Abstract
As self-incompatibility is a major issue in pummelo breeding and production, its mechanism in citrus was analyzed to improve breeding efficiency and reduce production costs. Rutaceae belongs to S-RNase type of gametophytic self-incompatibility. While the function of S-RNase/SLF and the mechanism of self-incompatibility have been studied extensively, the transcriptional regulation of S-RNase has been less studied. We performed transcriptome sequencing with the styles of ‘Shatian’ pummelo on the day of anthesis and 1–5 days before anthesis, and found that the transcript level of S-RNase gradually decreased with flower development. By analyzing differentially expressed genes and correlation with the expression trend of S-RNase, we identified a candidate gene, CgHSFB1, and utilized biochemical experiments such as yeast one-hybrid assay, electrophoretic mobility shift assay and dual-luciferase assay, as well as transient transformation of citrus calli and Citrus microcarpa and demonstrated that CgHSFB1 could directly bind to the S1-RNase promoter and repress the expression of S1-RNase, which is involved in the pummelo self-incompatibility response. In contrast, CgHSFB1 did not bind to the promoter of S2-RNase, and there was specificity in the regulation of S-RNase.
Key message
Transcription factor CgHSFB1 regulates the expression of S -RNase and is thus involved in self-incompatibility in citrus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Self-incompatibility (SI) is a prevalent mechanism of reproductive isolation that has evolved naturally in flowering plants. It allows plants to distinguish and reject self-incompatible pollen, while accepting intraspecific hybrid-compatible pollen. This mechanism serves to avert the negative effects of inbreeding depression, prevent inbreeding and species deterioration, and preserve the genetic diversity within species (Silva and Goring 2001; Hiscock and McInnis 2003).
Four molecular mechanisms of SI types have been reported for true dicotyledons. S-RNase-mediated gametophyte self-incompatibility (GSI) is phylogenetically more widespread and occurs in Solanaceae, Rosaceae, Plantaginaceae, Brassicaceae, and Cactaceae (Anderson et al. 1986; Lai et al. 2002; Okada et al. 2008; Liang et al. 2020; Ramanauskas and Igić 2021). The style determinant gene encodes a class of T2 family ribonucleases, also known as S ribonucleases (S-RNase), and the pollen determinant gene encodes a class of F-box proteins (SLF/SFB, S-locus F-box/S-haplotype-specific F-box). When the pollen S genotype matches one of the S genotypes carried by the style, proper growth of the pollen tube in the style does not occur. However, if the pollen S genotype differs from the style, the pollen tube grows properly, leading to successful fertilization of the pollen (Murfett et al. 1994; Kubo et al. 2010; Li et al. 2014). Brassicaceae represent a sporophytic self-incompatible (SSI) system in which the SSI response is regulated by two closely linked genes known as the S-determinant genes located at the S-locus. Additionally, the androgynous determinant genes encode a specific class of small secretory peptides known as SP11 (S-locus protein 11) or SCR (S-locus cysteine-rich protein) (Schopfer et al. 1999; Takayama et al. 2000). These peptides are localized on the surface of the pollen. The pistil S-determinant gene encodes a transmembrane serine/threonine receptor kinase (SRK, S-locus receptor kinase), which is localized to the plasma membrane of papillocytes in the stylar tissue (Takasaki et al. 2000). In SI reactions, SCR from the same S haplotype directly and specifically interacts with SRK proteins to inhibit the growth of self-pollen tubes (Kachroo et al. 2001; Shimosato et al. 2007). In the GSI type mediated by PrsS (Papaver rhoeas stigma S determinant) proteins, represented by the Papaveraceae, the style determinant encodes a class of small-molecule hydrophilic secreted proteins (Wheeler et al. 2009), whereas the pollen determinant gene encodes a class of small-molecule transmembrane proteins (PrpS, Papaver rhoeas pollen S). Stylar PrsS proteins undergo mutual recognition with PrpS proteins in self-pollen grains and undergo SI reactions in pollen grains, preventing pollen germination (de Graaf et al. 2006; Wilkins et al. 2015). The affinity of heterozygous SI of styles, represented by Primulaceae, is related to different flower morphologies (Li et al. 2016).
S-RNase type GSI have mainly focused on the function of S-RNase/SLF at the S-locus and the mechanism of SI. However, little has been reported on S-RNase transcriptional regulation. The MDF modification site, which is not chained to the S-locus, can specifically affect the expression of S13-RNase (Tsukamoto et al. 2003). The S1 specific modifier that is not linked to the S-locus leads to self-compatibility by functionally regulating S1 expression and is most likely a transcription factor (Li et al. 2023). The transcription factor CgHB40 was found to be involved in the regulation of S2-RNase in ‘Guiyou No.1’ pummelo, which in turn is involved in the affinity mutation of ‘Guiyou No.1’ pummelo (Hu et al. 2021). Environmental factors that are closely related to transcription factors have been studied frequently for the transformation of SI in plants (Gray et al. 1991). However, little is known about the upstream transcriptional regulation of the female determinant S-RNase.
Citrus is a perennial woody fruit tree of the Rutaceae family and is recognized as one of the best fruits worldwide. Rutaceae belongs to the GSI of S-RNase types, and the S-locus usually contains one S-RNase specifically expressed in the styles and about 14 SLFs specifically expressed in the anthers. ‘Shatian’ pummelo (Citrus maxima) has the typical gametophytic self-incompatibility characteristics, and the self-fertilization rate was only 0.3%. After self-pollination, the pollen tube growth stops at 1/3 of the style (Fig. 1a), and no seeds were observed in the transverse and longitudinal sections of fruits obtained from self-pollination of ‘Shatian’ pummelo (Fig. 1b) (Hu et al. 2021). S-RNase is specifically expressed in the style, and its transcript level is correlated with the period of floral development. It was found to be the lowest in Solanaceae at the bud stage and gradually increased with floral development. It gradually increases with flower development (Roldán et al. 2010).
