Abstract
Amylose biosynthesis is strictly associated with granule-bound starch synthase I (GBSSI) encoded by the Waxy gene. Mutagenesis of single bases in the Waxy gene, which induced by CRISPR/Cas9 genome editing, caused absence of intact GBSSI protein in grain of the edited line. The amylose and amylopectin contents of waxy mutants were zero and 31.73%, while those in the wild type were 33.50% and 39.00%, respectively. The absence of GBSSI protein led to increase in soluble sugar content to 37.30% compared with only 10.0% in the wild type. Sucrose and β-glucan, were 39.16% and 35.40% higher in waxy mutants than in the wild type, respectively. Transcriptome analysis identified differences between the wild type and waxy mutants that could partly explain the reduction in amylose and amylopectin contents and the increase in soluble sugar, sucrose and β-glucan contents. This waxy flour, which showed lower final viscosity and setback, and higher breakdown, could provide more option for food processing.
Key message
The new zero amylose barley was obtained via the CRISPR/Cas9 genome editing, showing higher soluble sugar content and lower final viscosity and setback.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Barley (Hordeum vulgare L.) is an ancient cereal crop that currently ranks fourth in production of grain crops worldwide after maize, wheat and rice. Approximately two-thirds of barley is used for feed, one-third is used for brewing and only a tiny fraction (2%) is used for food directly (Baik & Ullrich 2008). Barley is increasingly becoming a desirable food grain for human diets because of potential health benefits such as lowering glycemic load related to dietary fiber, high soluble fiber (β-glucan) content and nutritional value (Pins et al. 2007). Moreover, barley can be produced at high altitudes and in deserts with extreme climates, where wheat and most other cereal crops cannot grow; thus, barley is always a staple food source in harsh landscapes.
Starch is one of the main components (60–70%) of barley grains and usually contains about 25% amylose and 75% amylopectin. Amylose is essentially a linear molecule composed of glucosyl units connected by α-(1–4)-linkages together with a few branches. Amylopectin has many branches formed by α-(1,6)-bonds in addition to the α-(1,4)-linkages in the linear chains (Evers et al. 1999; Hebelstrup et al. 2017; Chen et al. 2020; Punia 2020). Barley can be classified into normal, waxy, and high amylose barley based on the amylose content of its grains. Normal barley typically contains 20–30% amylose, while waxy barley has minimal or no amylose (< 10%), and high amylose barley always possesses a higher proportion (> 30%) of amylose (Punia 2020). The content and ratio of amylose and amylopectin determine starch properties and characteristics and have a considerable influence on the processing, malting and food quality of barley grain (Chen et al. 2020). Waxy barley has many merits, such as high swelling power, pasting viscosity, freeze-thaw stability, breakdown value and thickening properties, and low pasting temperature and setback value (Jobling 2004; Sparla et al. 2014; Li et al. 2019). Furthermore, previous research has shown that waxy barley contains more β-glucan compared with non-waxy barley. Barley β-glucan is a soluble dietary fiber regarded as an important functional ingredient for lowering blood cholesterol, reducing glycemic response, regulating intestinal flora and maintaining weight (Izydorczyk & Dexter 2008; Thondre et al. 2011). Starch is usually stored in large, lens-shaped A-type granules (diameter 10–35 μm) and small, spherical B-type granules (diameter less than 2–7 μm) in barley endosperm. The small starch granules comprise 6–30% of the total starch mass of barley. Waxy barley contains fewer small B-granules by weight than high-amylose barley (with a higher amylose / amylopectin ratio) (Chen et al. 2020; De et al. 2020). During beer production, small B-type granules in barley grains are protected by a heterogeneous matrix (proteins and cell walls), allowing them to more easily escape degradation and cause technical problems (De et al. 2020). Therefore, waxy barley has extensive prospects for application in the functional food processing and brewing industries.
Starch biosynthesis involves a series of coordinated enzymes. Amylopectin biosynthesis requires ADP glucose pyrophosphorylase, soluble starch synthase, starch branching enzyme and starch debranching enzyme (Li et al. 2014). Amylose biosynthesis is strictly associated with granule-bound starch synthase I (GBSSI) encoded by the Waxy gene (Domon et al. 2002; Zeeman et al. 2010; Guzman & Alvarez 2016). The barley Waxy gene is located on the chromosome 7 H (17,089,965 − 17,094,192) reverse strand (MorexV3_pseudomolecules_assembly), contains 12 exons and 11 introns, and encodes the 60-KDa GBSSI protein. Variations in the Waxy gene or its regulatory sequences, such as point, insertion and deletion mutations, influence the function or catalytic activity of GBSSI protein and further affect amylose biosynthesis (Domon et al. 2002; Ma et al. 2013; Li et al. 2014, 2019; Hebelstrup et al. 2017). Only four allelic variations of the barley Waxy gene have been found to be relative to the low or zero amylose content. An approximately 400 bp deletion within the 5ʹuntranslated region of the barley Waxy gene, encompassing the transcriptional start site, TATA box and exon 1, results in a significant reduction in both Waxy gene expression levels and grain amylose content (Asare et al. 2012; Ma et al. 2013; Li et al. 2014). The C2453-to-T mutation in the Waxy gene halts peptide chain elongation of GBSSI, thereby impeding amylose biosynthesis in barley (Domon et al. 2002). Zero amylose barley ‘CDC Alamo’ exhibits normal Waxy transcription and GBSSI protein levels, but displays significantly reduced GBSSI activity due to three single-nucleotide polymorphism (SNP) mutations in the Waxy gene, which decrease GBSSI activity by 90% (Hebelstrup et al. 2017). A G3935-to-T mutation in the Waxy gene, resulting in the substitution of glycine at position 513 with tryptophan, is identified in zero-amylose barley cultivars ‘CDC Fibar’ and ‘Z999’ (Asare et al. 2012; Li et al. 2014). The alleles were rare relatively in barley improvement.
