Abstract
Invasive giant hogweeds are infamous in Europe for causing ecological and economic damage, but genetic resources for their study are limited. We used next-generation sequencing to develop a microsatellite library for Heracleum persicum, a widespread invasive in Nordic countries. In addition, these markers were cross amplified with the closely related Heracleum mantegazzianum, Heracleum sosnowskyi, Heracleum sphondylium, and the putative hybrid H. persicum × H. sphondylium, as well as the more distantly related Anthriscus sylvestris. We designed and validated 164 primer pairs. A cost-effective PCR approach with modified forward primer, reverse primer, and fluorescently labeled universal tail was used to test the functionality of each marker. Twenty-five of thirty markers screened on eight geographically distant samples of H. persicum were polymorphic. The number of alleles was 2–4 whereas the expected and observed heterozygosity varied from 0.06 to 0.84 and 0.0 to 1.0 respectively. The cross-species amplification efficiency was 84–100 %, in which 60–76 % of the cross-species amplified markers were polymorphic for Heracleum taxa including H. persicum × sphondylium. Three out of eight of the cross-amplified markers were polymorphic in Anthriscus sylvestris. Ordination revealed a clear genetic structure of Heracleum taxa. Thus, these markers can serve as important genetic resources for understanding taxonomy, population genetics, and phylogeny of giant hogweeds and their hybrids, which in turn, is expected to contribute to invasive species management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive giant hogweeds and their hybrids can pose serious threats to the native biodiversity and need to be managed at local and regional scales. However, morphological similarity, taxonomic uncertainty, and hybridization impede the management process. In such cases, understanding population genetics of the species is helpful. Studies of the population genetics of giant hogweeds are limited due to paucity of genetic resources (Henry et al. 2009; Jahodova et al. 2007; Walker et al. 2003). Among different genetic markers, microsatellites are one of the most popular markers for the study of genetic structure and mixture analysis due to their high allelic diversity and possibility of amplification in other closely related species (Guichoux et al. 2011). On the other hand, the lack of sequence information for non-model organisms has stymied microsatellite development, but the emergence of next-generation sequencing has now opened the door for genetic exploration of organisms (Guichoux et al. 2011) like giant hogweeds which have a broad impact on biodiversity, ecology, and socioeconomy of the invaded country (Hejda et al. 2009; Pimentel et al. 2005; Thiele and Otte 2007).
The invasive giant hogweeds include Heracleum persicum Desf. ex Fisch., Heracleum mantegazzianum Sommier & Levier, and Heracleum sosnowskyi Mandenova (Nielsen et al. 2005). The taxonomy of giant hogweeds is disputed, for example, the northern Norwegian giant hogweed has been classified as Heracleum panaces, Heracleum giganteum, Heracleum laciniatum, and Heracleum tromsoensis (Alm 2013). Recently, Jahodova et al. (2007) suggested H. persicum as the name for the northern Norwegian giant hogweed based on amplified fragment length polymorphism (AFLP). Heracleum persicum was further used by Fröberg (2010) in the ‘Flora Nordica’; however, taxonomy of the northern Norwegian giant hogweed is still unclear (Alm 2013). Similarly, the taxonomy of English giant hogweed is controversial. In a recent flora, Sell and Murrell (2009) described Heracleum trachyloma Fisch. & C. A. Mey., Heracleum lehmanianum Bunge, and Heracleum grossheimii Manden. ex. Grossh. from the UK and have treated H. mantegazzianum as a synonym. However, Stace (2010) retained H. mantegazzianum as the valid name for giant hogweed of the British Isles. These uncertainties clearly show that Heracleum is a taxonomically complex genus.
Giant hogweeds were famous in the European gardens during the 19th century due to their spectacular appearance. However, within less than two centuries of their introduction, they have become prominent and problematic invasive species (Alm 2013; Elvebakk 1992; EPPO 2009). Heracleum persicum, commonly known in Norway as tromsøpalme, is a native of Persia and invasive in Nordic countries. This herbaceous polycarpic species is generally 2 m tall and sometimes reaches up to 3 m with large leaves of up to 2.5 m long. The giant hogweed, H. mantegazzianum, derives from western Caucasus and has aggressively colonized most of the Europe (EPPO 2009). H. mantegazzianum is monocarpic and can grow up to 5 m with leaves reaching up to 3 m. Similarly, H. sosnowskyi, commonly known as Sosnowsky’s hogweed, is a native of Caucasus and Transcaucasia which has colonized most of the Baltic region (EPPO 2009; Nielsen et al. 2005). This species, which grows up to 3 m tall, is also monocarpic and morphologically closer to H. mantegazzianum. All hogweed species are perennial, seed-propagated plants. These three giant hogweeds have damaged natural systems of geographically distinct areas (EPPO 2009), and there is still risk of further expansion and invasion into adjacent areas.