Styles stained with aniline blue after self-pollination and observations on fruits. a Representative fluorescence images of aniline blue staining of styles collected 5 days after pollination of ‘Shatian’ pummelo. b Representative images of fruits after self-crossing of Shatin pomelo. Pollen tubes (pt) and vascular bundles (vb) are indicated by different arrows
The gene expression of S-RNase, a female determinant, plays an important role in species affinity, and its transcript levels are spatio-temporal and tissue-specific. In this study, we analyzed the stylar transcriptome data of ‘Shatian’ pummelo at different developmental periods and identified an HSF family gene, CgHSFB1, as a candidate gene to regulate S1-RNase in ‘Shatian’ pummelo, which was significantly correlated with the expression of S1-RNase and whose function was confirmed by transient transformation of citrus calli. Exploring the upstream transcriptional regulation of S-RNase can deepen our understanding of the genetic basis of the citrus SI system and provide theoretical guidance for solving the agricultural problem of flower pollination in ‘Shatian’ pummelo.
Experimental materials and methods
Plant material
‘Shatian’ pummelo (Citrus maxima) is the main cultivar in Rong County, Guangxi Zhuang Autonomous Region, with typical GSI traits. The pummelo used in the experiment was grown in Chuangtong Pummelo Farm, Ziliang Town, Rong County (110° 39′ 48.679ʺ E, 23° 4′ 38.334ʺ N), and the age of the trees was approximately 8 years. The flowering period of ‘Shatian’ pummelo is usually in early March, and it enters the ripening period after the frost season.
Sample collection
At the beginning of March, during the full blooming period of the ‘Shatian’ pummelo, we collected pairs of styles at different stages of development. These stages included Stage 1 (S1) which was 5 days before flowering, Stage 2 (S2) which was 4 days before flowering, Stage 3 (S3) which was 3 days before flowering, Stage 4 (S4) which was 2 days before flowering, Stage 5 (S5) which was 1 day before flowering, and Stage 6 (S6) which was the day of flowering. The collected styles were immediately snap-frozen in liquid nitrogen and then stored in a − 80 °C freezer for subsequent sequencing.
RNA-Seq and data processing
A total of 18 samples were collected from six treatments that included stylar tissue samples from S1, S2, S3, S4, S5 and S6. Total RNA from plant tissues was treated with DNase I, mRNA was enriched using oligo (dT) magnetic beads, mRNA was mixed with fragmentation buffer to fragment into short fragments, and cDNA was synthesized using the mRNA fragments as templates. The sample libraries were sequenced using the BGISEQ-500 sequencing platform (BGI, China) after quality control and quantification, and the raw data obtained by the downstream machine may have affected subsequent sequencing of the libraries, subsequent analyses, and assembly. Raw reads were filtered using fastp to obtain high-quality clean reads. The published HWB pummelo (Citrus maxima) genome was used as reference. The reproducibility between samples was assessed using Spearman’s rank correlation, and expression levels were calculated as transcripts per million mapped reads per thousand bases (TPM). Differentially expressed genes (DEGs) in the transcriptome data were analyzed using DEseq2 in the R package. The threshold for determining significant DEGs was a log2FoldChange > 1, with an adjusted P-value of less than 0.01. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment of DEGs were performed using TBtools-II v2.037 to identify the biological processes and metabolic pathways associated with the DEGs (Chen et al. 2023a).
Phylogenetic analysis and multiple sequence comparison of HSFB1
HWB pummelo (Citrus maxima) genomic data was downloaded from the Citrus Pan-genome to Breeding Database (CPBD) (http://citrus.hzau.edu.cn/), and Arabidopsis thaliana HSF protein sequences were downloaded from The Arabidopsis Information Resource (TAIR) database (https://www.arabidopsis.org/). Multi-protein sequence comparison analysis of HSF was performed using the MEGA 11.0 software and Cluster W tool, and a phylogenetic evolutionary tree of HSF protein sequences of candidate genes and Arabidopsis was constructed using default parameter values and the maximum likelihood method. Multiple comparisons of the amino acid sequences of HSFB1 from Arabidopsis thaliana, Oryza sativa, Prunus persica, Malus domestica, and HWB pummelo (Citrus maxima) were analyzed using Cluster W.
Subcellular localization assay
Full-length cDNA of CgHSFB1 without a stop codon was amplified using PCR with the primers listed in Supplementary Table 1 and inserted into plasmid pRI121. Recombinant plasmid (CgHSFB1::eGFP) and pRI121 empty vector (35S::eGFP) were transformed into strain GV3101, respectively. 35S::eGFP or CgHSFB1::eGFP was mixed with H2B-mCherry in equal volumes and injected into Nicotiana benthamiana leaves, which was observed 3 days later under a laser confocal microscope (Leica SP8).
Yeast one-hybrid assay
The promoter fragment of S1-RNase from ‘Shatian’ pummelo was cloned into the pAbAi vector to generate the bait plasmid pAbAi-S1-RNasepro. The CDS of CgHSFB1 was fused to the GAL4 activation structural domain (AD) in the pGADT7 vector to generate the prey construct AD-CgHSFB1. The detailed methodology for the yeast one-hybrid (Y1H) assay has been described previously (Duan et al. 2020).
Dual-luciferase transient expression assay
Promoter sequences were inserted into pGreen0800-LUC upstream of the LUC CDS to generate a promoter-LUC reporter vector. The CDS of CgHSFB1 was cloned into the overexpression vector pK7WG2D using a Gateway Cloning System (Invitrogen). Empty vector (EV) pK7WG2D was used as a negative control. Assays were performed as previously described (Lu et al. 2018).
Electrophoretic mobility shift assay
The CDS of CgHSFB1 without a stop codon was cloned into pET-32a to generate the recombinant vector, CgHSFB1-His. The recombinant construct was confirmed by sequencing and transformed into E. coli strain BL21 (DE3). The promoter fragment containing the binding motif was synthesized and labeled using Biotin (Sangon Biotech). Complementary oligonucleotides were annealed at 100 °C and used for electrophoretic mobility shift assay (EMSA). Unlabeled probes with identical or mutant oligonucleotides were used as cold and mutant competitors. The detailed methods have been described previously (Zhu et al. 2021).