The recent emergence of CRISPR/Cas9 genome editing technology offers the potential to obtain novel genotypes in a much faster manner than traditional approaches, significantly accelerating crop breeding (Ma et al. 2016; Es et al. 2019; Huang et al. 2020; Qin et al. 2020; Xie et al. 2020). Here, we generated a novel waxy barley using the CRISPR/Cas9 system. The waxy mutants possessed zero amylose, higher soluble sugar, fewer small B-granules and increased β-glucan content in grains compared with the wild type barley.
Materials and methods
Plant materials
The non-waxy spring barley (Hordeum vulgare L.) ‘Golden Promise’ was grown in a greenhouse under controlled conditions with 18 °C during 16 h of light and 15 °C during 8 h of darkness. When immature embryos were 1.5 to 2 mm in diameter, immature grains were collected from the center of spikes. Immature embryos were used for Agrobacterium-mediated transformation.
SgRNA design and plasmid construction
Exon 1 of the Waxy gene in ‘Golden Promise’ was cloned and sequenced using the specific primers wxF/R (Table S1). Two target sites located in exon 1 of the Waxy gene were selected using the online software CRISPR-GE (http://skl.scau.edu.cn/home/). The CRISPR/Cas9 system consisting of PMED and pLGYE001 was a gift from the Crop Research Institute, Shandong Academy of Agricultural Sciences. A construct containing two sgRNAs driven by TaU3 was constructed according to previously published protocols (Romualdi et al. 2003; Qin et al. 2020). In brief, four primers, T1F, T1F0, T2R and T2R0 (Table S1), containing target sequences were synthesized, and the T1-gRNA-Ter-TaU3-T2 cassette was amplified using plasmid PMED as template in a 50 µL reaction (Sangon Biotech, Shanghai, China). This cassette was then digested using BsaI and ligated to pLGYE001 (NEbiolabs, Beijing, China). The recombinant vector pLGYE001-wx was transferred into Agrobacterium EHA105 competent cells (Huayueyang, Beijing, China).
Agrobacterium-mediated transformation of barley
Agrobacterium-mediated transformation of barley (Hordeum vulgare L.) ‘Golden Promise’ was performed as described (Sparla et al. 2014). In brief, immature embryos were separated from sterile grains and co-cultivated with Agrobacterium EHA105 harboring the CRISPR/Cas9 vector pLGYE001-wx at 24 °C for 3 days in the dark. Embryos were subsequently transferred to selective medium containing 5 mg/L glufosinate ammonium (PPT) (Coolaber, Beijing, China) and cultured at 24 °C for 6 to 8 weeks in the dark. Proliferating calli were transferred to transition medium containing 3 mg/L PPT and cultured at 24 °C under 16 h low light/8 h dark photoperiod. Small plantlets were transferred to regeneration medium containing 3 mg/L PPT and cultured at 24° under 16 h light/8 h dark photoperiod for 2 weeks. Putative transgenic plants were then transferred to soil and grown to maturity in the greenhouse under controlled conditions of 18 °C during 16 h light and 15 °C during 8 h darkness.
Detection of transgenic plants and mutations
Transgenic plants were identified using a PAT/bar test strip resistance kit for Bar protein (Youlong, Shanghai, China). Genomic DNA was isolated from leaves of putative transgenic plants using a TaKaRa MineBEAT Plant Genomic DNA Extraction Kit (TaKaRa, Beijing, China). For detection of T-DNA insertions, PCR with primers ubiF/R (Table S1) was performed using 100 ng of DNA in a 20 µL reaction (Sangon Biotech, Shanghai, China). For detection of mutations in transgenic plants, the specific PCR primers wxF/R were used for PCR amplification. Amplified DNA fragments of target genes were ligated into pGEM-Teasy Vector (Promega, Shanghai, China), and 20 positive clones were sequenced. Nucleotide sequencing results were analyzed using the AlignX program (Invitrogen, California, USA).
SDS-PAGE analysis of GBSSI protein
GBSSI protein was extracted according to previous reports with some modifications (Takeuchi et al. 2001; Hebelstrup et al. 2017). Fine powder was scraped from the endosperm of the grain, then homogenized in washing extraction buffer (55 mmol/L Tris-HCl, pH 6.8; 23 g/L SDS; 5% (v/v) β-mercaptoethanol). The mixture was incubated at 4 °C overnight and centrifuged at 9000 g for 10 min. The supernatant was discarded, and the sediment was washed twice with distilled water and finally dried overnight. GBSSI protein was released from the starch by boiling (10 mg) in 400 µL of protein extraction buffer (containing 62 mmol/L Tris-HCl, pH 6.8; 23 g/L SDS; 5% (v/v) β-mercaptoethanol; 10% (v/v) glycerol; 0.05 g/L bromophenol blue) for 5 min and centrifuged at 9000 g for 10 min. Protein supernatant (20 µL) was separated using SDS-PAGE gels (Sangon Biotech, Shanghai, China) for 3 h at 50 mA. SDS-PAGE gels were stained using a solution of 0.05% (w/v) Coomassie Blue R-250, 5% (v/v) ethanol and 12% (w/v) acetic acid for 3 h and then immersed in 10% (w/v) acetic acid overnight to remove excess stain.
