Abstract
The Monte Carlo method was used for QSAR modeling of dimeric pyridinium compounds as acetylcholine esterase inhibitors. QSAR model was developed for a series of 39 dimeric pyridinium compounds. QSAR models were calculated with the representation of the molecular structure by the simplified molecular-input line-entry system. One split into the training and test set have been examined. The statistical quality of the developed model is very good. The calculated model for dimeric pyridinium derivatives had following statistical parameters: r 2 = 0.9477 for the training set and r 2 = 0.9332 the test set. Structural indicators considered as molecular fragments responsible for the increase and decrease in the inhibition activity have been defined. The computer-aided design of new dimeric pyridinium compounds potential acetylcholine esterase inhibitors with the application of defined structural alerts has been presented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
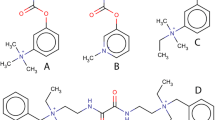
The treatment of disorders with impaired cholinergic transmission usually involves the inhibition of acetylcholinesterase (AChE), and this approach is prime strategy in therapy of Alzheimer’s disease, the most common form of dementia in the elderly population [1]. Further, various conditions, such as glaucoma, constipation, spasmolysis, and also antagonise muscle relaxation in anesthesiology, can be treated with the application of peripherally acting AChE inhibitors [2, 3]. Also the first-line treatment in early stages and in the mild forms of myasthenia gravis (MG) is based on peripheral inhibitors of AChE [4–6]. MG is an autoimmune disorder which is caused by antibody-mediated destruction of postsynaptic membrane in neuromuscular junction. Most commonly, antibodies target the nicotinic acetylcholine receptors (nAChRs), but other components of postsynaptic membrane can be affected as well, leading to painless weakness and fatigue of striated muscles resulting from impaired neuromuscular transmission [7, 8]. These compounds enhance cholinergic transmission by inhibiting the hydrolysis of acetylcholine (ACh), and therefore, they increase its concentration in the synaptic cleft. The treatment of patients with MG is a complex task. Quality of life of patients with MG can be improved with the application of mild physical exercise, and therefore, physicians specialized in sport medicine, physiatrist, and experts in sports in general can ease life of MG patients and help them in the continuous fight against this severe disorder [9, 10]. In current MG therapy, pyridostigmine bromide or neostigmine bromide (carbamate-based AChE inhibitors) are used (Fig. 1). Unfortunately, the usage of these compounds often leads to development of serious gastrointestinal side effects, increased bronchial secretion, and cardiac arrhythmia. Also high intake of these compounds may lead to cholinergic crisis, characterized by even more severe weakness [11–14]. Having in mind stated facts, a search is being continued for a far more effective AChE inhibitor that can be used in the treatment of MG. One of the main issues related to drug development for myasthenia gravis treatment is that myasthenia gravis is a rare disease and that all drugs used for its treatment are considered as orphan drugs. That term is related to the fact that many pharmaceutical industries do not want to invest in their development since it is a very costly process and they do not have economical benefits. For this reason, also, there is a growing need for alternative drug development process that will lead to new and better drugs.
Quantitative structure–activity relationship (QSAR) modeling is widely used in modern drug design because these methods can make the early prediction of activity-related characteristics of drug candidates and eliminate molecules with undesired properties [15–17]. The main aim of QSAR modeling is developing a simple mathematical equation that correlates a molecule’s studied biological activity with its properties and molecular characteristics calculated as quantitative parameters—descriptors. Therefore, descriptors can be defined as entities that characterize specific information of the studied molecule in terms of numerical values associated with the chemical constitution for correlation of chemical structure with biological activity. The contribution of essential structural requisites of the molecule to its biological activity must be defined with developed QSAR model represented with appropriate equation [18]. QSAR model built on geometry-dependent molecular descriptors usually involves a relatively difficult calculation of the optimum molecular geometry which involves application of high computational resources and a long time for computational experiments. For these reasons, the conformation-independent 0D, 1D, and 2D QSAR methods emerge as an alternative approach for developing models based on the constitutional and topological molecular features of molecules [19, 20]. Further, in QSAR modeling, descriptors calculated with molecular graphs are often used [20–23]. The simplified molecular-input line-entry system (SMILES) can be considered as an alternative to molecular graph, and it can be used for defining of molecular structures [24–26]. One-variable QSAR models built up by the Monte Carlo optimization method based on SMILES descriptors have been published recently [27–29]. One of the main advantages of this method in comparison with most commonly used QSAR models is its independence on molecule conformation since it is based on constitutional and topological features of molecules. Also QSAR model based on SMILES notation descriptors has similar or better statistical characteristics in comparison with 3D descriptor-based QSAR models [29]. All above stated facts make QSAR modeling based on SMILES notation descriptors attractive alternative to commonly used methods in drug design and discovery.