Transient transformation of citrus calli and Citrus microcarpa styles
The S1-RNasepro::S1-RNase::GFP-CAM35S::CgHSFB1 fragment was cloned into the pGGK A-G vector using the golden gate cloning system, the S1-RNasepro::S1-RNase::GFP fragment was cloned into the pGGK A-G vector to serve as a negative control (Lampropoulos et al. 2013), and citrus calli tissues were transformed in detail as described previously (Zhu et al. 2021). Agrobacterium tumefaciens transiently infiltrated the Citrus microcarpa styles twice, each time by vacuum infiltration for 3 min, washed with sterile water 3–5 times, and then placed in co-culture medium for 3 days. Whether GFP was expressed in the styles was detected using the excitation light of 440–460 nm, and then the styles that had been successfully transformed transiently were screened.
Results
Transcriptome analysis of stylar developmental stages in ‘Shatian’ pummelo
We performed RNA-seq sequencing of 18 different samples of stylar tissues (S1, S2, S3, S4, S5, and S6) for transcriptomic study of pre-flowering stylar of ‘Shatian’ pummelo. After quality assessment of the RNA-Seq data, a total of 97.7 GB of high-quality clean reads, each containing at least 5 GB of clean data with a Q 30 quality score of ≥ 91.7%, were obtained for the 18 samples (Supplementary Table 2), which were mapped to the reference genome of HWB C. maxima.
We used TPM to calculate and analyze gene expression abundance. To detect relationships between samples, Spearman correlation coefficients were calculated for each group of three replicate libraries, and the R2 values between different biological replicates were found to be greater than 0.98 (Supplementary Fig. 1). Principal component analysis (PCA) based on the expression in different samples showed that the three biological replicates in each group were highly similar, and these data could be used for further analysis. The results of the PCA showed that the first two principal components explained 89% and 7% of the variance between the samples (Fig. 2a).
Transcriptome analysis of styles at different developmental periods. a Principal component analysis (PCA) of RNA-Seq data from styles. The data show the consistency among the three biological replicates. b DEGs in the style tissues at different developmental stages. The numbers of DEGs are indicated. c Differentially expressed transcription factors in the style tissues at different developmental stages. The numbers of DEGs are indicated. d representative GO terms enriched in 418 co-DEGs. e Significantly enriched KEGG pathways in the 418 co-DEGs
Comparison of DEGs at different developmental stages
S1 and S2–S6 were compared to identify the DEGs associated with S-RNase expression in ‘Shatian’ pummelo. In the comparison of “S1_vs_S2,” “S1_vs_S3,” “S1_vs_S4,” “S1_vs_S5,” and “S1_vs_S6,” 724, 2 433, 3 514, 4 138, and 6 187 DEGs and 418 co-DEGs were identified, respectively (Fig. 2b). 84, 183, 251, 276, and 416 differentially expressed transcription factors were identified in each combination (Fig. 2c), and a total of 549 differentially expressed transcription factors were identified in the five combinations. GO analysis showed that 418 co-DEGs were involved in biological processes, such as “response to abiotic stimulus,” “secondary metabolic processes,” “response to inorganic substances,” “response to temperature stimulus,” and “response to heat” (Fig. 2d). KEGG analysis showed that the 418 co-DEGs were significantly enriched in “plant hormone signal transduction,” “flavonoid biosynthesis,” “signal transduction,” and “environmental information processing” (Fig. 2e).
Identification of candidate TFs involved in the regulation of S-RNase
The expression of S1-RNase and S2-RNase transcript levels in ‘Shatian’ pummelo decreased gradually with style development, and the highest expression of the transcript levels is in the S1 (5 days before flowering) (Fig. 3a). The results are consistent with Liang et al. (2020). A total of 549 differentially expressed transcription factors were subjected to trend analysis to identify candidate TFs that regulate the expression of S-RNase in ‘Shatian’ pummelo style development. Modules 8, 10, and 15 showed opposite expression trends to those of S-RNase, whereas modules 6, 13, and 19 showed similar expression trends (Fig. 3b). A total of 21 TFs were identified based on the expression levels and trends of the genes, including TFs of the MYB, HSF, and NAC families (Fig. 3c). Among them, “response to temperature stimulus” and “response to heat” biological processes were significantly enriched, and HSF was strongly linked to ambient temperature, which influences plant affinity. Therefore, it was hypothesized that this CgHSF might be involved in the regulation of S-RNase in ‘Shatian’ pummelo.
Screening candidate genes and gene sequence analysis. a Expression of stylar S-RNase at each developmental stage. b Expression modules that are consistent with or opposite to the expression trend of S-RNase. c Expression analysis of candidate genes. d Phylogenetic analysis of candidate genes and Arabidopsis HSF family. The scale bar represents 0.5 substitutions per site. e Relative expression of CgHSFB1 in different tissues. The data are expressed as means ± SD of three replications. f Multiple sequence comparison of CgHSFB1 and its related proteins in Arabidopsis thaliana, Oryza sativa, Prunus persica, Malus domestica. The horizontal lines mark the three conserved structural domains
Sequence analysis and expression analysis of CgHSFB1
Plant HSFs are classified into A, B, and C based on the structural properties of the DNA-binding domain oligomerization domain, and the presence of class-specific motifs (each class has been characterized) (Scharf et al. 2012). To understand the class to which the candidate CgHSF belongs, a phylogenetic tree of CgHSF and Arabidopsis HSFs proteins was constructed. We identified Cg1g002680 as CgHSFB1, which belongs to class B (Fig. 3d). Class B HSFs contain the conserved amino acid motif LFGV in their CTD, which confers deterrent protein functions (Czarnecka-Verner et al. 2004; Ikeda and Ohme-Takagi 2009). Multiple sequence comparisons with homologous genes from Arabidopsis thaliana, Oryza sativa, Prunus persica, and Malus domestica showed that these proteins contained three conserved regions: the HSF DNA-binding domain, oligomerization domain (HR-A/B), and repressor domain (RD) (Fig. 3f). We successfully cloned CgHSFB1 from ‘Shatian’ pummelo, which encodes 270 amino acids, with a calculated molecular weight of 30.12 kDa and a theoretical pI of 6.85. To investigate the spatio-temporal expression pattern of CgHSFB1, RT-qPCR was performed to determine transcript levels of CgHSFB1 in different tissues. CgHSFB1 was expressed in various tissues and showed relatively high expression in the leaves and filaments (Fig. 3e). The transcript levels of CgHSFB1 gradually increased during style development, and its expression trend was opposite to that of S-RNase.