Iodine staining and scanning electron microscopy of endosperm
Grains of wild type and edited lines were randomly selected. Transverse sections of grains were stained using 0.01% (w/v) I2–0.1% (w/v) KI solution. Stained starch granules were examined and images were captured using an Olympus BX53 microscope (Olympus Corp, Tokyo, Japan). The transverse sections of dried grain were attached to metallic stubs using carbon stickers and sputter-coated with gold for 30 s. The images were observed and captured using an SU8100 scanning electron microscope (Hitachi, Tokyo, Japan). Subsequently, the long axis and short axis of the granules were measured utilizing Image-pro plus 6.0 software (Media Cybernetics, Inc., Rockville, MD, USA). A total of twenty intact granules were measured in each category.
Expression analysis using qRT-PCR
Total RNA was extracted from developing grains of wild type and edited lines at 8, 16, 24 and 32 days post-anthesis (dpa) using an RNAprep Pure Plant Kit (Tiangen, Beijing, China). The cDNAs were synthesized using a PrimeScript™ RT reagent Kit with gDNA Eraser (TaKaRa, Tokyo, Japan). The gene-specific qPCR primers wxQF/R (Table S1) were designed according to the barley ‘Golden Promise’ Waxy mRNA sequence. The housekeeping gene α-Tubulin was co-amplified as a control for normalizing cDNA templates. qRT-PCR was performed in a 7500 Fast Real-Time PCR System (ABI, Carlsbad, USA) using TB Green™ Premix Ex Taq™ II (TaKaRa, Tokyo, Japan). Every sample was analyzed with three replicates.
Transcriptome analysis
Total RNA of immature grains of the wild type and edited line at 16 dpa was used for transcriptome analysis. Six libraries, with three biological replicates, were prepared and sequenced using an Illumina HiSeq 2000 system (Novogene, China). Raw reads were processed to obtain clean reads by removing low quality reads, and assembly of the clean reads was performed using Trinity. Functional annotation was performed by comparing all unigenes with the following databases: NCBI nonredundant protein sequences (Nr), Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO). Unigene expression levels were calculated using fragments per kb per million reads (FPKM) values. Unigenes with differential expression levels between the wild type and edited line were analyzed using a chi-square test with IDEG6 software (Thondre et al. 2011). The false discovery rate method was employed to determine the threshold P-value at a false discovery rate ≤ 0.001, and the absolute value of|log2ratio| ≥ 2 was used as the threshold to determine the significance of differential expression levels of unigenes. Significantly enriched GO and KEGG terms were obtained from the set of differentially abundant unigenes using the hypergeometric test.
Determination of grain quality and agronomic traits
T1 lines were generated from edited plants grown in the greenhouse under controlled conditions of 18 °C during 16 h light and 15 °C during 8 h darkness. Analysis of the agricultural traits of wild type and edited lines was performed using 20 plants per T1 line. Plant height, number of tillers, spike length and grains per spike were recorded. Six repetitions of thousand-grain weight for the wild type and edited lines were also recorded. The contents of amylose/amylopectin, soluble sugar, sucrose, β-glucan and protein contents of grains were determined, respectively, via the dual-wavelength iodine binding method (Zhu et al. 2008), phenol-sulfuric acid colorimetry method, high-performance liquid chromatography (Xing et al. 2014), streamlined enzymatic method (Yamamori & Endo 1996) and the Kjeldahl method (Zeeman et al. 2010). Each sample was measured with three biological replicates. The contents of amylose/amylopectin, soluble sugar, sucrose, β-glucan, and protein in the grain were expressed as a percentage of the dry weight.
Pasting properties of barley flour
The pasting properties of barley flour were detected using a Rapid Visco Analyser (RVA, Model 4D, Newport Scientific, Australia). Barley flour (1.5 g, dry basis) was accurately weighed into an aluminium RVA canister and distilled water was added to reach a total of 28 g. The samples were first heated at 50 °C for 1 min, then heated to 95 °C for 4.5 min, and then held at 95 °C for 2 min. Afterwards the samples were cooled to 50 °C and held at 50 °C for 2 min. The rotating speed of 160 rpm was maintained throughout the process. Pasting parameters including peak viscosity, trough viscosity, final viscosity, breakdown, setback, pasting temperature and pasting time were determined. Each sample was measured in three biological replicates.
Cooking quality of dried noodles
The 300 g mixed flour consisting of 51% barley flour and 49% wheat flour (w/w) was added into 120 g distilled water to produce dough. The noodles were made using noodle extruder (Joyoung, Jinan, China). The noodles were dried in an oven at 40 °C for 10 h to obtain dried noodles. Dried noodles were boiled in boiling water until there was no hard core observed in the center of the noodles. The boiled noodles were cooled to room temperature and then used for analysis. Sensory evaluation and cooking properties of dried noodles were carried out according to LS/T3202-1993 and LS/T 3212 − 2021 (Industry Standard of the People’s Republic of China). The texture properties and tensile strength of cooked noodles were determined using TMS-PRO texture analyzer (FTC, Virginia, USA). The texture properties were measured by P75 probe with a condition of 1.5 mm/s test speed, 70% compression ratio, 0.5 N trigging force. The tensile strength was measured by A/SPR probe at a test speed of 1.5 mm/s, tensile distance of 80 mm and 0.15 N initial force. Each sample was measured in three biological replicates.
Statistical analysis
The statistical software PASW Statistics 18 (IBM SPSS, Chicago, USA) was used for data analysis. A value of p < 0.05 was considered to be statistically significant, and all values from a sample were expressed as the mean ± standard deviation.