The aims of this study are to build a QSAR model based on the SMILES notation optimal descriptors using the Monte Carlo method for dimeric pyridinium compounds as AChE inhibitors and attempt to define the molecular fragments responsible for the stated inhibitory effect. Further, we used the built model and defined molecular fragments for the computer-aided drug design of new potentially promising AChE inhibitors.
Method
The dataset
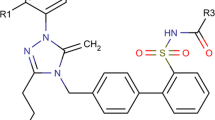
QSAR model was developed for a series of 39 dimeric pyridinium compounds acting as AChE inhibitors [30]. General structures of dimeric pyridinium compounds are presented in Fig. 2. Molecular structures of studied compounds were transformed into canonical SMILES with the ACD/ChemSketch program (ACD/ChemSketch version 11.0). For representing the pharmacological activity, pIC50 (−log IC50) was used as a dependent variable for building both QSAR model (Table S1). The QSAR model for was built up for one random split (30 % of compounds were used in the test set). The normality distribution plot of the whole dataset was checked according to literature [31].
QSAR model development
The main concept of the QSAR modeling can be defined as the following:
Simplified molecular-input line-entry system (SMILES) notation is one of the most convenient molecule structure representations. In presented QSAR modeling, SMILES notation is used for defining a molecular optimal descriptor (DCW) which is calculated as a function of the molecule’s SMILES notation as presented with Eq. 2:
On the basis of SMILES notation, two types of optimal descriptors defined can be defined. First type of optimal descriptors is a local SMILES attribute, defined as SMILES atoms, a fragment of the SMILES notation which contains one (‘C,’ ‘O,’ ‘#,’ etc.) or two symbols (‘Cl,’ ‘Br,’ ‘@@,’ etc.) which cannot be examined separately. The simplest way for defining the molecular DCW as a mathematical function of SMILES atoms can be achieved with the application of local SMILES attributes, as a mathematical function of each character of the SMILES. Therefore, the first and the simplest one SMILES notation-based descriptor is Sk (SMILES atom), a descriptor related to one SMILES symbol (or two that cannot be separated). A linear combination of two or three SMILES atoms are descriptors defined as SSk and SSSk [29, 32]. Linear combination of presented SMILES notation-based descriptors is summarized in Eq. 3.
The inclusion of SMILES notation-based descriptors in the QSAR model development can be achieved with the application of a simple rule—if the value of a coefficient is 1, then an appropriate descriptor is included, or if it is 0, then an appropriate descriptor is discarded form model building. For this reason, coefficients α, β, and γ from Eq. 3 are either 1 (yes) or 0 (no). In presented QSAR model for AChE inhibition, all coefficients were 1, and therefore, all local SMILES notation-based descriptors were used in model development.