CgHSFB1 localizes to the nucleus and inhibits S 1 -RNase expression
To detect the subcellular localization of CgHSFB1, we constructed a CgHSFB1-eGFP fusion vector and co-injected it into Nicotiana benthamiana leaves, using H2B as a nuclear localization marker. Laser confocal microscopy revealed that the green fluorescence of CgHSFB1-eGFP colocalized with the red fluorescence of the nuclear localization marker, indicating that CgHSFB1-eGFP was localized in the nucleus (Fig. 4a). To confirm the interaction between CgHSFB1 and the S-RNase promoter, we performed a Y1H assay. Yeast cells co-transformed with the positive control (pGADT7-p53 + p53-AbAi), CgHSFB1 (pGADT7-CgHSFB1 + pAbAi-S1-RNasepro), and negative control (empty vector pGADT7 + pAbAi-S1-RNasepro) on defective nutrient medium (SD/-Leu) grew well. However, only yeast cells co-transformed with the positive control or CgHSFB1 could survive on selective medium supplemented with 200 ng ml−1 aureobasidin A (SD/-Leu/AbA200) (Fig. 4b), whereas yeast cells co-transformed with pGADT7-CgHSFB1 + pAbAi-S2-RNasepro did not survive on selective medium supplemented with 500 ng ml−1 aureobasidin A (SD/-Leu/AbA500) (Fig. 4h).
Interaction of CgHSFB1 protein with S1-RNase promoter. a Subcellular localization of CgHSFB1 protein. 35S::eGFP, pRI121 empty vector; CgHSFB1::eGFP, pRI121 vector containing CgHSFB1; eGFP, green fluorescence; mCherry, red fluorescence; Bright Field, bright field; Merged, overlaid map of eGFP and mCherry signals. Scale bar is 100 μm. b Y1H shows CgHSFB1 binding to the S1-RNase promoter. Positive control (p53-AbAi + pGADT-p53), negative control (pAbAi-S1-RNasepro + PGADT7); SD/-Leu, leucine-free SD medium; SD/-Leu/AbA200, leucine-free SD medium supplemented with AbA at a concentration of 200 ng ml−1. c Schematic of vectors used for dual luciferase assay. EV, pK7WG2D empty vector; CgHSFB1, pK7WG2D vector containing CgHSFB1. d Dual luciferase showing the relative inhibitory activity of CgHSFB1 on the S1-RNase promoter compared with EV. e Transient luciferase imaging assay showing co-injection of S1-RNasepro-LUC compared with EV region shows a relatively weak fluorescent signal. f EMSA shows that the CgHSFB1 binds to the S1-RNase promoter sequence. 2 μL of protein for each lane. EMSA was performed with a biotin-labeled S1-RNase promoter fragment containing TATAATCTTCC. The unlabeled S1-RNase promoter fragment of the same sequence and the mutated sequence (AAAAAAAAAAAA) were used as competition probes, respectively. Empty proteins were used as negative controls. ‘+’ and ‘−’ symbols indicate the presence and absence of specific probes or proteins, respectively. g No difference in fluorescence signal between the co-injected regions of CgHSFB1 and S2-RNasepro-LUC and the co-injected regions of control EV and S2-RNasepro-LUC as detected by transient luciferase imaging. h Y1H shows CgHSFB1 not binding to the S2-RNase promoter. SD/-Leu/AbA500, leucine-free SD medium supplemented with AbA at a concentration of 500 ng ml.−1. Error lines indicate the standard deviation of three biological replicates. Asterisks indicate statistically significant differences (Student’s t test, *P < 0.05)
Additionally, we performed a dual-luciferase assay to investigate the effects of CgHSFB1 on S1-RNase promoter activity. The promoter of S1-RNase was inserted into the LUC reporter vector, and the CDS of CgHSFB1 was cloned into the effector vector (Fig. 4c). In the presence of CgHSFB1, the relative luciferase expression driven by the S1-RNase promoter was significantly lower than that of the control (empty vector, EV) (~ threefold), suggesting that CgHSFB1 repressed the S1-RNase promoter (Fig. 4d). The binding of CgHSFB1 to the S1-RNase promoter was further verified using a transient luciferase imaging assay, in which strong fluorescent signals were detected in the co-injected region of CgHSFB1 and S1-RNasepro-LUC, whereas the co-injected region of the control EV and S1-RNasepro-LUC showed relatively weak fluorescent signals (Fig. 4e). These results suggested that CgHSFB1 represses the S1-RNase expression by directly binding to motifs in the promoter. No binding between CgHSFB1 and S2-RNase promoter was detected by transient luciferase imaging, and there was no difference in fluorescence signals between the co-injected regions of CgHSFB1 and S2-RNasepro-LUC and the co-injected regions of control EV and S2-RNasepro-LUC (Fig. 4g). These results indicate that CgHSFB1 directly binds to the S1-RNase promoter and represses S1-RNase expression, but not to the S2-RNase promoter, suggesting that there is specificity in the regulation of citrus S-RNases.
HSF proteins can bind to specific DNA sequences such as GAA(G)NTTC(T), and we identified a DNA sequence in the S1-RNase promoter that could potentially bind. We then performed an EMSA to test whether the CgHSFB1 protein could bind to the S1-RNase promoter at this potential binding site. Recombinant CgHSFB1-His showed bands with labeled DNA probes, indicating the presence of protein-DNA complexes, whereas empty protein controls did not. Upon addition of excess unlabeled DNA probes, the shifted bands dramatically diminished. In contrast, CgHSFB1-His still showed bands with labeled DNA probes when excess mutant unlabeled DNA probes were added (Fig. 4f). These data suggest that CgHSFB1 specifically binds to sites in the S1-RNase promoter.