In addition, giant hogweeds often hybridize with H. sphondylium L. (common hogweed), a smaller plant which reaches up to 1.4 m in height (Fröberg 2010). Heracleum sphondylium is considered indigenous to most European countries. Scattered natural hybrids of giant hogweeds with native H. sphondylium have been reported from the British Isles (Sell and Murrell 2009; Stace 2010) as well as in Scandinavia (Fröberg 2010). Several well-established populations of natural hybrids have been observed in northern Norway which are sometimes morphologically very similar to H. persicum, but without its characteristic anise smell (Alm 2013). Norwegian hybrids are as invasively vigorous as H. persicum (Alm 2013), and biological systems face even stronger challenges due to the combined threat of giant hogweeds and hybrids.
As a consequence of their large size and toxic sap, giant hogweeds have rarely received priority during plant collection, which in turn, has always been hindering the plant identification process (Fröberg 2010). In such cases, microsatellite markers can serve as a useful tool to resolve taxonomy. However, microsatellite markers have not been reported so far for H. persicum, H. sosnowskyi, and the putative hybrid H. persicum × H. sphondylium. Our aims were to design and test a suite of novel microsatellite markers for identification, population genetic studies, and hybrid characterization of giant hogweeds. Specifically, we consider H. persicum as a model species for designing and testing microsatellite markers that also cross-amplify H. sosnowskyi, H. mantegazzianum, H. sphondylium, and hybrids. To extend the application of this approach, we also test cross-generic transferability to Anthriscus sylvestris from Apiaceae. Finally, we evaluate taxonomic applications of the validated markers.
Materials and Methods
Plant Material
Leaf samples were collected from up to eight individuals per population, if possible at 5–10 m intervals, and dried in silica gel (Tables 1 and 2). The leaf samples, DNA extracts, and voucher specimens are deposited at the Tromsø Museum (TROM).
DNA Extraction and Standardization
DNA was extracted using a DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. The quality of DNA was checked by running 2 % agarose gels. DNA concentrations were measured by NanoDrop 2000 (Thermo Scientific, Waltham, USA), and all the samples were normalized to 10 ng/μl for PCR.
Microsatellite Library Preparation and Primer Design
The facility available at GenoScreen (Lille, France) was used to prepare libraries and design primers. High-quality DNA of ten individuals from Norwegian, Danish, and English populations of H. persicum were pooled for microsatellite library preparation. The pooled DNA was fragmented and hybridized with eight probes namely: TG, TC, AAC, AAG, AGG, ACG, ACAT, and ACTC to enrich the DNA fragments containing microsatellite repeats. Following the protocol described in Malausa et al. (2011), the enriched microsatellite DNA libraries were further amplified by PCR using high-fidelity Taq (Roche, Indiana, USA) to increase their concentrations. The concentrations of the libraries were measured for quality assurance prior to sequencing by 454-GS-FLX Titanium chemistry (Malausa et al. 2011). QDD software (Meglécz et al. 2010) with default settings was used to design the microsatellite primers.