Each calculated optimal descriptor receives a numerical value defined as correlation weight (CW) with the application of Monte Carlo method. The applied Monte Carlo method is based on a principle that suitable random numbers are generated and further observed how that fraction of numbers obeys some property/properties. The application of iterative algorithms is used for the computation for obtaining and distribution of an unknown probabilistic entity [33]. Therefore, the main purpose of the Monte Carlo optimization process is the calculation of the numerical data for CW, which gives the maximal value of the correlation coefficient between the endpoint and the optimal descriptor. The Monte Carlo method is based on two parameters for the computation of stated numerical values—threshold (T) and the number of epochs (Nepoch). Threshold can be defined as the separator of calculated SMILES notation-based descriptors into active and inactive with application of following principle: If the descriptor is active, then T has some numerical value, but if the descriptor is inactive, then T has a fixed value of zero. The number of epochs (Nepoch) is related to the computational iterative process with one aim—to obtain the best statistical quality for the training set [29, 32].
Nepoch can be defined as the number of epochs of the Monte Carlo optimization. The target function (TF) of the optimization is given in Eq. 4:
where R is correlation coefficient between the optimal SMILES notation-based descriptor and an endpoint (EP) for training set; W R = 0.1 and W C = 0.01 are empirical parameters; C 0 and C 1 are coefficients from equation obtained by the least squares method:
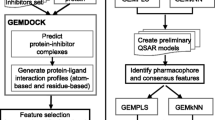
Flowchart of one epoch of the Monte Carlo optimization is presented in Fig. 3.
For development of presented QSAR model, CORAL software was used (http://www.insilico.eu/coral). Building QSAR models took into consideration all local SMILES descriptors and global attributes (S k, SSk and SSSk). The search for the most predictive combination of T and N epoch for all splits was performed from values 0–10 for T and 0–60 for N epoch.
Validation of developed QSAR model
The main purpose of any QSAR modeling is developing a robust model capable of predicting the properties of new molecules in an objective, reliable, and precise manner [34, 35]. Validation methods are needed to establish the predictive power of a model on unseen data and to help determine the complexity of an equation that the amount of data justifies validation methods that are necessary. The data obtained from the created model (an internal method) or using a separate dataset (an external method) can help to validate the developed QSAR model. The methods of least squares fit (R 2), cross-validation (Q 2), root-mean-squared error (RMSE), and Y-randomization are some of the most important methods for validating a model [36, 37]. The application of three methods can be used for the assessment of robustness and reliability of a developed QSAR model [29]: (a) internal validation or cross-validation using the training set compounds, (b) external validation using the test set compounds, and (c) data randomization or Y-scrambling.
Leave-one-out (LOO) cross-validation method is used to develop models as an internal validation. LOO is based on principle that one molecule is randomly omitted from dataset in each cycle and then the rest of molecules are used for model development. The process is repeated until all the compounds are eliminated once. Cross-validated coefficient Q 2 demonstrates the predictive ability of the model [29], where higher value of Q 2 means better model prediction. The cross-validated Q 2 is defined as:
In Eq. 7, Y obs is observed property of the training set compounds, Y pred is LOO-predicted property of the training set compounds, and \({\bar{\text{Y}}}_{\rm train}\) is mean observed property of the training set compounds.
Same principles and statistical methodology can be applied for external validation. The predictive ability of a model is determined by calculating R 2ext which is defined as:
In Eq. 8, Y obs(test) is the observed property of the test set compounds, Y pred(test) is the predicted property of the test set compounds, and \({\bar{\text{Y}}}_{\rm train}\) is mean observed property of the training set compounds.
True predictive potential of developed QSAR models can be defined with novel statistical metric (R 2 m ) [38, 39]. This metric is calculated based on the correlations between the observed and predicted values with (R 2) and without (R 20 ) intercept for the least squares regression lines as represented with equation X:
Main advantage of this metric is that it avoids overestimation of the quality of prediction due to a wide response range (Y-range) since it does not consider the differences between individual responses and the training set mean. For calculating R 2 m metric, an open-access Web application ‘R 2 m calculator’ is available at http://aptsoftware.co.in/rmsquare/.
