational demands, the need for availability of enough training data and some expert knowledge is still required to make the most of them. This fact is limiting their practical applicability across the community of biologists in the ET field.
Membranes constitute the natural boundaries of cells and the organelles therein, so they turn out to be an ideal target for segmentation [12,13,14,15]. A few years ago, we developed a robust method for membrane segmentation [14] that is being very successfully used in ET [16, 17] and provides a basis for further quantitative analysis of membranous structures [5, 18,19,20]. The method provides useful solutions even under very low SNR scenarios. Nonetheless, it may be slow when dealing with huge 3D volumes, as typically obtained in ET.
In this work, we have used High Performance Computing (HPC) techniques to develop efficient implementations of the membrane segmentation method with the aim to take full advantage of the resources available in modern multi-core processors and GPUs and provide solutions in reasonable time. Both types of platforms are representative examples of HPC machines that are currently available in structural biology laboratories.
2 Membrane segmentation with steerable tensor voting
2.1 Membrane segmentation and tensor voting
Our robust method for membrane segmentation relies on a Gaussian model for membrane profile and a local structure detector based on the Hessian tensor that finds potential membrane-like features [14]. To reduce the noise and ensure the Gaussian profile of membranes, the original tomogram is subjected to Gaussian filtering using a standard deviation according to the thickness of the membrane to detect [12]. The local detector is then applied to the Gaussian-filtered tomogram.
One key aspect for the efficiency of the local structure detector is that the method is to be applied to two-dimensional (2D) planes of the 3D volumes. This is supported by the fact that membranes in 3D volumes appear as curves in 2D planes [12]. This greatly simplifies the complexity with respect to a pure 3D implementation of the whole procedure, in particular the tensor voting algorithm that is described below [14].
Therefore, for a given 2D plane, the Hessian tensor is constructed from the second order derivatives and can act as a local curve detector from its eigenvalues (\(\lambda _{1}\) and \(\lambda _{2}\), with \(\vert \lambda _{1}\vert \ge \vert \lambda _{2}\vert \ge 0\)), and the corresponding eigenvectors (\(\vec {v}_{1}\) and \(\vec {v}_{2}\)):
where \(\hbox {t}_{\mathrm{xx}} = \frac{\partial ^{2}L}{\partial x^{2}}\), \(\hbox {t}_{\mathrm{yy}} = \frac{\partial ^{2}L}{\partial y^{2}}\) and \(\hbox {t}_{\mathrm{xy}} = \frac{\partial ^{2}L}{\partial x\partial y}\) are the second order derivatives with respect to the axes x and y of the 2D plane, and L denotes the 2D plane from the Gaussian-filtered tomogram.
The first eigenvector \(\vec {v}_{1}\), that is, the one whose eigenvalue exhibits the largest value in absolute value \(\vert \lambda _{1}\vert\), points to the direction of the maximum variation. In a point belonging to a 2D curve, this direction is the normal to the curve. Accordingly, the second eigenvector \(\vec {v}_{2}\) points to the tangent of the curve. Consequently, a local detector can be derived from the eigenvectors and eigenvalues of the Hessian tensor [14].
Therefore, voxels belonging to a local curve have \(\vert \lambda _{1}\vert>>\vert \lambda _{2}\vert\), with \(\vec {v}_{1}\) in the direction perpendicular to the curve, and the term \(\vert \lambda _{1}-\lambda _{2}\vert\) represents the curve saliency (i.e. the likelihood of a voxel to belong to a curve). The orientation of \(\vec {v}_{1}\) with respect to the X axis is given by \(\arccos {(\vec {v}_{1}\cdot {\hat{e}}_{x})}\). As a consequence, a tensor field is obtained where all voxels are described by their saliency and orientation:
This information, \(S_{\mathrm{in}}({\mathbf {x}})\) and \(\alpha _{\mathrm{in}}({\mathbf {x}})\), represents the input tensor field that will be fed to the following stage.
Unfortunately, the performance of local detectors is limited because they are susceptible to artefacts and noise, thereby producing gaps or false positives. Therefore, procedures that provide robustness to the local detection are needed. For that purpose, we use the Tensor Voting algorithm [14, 21].