Biological Validation and Multiplex Optimization
We selected 30 pairs of primers from the validated set with wide ranges of expected PCR products to screen Heracleum samples. We chose primers targeting di-, tri-, and tetranucleotide microsatellite repeats with less than 15 total repeats. Multiplex manager (Holleley and Geerts 2009) was used to assess the minimum number of primers for multiplexed reactions. Eight individuals of H. persicum from different populations covering its entire range were used for the initial screening of the markers (Table 1). Four universal primers suggested by Blacket et al. (2012) were fluorescently labeled with PET, NED, HEX, and 6-FAM, and all forward primers were modified in their 5′ end with universal tails (Table 3). The universal primer remains inactive during the 1st and 2nd cycle of the PCR; however, it acts as a locus-specific primer from the 3rd PCR cycle onwards (Blacket et al. 2012). Each reaction contained 1 pmol of forward primer and 10 pmol each of reverse and universal primers. The total volume of each reaction was 6 μl including 1 μl of standardized template DNA, 1 μl of primer mix, 3 μl of 2× Type-it Master Mix, and 1 μl of RNA-free water (Type-it Microsatellite PCR Kit, Qiagen). The cycling conditions for singleplex PCR were as follows: initial denaturation at 95 °C for 15 min followed by 15 cycles of 95 °C for 30s, 64–56.5 °C of touchdown PCR for 1 min with 0.5 °C decrease per cycle, and 72 °C for 40s; 17 cycles of 95 °C for 30s, 57 °C for 45 s, 72 °C for 40s; 8 cycles of 95 °C for 30s, 53 °C for 45 s, 72 °C for 40s; and a final extension of 60 °C for 30 min. The overnight holding temperature was set as 4 °C. To find the optimal dilution level, undiluted, 1:10, 1:20, and 1:30 diluted PCR product were used in capillary electrophoresis. Finally, a mixture of 2 μl of 1:20 diluted PCR product, 7.8 μl of HiDi Formamide, and 0.2 μl of LIZ 600 (Applied Biosystems, Foster City, CA, USA) was denatured at 95 °C for 5 min, and electrophoresis was performed on 3130xl Genetic Analyzer (Applied Biosystems).
All markers which were successfully amplified in single reactions (see examples in Fig. S1) were further accommodated in three multiplex reactions by using multiplex manager (Holleley and Geerts 2009). The modified forward primer, reverse primer, and fluorescently labeled universal tail were mixed in a 1:2:1 ratio (Blacket et al. 2012). The concentrations of forward primers were 0.1 μM for the first screening. Each multiplex PCR reaction consisted of 3 μl of Master Mix and 0.5 μl of RNA-free water (Type-it Microsatellite PCR Kit, Qiagen), 1 μl of primer mix, and 1.5 μl of template DNA. The initial multiplex PCR amplification was performed with identical thermal conditions listed above. We observed high variation in amplification efficiency between markers. Thus, concentrations of the weaker markers were increased in 0.05 μM increments until a satisfactory amplification was observed. The final concentration of the primers is given in Table 3. This step increased the peak height of weakly amplified loci; however, the amplification was still not uniform across all loci within each multiplex reaction. Several troubleshooting strategies mentioned in the Type-it Microsatellite PCR handbook (Anonymous 2009), for example, increasing the annealing time, decreasing the number of PCR cycles, increasing the time for final extension, etc. were attempted. Finally, PCR conditions of the multiplex were as follows: initial denaturation at 95 °C for 10 min followed by 10 cycles of 95 °C for 30s, 60–50 °C of touchdown PCR for 1 min with 1 °C decrease per cycle, and 72 °C for 45 s; 25 cycles of 95 °C for 30s, 50 °C for 1 min, 72 °C for 45 s; and a final extension of 60 °C for 15 min. Five samples of H. persicum each from Norway, Iran, England, Finland, Sweden, and Denmark (Table 1) were screened for functional markers in three multiplex reactions as indicated in Table 3 (also see Fig. S2-S4).
Cross-Species Amplification
Cross-species amplification was performed on eight distant samples for each species (Table 2). Initial screening revealed that the alleles were very close to each other among species. Thus, final cross-species amplification was performed with the conditions as described for H. persicum. Cross-generic amplification tests were also performed on eight samples of Anthriscus sylvestris collected from Tromsø, Norway. We selected 25 markers, which were well amplified in Heracleum, for this test keeping all other PCR conditions identical with H. persicum. The details on the number of samples for cross-amplification are given in the Table 2.
Data Analysis
The fragments were further analyzed in Geneious (v. 6.1.6, Biomatters) for allele calling. The data were checked for genotyping errors due to null alleles, large allele drop outs, and stutter peaks in Micro-Checker v. 2.2.3 (Van Oosterhout et al. 2004). Allele size range was calculated based on the distant samples of each taxa used in the initial screening. GeneAlEx v. 6.5 (Peakall and Smouse 2012) was used for molecular diversity analysis, i.e., the number of allele (N A), expected (H E), and observed heterozygosity (H O) as well as Shannon’s information index (I) of eight samples from a single population of each taxon, except A. syslvestris which was based on four samples. Arlequin v. 3.5.1.3 (Excoffier and Lischer 2010) was used to test Hardy-Weinberg equilibrium. The potential role of validated primers for resolving Heracleum taxonomy was assessed by principal coordinate analysis (PCoA) in GenAlEx v. 6.5 (Peakall and Smouse 2012).