Y-randomization test is used for testing the robustness of the developed QSAR model. For an appropriate QSPR model, the average correlation coefficient (R r ) of randomized models should be less than the correlation coefficient (R) of non-randomized model. A parameter C R 2 p penalizes the model R 2 for a small difference between squared mean correlation coefficient (R 2 r ) of randomized models and squared correlation coefficient (R 2) of the non-randomized model [40]. The parameter C R 2 p is defined as:
In relation to defined metrics an acceptable QSPR model must have values of R 2ext , Q 2, C R 2 p should be > 0.5. Value for ∆R 2 m should be lower and average value for R 2 m > 0.5. In addition, values for standard error of estimate (s), mean absolute error (MAE), and root-mean-square error (RMSE) should be small, and the value of Fischer’s F ratio should be large. Also R 2 and Q 2 values should be close to each other [36, 39]. Methodology for validating developed QSAR model used in presented research was already successfully applied for the validation of SMILES notation optimal descriptor-based QSAR models [29, 32, 33].
Applicability domain
One of the main goals of a developed QSAR model is its applicability domain, defined with compounds from the training set. Defined AD is further used for the assessment of the reliability of the developed QSAR model. AD of the developed QSAR model is defined as the biological, structural, or physicochemical space, knowledge, or information on which the model of the training set is developed and which can be used for predicting whether the developed QSAR model can be used on compounds which are not used in model developing. When a compound is very different in comparison with all compounds of the modeling set (this assessment is made with the application of AD), then a reliable prediction of its property/activity is uncertain [36].
Difference between experimental and calculated values for studied endpoint can be used for defining AD with the application of Delta(obs), d and \({\bar{\text{d}}}\) [37]. For each molecule used in QSAR study, Delta(obs) is calculated as difference between experimental and calculated values from studied end point. d and \({\bar{\text{d}}}\) are defined in Eqs 11 and 12, respectively, where n is the number of studied compounds.
If for studied compound its Delta(obs) belongs inside of range \({\bar{\text{d}}} - {\text{d }}\) and \({\bar{\text{d}}} + {\text{d}}\), then it falls in defined model applicability domain. However, if its Delta(obs) does not belong inside defined range, then that compound does not belong in defined AD; it is considered as outlier, and it is discarded from development of QSAR model. This relation is given in Eq. 12:
Defined methodology for the determination of AD was successfully applied in development of various QSAR models based on Monte Carlo method [29, 37].
Results and discussion
The results from the applied methodology for defining AD show that all molecules are within the defined AD, and therefore, all studied compounds have typical behavior and all were taken into consideration when QSAR model were developed. Table 1 shows the statistical quality of the built QSAR model for dimeric pyridinium compounds acting as AChE inhibitors. The presented results reveal that there is the reproduction of the statistical quality for the calculated model in three independent runs of the Monte Carlo optimization. Further, the results from Table 1 show that the predictability is very good. For further assessment of developed QSAR model quality, novel statistical metric was used according to published methodologies and results are presented in Table 2. Presented QSAR model for IC50 is satisfactory from the point of view of new criteria [38, 39] (Table 3). Table S2 (supplementary material) shows Y-randomization [40], which also confirms the robustness of suggested models. The search for preferable T and N epoch revealed that preferable T is 1 and preferable N epoch 8. Figure 4 graphically presents the best Monte Carlo optimization run (the highest value for r 2) for developed QSAR model.