Tensor Voting (TV) allows anisotropic propagation of the local structural information derived from Hessian-tensor [14, 21] and encoded by \(S_{\mathrm{in}}({\mathbf {x}})\) and \(\alpha _{\mathrm{in}}({\mathbf {x}})\). In this process, the local structure at each voxel is refined according to the information received from neighbour voxels. As a result, voxels belonging to the same membrane will finally end up with coherent structural information, thereby strengthening the underlying global structure. The resulting 3D map represents how well every point in the tomogram fits a membrane model. Figure 1 illustrates the TV algorithm. Figure 2 shows application of the whole procedure to an experimental 3D volume.
Tensor Voting in 2D. A Model for vote casting. Votes include information about saliency and orientation. The voter at the origin \(\mathbf {O}\) is shown with its normal in green. The voxel \(\mathbf {x}\) is the receiver. The dashed arc represents the osculating circle passing through \(\mathbf {O}\) and \(\mathbf {x}\), which is the most likely smooth path between the two points. The vote cast from \(\mathbf {O}\) to \(\mathbf {x}\) is shown in red. Note that it turns out to be a transformed version of the normal at \(\mathbf {O}\) (in green) following the smooth path connecting \(\mathbf {O}\) and \(\mathbf {x}\). B 2D voting field, which is the collection of the votes cast by a voter located at the origin and has unit saliency and orientation along the x-axis. The centre of the field is placed at the origin, and the normal runs along the Y axis. The \(\infty\)-shape encompasses the votes with most significant saliency. C Tensor voting mechanism. For each voxel, the voting field is placed at its position, with the orientation of its normal. Then, the votes (red dotted arrows) are cast to all voxels in the neighbourhood. The procedure is repeated for all voxels in the plane, as sketched here for two voxels (left and right panels). At the end of the voting process, voxels belonging to a perceptual feature (e.g. the solid black curve here) will have been strengthened each other, hence enhancing the feature. The other voxels will have received divergent information, which will smear them out
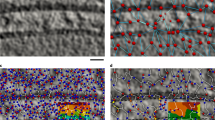
Application of membrane segmentation based on Hessian-tensor local detector and Tensor Voting on an experimental ET volume containing HIV-1 virions [22]. From left to right: a slice of the original volume, result from the Gaussian filtering operation and the resulting membrane detection (saliency). Right-most panel: 3D visualization of the segmented membranes, obtained by a simple thresholding operation on the saliency followed by extraction of connected components. The open membranes at the top/bottom of the volumes are features inherent to the ET technique
2.2 Steerable tensor voting
Tensor voting is a computationally demanding procedure. The standard implementation consists of pre-computing and storing the voting field [21] (Fig. 1(B)). Translation and rotation of the voting field throughout the image space is needed for casting votes, which is done by interpolation (Fig. 1(C)). There exists, however, a more efficient implementation that takes advantage of the theory of steerable filters [23]. A steerable filter is a filter that can be oriented in an arbitrary direction just by a linear combination of a finite number of predefined rotations of the filter (so-called basis functions or filters) [24]. If the number of basis filters is sufficiently small, this turns out to be a very efficient strategy for arbitrary oriented filtering of images.
The following steerable expression for the TV algorithm was derived to yield the final, refined saliency \(S_{\mathrm{out}}({\mathbf {x}})\) representing the likelihood of a voxel to belong to a curve. The derivation details can be found elsewhere [14, 23]:
where \(k_{m}\) are the linear coefficients:
and \(V_{m}({\mathbf {x}})\) are the basis filters given by:
where \(\gamma _{m}\) has constant values: \(\{1,4,6,4,1\}\) for \(m=0\dots 4\), respectively. \(\sigma _{v}\) denotes the length scale of the analysis, which determines the effective neighbourhood size (expressed in voxels).
As a result, the TV algorithm is reduced to just five convolutions followed by a linear combination. Moreover, computation of these convolutions in Fourier space speeds up the calculation significantly [23].