Results
RNA-guided Cas9-induced genome editing of Waxy in barley
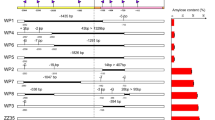
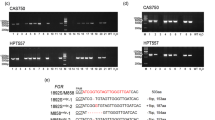
In barley grain, amylose biosynthesis is strictly associated with GBSSI encoded by the Waxy gene. We employed CRISPR/Cas9 technology to knock-out the Waxy gene of barley cultivar ‘Golden Promise’. Two target sites were designed for exon 1 of the Waxy gene (Fig. 1a) and assembled into a single vector (Fig. 1b). A total of 15 plantlets were regenerated from 355 transformed calli, and nine independent transgenic plants were identified using a PAT/bar test strip resistance kit and PCR amplification of the Ubi promoter (Fig. S1a and b).
CRISPR/Cas9-based strategy to generate waxy barley. Schematic diagram of target sites in the Waxy gene (a) and the genome editing vector (b). (c) Mutation types in transgenic plants revealed by DNA sequencing. (d) SDS-PAGE analysis of GBSSI protein. T1 and T2 are located in coding exon 1; PCas9: Cas9 gene modified with plant-optimized codons; TaU3: snRNA gene promoter from wheat; gRNA: guide RNA; T: target site; WT: wild type; black arrow: GBSSI protein; white arrow: deletion of GBSSI protein
Targeted mutagenesis in transgenic plants was examined by amplification of the exon 1 region of the Waxy gene and DNA sequencing. We identified two edited plants, wx13 and wx14, with biallelic mutation at the T1 site among nine transgenic plants. No plants edited at the T2 site were identified among the transformation events. The wx13 harbored two types of 1-bp (-G and -T) deletion, while the wx14 had a 1-bp (-G) deletion and a 1-bp (+ A) insertion at the T1 site (Fig. 1c). The mutagenesis caused frameshifts, and premature termination codons appeared at 124 bp and 412 bp, respectively. This meant that the wx13-1, wx13-2 and wx14-1 alleles only encoded 41 amino acids while the wx14-2 allele encoded 137 amino acids, and all lost the functional domains of the starch synthase catalytic domain and glycosyl transferases group 1 (Fig. S2). Two allelic genes were simultaneously knocked out in wx13 and wx14 lines. For further evaluation of translation of the Waxy gene, we extracted total GBSSI proteins from grains of wild type, wx13 and wx14 plants. SDS-PAGE was employed to determinate GBSSI protein content. The 60-KDa GBSSI protein was detected in the wild type, but it was absent in wx13 and wx14, consistent with amino acid sequence analysis of the new Waxy alleles (Fig. 1d). Homozygous waxy mutants were screened from T1 plants derived from wx14 edited plant for further analysis.
Starch character and the expression levels of the Waxy gene during the grain development
We employed the I2/KI stain method to differentiate waxy and non-waxy endosperms in barley. The endosperm and starch granules of wild type grains displayed the characteristic blue coloration after the I2/KI stain, while endosperms and starch granules from the waxy mutants stained brownish red, indicating that the wile type was non-waxy barley while waxy mutants contained very low amounts of amylose or were amylose-free (Fig. 2a). Starch granules were also observed at 8, 16, 24 and 32 day post anthesis (dpa). Interestingly, the starch granules of the wild type consisted of A-type and lots of B-type granules, whereas the starch granules of the waxy mutants were mainly A-type and rarely B-type granules. Only A-type granules were detected at 8 and 16 dpa stages, while abundant accumulation of B-type granules appeared at 24 and 32 dpa stages in wild type endosperms; in contrast, the endosperms of waxy mutants contained very few B-type granules throughout the developmental process (Fig. 2a). We performed scanning electron microscopy to identify structural differences between the wild type and waxy mutants. A-type granules of the wild type exhibited a plump morphology, whereas those of the waxy mutants appeared shriveled. The long axis of A-type granules did not show a significant difference between the wild type and waxy mutants; however, the short axis exhibited a 14.57% decrease compared to that of the wild type (Fig. 2b and Table S2). In contrast to the abundant presence of numerous small spherical B-type granules observed in the wild type endosperm, the waxy mutant exhibited a dramatic decrease in the abundance of B-type granules. This result was consistent with the microscopic observation of starch granules (Fig. 2b).
Identification of starch granules and the expression levels of the Waxy gene during grain development in both wild-type and mutant grains. (a) Starch granules at different developmental stages stained with I2/KI. (b) Scanning electron microscopy images of endosperm ultrastructure and starch granules. (c) The relative expression levels of Waxy gene in developing grains. Black arrow: A-type granules; white arrow: B-type granules; dpa: day post anthesis; bars: standard errors; values followed by the same letters in each sample are not significantly different at p < 0.01 level
Then, the transcript abundance of the Waxy gene was quantified at different developmental stages (8, 16, 24 and 32 dpa) using qRT-PCR. The expression level of the Waxy gene peaked at 16 dpa in both the wild type and waxy mutants. Its expression level was always higher in the wild type than in waxy mutants, except at 32 dpa. Waxy gene expression level fluctuated more in the wild type than in waxy mutants (Fig. 2c).
Grain compounds and agronomic performance of waxy mutants
Although the I2/KI stain could differentiate wild type and waxy mutants based on relative contents of amylose and amylopectin, we used chemical methods to measure the absolute contents of amylose, amylopectin, soluble sugar, sucrose, β-glucan and protein more specifically. The content of protein (10.88% in wild type and 11.94% in waxy) was relatively stable, however the content of amylose, amylopectin, soluble sugar, sucrose and β-glucan was significantly changed in both the wild type and waxy mutants. The wild type exhibited an amylose content of 33.50%, whereas the waxy mutants showed a complete absence of this component. Furthermore, the amylopectin content in the waxy mutants decreased by 18.64% (39.00% in wild type and 31.73% in waxy). So waxy mutants had a relatively low contents of amylopectin and amylose. The soluble sugar content of waxy mutants was 37.3% while that of the wild type was 10.2%. The amount of soluble sugar in waxy mutants was increased by almost as much as the amount of amylose was reduced. In addition, sucrose and β-glucan in waxy mutants were 39.16% and 35.40% higher than that in the wild type, respectively (Fig. 3a and Table S3).