Results and Discussion
Microsatellite Library and Primer Design
The concentration of the library was 1.03 × 1010 molecule/μl, thus, much higher than the threshold value of 1.46 × 108 molecule/μl (GenoScreen, France). A bead recovery value of 88 % was also higher than the minimum required value of 65 % (Anonymous 2011). The enrichment value was 10 %, well within the expected range (Anonymous 2011). Out of the 25,951 raw sequences, 3904 sequences contained microsatellite motifs (Online Resource 1). A total of 164 primer pairs (Online Resource 2) was designed for H. persicum from sequences which varied from 118 to 580 bp with an average sequence length of 345.78 bp. The expected product length for the designed primers ranged from 90 to 320 bp. About 80 % of the microsatellite motifs were dinucleotide repeats with 5–15 repeats followed by trinucleotide repeats (17 %) with repeat numbers ranging from 5 to 9. Only four of the loci were tetranucleotide with 5–7 repeats. There was a single motif with six nucleotides repeated five times.
The differences in salt unadjusted melting temperature (T m) of 15 primer pairs were between 5 and 8 °C (Online Resource 2); however, 91 % of the primer pairs had a melting temperature difference below 5 °C, an important multiplexing criterion (Butler 2005). Similarly, a high variation in the range of expected PCR products creates possibilities of accommodating several markers in a single reaction, allowing substantial reduction of project costs (Butler 2005; Guichoux et al. 2011).
Biological Validation of Primers
Out of the 30 markers selected for biological validation on H. persicum, 26 (87 %) amplified in singleplex PCR, yielding 83 % polymorphic markers. One of the markers did not amplify in the multiplex reaction and was thus discarded from further analysis (Hp_04, Online Resource 2). We successfully amplified ten markers in a single reaction. However, the number of loci per multiplex reaction could be increased if the forward primer is carefully modified. For example, multiplex manager accommodated 17–20 markers in a singleplex reaction. The in silico estimation is tractable as designed primers target PCR products of varying size and have similar T m values. In addition, the use of fluorescently labeled universal tails could be highly flexible in terms of re-modification of forward primers and cost reduction (Blacket et al. 2012). If some markers do not amplify with the given modification, one can re-modify the forward primer using another universal tail. Sometimes this strategy improves the primer amplification. In such cases, one can reorder the newly modified forward primer and continue to use the previously ordered reverse primer thus reducing the extra cost for reverse primers.
Polymorphism in Heracleum persicum
Out of the 25 functional markers, 21 were polymorphic for the Danish population of H. persicum. However, all markers were polymorphic for more than one population. The Shannon’s information index ranged from 0.23 to 1.35 among polymorphic loci (Table 4). The number of alleles varied from 2 to 4. The expected and observed heterozygosities ranged from 0.12 to 0.73 and 0.13 to 1.0, respectively. All the loci were found to be in Hardy-Weinberg equilibrium (HWE). There was no evidence of null alleles, large allele dropouts, and scoring errors due to stutter bands. Observed and expected heterozygosities for H. persicum are comparable with the closely related H. mantegazzianum (Henry et al. 2008; Walker et al. 2003) except for a few loci which are completely heterozygous in this study. The number of alleles that we found for Danish H. persicum is remarkably lower than that for the English population of H. mantegazzianum (Walker et al. 2003).