CgHSFB1 inhibits transcription of S 1 -RNase in transgenic citrus calli and Citrus microcarpa
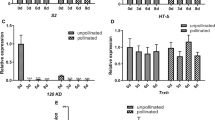
To validate the function of CgHSFB1 in citrus ontogeny, we transiently overexpressed it in citrus calli. Citrus calli have been shown to be the best endogenous expression system for studying the functions of TFs in citrus (Lu et al. 2018). As citrus calli do not contain S1-RNase, a binary expression vector was constructed and transformed into Agrobacterium tumefaciens to transiently infiltrate citrus calli (Fig. 5a). The citrus calli after 3 d of co-cultivation did not undergo significant changes (Fig. 5b). The DNA of the infested citrus calli was extracted and the target fragment was detected by PCR, and it was found that all infested citrus calli contained the target fragment. The expression of S1-RNase was verified by RT-qPCR in the S1-RNasepro::S1-RNase transgenic line versus the S1-RNasepro::S1-RNase-CAM35S::CgHSFB1 transgenic line. Three replicate experiments revealed that CgHSFB1 suppressed S1-RNase expression (Fig. 5c–e). After Agrobacterium tumefaciens containing the target gene infested the Citrus microcarpa styles, the post-infested styles were observed using a 440–460 nm excitation light source, and it was found that there was no significant difference in GFP fluorescence between EV and CgHSFB1 (Fig. 5f). Stigmas with GFP fluorescence were selected, and gene expression in transiently infiltrated styles was verified by RT-qPCR. It was observed that, consistent with the experimental results of transiently transformed citrus calli, the expression of S1-RNase was significantly reduced after overexpression of CgHSFB1 (Fig. 5g, h). The inhibition of S1-RNase expression by CgHSFB1 was verified in plants by transient transformation of Citrus microcarpa.
CgHSFB1 inhibits S1-RNase expression. a Schematic diagram of vectors used for transient transformation. EV, pGKK vector containing only S1-RNasepro::S1-RNase. CgHSFB1, pGGK vector containing S1-RNasepro::S1-RNase-CAMV35S::CgHSFB1. b Transient infestation of citrus calli. c Detection of the presence of target genes in citrus calli after transient infestation using PCR. 1. Maker: DL5 000; 2. \(\text{EV}-_{\text{s}_{1}}-\text{RNase}\): Agrobacterium tumefaciens containing EV vectors infiltrated the citrus calli and detected the S1-RNase gene (positive fragment size: 521 bp); 3. \(\text{CgHSFB1}-_{\text{s}_{1}}-\text{RNase}\): Agrobacterium tumefaciens containing CgHSFB1 vectors infiltrated the citrus calli and detected the S1-RNase gene (positive fragment size: 521 bp); 4. CgHSFB1-CgHSFB1: Agrobacterium tumefaciens containing CgHSFB1 vectors infiltrated the citrus calli and detected the CgHSFB1 gene (positive fragment size: 753 bp). d RT-qPCR assay for relative expression of CgHSFB1 after transient transformation of EV, CgHSFB1 in citrus calli. e RT-qPCR assay for relative expression of S1-RNase after transient transformation of EV, CgHSFB1 in citrus calli. f GPF assay after transient transformation of Citrus microcarpa styles, styles with GFP fluorescence are shown in the dashed ellipse. g RT-qPCR assay for relative expression of CgHSFB1 after transient transformation of EV, CgHSFB1 in Citrus microcarpa. h RT-qPCR to detect the relative expression of S1-RNase after transient transformation of EV and CgHSFB1 in Citrus microcarpa. Error lines indicate the standard deviation of three biological replicates. Asterisks indicate statistically significant differences (Student’s t test, *P < 0.05, **P < 0.01)
Discussion
‘Shatian’ pummelo has typical GSI. The female determinant S genotypes of ‘Shatian’ pummelo are S1-RNase and S2-RNase. We found that CgHB40 was involved in regulating the expression of S2-RNase (Hu et al. 2021). However, few studies have reported the regulation of S-RNase. As a female determinant of GSI, the expression of S-RNase has a decisive role in the SI of species. The deletion or inactivation of S-RNase results in the loss of SI. For example, the early termination of the gene due to S-RNase shift mutation results in the loss of pollen repressor activity of the short-truncated S-RNase protein, resulting in the loss of SI in citrus (Liang et al. 2020); the low expression of S-RNase is associated with the loss of SI in cultivated tomato (Kondo et al. 2002), and changes in S-RNase expression affect plant SI. The low expression of S2-RNase resulted in self-compatibility of ‘Guiyou No.1’ pummelo (Hu et al. 2021). In the present study, we used RNA-Seq data to identify a citrus HSF transcription factor, CgHSFB1, which has a potential role in regulating the expression of S1-RNase and modulating SI in citrus.
CgHSFB1 is involved in the regulation of SI in citrus
Heat shock factors play a key conservation role in the plant transcriptional network that regulates the expression of heat-responsive genes and plant HSFs are categorized as classes A, B, and C. Phylogenetic analysis showed that the candidate CgHSF is a homolog of AtHSFB1, which belongs to the class B HSF genes. The highly conserved LFGV tetrapeptide in class B HSFs constitutes the core structural domain of deterrent proteins. Currently, HSFB1 is mainly reported to be a transcriptional deterrent involved in heat tolerance response in plants. Analysis of Arabidopsis hsfB1/hsfB2b double KO plants showed that class B HSFs repress heat stress gene expression during recovery and suppress pathogen resistance by controlling defensin Pdf1.2 gene expression (Kumar et al. 2009; Ikeda et al. 2011). Overexpression of HSFB1 increases heat tolerance in heat-sensitive plants, and inhibition of HSFB1 decreases heat tolerance in grapevines (Chen et al. 2023b). In tomatoes, HsfB1 overexpression stimulates the coactivator function of HsfB1 and the accumulation of heat stress-related proteins. Plants overexpressing HsfB1 show abnormal growth and development and increased heat tolerance (Fragkostefanakis et al. 2019).