Therefore, the robust method for membrane segmentation based on steerable TV consists in two steps. First, local curve descriptors are calculated, which encode saliency and orientation of the local curve for each point. This is followed by an efficient TV algorithm that propagates the local information among neighbours so that membrane information from points belonging to the same underlying feature are strengthening each other. This process is applied to the 2D planes of the 3D tomograms along the three major axes: first, along the Z axis (i.e. XY planes), then along the Y axis (XZ planes), finally along X axis (YZ planes). The final output saliency for each voxel is taken as the average of the three curve saliency values available [14].
3 HPC Implementations
3.1 Membrane segmentation with steerable tensor voting in fourier space
The method for 3D membrane segmentation comprises three rounds of 2D curve segmentation by using the 2D steerable TV algorithm. In each round, the volume is swept across one of the three major axes (X, Y or Z) and the individual \(N_p\) 2D planes, with coordinates denoted by (x, y), are then processed. This 2D TV algorithm consists of the linear combination of five convolutions, as given by Equation 4. These convolutions are performed in Fourier space, which can be mathematically expressed as:
where \({\mathcal {F}}\) and \({\mathcal {F}}^{-1}\) denote direct and inverse Fourier transforms (FT), respectively. This expression clearly shows that ten direct FTs and one inverse FT are required to process a single 2D plane. It is important to note that the basis filters \(V_{m}({\mathbf {x}})\) (Equation 6) do not depend on the actual density values present in the 2D planes. Instead, the computation of the basis filters only depend on the coordinates (x, y). As a consequence, the five basis filters \(V_{m}({\mathbf {x}})\), with \(m=0\dots 4\) and their FT can be precomputed and re-used for all 2D planes along an axis. This reduces the computation involved for a 2D plane to five direct FTs and one inverse FT, together with the computation of the saliency \(S_{\mathrm{in}}({\mathbf {x}})\), orientation \(\alpha _{\mathrm{in}} ({\mathbf {x}})\) and linear coefficients \(k_{m}(\alpha _{\mathrm{in}} ({\mathbf {x}}))\), with \(m=0\dots 4\). In order to use the most optimized library for FT calculations, we used the FFTW [25] compiled to use vector instructions currently available in current CPU processors, and the CUDA CuFFT library optimized for NVIDIA GPUs.
3.2 Multithreaded CPU implementation
Modern computers are equipped with multi-core processors [26]. The use of multithreading techniques turns out to be important in ET as they make it possible to fully exploit the computational capabilities of state-of-the-art computers and reduce the typically long processing time of the image processing procedures in this field. These techniques have been paramount to accelerate tomographic reconstruction and denoising methods, among others [4, 27,28,29,30,31].
Multithreaded implementation of the steerable Tensor Voting algorithm. The volume is swept across the direction of one of the major axes (X, Y or Z). Let us denote \(N_p\) the number of planes in that direction, with local coordinates (x, y) within each plane. The 2D planes are distributed across the multiple \(N_t\) threads running in parallel. The basis filters can be computed only once and their Fourier components are shared by all threads. The processing of each individual 2D plane consists of the computation of the Hessian tensor, from which the local saliency and orientation are obtained. This is followed by the five convolutions computed in Fourier space, as described in the main text. This multithreaded implementation is run three times: first, by sweeping the 2D planes in the direction of the Z axis of the volume, then in the Y direction and finally X
To make the most of the power of modern multi-core computers, we have developed a multithreaded implementation of the steerable TV algorithm using POSIX Threads (PThreads) [32]. The \(N_p\) 2D planes of the volume along an axis are distributed across the multiple threads so that they can be processed in parallel. Within each thread, its subset of 2D planes is processed sequentially, one after the other, by running the steerable TV algorithm in Fourier space, as described above. The processing involved in a single plane thus consists of the computation of the Hessian tensor, from which the local saliency and orientation are obtained, followed by the five convolutions performed in Fourier space. Note that the basis filters \(V_{m}({\mathbf {x}})\) and their FTs are computed only once, and they are shared by all threads. Figure 3 sketches this multithreaded implementation. Note that each 2D plane has to be extracted from the input volume before its processing and, once processed, the result has to be inserted into the output volume.