Cross-Species Amplification
Ten loci were polymorphic for H. mantegazzianum collected from a single Norwegian population (Table 5). Additionally, 15 markers were polymorphic for more than one population (see allele size range, Table 5). The number of alleles varied from 2 to 3 depending on the polymorphic marker. The expected and observed heterozygosities ranged from 0.22 to 0.51 and 0.13 to 1.0, respectively. The number of alleles for H. sphondylium ranged from 2 to 3 among ten polymorphic loci. Observed heterozygosity ranged from 0.25 to 1.0 whereas expected heterozygosity varied from 0.22 to 0.53. Similarly, the number of alleles in H. sosnowskyi ranged from 2 to 6 for 19 polymorphic loci. The expected and observed heterozygosities ranged from 0.12 to 0.76 and 0.13 to 1.0, respectively (Table 5). The number of alleles was 2–4 for the putative hybrid H. persicum × H. sphondylium for 15 polymorphic loci with observed heterozygosity ranging from 0.13 to 1.0. The expected heterozygosity ranged from 0.12 to 0.7. Similarly, 2–7 alleles were present in A. sylvestris. Expected and observed heterozygosities varied from 0.36 to 0.76 and 0.29 to 0.86, respectively. All the markers are bi- or tri-allelic for H. mantegazzianum and H. sphondylium, which contrasts previous studies that have reported 2–10 alleles for H. mantegazzianum and 2–8 alleles for H. sphondylium for single populations (Henry et al. 2008; Walker et al. 2003). The differences may be due to the higher number of samples used in those studies.
The cross-species amplification efficiencies for multiple populations varied from 84 to 100 % for functional markers, and the polymorphism ranged from 60 % (H. sphondylium and H. mantegazzianum) to 76 % (H. sosnowskyi and hybrid). These results indicate that the set of designed primers could be used for morphologically distant species of Heracleum. There are about 70 species of Heracleum distributed primarily in Asia and Europe. Some species are important ingredients of traditional medicines in China (Fading and Watson 2005) and Persia (Alm 2013). Thus, the given set of primers could possibly be used to evaluate the genetic status, e.g., of medicinally important plants.
Out of the 25 functional markers, eight loci were amplified in A. sylvestris (Table 5). However, only three of the loci were polymorphic. This low level of polymorphism was expected as we used samples from a range of only few kilometers. Nevertheless, the proportion of polymorphic loci was 37 % of the amplified markers. Heracleum belongs to the family Apiaceae which includes between 250 and 440 genera and numerous economically important plants (Fading and Watson 2005). Microsatellite markers have also been used for cross-genus amplification (Fan et al. 2013), in our case, 32 % of the functional markers amplified in A. sylvestris. As Heracleum is genetically closer to several other genera than Anthriscus (Downie et al. 2000), we assume there is a high possibility to amplify several markers in other genera of Apiaceae.
Taxonomic Application
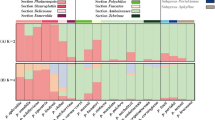
Out of the 25 markers, six were highly polymorphic with the number of alleles ranging from 3 to 6 (Table 6). These polymorphic markers explain most of the genetic variation in the data. The tested markers provided clear taxonomic resolution as all five taxa were noticeably separated in the PCoA analysis (Fig. 1). The genetic structure of the taxa revealed by PCoA also mimics the general morphological similarity demonstrated by the close position of the morphologically similar H. mantegazzianum and H. sosnowskyi. The controversial samples from England, which have been named either H. trachyloma (Sell and Murrell 2009) or H. mantegazzianum (Stace 2010), indeed clustered with other H. mantegazzianum. Thus, the tested markers can readily be used to resolve the Heracleum taxonomy.
In addition, the tested markers could be used for studying hybridization among different species of Heracleum. The putative hybrid H. persicum × H. sphondylium was clustered between the two parental species (Fig. 1), which corroborates other studies (Alm 2013). The quick replacement of local flora by a stable hybrid has been observed previously (Alm 2013). This could happen anywhere if different species of Heracleum grow in the vicinity, as has, e.g., been observed for H. mantegazzianum (Fröberg 2010). Our designed primers provide additional applications in tracking invasive hybrids. We observed that 72 % of the tested markers were polymorphic for hybrids, which implies that a good number of markers could be developed from the given set of untested markers (Online Resource 2).
Conclusions
Identification and population genetics of invasive species are crucial for formulating management plans. This study provides a suite of 25 biologically validated markers as well as 134 newly designed primer pairs which could be used to discriminate Heracleum taxa, which has the potential to illuminate population genetics of giant hogweeds and their putative hybrids. This suite of markers could also be used for QTL analysis or genetic mapping to explore new adaptive traits in giant hogweeds as well as in their hybrids. Thus, these markers can contribute to biodiversity conservation through invasive species management. In addition, cross-generic amplification of a few markers indicates the possibility of wider application of the designed markers within the Apiaceae, for example, for phylogenetic studies. Future studies on several economically important species could benefit from these markers and primers.