DCW (T, N epoch) for compounds in the training and test set is calculated as described in the Method part of this manuscript. The application of the above-mentioned T and the N epoch gives the following model for the pIC50 calculated according to Eq. 5:
According to calculated correlation weight (CW) values of molecular features (SAk) from three Monte Carlo optimization runs, SAk can be classified as promoters of increase or promoters of decrease in a studied activity, and as undefined molecular features. A simple rule is applied for this classification: If CW (SAk) obtained from three independent Monte Carlo optimization runs is positive, then that SAk is the promoter of increase; likewise, if CW (SAk) from three independent Monte Carlo optimization runs is negative, then that SAk is the promoter of decrease; if there are both positive and negative values of CW(Sk) in three runs of the Monte Carlo optimization process, then that SAk is undefined [29, 41, 42]. The list of all SAk, with the correlation weights for three runs of the Monte Carlo optimization process of the built QSAR model for maleimide derivatives, is given in Table S3. The built QSAR model represented with Eq. 6 has the mechanistic interpretation as well as calculated SAk, which can be further defined as molecular fragments. According to the presented data, several important SAks that can be classified as promoters of pIC50 value decrease are C……….. (methyl group), C…C……. (ethyl group), C…C…C… (propyl group), and C…C…(… (ethyl group connected to carbon atom with molecular branching). Several important SAks that can be classified as promoters of pIC50 value increase are (……….., C…(……. and (…C…(… all SAks related to molecular branching, O……….. (sp3 hybridized oxygen atom), N……….. (sp3 hybridized nitrogen atom), Cl………. (chlorine atom) O…C……. (methoxy group), O…C…C… (ethoxy group), etc.
The analysis of SAk presented in Table S3 can be useful in the search and computer-aided design of novel maleimide derivatives with desired pIC50 values. The structures of novel dimeric pyridinium compounds obtained by molecular modeling are presented in Fig. 5.
Molecular design of perspective acetylcholine esterase inhibitors using the QSAR model calculated with Eq. (6) and SAks calculated using the Monte Carlo method
As a template for molecular design, the molecule 1 from dataset was selected. One of the goals of the presented molecular design was to obtain molecules with higher pIC50 values in comparison with the template molecule, since higher pIC50 values mean that a lower concentration is needed to inhibit 50 % of an enzyme. Molecular structures presented as the SMILES notation of designed molecules and their calculated pIC50 values obtained with the application of the built QSAR model for dimeric pyridinium compounds (Eq. 13) are presented in Table 3.
Table S4 presents an example of calculation DCW(1,5) for the template molecule A. When Eq. 13 is applied for calculating DCW, the resulting endpoint value (pIC50) is 0.3362. In molecule A1+ on hydrogen from CH3 bonded to amino, nitrogen atom was substituted with chlorine atom. As already stated, chlorine atom is a promoter of endpoint increase, and therefore, molecule A1+ calculated IC50 value was 0.4137. Molecule A2+ has additional molecular branching in amino group, and since most branching descriptors are defined as promoters on endpoint increase, calculated IC50 value was 1.114. In molecule A3+, methoxy group was inserted in amino group part of molecule. Both oxygen atom and methoxy group are promoters on endpoint increase, and calculated IC50 value was 0.3755. In presented research, methyl , ethyl , and propyl groups are defined as promoters on endpoint decrease. Molecules A1− and A2− have inserted additional carbon atom in amino part of molecule. With this, additional descriptors were calculated for molecules A1− and A2− (C……….. and C…C……. for molecule A1− and C……….., C…C……. and C…C…C… for molecule A2−). Since all of these descriptors are promoters of on endpoint decrease, calculated IC50 values for molecules A1− and A2− were 0.1448 and 0.0155, respectively.
Conclusion
QSAR models for dimeric pyridinium compounds as acetylcholine esterase inhibitors were built. The Monte Carlo optimization process incorporated within CORAL software was capable to be an efficient tool to build up robust model with good statistical quality. The predictive potential of the applied approach was tested and the robustness of the model was proven with different methods. The SMILES attributes, defined as SMILES notation-based molecular descriptors, which are promoters of IC50 increase/decrease, are identified. The suggested modeling process and computer-aided drug design are based on computational experiments with the application of statistically stable structural alerts (promoters of increase or decrease of IC50). This approach can be applied in the search for new potential acetylcholine esterase inhibitors which can be used further for the treatment of myasthenia gravis.