3.3 GPU implementation
Most HPC platforms and modern servers include GPUs to accelerate specific procedures (kernels) which fit with the SIMT programming model. CUDA (Compute Unified Device Architecture) is a well-known parallel interface developed by NVIDIA to program such devices. In the CUDA programming model, the CPU performs a succession of kernels invocations to accelerate the corresponding computation on GPU. The input/output data of the GPU kernels is communicated between the CPU and the ‘global’ GPU memories. Successive generations of NVIDIA GPUs have increased resources and features supported by their hardware (Compute Capability). For example, asynchronous concurrent kernels/streams executions are supported on GPUs with Compute Capability 3.5 and higher.
The parallel steerable TV of 3D volume in every spatial dimension (X, Y and Z) can be organised by 2D planes without synchronisation points. To get a high acceleration on GPUs the first step is to extract every 2D plane to store it on the GPU memory. This way, the memory accesses to process the extracted plane on GPU are almost fully coalescent, therefore the corresponding computation is very efficient. When it finishes, the resulting segmented plane is inserted into the 3D data structure.
CuTV-Planes, a GPU implementation of the steerable TV algorithm. The volume is swept across the direction of one of the major axes (X, Y or Z). Let us denote \(N_p\) the number of planes in that direction, with local coordinates (x, y) within each plane. CuTV-Planes communicates the input/output volume between GPU-CPU by planes. \(N_t\) CPU threads are spawned, each of them extracts one plane of the volume, sends it to GPU and controls one GPU stream. \(N_t\) GPU streams concurrently launch the processing of \(N_t\) planes on GPU. Each stream sends the output plane to the CPU
Two GPU implementations have been developed according to two different communications schemes:
In the GPU version named CuTV-Planes, the CPU-GPU communication is organised by planes. A set of \(N_t\) CPU threads are created to process a subset of planes with the same distribution of planes as the multithreaded implementation. Every CPU-thread extracts one plane from the volume data, creates a GPU-Stream and sends the plane to the GPU memory. Then, the sequence of GPU kernels is executed to process the plane stored on the GPU memory with the steerable TV. When it finishes, the CPU receives a processed plane and inserts it into the segmented volume. This procedure is completed for all planes in the subset assigned to every CPU-thread/GPU-Stream. This way, the computation of every plane is accelerated on GPU and the planes of different streams are concurrently processed on GPU, as shown in Fig. 4. Therefore, to allow the concurrent processing, it is necessary to store \(N_t\) planes on GPU memory, whose content is updated as computing advances by GPU-CPU communications. Therefore, the GPU memory requirements of CuTV-Planes linearly depends on the number of GPU-Streams activated (\(N_t\)) and it can be easily adapted to the available memory of different GPUs using this parameter.
CuTV-Volume is the second GPU implementation of the steerable Tensor Voting algorithm. The whole volume is transferred between CPU-GPU with only two communications. The GPU extracts, processes and inserts the results in the device memory. CuTV-Volume processes asynchronously all planes in the volume, in the same way as CUTV-Planes
The second GPU version, named CuTV-Volume, creates only one CPU-thread. It starts with the communication of the whole volume to the GPU memory. Then, the GPU extracts planes from the volume as it is swept across the axes. Memory coalescing during these operations is maintained. The X axis (YZ planes) is the most challenging, as adjacent voxels are stored far apart in memory. Coalesced memory access is maintained by performing plane operations in batches of 32 planes (the CUDA warp size) and using block shared memory. Next, a sequence of kernels is launched to process the stages of the steerable TV for each plane. Therefore, the computation for all planes in the volume is asynchronously processed in parallel on GPU. As every plane is segmented, it is inserted into the GPU data structure to store the volume. When the GPU concludes, the processed volume is communicated from GPU to CPU memory. Figure 5 represents this process. CuTV-Volume can efficiently accelerate the computation on GPU. However, as counterpart, it is necessary to store the whole volume twice on the GPU memory. This could be a serious drawback when the memory requirements to store the volume are higher than the available GPU memory.