Grain compounds and agronomic traits between the wild type and waxy mutants. (a) Content of amylose, amylopectin, soluble sugar, sucrose, β-glucan and protein. (b) Plant height, number of tillers, panicle length, grains per spike and thousand-grain weight. Grain compounds were quantified with three replicates, and agronomic traits were assessed with six replicates. *P < 0.05; **P < 0.01
Additionally, the plant height, number of tillers, spike length and grains per spike of the waxy mutants were similar to those of the wild type. However, the thousand-grain weight of the waxy mutants was 10.42% lower than that of the wild type (Fig. 3b and Table S4).
Transcriptome differences between the wild type and waxy mutants
High-throughput RNA sequencing (RNA-Seq) has emerged as a powerful and cost-efficient tool for transcriptome analysis and transcript profiling in various plant species. RNA-Seq has a large advantage in providing relatively comprehensive information on the nucleotide sequences of all genes expressed in the transcriptome (Zhang et al. 2014). Grains of wild type and waxy mutants were collected at 16 dpa for transcriptome analysis, based on the expression level of the Waxy gene (Fig. 2c). 1,204 differentially expressed unigenes (DEGs) were identified, with 386 down-regulated and 818 up-regulated in waxy mutants compared with the wild type based on FPKM value (Fig. S3a). Of these, 526 DEGs could be categorized into cellular process, environmental information processing, genetic information processing, metabolism, and organismal systems functions, with almost all of the DEGs associated with metabolism. Within the metabolism category, 95 and 80 DEGs were enriched in biosynthesis of other secondary metabolites and carbohydrate metabolism, respectively (Fig. S3b); 22 structural genes in starch and sucrose metabolism showed different expression levels. Similar to the result of qRT-PCR, expression levels of the Waxy gene were down in waxy mutants, and this is probably due to the absence of GBSSI protein. A block in amylose biosynthesis promoted the biosynthesis of amylopectin and other polysaccharides sharing the same substrate, D-glucose. Expression levels of genes encoding two ADP glucose pyrophosphorylase, two starch synthases and two 1,4-alpha-glucan branching enzymes, related to amylopectin biosynthesis, were higher in waxy mutants than in the wild type, and their log2FoldChange values were 2.05–3.08. Genes encoding four glucan endo-1,3-beta-glucosidases had different expression levels between the two genotypes, with three up-regulated in waxy mutants (Fig. 4; Table 1). The genes encoding beta-amylase and alpha-amylase showed lower expression because of the low starch content in waxy mutants.
Pasting properties of flour and cooking quality of dried noodles in waxy mutants
The pasting properties of barley flour, mainly determined by the content and ratio of amylose and amylopectin, have a considerable influence on the processing quality. The pasting properties of the wild type and waxy mutants were presented in Table 2. Compared to wild type, the pasting temperature of the waxy mutants decreased from 84.50 °C (wild type) to 69.38 °C. And the peak viscosity, trough viscosity, final viscosity and setback of the waxy mutants were 16.40%, 85.25%, 85.38% and 86.76% lower than those of the wild type, respectively, while the breakdown of the waxy mutants was 53.93% higher than that of the wild type. However, pasting time were not significantly different among the wild type and waxy mutants (Table 2).
For evaluating the effects of the waxy flour on processing product, the sensory evaluation, cooking properties and texture profile of dried noodles were determined. The dried noodles made by the wild type and waxy flour showed the similar sensory feature, e.g. color, appearance, palatability, elasticity, sticky, slipperiness and taste (Table S5). However, the water absorption rate and cooking time of the waxy dried noodles were 40.98% and 38.64% lower than those of the wild type dried noodles, but the waxy dried noodles had higher percentage of weight loss after cooking. Furthermore, the texture properties analysis showed the hardness, springness, cohesiveness, chewiness, gumminess, tensile strength and tensile size of the waxy dried noodles decrease by 49.28%, 28.75%, 16.67%, 70.58%, 58.69%, 17.39% and 45.99%, respectively, compared with the wild type dried noodles (Table 2).
Create waxy Qingke with new allelic variation
Finally, the waxy mutants pollen donor was crossed with the Qingke elite variety (the six-row hull-less barley) ‘Dulihuang’. We screened the waxy Qingke versions from the F3 lines according to the performance of the spike, I2/KI staining of grains and genotype detection of Waxy. And the marker-free lines were selected using the PAT/bar test strip test and Ubi PCR screening (Figure S4). This waxy Qingke versions would be a new germplasm resource for quality improvement.
Discussion
In this study, we created a new zero amylose barley through genome editing of the Waxy gene and evaluated its chemical and transcriptome characteristics.
The CRISPR/Cas9 system produced the insertion or deletion of single nucleotide in the coding sequence of the Waxy gene. The SNP modification of the Waxy gene in waxy mutants should have little impact on its expression. In our case, the expression level of Waxy gene was changed in waxy mutants compared with the wild type. The phenotype was also reported in Ma’s research (Ma et al. 2013). Waxy accessions (GSHO908, GSHO1828 and NA40 had lower Waxy expression levels than those of non-waxy accessions (PI48323 and CIho15773) (Ma et al. 2013). It implied that expression of the Waxy gene was possible to be regulated positively by its product.