The transcript level of CgHSFB1 was closely related to the expression level of S-RNase. We found that CgHSFB1 repressed the expression of S1-RNase and participated in the regulation of citrus SI, linking citrus SI to heat shock family genes. CgHSFB1 is expressed in various tissues, and its transcript levels tend to be opposite to S-RNase during style development. The Y1H and LUC assays demonstrated an interaction with S1-RNase. CgHSFB1 directly bound to and repressed the S1-RNase promoter, as determined by EMSA and dual-luciferase assays (Fig. 4). Consistent with this result, the expression of S1-RNase was significantly reduced in the transiently transgenic citrus calli and Citrus microcarpa styles. (Fig. 5), which would subsequently be involved in the regulation of SI in citrus. Unfortunately, because of the difficulty and long duration of transgenesis in this citrus species, no stable transformation of CgHSFB1 was performed to verify its function.
HSF and environmental temperature are closely linked. While temperature affects plant affinity, high-temperature treatment of weedy clover can cause the temperature-sensitive T gene to interact with the SI S gene, thus inducing high-temperature SI (Townsend 1971). In Asteraceae 35–40 ℃ high-temperature treatment for 24–48 h can make them obtain self-compatible by affecting the SI of the style, and the style CLE45 induces self-compatibility by protecting the growth of pollen tube under high-temperature (Endo et al. 2013). Lilium longiflorum ‘Aral No. 5’ pistil immersed in water at 50 °C for 4–5 min could break the SI (Campbell and Linskens 1984). S-RNase activity was significantly reduced by heating ‘Somei-yoshino’ styles in a 50 °C water bathing. Furthermore, S-RNase activity decreased with increasing incubation time (e.g., 1, 3, 5 min). In addition, heat treatment at 50 °C promoted the growth of self-pollen tubes in the styles. Some self-pollen tubes even reached the bottom of the style. High-temperature treatments that reduced S-RNase activity could overcome Prunus stylar SI (Tsuruta et al. 2020). These findings imply that high temperatures have the potential to disrupt the SI of certain species, thereby raising the hypothesis that high temperatures may similarly impact the SI of citrus.
Upstream transcriptional regulation of S-RNase
SI in most flowering plants is mediated by S-RNase. Currently, there are more in-depth studies on the function of S-RNase/SLF and the mechanism of SI in several species such as petunia, potato, and pear. However, owing to the high polymorphism in the sequence of the S-locus, there are few studies on the upstream transcriptional regulation of S-RNase. The modification site MDF, which is not chained to the S site, was found in Petunia axillaris, and it can specifically affect the expression of S13-RNase, and the modification site MDF is likely to be a transcription factor (Tsukamoto et al. 2003). In Arabidopsis, the self-compatibility of the S1S1-homozygotes in the selfing population is not due to mutation of the S-locus, but rather to the functional disruption of S1 by a specific modifier that is not linked to the S-locus, and this leads to the loss of SI (Li et al. 2023). S-RNase expression level affects the SI of plants, and in ‘Guiyou No.1’ SI was lost due to the non-expression of S2-RNase. CgHB40 is involved in the regulation of S2-RNase (Hu et al. 2021). We found that CgHSFB1 can inhibit the expression of S1-RNase without binding to the promoter of S2-RNase, suggesting that the transcriptional regulation of S-RNase polymorphism is specific. CgHSFB1 and CgHB40 do not belong to the same gene family and have no sequence similarity, but the expression trend of both is opposite to that of S-RNase, and the expression of CgHSFB1 and CgHB40 gradually increased with pistil development. The presence of large variations in the S1-RNase and S2-RNase promoter sequences may be the reason for the specific regulation of S-RNase in ‘Shatian’ pummelo. Moreover, S-RNase expression was characterized by tissue-specific expression and spatial and temporal differential expression, suggesting that it is of great significance to analyze the regulatory characteristics of S-RNase and conduct research on the regulation of S-RNase. The results of this study deepen our understanding of the genetic basis of the SI system in citrus and lay the foundation for the transcriptional regulation of S-RNase.
Data availability
RNA-Seq data have been deposited in the CNGB Sequence Archive (CNSA) of the China National GeneBank DataBase (CNGBdb) with accession numbers CNP0005375 (https://db.cngb.org/). S1-RNase, S2-RNase accession numbers are MN652897, MN652898. The sequence data of the CgHSFB1 are available in CPBD (http://citrus.hzau.edu.cn) under the gene ID Cg1g002680.