References
Alm T (2013) Ethnobotany of Heracleum persicum Desf. ex Fisch., an invasive species in Norway, or how plant names, uses, and other traditions evolve. J Ethnobiol Ethnomed 9(42):1–12. doi:10.1186/1746-4269-9-42
Anonymous (2009) Type-it Microsatellite PCR hand book: for reliable multiplex PCR-based analysis of microsatellites without the need for optimization. Qiagen, Hilden, Germany. http://www.qiagen.com/resources/resourcedetail?id=d6135896-0466-4d9d-aeef-1c4f23f8964e&lang=en. Accessed 15 December 2012
Anonymous (2011) emPCR amplification method manual - Lib L LV. 454 Sequencing, Branford, Connecticut, USA. http://454.com/downloads/my454/documentation/gs-flx-plus/emPCR-Amp-Lib-L-LV-Method-Manual_XLPlus_May2011.pdf. Accessed 07 November 2014
Blacket MJ, Robin C, Good RT, Lee SF, Miller AD (2012) Universal primers for fluorescent labelling of PCR fragments—an efficient and cost-effective approach to genotyping by fluorescence. Mol Ecol Resour 12:456–463. doi:10.1111/j.1755-0998.2011.03104.x
Butler J (2005) Constructing STR multiplex assays. In: Carracedo A (ed) Forensic DNA Typing Protocols, vol 297. Methods in Molecular Biology. Humana Press, Totowa, New Jersey, pp 53–65
Downie SR, Katz-Downie DS, Watson MF (2000) A phylogeny of the flowering plant family Apiaceae based on chloroplast DNA rpl16 and rpoC1 intron sequences: towards a suprageneric classification of subfamily Apioideae. Am J Bot 87:273–292
Elvebakk A (1992) Has Heracleum laciniatum auct. scand. been introduced to North Norway through England? Polarflokken 16:313–316
EPPO (2009) Heracleum mantegazzianum, Heracleum sosnowskyi and Heracleum persicum. EPPO Bull 39:489–499. doi:10.1111/j.1365-2338.2009.02313.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. doi:10.1111/j.1755-0998.2010.02847.x
Fading P, Watson MF (2005) Heracleum Linnaeus. In: Zhengyi W, Raven P (eds) Flora of China (Apiaceae through Ericaceae), vol 14. Missouri Botanical Garden Press, St. Louis, pp 194–202
Fan L et al (2013) Transferability of newly developed pear SSR markers to other Rosaceae species. Plant Mol Biol Rep 31:1271–1282. doi:10.1007/s11105-013-0586-z
Fröberg L (2010) Heracleum L. In: Jonsell B, Karlsson T (eds) Flora Nordica (Thymelaeaceae to Apiaceae), vol 6. The Swedish Museum of Natural History, Stockholm, pp 224–234
Guichoux E et al (2011) Current trends in microsatellite genotyping. Mol Ecol Resour 11:591–611. doi:10.1111/j.1755-0998.2011.03014.x
Hejda M, Pyšek P, Jarošík V (2009) Impact of invasive plants on the species richness, diversity and composition of invaded communities. J Ecol 97:393–403. doi:10.1111/j.1365-2745.2009.01480.x
Henry P, Provan J, Goudet J, Guisan A, JahodovÁ Š, Besnard G (2008) A set of primers for plastid indels and nuclear microsatellites in the invasive plant Heracleum mantegazzianum (Apiaceae) and their transferability to Heracleum sphondylium. Mol Ecol Resour 8:161–163. doi:10.1111/j.1471-8286.2007.01911.x
Henry P, Le Lay G, Goudet J, Guisan A, JahodovÁ Š, Besnard G (2009) Reduced genetic diversity, increased isolation and multiple introductions of invasive giant hogweed in the western Swiss Alps. Mol Ecol 18:2819–2831. doi:10.1111/j.1365-294X.2009.04237.x
Holleley CE, Geerts PG (2009) Multiplex Manager 1.0: a cross-platform computer program that plans and optimizes multiplex PCR. BioTechniques 46:511–517. doi:10.2144/000113156
Jahodova S, Trybush S, Pysek P, Wade M, Karp A (2007) Invasive species of Heracleum in Europe: an insight into genetic relationships and invasion history. Divers Distrib 13:99–114. doi:10.1111/j.1472-4642.2006.00305.x
Malausa T et al (2011) High-throughput microsatellite isolation through 454 GS-FLX Titanium pyrosequencing of enriched DNA libraries. Mol Ecol Resour 11:638–644