The SNP modification caused frame-shifts, and premature termination. The incomplete GBSSI protein lost the functional domains of the starch synthase catalytic domain and glycosyl transferases group 1, and couldn’t produce amylose thoroughly. Consequently, the amylose can’t be detected in the grain of the mutants, which were similar to the previous researches about waxy mutants (Domon et al. 2002; Asare et al. 2012; Ma et al. 2013; Li et al. 2014; Hebelstrup et al. 2017). The GBSSI encoded by the Waxy gene is strictly associated with the amylose content in the grain of barley. The similar phenotypes also contained the lower B-type granules and higher β-glucan content. But the surface holes and irregular morphology in starch granules can’t be detected in our research (Ma et al. 2013; Li et al. 2019). Previous research has usually compared the characteristics of different cultivars or near-isogenic lines to draw conclusions, which cannot thoroughly remove the influences of background differences. In our case, the waxy mutants should have the same genetic background as the wild type except for the Waxy gene.
Accompanied by amylose, which occupied 33.50% of the grain and disappears in the mutants, there is a clear increase in soluble sugar content. The soluble sugar content of modified lines was 37.30%, while the wild type only possessed 10.20%. It was highly probable that the waxy mutants had lost their ability to convert soluble sugars into amylose, resulting in direct storage of these sugars in the seeds. The high content of soluble sugars also upregulated structural genes involved in biosynthesis of other polysaccharides, leading to an increase in β-glucan content. Even though the structural genes for amylopectin biosynthesis showed high expression levels in the mutant lines, the amylopectin content decreased by 18.64% in the wild type. This can be explained by the ability of GBSSI to catalyze the elongation of existing side chains in amylopectin and facilitate the formation of exceptionally long chains within the amylopectin structure (Hanashiro et al. 2008; Zeeman et al. 2010). The high expression levels of structural genes involved in amylopectin biosynthesis may not compensate for the functional deficiency of GBSSI.
Usually, waxy starches are utilized as stabilizers and thickeners in food products. For example, waxy starches had improved freeze-thaw stability compared to normal starches (Zheng & Sosulski 1998; Kim et al. 2003). Our new zero amylose starches showed reduced peak viscosity, trough viscosity, final viscosity and setback, as well as increased breakdown. The cooking time of the dried noodles was shorter than wild types, and the waxy flour required less energy in food processing. The reduction in springiness and chewiness indicated that this type of flour was unsuitable for dried noodles directly. It may be more appropriate for use as a food additive or in frozen foods and Japanese white salted noodles due to its unique advantages (Zheng & Sosulski 1998; Kim et al. 2003). After modifying the Waxy gene, the new waxy barley showed 35.40% higher β-glucan content than the wild type, which could provide higher nutritional value. β-glucan is a soluble dietary fiber and regarded as an important functional ingredient for lowering blood cholesterol, reducing glycemic response, regulating intestinal flora and maintaining weight (Izydorczyk & Dexter 2008; Thondre et al. 2011). In some parts of East Asia, immature barley grains are used to produce sweet, fermented grains. The waxy barley grains with higher soluble sugar content will allocate them to richer taste and nutrition. Our new waxy barley possessed very different chemical compounds from currently available cultivars, providing more choice in food product.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request. The transcriptomic data has been successfully uploaded to CNGBdb (https://db.cngb.org/), Submission ID: sub027210; BioProject ID: CNP0002515.
References
Asare EK, Båga M, Rossnagel BG, Chibbar RN (2012) Polymorphism in the barley granule bound starch synthase 1 (gbss1) gene associated with grain starch variant amylose concentration. J Agr Food Chem 60:10082–10092. https://doi.org/10.1021/jf302291t
Baik BK, Ullrich S (2008) Barley for food: characteristics, improvement, and renewed interest. J Cereal Sci 48:233–242. https://doi.org/10.1016/j.jcs.2008.02.002
Ban X, Dhoble AS, Li C, Gu Z, Hong Y, Cheng L, Holler TP, Kaustubh B, Li Z (2020) Bacterial 1,4-α-glucan branching enzymes: characteristics, preparation and commercial applications. Crit Rev Biotechnol 40:380–396. https://doi.org/10.1080/07388551.2020.1713720
Chen X, Shao S, Chen M, Hou C, Yu X, Xiong F (2020) Morphology and Physicochemical Properties of Starch from Waxy and Non-waxy Barley. Starch - Stärke 72:5–6. https://doi.org/10.1002/star.201900206
Damaris RN, Lin Z, Yang P, He D (2019) The Rice Alpha-Amylase, Conserved Regulator of Seed Maturation and Germination. Int J Mol Sci 2010.3390/ijms20020450. https://doi.org/10.3390/ijms20020450
De Schepper CF, Michiels P, Langenaeken NA, Courtin CM (2020) Accurate quantification of small and large starch granules in barley and malt. Carbohydr Polym 227:115329. https://doi.org/10.1016/j.carbpol.2019.115329