References
Anderson MA, Cornish EC, Mau SL, Williams EG, Hoggart R, Atkinson A, Bonig I, Grego B, Simpson R, Roche PJ, Haley JD, Penschow JD, Niall HD, Tregear GW, Coghlan JP, Crawford RJ, Clarke AE (1986) Cloning of cDNA for a stylar glycoprotein associated with expression of self-incompatibility in Nicotiana alata. Nature 321:38–44. https://doi.org/10.1038/321038a0
Campbell RJ, Linskens HF (1984) Temperature effects on self incompatibility in Lilium longiflorum. Theor Appl Genet 68:259–264. https://doi.org/10.1007/BF00266900
Chen C, Wu Y, Li J, Wang X, Zeng Z, Xu J, Liu Y, Feng J, Chen H, He Y, Xia R (2023a) TBtools-II: a “one for all, all for one” bioinformatics platform for biological big-data mining. Mol Plant 16:1733–1742. https://doi.org/10.1016/j.molp.2023.09.010
Chen H, Liu X, Li S, Yuan L, Mu H, Wang Y, Li Y, Duan W, Fan P, Liang Z, Wang L (2023b) The class B heat shock factor HSFB1 regulates heat tolerance in grapevine. Hortic Res. https://doi.org/10.1093/hr/uhad001
Czarnecka-Verner E, Pan S, Salem T, Gurley WB (2004) Plant class B HSFs inhibit transcription and exhibit affinity for TFIIB and TBP. Plant Mol Biol 56:57–75. https://doi.org/10.1007/s11103-004-2307-3
de Graaf BHJ, Rudd JJ, Wheeler MJ, Perry RM, Bell EM, Osman K, Franklin FCH, Franklin-Tong VE (2006) Self-incompatibility in Papaver targets soluble inorganic pyrophosphatases in pollen. Nature 444:490–493. https://doi.org/10.1038/nature05311
Duan Z, Zhang Y, Tu J, Shen J, Yi B, Fu T, Dai C, Ma C (2020) The Brassica napus GATA transcription factor BnA5.ZML1 is a stigma compatibility factor. J Integr Plant Biol 62:1112–1131. https://doi.org/10.1111/jipb.12916
Endo S, Shinohara H, Matsubayashi Y, Fukuda H (2013) A novel pollen-pistil interaction conferring high-temperature tolerance during reproduction via CLE45 signaling. Curr Biol 23:1670–1676. https://doi.org/10.1016/j.cub.2013.06.060
Fragkostefanakis S, Simm S, El-Shershaby A, Hu Y, Bublak D, Mesihovic A, Darm K, Mishra SK, Tschiersch B, Theres K, Scharf C, Schleiff E, Scharf KD (2019) The repressor and co-activator HsfB1 regulates the major heat stress transcription factors in tomato. Plant Cell Environ 42:874–890. https://doi.org/10.1111/pce.13434
Gray JE, McClure BA, Bonig I, Anderson MA, Clarke AE (1991) Action of the style product of the self-incompatibility gene of Nicotiana alata (S-RNase) on in vitro-grown pollen tubes. Plant Cell 3:271–283. https://doi.org/10.1105/tpc.3.3.271
Hiscock SJ, McInnis SM (2003) Pollen recognition and rejection during the sporophytic self-incompatibility response: Brassica and beyond. Trends Plant Sci 8:606–613
Hu J, Xu Q, Liu C, Liu B, Deng C, Chen C, Wei Z, Ahmad MH, Peng K, Wen H, Chen X, Chen P, Larkin RM, Ye J, Deng X, Chai L (2021) Downregulated expression of S2-RNase attenuates self-incompatibility in “Guiyou No. 1” pummelo. Hortic Res 8:199
Ikeda M, Ohme-Takagi M (2009) A novel group of transcriptional repressors in arabidopsis. Plant Cell Physiol 50:970–975. https://doi.org/10.1093/pcp/pcp048
Ikeda M, Mitsuda N, Ohme-Takagi M (2011) Arabidopsis HsfB1 and HsfB2b act as repressors of the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance. Plant Physiol 157:1243–1254. https://doi.org/10.1104/pp.111.179036
Kachroo A, Schopfer CR, Nasrallah ME, Nasrallah JB (2001) Allele-specific receptor-ligand interactions in Brassica self-incompatibility. Science 293:1824–1826
Kondo K, Yamamoto M, Matton DP, Sato T, Hirai M, Norioka S, Hattori T, Kowyama Y (2002) Cultivated tomato has defects in both S-RNase and HT genes required for stylar function of self-incompatibility. Plant J 29:627–636. https://doi.org/10.1046/j.0960-7412.2001.01245.x
Kubo K-i, Entani T, Takara A, Wang N, Fields AM, Hua Z, Toyoda M, Kawashima S-i, Ando T, Isogai A, Kao T-h, Takayama S (2010) Collaborative non-self recognition system in S-RNase–based self-incompatibility. Science 330:796–799
Kumar M, Busch W, Birke H, Kemmerling B, Nürnberger T, Schöffl F (2009) Heat shock factors HsfB1 and HsfB2b are involved in the regulation of Pdf1.2 expression and pathogen resistance in Arabidopsis. Mol Plant 2:152–165. https://doi.org/10.1093/mp/ssn095
Lai Z, Ma W, Han B, Liang L, Zhang Y, Hong G, Xue Y (2002) An F-box gene linked to the self-incompatibility (S) locus of Antirrhinum is expressed specifically in pollen and tapetum. Plant Mol Biol 50:29–42. https://doi.org/10.1023/a:1016050018779
Lampropoulos A, Sutikovic Z, Wenzl C, Maegele I, Lohmann JU, Forner J (2013) GreenGate—a novel, versatile, and efficient cloning system for plant transgenesis. PLoS ONE 8:e83043. https://doi.org/10.1371/journal.pone.0083043
Li S, Sun P, Williams JS, Kao T-H (2014) Identification of the self-incompatibility locus F-box protein-containing complex in Petunia inflata. Plant Reproduction 27:31–45. https://doi.org/10.1007/s00497-013-0238-3
Li J, Cocker JM, Wright J, Webster MA, McMullan M, Dyer S, Swarbreck D, Caccamo M, Cv O, Gilmartin PM (2016) Genetic architecture and evolution of the S locus supergene in Primula vulgaris. Nat Plants 2:16188. https://doi.org/10.1038/nplants.2016.188
Li Y, Mamonova E, Köhler N, van Kleunen M, Stift M (2023) Breakdown of self-incompatibility due to genetic interaction between a specific S-allele and an unlinked modifier. Nat Commun 14:3420. https://doi.org/10.1038/s41467-023-38802-0
Liang M, Cao Z, Zhu A, Liu Y, Tao M, Yang H, Xu Q, Wang S, Liu J, Li Y, Chen C, Xie Z, Deng C, Ye J, Guo W, Xu Q, Xia R, Larkin RM, Deng X, Bosch M, Franklin-Tong VE, Chai L (2020) Evolution of self-compatibility by a mutant Sm-RNase in citrus. Nature Plants 6:131–142. https://doi.org/10.1038/s41477-020-0597-3