Application of membrane segmentation based on Hessian-tensor local detector and Tensor Voting to dataset EMD-3977. A slice of the original volume A, result from the Gaussian filtering operation B, the resulting membrane detection -saliency- C, and 3D visualization of the segmented membranes by a simple thresholding operation on the saliency D. Note that the contrast in this tomogram (black foreground over a lighter background, see panels A and B) is the opposite as that in Fig. 2
4 Results
4.1 Datasets
The HPC implementations of the TV-based membrane segmentation method were evaluated using datasets from the public databases Electron Microscopy Data Resource (EMD, http://emdataresource.org) [33] and Electron Microscopy Public Image Archive (EMPIAR, http://www.ebi.ac.uk/empiar) [34].
Two datasets, denoted by EMD-3977 and EMPIAR-10442, with representative sizes of the current structural studies from different modalities of ET were used. They were obtained from Chlamydomonas reinhardtii [35] and Arabidopsis thaliana [36], which are model organisms in biological studies. The 3D volumes had sizes of \(928\times 928 \times 464\) and \(2596 \times 1731 \times 717\) voxels (around 1.5 GB and 12 GB size using single precision floating point numbers), respectively. Figure 6 illustrates the result of the membrane segmentation method applied to the dataset EMD-3977.
4.2 Evaluation platforms
To evaluate the performance of the multicore and GPU implementations, two representative HPC platforms have been selected. The first platform contains two AMD EPYC 7642 processors from the microarchitecture Zen 2 (launched in 2019) for a total of 96 CPU cores, 512 GB of DDR4 RAM at 3200 MHz and one NVIDIA Tesla V100 (32 GB) of the microarchitecture Volta (launched in 2017). This platform is a good example of the compute nodes used in modern HPC clusters. The second platform contains two Intel Xeon E5-2620v3 from the microarchitecture Haswell (launched in 2013) for a total of 12 CPU cores, 64 GB of DDR3 RAM at 1866 MHz and one NVIDIA Kepler K80 (12 GB) of the microarchitecture Kepler (launched in 2012). Although this platform is slightly outdated by HPC standards, it performs akin to current high-end desktop computers. Therefore, it is a good benchmark for the real-world performance of our implementations.
4.3 Experimental evaluation
The HPC implementations have been applied to the two test datasets on the computing platforms. The global runtime and, if applicable, the CPU-GPU communication times have been measured under different configurations and the speedup has been computed. The results are shown in Tables 1 and 2 for the datasets EMD-3977 and EMPIAR-10442, respectively.
The multithreaded version yields monotonically increasing speedup factors as a function of the number of threads, with a remarkable maximum value approaching \(45 \times\) in the AMD platform. The tables also show that the speedup moves away from the ideal linear behaviour, particularly for high number of threads, especially beyond 16 cores on the AMD platform. The Intel platform shows similar speedup values, but limited up to 12 threads (maximum number of cores). Interestingly, it is not observed a significant influence of the volume size on the acceleration obtained on these multi-core platforms. Thus, the processing of both datasets brings similar speedup factors in general, though with some decrease in the case of EMPIAR-10442 on the AMD platform.
The GPU implementations achieve outstanding acceleration factors that, overall, outperform the multi-core implementation using the largest amounts of threads. This is particularly striking in the case of the Tesla V100 GPU. Tables 1 and 2 demonstrate that the CuTV-Volume version is faster than CuTV-Planes, reaching speedups higher than \(100 \times\) for both test cases on Tesla V100. Although plane extraction on CPU is significantly slower than on GPU, CuTV-Planes keeps the pace better than expected, as it leverages both the CPU and GPU computing power and overlaps memory transfers and computation, taking advantage of the multiple copy and kernel engines available on current GPUs.
Also noteworthy is that the superiority of CuTV-Volume comes at the expense of significant memory consumption, which may turn out to be a limiting factor for its applicability. This is the case for the largest dataset (EMPIAR-10442), where application of the CuTV-Volume version to run on the Tesla K80 GPU was not possible.
The dataset EMD-3977 is representative of the sizes most widely used now in the ET field for segmentation. Table 1 indicates that multithreading reduces the processing time to less than a minute beyond 8 cores. This is an important result because it suggests that current datasets can be processed efficiently in standard desktop/laptop computers. Moreover, the use of GPU computing allows further reduction of the processing time to just seconds.