References
Giacobini E (2008) Cholinesterase inhibitors: new roles and therapeutic alternatives. Pharmacol Res 50:433–440
Millard CB, Broomfield CA (1995) Anticholinesterases: medical applications of neurochemical principles. J Neurochem 64:1909–1918
Bevan DR, Donati F, Kopman AF (1992) Reversal of neuromuscular blockade. Anesthesiology 77:785–805
Drachman DB (1994) Myasthenia gravis. N Engl J Med 330:1797–1810
Richman DP, Agius MA (2003) Treatment of autoimmune myasthenia gravis. Neurology 61:1652–1661
Lindstrom JM (2000) Acetylcholine receptors and myasthenia. Muscle Nerve 23:453–477
Vincent A, Palace J, Hilton-Jones D (2001) Myasthenia gravis. Lancet 357:2122–2128
Conti-Fine BM, Milani M, Kaminski HJ (2006) Myasthenia gravis: past, present, and future. J Clin Invest 116:2843–2854
Rubin DI, Hentschel K (2007) Is exercise necessary with repetitive nerve stimulation in evaluating patients with suspected myasthenia gravis? Muscle Nerve 35:103–106
Lucia A, Maté-Muñoz JL, Pérez M, Foster C, Gutiérrez-Rivas E, Arenas J (2007) Double trouble (McArdle’s disease and myasthenia gravis): How can exercise help? Muscle Nerve 35:125–128
Leigh P, Abrahams S, Al-Chalabi A, Ampong M, Goldstein L, Johnson J, Lyall R, Moxham J, Mustfa N, Rio A, Shaw C, Willey E (2003) The management of motor neurone disease. J Neurol Neurosurg Psychiatry 74(Suppl 4):iv32–iv47
Juel VC, Massey JM (2005) Autoimmune Myasthenia Gravis: recommendations for treatment and immunologic modulation. Curr Treat Options Neurol 7:3–14
Froelich J, Eagle CJ (1996) Anaesthetic management of a patient with myasthenia gravis and tracheal stenosis. Can J Anaesth 43:84–89
Juel VC, Massey JM (2007) Myasthenia gravis. Orphanet J Rare Dis 2:44
Hansch C, Hoekman D, Gao H (1996) Comparative QSAR: toward a deeper understanding of chemicobiological interactions. Chem Rev 96:1045–1076
Cherkasov A, Muratov EN, Fourches D, Varnek A, Baskin II, Cronin M, Dearden J, Gramatica P, Martin YC, Todeschini R, Consonni V, Kuz’Min VE, Cramer R, Benigni R, Yang C, Rathman J, Terfloth L, Gasteiger J, Richard A, Tropsha A (2014) QSAR modeling: Where have you been? Where are you going to? J Med Chem 57:4977–5010
Tropsha A, Golbraikh A (2007) Predictive QSAR modeling workflow, model applicability domains, and virtual screening. Curr Pharm Des 13:3494–3504
Cronin MTD, Schultz TW (2003) Pitfalls in QSAR. J Mol Struct-THEOCHEM 622:39–51
Duchowicz PR, Comelli NC, Ortiz EV, Castro EA (2012) QSAR study for carcinogenicity in a large set of organic compounds. Curr Drug Saf 7:282–288
Talevi A, Bellera CL, Ianni MD, Duchowicz PR, Bruno-Blanch LE, Castro EA (2012) An integrated drug development approach applying topological descriptors. Curr Comput Aided Drug Des 8:172–181
Randic M, Basak SC (2010) New descriptor for structure-property and structure-activity correlations. J Chem Inf Comput Sci 41:650–656
da Silva Junkes B, Arruda ACS, Yunes RA, Porto LC, Heinzen VEF (2005) Semi-empirical topological index: a tool for QSPR/QSAR studies. J Mol Model 11:128–134
Ivanciuc O (2013) Chemical graphs, molecular matrices and topological indices in chemoinformatics and quantitative structure–activity relationships. Curr Comput Aided Drug Des 9:153–163
Weininger D (1988) SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Inf Comput Sci 28:31–36
Weininger D, Weininger A, Weininger JL (1989) SMILES. 2. Algorithm for generation of unique SMILES notation. J Chem Inf Comput Sci 29:97–101