Lu S, Zhang Y, Zhu K, Yang W, Ye J, Chai L, Xu Q, Deng X (2018) The citrus transcription factor CsMADS6 modulates carotenoid metabolism by directly regulating carotenogenic genes. Plant Physiol 176:2657–2676. https://doi.org/10.1104/pp.17.01830
Murfett J, Atherton TL, Mou B, Gassert CS, McClure BA (1994) S-RNase expressed in transgenic Nicotiana causes S-allele-specific pollen rejection. Nature 367:563–566. https://doi.org/10.1038/367563a0
Okada K, Tonaka N, Moriya Y, Norioka N, Sawamura Y, Matsumoto T, Nakanishi T, Takasaki-Yasuda T (2008) Deletion of a 236 kb region around S4-RNase in a stylar-part mutant S4sm-haplotype of Japanese pear. Plant Mol Biol 66:389–400. https://doi.org/10.1007/s11103-007-9277-1
Ramanauskas K, Igić B (2021) RNase-based self-incompatibility in cacti. New Phytol 231:2039–2049. https://doi.org/10.1111/nph.17541
Roldán JA, Quiroga R, Goldraij A (2010) Molecular and genetic characterization of novel S-RNases from a natural population of Nicotiana alata. Plant Cell Rep 29:735–746. https://doi.org/10.1007/s00299-010-0860-6
Scharf KD, Berberich T, Ebersberger I, Nover L (2012) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. Biochim Biophys Acta 1819:104–119. https://doi.org/10.1016/j.bbagrm.2011.10.002
Schopfer CR, Nasrallah ME, Nasrallah JB (1999) The male determinant of self-incompatibility in Brassica. Science 286:1697–1700. https://doi.org/10.1126/science.286.5445.1697
Shimosato H, Yokota N, Shiba H, Iwano M, Entani T, Che F-S, Watanabe M, Isogai A, Takayama S (2007) Characterization of the SP11/SCR high-affinity binding site involved in self/nonself recognition in Brassica self-incompatibility. Plant Cell 19:107–117. https://doi.org/10.1105/tpc.105.038869
Silva NF, Goring DR (2001) Mechanisms of self-incompatibility in flowering plants. Cell Mol Life Sci 58:1988–2007. https://doi.org/10.1007/PL00000832
Takasaki T, Hatakeyama K, Suzuki G, Watanabe M, Isogai A, Hinata K (2000) The S receptor kinase determines self-incompatibility in Brassica stigma. Nature 403:913–916. https://doi.org/10.1038/35002628
Takayama S, Shiba H, Iwano M, Shimosato H, Che FS, Kai N, Watanabe M, Suzuki G, Hinata K, Isogai A (2000) The pollen determinant of self-incompatibility in Brassica campestris. Proc Natl Acad Sci USA 97:1920–1925. https://doi.org/10.1073/pnas.040556397
Townsend CE (1971) Further studies on the inheritance of a self-compatibility response to temperature and the segregation of S alleles in diploid alsike clover. Crop Sci 11:860–863. https://doi.org/10.2135/cropsci1971.0011183X001100060028x
Tsukamoto T, Ando T, Kokubun H, Watanabe H, Sato T, Masada M, Marchesi E, Kao T-h (2003) Breakdown of self-incompatibility in a natural population of Petunia axillaris caused by a modifier locus that suppresses the expression of an S-RNase gene. Sex Plant Reprod 15:255–263. https://doi.org/10.1007/s00497-002-0161-5
Tsuruta M, Iwaki R, Lian C, Mukai Y (2020) Decreased RNase activity under high temperature is related to promotion of self-pollen tube growth in the pistil of the Japanese Flowering Cherry, Prunus × yedoensis ‘Somei-yoshino.’ Hortic J 89:306–310
Wheeler MJ, de Graaf BHJ, Hadjiosif N, Perry RM, Poulter NS, Osman K, Vatovec S, Harper A, Franklin FCH, Franklin-Tong VE (2009) Identification of the pollen self-incompatibility determinant in Papaver rhoeas. Nature 459:992–995. https://doi.org/10.1038/nature08027
Wilkins KA, Bosch M, Haque T, Teng N, Poulter NS, Franklin-Tong VE (2015) Self-incompatibility-induced programmed cell death in field poppy pollen involves dramatic acidification of the incompatible pollen tube cytosol. Plant Physiol 167:766–779. https://doi.org/10.1104/pp.114.252742
Zhu K, Sun Q, Chen H, Mei X, Lu S, Ye J, Chai L, Xu Q, Deng X (2021) Ethylene activation of carotenoid biosynthesis by a novel transcription factor CsERF061. J Exp Bot 72:3137–3154. https://doi.org/10.1093/jxb/erab047
Acknowledgements
This research was financially supported by the National Natural Science Foundation of China (Grant Nos. 32122075, 32072523, 32302489), the Key-Area Research and Development Program of Guangdong Province (Grant No. 2022B0202070002), the China Agriculture Research System (Grant No. CARS-27), and Hubei Provincial Natural Science Foundation of China (Grant No. 2023AFB094).
Funding
Funding was provided by National Natural Science Foundation of China (Grant Nos. 32122075, 32072523, 32302489), the Key-Area Research and Development Program of Guangdong Province (Grant No. 2022B0202070002), the China Agriculture Research System (Grant No. CARS-27), and Hubei Provincial Natural Science Foundation of China (Grant No. 2023AFB094).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Lijun Chai conceived and coordinated the project; Chenchen Liu, Xin Zheng and Jianbing Hu designed the experiments; Chenchen Liu performed most of the experiments with contributions from Qiang Xu, Hao Wen, Zhezhong Zhang and Ran Liu; Xiangling Chen provided the plant materials; Chenchen Liu wrote the manuscript with help from Zongzhou Xie, Junli Ye and Xiuxin Deng.
Corresponding author
Ethics declarations
Conflict of interest
There are no competing interests in this paper, and the authors do not have any possible conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11103_2024_1475_MOESM2_ESM.jpg
Supplementary Fig. 2. Enrichment analysis between differentially expressed transcription factors. Supplementary file2 (JPG 771 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, C., Zheng, X., Hu, J. et al. Involvement of CgHSFB1 in the regulation of self-incompatibility in ‘Shatian’ pummelo. Plant Mol Biol 114, 77 (2024). https://doi.org/10.1007/s11103-024-01475-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01475-4