The dataset EMPIAR-10442 can be considered as an example of the sizes that are expected in the short term, owing to the increasing resolution demands. Table 2 demonstrates that these 3D volumes can be processed in a matter of 5-10 minutes in standard computers equipped with 4-8 CPU cores. These large volumes are especially well suited for exploitation of GPUs, as corroborated by the exceptional acceleration factors obtained in both GPUs tested. Therefore, depending upon the GPU architecture, there is potential to process these volumes even in less than a minute.
5 Conclusions
We have presented and evaluated efficient implementations of a membrane segmentation method for their application to large 3D volumes in structural studies by electron tomography. The implementations rely on the steerable Tensor Voting algorithm performed in Fourier space as well as the use of HPC techniques to exploit CPUs and GPUs. First, multithreading techniques have been used to make the most of the state-of-the-art CPU-based multicore processors. Second, we have further elaborated the implementation to exploit the fine-grained parallelism levels in the advanced GPU architectures, and we have developed two GPU versions with different memory demands. All HPC implementations faithfully proceed as the original sequential version and reproduce the same segmentation results.
Outstanding acceleration rates, even reaching 45-\(100\times\), have been obtained on powerful platforms equipped with substantial number of CPU cores or on modern GPUs. Remarkably, our results demonstrate that our implementations allow segmentation of membranes present in 3D volumes of representative size in a matter of seconds or a few minutes, even with standard computers equipped with relatively modest number of CPU cores (4-8).
The GPU implementations that we have presented are particularly interesting. Both versions have demonstrated capabilities to achieve high acceleration factors. The one that maintains the whole volume in the GPU memory shows an exceptional performance, with speedup values around 100\(\times\), but its application may be restricted to high-end GPUs. The GPU version working on a plane-basis obtains lower speedup values, with the advantage that the memory demands are limited to those required for processing a relatively small subset of planes. This memory consumption ensures its practical applicability in a wide range of GPUs, even modest ones. The availability of the two GPU versions makes our program versatile in the sense that, depending on the memory demands and the GPU platform, the proper version is selected.
The speed of our implementations paves the way for running the method on standard desktop/laptop computers, which are the machines usually available in most laboratories of life sciences. Moreover, this implementation will facilitate the processing of the huge 3D volumes (e.g. 4096x4096x2048 or larger) that will shortly be required by the increasing resolution needs in the electron tomography field. Our future plans include to exploit other parallelism levels of the algorithm and explore hybrid implementations that jointly take advantage of CPU and GPUs available in the computing platforms.
References
Turk M, Baumeister W (2020) The promise and the challenges of cryo-electron tomography. Federation Eur Biochem Soc (FEBS) Lett 594:3243–3261
Herman GT (2009) Image reconstruction from projections: the fundamentals of computerized tomography, 2nd edn. Springer, London
Fernandez JJ (2012) Computational methods for electron tomography. Micron 43:1010–1030
Moreno JJ et al (2018) Tomoeed: Fast edge-enhancing denoising of tomographic volumes. Bioinformatics 34:3776–3778
Martinez-Sanchez A et al (2020) Template-free detection and classification of membrane-bound complexes in cryo-electron tomograms. Nat Method 17:209–216
Tasel SF et al (2016) A validated active contour method driven by parabolic arc model for detection and segmentation of mitochondria. J Struct Biol 194:253–271
Luengo I et al (2017) SuRVoS: Super-region volume segmentation workbench. J Struct Biol 198:43–53
Chen M et al (2017) Convolutional neural networks for automated annotation of cellular cryo-electron tomograms. Nat Methods 14:983–985
Li R et al (2019) Automatic localization and identification of mitochondria in cellular electron cryo-tomography using faster-RCNN. BMC Bioinformatics 20(Suppl 3):132