Domon E, Fuijita M, Ishikawa N (2002) The insertion deletion polymorphisms in the waxy gene of barley genetic resources from East Asia. Theor Appl Genet 104:132–138. https://doi.org/10.1007/s001220200016
Doxey AC, Yaish MW, Moffatt BA, Griffith M, McConkey BJ (2007) Functional divergence in the Arabidopsis beta-1,3-glucanase gene family inferred by phylogenetic reconstruction of expression states. Mol Biol Evol 24:1045–1055. https://doi.org/10.1093/molbev/msm024
Es I, Gavahian M, Marti-Quijal FJ, Lorenzo JM, Mousavi Khaneghah A, Tsatsanis C (2019) The application of the CRISPR-Cas9 genome editing machinery in food and agricultural science: current status, future perspectives, and associated challenges. Biotechnol Adv 37:410–421. https://doi.org/10.1016/j.biotechadv.2019.02.006
Evers AD, Blakeney AB, Brien LO (1999) Cereal structure and composition. Aust J Agric Res 50:629–650. https://doi.org/10.1071/AR98158
Figueroa CM, Asencion Diez MD, Ballicora MA, Iglesias AA (2022) Structure, function, and evolution of plant ADP-glucose pyrophosphorylase. Plant Mol Biol 108:307–323. https://doi.org/10.1007/s11103-021-01235-8
Guzman C, Alvarez JB (2016) Wheat waxy proteins: polymorphism, molecular characterization and effects on starch properties. Theor Appl Genet 129:1–16. https://doi.org/10.1007/s00122-015-2595-9
Hanashiro I, Itoh K, Kuratomi Y, Yamazaki M, Igarashi T, Matsugasako J, Takeda Y (2008) Granule-bound starch synthase I is responsible for biosynthesis of extra-long unit chains of amylopectin in rice. Plant Cell Physiol 49(6):925–933. https://doi.org/10.1093/pcp/pcn066. Epub 2008 Apr 22. PMID: 18430767
Hawkins E, Chen J, Watson-Lazowski A, Ahn-Jarvis J, Barclay JE, Fahy B, Hartley M, Warren FJ, Seung D (2021) STARCH SYNTHASE 4 is required for normal starch granule initiation in amyloplasts of wheat endosperm. New Phytol 230:2371–2386. https://doi.org/10.1111/nph.17342
He S, Hao X, Wang S, Zhou W, Ma Q, Lu X, Chen L, Zhang P (2022) Starch synthase II plays a crucial role in starch biosynthesis and the formation of multienzyme complexes in cassava storage roots. J Exp Bot 73:2540–2557. https://doi.org/10.1093/jxb/erac022
Hebelstrup KH, Nielsen MM, Carciofi M, Andrzejczak O, Shaik SS, Blennow A et al (2017) Waxy and non-waxy barley cultivars exhibit differences in the targeting and catalytic activity of GBSS1a. J Exp Bot 68:931–941. https://doi.org/10.1093/jxb/erw503
Huang S, Xin S, Xie G, Han J, Liu Z et al (2020) Mutagenesis reveals that the rice OsMPT3 gene is an important osmotic regulatory factor. Crop J 8:465–479. https://doi.org/10.1016/j.cj.2020.02.001
Izydorczyk MS, Dexter JE (2008) Barley β-glucans and arabinoxylans: molecular structure, physicochemical properties, and uses in food products–a review. Food Res Int 41:850–868. https://doi.org/10.1016/j.foodres.2008.04.001
Jedrzejczak-Krzepkowska M, Kalinowska H, Bielecki S (2011) Beta-fructofuranosidase–properties, structure and applications. Postepy Biochem 57:401–410. https://www.researchgate.net/publication/224924191
Jobling S (2004) Improving starch for food and industrial applications. Curr Opin Plant Biol 7:210–218. https://doi.org/10.1016/j.pbi.2003.12.001
Kannan P, Shafreen MM, Achudhan AB, Gupta A, Saleena LM (2023) A review on applications of β-glucosidase in food, brewery, pharmaceutical and cosmetic industries. Carbohydr Res 530:108855. https://doi.org/10.1016/j.carres.2023.108855
Kebede A, Kebede M (2021) In silico analysis of promoter region and regulatory elements of glucan endo-1,3-beta-glucosidase encoding genes in Solanum tuberosum: cultivar DM 1–3 516 R44. J Genetic Eng Biotechnol 19:145. https://doi.org/10.1186/s43141-021-00240-0
Kim W, Johnson JW, Graybosch RA, Gaines CS (2003) Physiochemical properties and end-use quality of wheat starch as a function of waxy protein alleles. J Cereal Sci 37:195–204. https://doi.org/10.1006/jcrs.2002.0494
Kundu S, Sharma R (2018) Origin, evolution, and divergence of plant class C GH9 endoglucanases. BMC Evol Biol 18:79. https://doi.org/10.1186/s12862-018-1185-2
Li Q, Pan Z, Deng G, Long H, Li Z, Deng X (2014) Effect of wide variation of the Waxy gene on starch properties in hull-less barley from Qinghai-Tibet plateau in China. J Agric Food Chem 62:11369–11385. https://doi.org/10.1021/jf5026746
Li Q, Pan Z, Liu J, Deng G, Long H, Zhang H et al (2019) A mutation in Waxy gene affects amylose content, starch granules and kernel characteristics of barley (Hordeum vulgare). Plant Breed 138:513–523. https://doi.org/10.1111/pbr.12695
Ma J, Jiang QT, Zhao QZ, Zhao S, Lan XJ, Dai SF (2013) Characterization and expression analysis of waxy alleles in barley accessions. Genetica 141:227–238. https://doi.org/10.1007/s10709-013-9721-x
Ma X, Zhu Q, Chen Y, Liu YG (2016) CRISPR/Cas9 platforms for genome editing in plants: developments and applications. Mol Plant 9:961–974. https://doi.org/10.1016/j.molp.2016.04.009
Monroe JD, Storm AR (2018) Review: the Arabidopsis β-amylase (BAM) gene family: diversity of form and function. Plant Science: Int J Experimental Plant Biology 276:163–170. https://doi.org/10.1016/j.plantsci.2018.08.016