Meglécz E, Costedoat C, Dubut V, Gilles A, Malausa T, Pech N, Martin J-F (2010) QDD: a user-friendly program to select microsatellite markers and design primers from large sequencing projects. Bioinformatics 26:403–404. doi:10.1093/bioinformatics/btp670
Nielsen C, Ravn HP, Nentwig W, Wade M (eds) (2005) The giant hogweed best practice manual. Guidelines for the management and control of an invasive weed in Europe. Forest and Landscape Denmark, Hoersholm, Denmark, p 44
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. doi:10.1093/bioinformatics/bts460
Pimentel D, Zuniga R, Morrison D (2005) Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol Econ 52:273–288. doi:10.1016/j.ecolecon.2004.10.002
Sell P, Murrell G (2009) Flora of Great Britain and Ireland. Volume 3 (Mimosaceae-Lentibulariaceae) vol 3. Cambridge University Press, New York
Stace C (2010) New flora of the British Isles, 3rd edn. Cambridge University Press, Cambridge
Thiele J, Otte A (2007) Impact of Heracleum mantegazzianum on invaded vegetation and human activities. In: Pysek P, Cock MJW, Nentwig W, Ravn HP (eds) Ecology and management of giant hogweed (Heracleum mantegazzianum). CABI, Wallingford, pp 144–156
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538. doi:10.1111/j.1471-8286.2004.00684.x
Walker NF, Hulme PE, Hoelzel AR (2003) Population genetics of an invasive species, Heracleum mantegazzianum: implications for the role of life history, demographics and independent introductions. Mol Ecol 12:1747–1756. doi:10.1046/j.1365-294X.2003.01866.x
Acknowledgments
We would like to thank Atefeh Pirany for collecting samples from Iran. We are grateful to Jens-Petter Kjemprud, Embassy of Norway in Iran, for delivering samples from Iran. We would also like to thank Sarka Jahodova for sample contribution. The authors are thankful to all the sample contributors listed in Tables 1 and 2. We would like to thank two anonymous reviewers for their constructive comments which were invaluable while preparing this manuscript. Finally, we would like to thank Matthew Perisin from Department of Ecology and Evolution, University of Chicago, for language correction. This project was funded by Tromsø Museum, UiT—the Arctic University of Norway.
Author Contributions
DPR, TA, and IGA designed the project, obtained funding, and performed the fieldwork. MFA instructed DPR during primer optimization and laboratory work. DPR performed the laboratory work and wrote the manuscript. All coauthors commented on the manuscript.
Conflict of Interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online resource 1
Microsatellite library of H. persicum with 3904 sequences containing microsatellite motifs (TXT 1173 kb)
Online resource 2
List of 30 biologically and 134 bioinformatically validated microsatellite markers (XLSX 47 kb)
Fig. S1
Screen shot of some of the markers amplified in H. persicum in a singleplex. PET labeled marker Hp_02 with two alleles at 131.5 and 135.6, Hp_08 with two alleles at 170.6 and 173.5, and Hp_18 with single allele at 256.8 bp (base pair); HEX labeled marker Hp_19 with two alleles at 170 and 180.1, and Hp_05 with one allele at 125 bp; 6-FAM labeled marker Hp_09 with single allele at 159 bp; and NED labeled marker Hp_10 and Hp_16 with single allele each at 187.6 and 135.6 bp respectively (see Tables 3 and 4 of the paper for details) (PDF 81 kb)
Rights and permissions
About this article
Cite this article
Rijal, D.P., Falahati-Anbaran, M., Alm, T. et al. Microsatellite Markers for Heracleum persicum (Apiaceae) and Allied Taxa: Application of Next-Generation Sequencing to Develop Genetic Resources for Invasive Species Management. Plant Mol Biol Rep 33, 1381–1390 (2015). https://doi.org/10.1007/s11105-014-0841-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0841-y