Fischer CA et al (2020) MitoSegNet: Easy-to-use deep learning segmentation for analyzing mitochondrial morphology. iScience 23(10):101601
Moebel E et al (2021) Deep learning improves macromolecule identification in 3D cellular cryo-electron tomograms. Nat Methods 18:1386–1394
Martinez-Sanchez A et al (2011) A differential structure approach to membrane segmentation in electron tomography. J Struct Biol 175:372–383
Martinez-Sanchez A et al (2013) A ridge-based framework for segmentation of 3D electron microscopy datasets. J Struct Biol 181:61–70
Martinez-Sanchez A et al (2014) Robust membrane detection based on tensor voting for electron tomography. J Struct Biol 186:49–61
Page C, Hanein D, Volkmann N (2015) Accurate membrane tracing in three-dimensional reconstructions from electron cryotomography data. Ultramicroscopy 155:20–26
Fernandez-Fernandez MR et al (2017) 3D electron tomography of brain tissue unveils distinct Golgi structures that sequester cytoplasmic contents in neurons. J Cell Sci 130:83–89
Chaikeeratisak V et al (2019) Viral capsid trafficking along treadmilling tubulin filaments in bacteria. Cell 177(7):1771–1780
Bäuerlein FJB et al (2017) In situ architecture and cellular interactions of polyq inclusions. Cell 171(1):179–187
Guo, Q.,. et al (2018) In situ structure of neuronal C9orf72 Poly-GA aggregates reveals proteasome recruitment. Cell 172(4):696–705
Salfer M et al (2020) Reliable estimation of membrane curvature for cryo-electron tomography. PLoS Comput Biol 16:1007962
Medioni G, Lee MS, Tang CK (2000) A Computational Framework for Segmentation and Grouping. Elsevier
Briggs JA et al (2006) The mechanism of hiv-1 core assembly: insights from three-dimensional reconstructions of authentic virions. Structure 14:15–20
Franken E et al (2006) An efficient method for tensor voting using steerable filters. Lect Notes Comput Sci 3954:228–240
Freeman WT, Adelson EH (1991) The design and use of steerable filters. IEEE Trans Pattern Anal Mach Intell 13:891–906
Frigo M, Johnson SG (2005) The design and implementation of fftw3. Proc IEEE 93:216–231
Hennessy JL, Patterson DA (2019) Computer Architecture. A Quantitative Approach, 6th edn. Morgan Kauffman Publishers, Elsevier, Cambridge, MA, USA
Fernandez JJ (2008) High performance computing in structural determination by electron cryomicroscopy. J Struct Biol 164:1–6
Tabik S et al (2007) High performance noise reduction for biomedical multidimensional data. Digit Signal Proc 17(4):724–736
Fernandez JJ, Martinez JA (2010) Three-dimensional feature-preserving noise reduction for real-time electron tomography. Digital Signal Proc 20:1162–1172
Agulleiro JI, Fernandez JJ (2011) Fast tomographic reconstruction on multicore computers. Bioinformatics 27(4):582–583
Agulleiro JI, Fernández JJ (2012) Evaluation of multicore-optimized implementation for tomographic reconstruction. PLoS ONE 7:48261
Butenhof DR (1997) Programming with POSIX Threads. Addison-Wesley Professional, Boston, MA, USA
Lawson CL et al (2016) EMDataBank unified data resource for 3DEM. Nucleic Acids Res 44:396–403
Iudin A et al (2016) EMPIAR: A public archive for raw electron microscopy image data. Nat Method 13:387–388
Bykov YS et al (2017) The structure of the copi coat determined within the cell. Elife 6:32493
Yan D et al (2019) Sphingolipid biosynthesis modulates plasmodesmal ultrastructure and phloem unloading. Nature Plants 5:604–615
Acknowledgements
This work has been supported by the projects and contracts: RTI2018-095993-B-I00, SAF2017-84565-R, PID2021-123278OB, PID2021-123424OB, TED2021-132020B (funded by MCIN/AEI/10.13039/501100011033/FEDER “A way to make Europe”); UAL18-TIC-A020020-B and P18-RT-1193 (both funded by Junta de Andalucía and FEDER); PAPI-21-GR-2011-0048 (funded by the University of Oviedo) and FUO-200-21 (funded by Thermo Fisher Scientific); and a FPU fellowship (FPU16/05946 funded by MCIN/AEI/10.13039/501100011033/ “El FSE invierte en tu futuro”) awarded to J.J. Moreno.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Moreno, J.J., Garzón, E.M., Fernández, J.J. et al. HPC enables efficient 3D membrane segmentation in electron tomography. J Supercomput 78, 19097–19113 (2022). https://doi.org/10.1007/s11227-022-04607-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11227-022-04607-z