Weininger D (1990) SMILES. 3. Depict. Graphical depiction of chemical structures. J Chem Inf Comput Sci 30:237–243
Toropov AA, Benfenati E (2007) SMILES in QSPR/QSAR modeling: results and perspectives. Curr Drug Discov Technol 4:77–116
Toropov AA, Benfenati E (2007) SMILES as an alternative to the graph in QSAR modelling of bee toxicity. Comput Biol Chem 31:57–60
Veselinović AM, Veselinović JB, Živković JV, Nikolić GM (2015) Application of SMILES notation based optimal descriptors in drug discovery and design. Curr Top Med Chem 15:1768–1779
Conejo-García A, Pisani L, Del Carmen Núñez M, Catto M, Nicolotti O, Leonetti F, Campos JM, Gallo MA, Espinosa A, Carotti A (2011) Homodimeric bis-quaternary heterocyclic ammonium salts as potent acetyl- and butyrylcholinesterase inhibitors: a systematic investigation of the influence of linker and cationic heads over affinity and selectivity. J Med Chem 54:2627–2645
Ojha PK, Roy K (2011) Comparative QSARs for antimalarial endochins: importance of descriptor-thinning and noise reduction prior to feature selection. Chemometr Intell Lab Syst 109:146–161
Toropova AP, Toropov AA, Benfenati E, Gini G, Leszczynska D, Leszczynski J (2011) CORAL: quantitative structure-activity relationship models for estimating toxicity of organic compounds in rats. J Comput Chem 32:2727–2733
Toropov AA, Toropova AP, Puzyn T, Benfenati E, Gini G, Leszczynska D, Leszczynski J (2013) QSAR as a random event: modeling of nanoparticles uptake in PaCa2 cancer cells. Chemosphere 92:31–37
Roy K (2007) On some aspects of validation of predictive quantitative structure activity relationship models. Expert Opin Drug Dis 2:1567–1577
Roy PP, Leonard JT, Roy K (2008) Exploring the impact of the size of training sets for the development of predictive QSAR models. Chemometr Intell Lab Syst 90:31–42
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701
Toropov AA, Toropova AP, Lombardo A, Roncaglioni A, Benfenati E, Gini G (2011) CORAL: building up the model for bioconcentration factor and defining it’s applicability domain. Eur J Med Chem 46:1400–1403
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H (2012) Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model 52:396–408
Ojha PK, Mitra I, Das R, Roy K (2011) Further exploring rm2 metrics for validation of QSPR models. Chemometr Intell Lab Syst 107:194–205
Ojha PK, Roy K (2011) Comparative QSARs for antimalarial endochins: importance of descriptor-thinning and noise reduction prior to feature selection. Chemometr Intell Lab 109:146–161
Toropova AP, Toropov AA, Veselinović JB, Miljković FN, Veselinović AM (2014) QSAR models for HEPT derivates as NNRTI inhibitors based on Monte Carlo method. Eur J Med Chem 77:298–305
Veselinović AM, Milosavljević JB, Toropov AA, Nikolić GM (2013) SMILES-based QSAR model for arylpiperazines as high-affinity 5-HT1A receptor ligands using CORAL. Eur J Pharm Sci 48:532–541
Acknowledgments
We would like to thank reviewers whose suggestions have improved our manuscript. This work has been supported by the Ministry of Education and Science, the Republic of Serbia, under Project Number 43012.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sokolović, D., Stanković, V., Toskić, D. et al. Monte Carlo-based QSAR modeling of dimeric pyridinium compounds and drug design of new potent acetylcholine esterase inhibitors for potential therapy of myasthenia gravis . Struct Chem 27, 1511–1519 (2016). https://doi.org/10.1007/s11224-016-0776-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-016-0776-z