Mueller M, Takemasa R, Schwarz A, Atomi H, Nidetzky B (2009) Short-chain alpha-1,4-glucan phosphorylase having a truncated N-terminal domain: functional expression and characterization of the enzyme from Sulfolobus solfataricus. Biochim Biophys Acta 1794:1709–1714. https://doi.org/10.1016/j.bbapap.2009.08.006
Pins JJ, Kaur H, Dodds E, Keenan JM (2007) The effects of cereal fibers and Barley Foods Rich in β-Glucan on Cardiovascular, Disease and Diabetes Risk. Whole Grains Health 75–86. https://doi.org/10.1002/9780470277607.ch7
Punia S (2020) Barley starch: structure, properties and in vitro digestibility - A review. Int J Biol Macromol 155:868–875. https://doi.org/10.1016/j.ijbiomac.2019.11.219
Qin R, Liao S, Li J, Li H, Liu X, Yang J (2020) Increasing fidelity and efficiency by modifying cytidine base-editing systems in rice. Crop J 89:396–402. https://doi.org/10.1016/j.cj.2019.04.007
Romualdi C, Bortoluzzi S, D’Alessi F, Danieli GA (2003) IDEG6: a web tool for detection of differentially expressed genes in multiple tag sampling experiments. Physiol Genomics 12:159–162. https://doi.org/10.1152/physiolgenomics.00096.2002
Sawada T, Nakamura Y, Ohdan T, Saitoh A, Francisco PB Jr, Suzuki E, Fujita N, Shimonaga T, Fujiwara S, Tsuzuki M (2014) Diversity of reaction characteristics of glucan branching enzymes and the fine structure of α-glucan from various sources. Arch Biochem Biophys 562:9–21. https://doi.org/10.1016/j.abb.2014.07.032
Solis-Badillo E, Agama-Acevedo E, Tiessen A, Lopez Valenzuela JA, Bello-Perez LA (2020) ADP-Glucose pyrophosphorylase is located in the Plastid and Cytosol in the Pulp of Tropical Banana Fruit (Musa acuminata). Plant foods for human nutrition. (Dordrecht Netherlands) 75:76–82. https://doi.org/10.1007/s11130-019-00788-w
Sparla F, Falini G, Botticella E, Pirone C, Talamè V, Bovina R (2014) New starch phenotypes produced by TILLING in barley. PLoS ONE 8:e107779. https://doi.org/10.1371/journal.pone.0107779
Takeuchi K, Ikuma T, Takahashi Y, Sagisaka K, Takasawa TS (2001) High sensitive phenol-sulfuric acid colorimetric method. Res Bull Obihiro Univ Nat Sci 22:103–107. https://www.researchgate.net/publication/277798941
Thondre PS, Ryan L, Henry CJK (2011) Barley β-glucan extracts as rich sources of polyphenols and antioxidants. Food Chem 126:72–77. https://doi.org/10.1016/j.foodchem.2010.10.074
Wang W, Chen Q, Xu S, Liu WC, Zhu X, Song CP (2020) Trehalose-6-phosphate phosphatase E modulates ABA-controlled root growth and stomatal movement in Arabidopsis. J Integr Plant Biol 62:1518–1534. https://doi.org/10.1111/jipb.12925
Xie C, XuY, Wan J (2020) Crop genome editing: a way to breeding by design. Crop J 8:379–383. https://doi.org/10.1016/j.cj.2020.05.001
Xing HL, Dong L, Wang ZP, Zhang HY, Yan HC, Bing L et al (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14:327. https://doi.org/10.1186/s12870-014-0327-y
Yamamori M, Endo T (1996) Variation of starch granule proteins and chromosome mapping of their coding genes in common wheat. Theor Appl Genet 93:275–281. https://doi.org/10.1007/BF00225757
Zeeman SC, Kossmann J, Smith AM (2010) Starch: its metabolism, evolution, and biotechnological modification in plants. Annu Rev Plant Biol 61:209–234. https://doi.org/10.1146/annurev-arplant-042809-112301
Zhang N, Liu B, Ma C, Zhang G, Chang J, Si H et al (2014) Transcriptome characterization and sequencing-based identification of drought-responsive genes in potato. Mol Biol Rep 41:505–517. https://doi.org/10.1007/s11033-013-2886-7
Zheng GH, Sosulski FH (1998) Determination of water separation from cooked starch and flour pastes after refrigeration and freeze–thaw. J Food Sci 63:134–139. https://doi.org/10.1111/j.1365-2621.1998.tb15693.x
Zhu T, Jackson DS, Wehling RL, Geera B (2008) Comparison of amylose determination methods and the development of a dual wavelength iodine binding technique. Cereal Chem 85(1):51–58. https://doi.org/10.1094/CCHEM-85-1-0051
Acknowledgements
Not Applicable.
Funding
This research was financially supported by the QingHai Science and Technology Department (2021-ZJ-958Q).
Author information
Authors and Affiliations
Contributions
Y.L. and B.L. conceived and designed the experiments. Y.J., F.T., and D.C. performed the experiments. X.Y, B.D. and Y.S. analyzed the data. G.L. provided the CRISPR/Cas9 system. Y.L. wrote the draft manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
: The determination of transgenic plants (Figure S1); the domains of GBSSI I protein and amino acids sequence alignment of Waxy allelic genes (Figure S2); DEGs in the grains and the KEGG pathway analysis of DEGs between the wild type and waxy mutants (Figure S3); selection of the marker-free waxy Qingke (Figure S4). primer sequences used in this research (Table S1); the disparity in starch granule size between the wild type and waxy mutants (Table S2); grain compounds between wild type and waxy mutants (Table S3); agronomic traits between wild type and waxy mutants (Table S4); sensory evaluation of dry noodles between the wild type and waxy mutants (Table S5)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Jiang, Y., Cao, D. et al. Creating a zero amylose barley with high soluble sugar content by genome editing. Plant Mol Biol 114, 50 (2024). https://doi.org/10.1007/s11103-024-01445-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01445-w