Abstract
Background
Durum wheat has a genetic capacity to accumulate toxic metals that can exceed the safety limit of the international standards, which may seriously affect human health. Identifying germplasms with low, nontoxic accumulated metal contents is important to select and develop new varieties. Thus, the objective of this study is to identify the levels of accumulated platinum in durum wheat and detect novel QTL.
Methods and Results
Platinum contents were determined using 130 durum genotypes. Results generally showed low values of accumulated Pt and significantly less than the maximum grain’s Pt content determined by international standards. Pt contents among genotypes varied from ≤ 0.001 to 0.72 µg/kg with an average of 0.02. Landraces showed the lowest average accumulated Pt. GWAS was then performed with 780 SSR markers. Five QTL were detected and explained 14.4–23.1% of the total phenotypic variation. Chromosomes 3 A, 3B, and 5B appear to be hotspots and may play a crucial role in accumulated Pt and were harbored in 1, 3, and 1 QTL, respectively.
Conclusions
This assessment of accumulated Pt within a unique panel included accessions mostly from Turkish regions, and GWAS used is the first study regarding accumulated Pt indices to reveal novel QTL. It will allow breeders to accelerate their selection of proper genotypes according to desired alleles and offer an opportunity to apply MAS to minimize Pt toxicity in durum wheat. Results indicated that the significance of genome (B) regions are likely related to the inheritance control of Pt content and may play a pivotal role regarding durum wheat’s Pt contents. Nonetheless, these novel QTL should be validated in independent populations in numerous environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Durum wheat (Triticum turgidum) is one of the most important food crops in South West Asia and North Africa (SWANA). It plays a pivotal role in the food of domestic people in SWANA, where about 75% of the world’s durum wheat is produced [1, 2]. Providing high-quality durum wheat varieties that meet international standards has become a focus, an increasing goal for breeders and consumers. Facing this shortage in availability has become an urgent necessity in wheat production areas for local and international markets. However, heavy metals are the highest risk of environmental pollutants that contribute significantly to soil contamination which causes damage to soil quality, plant health, and agricultural productivity. Moreover, these toxic elements may accumulate quickly and easily in the seeds and other parts of plants. They can be considered a universal problem is influencing food safety and human health [3]. Recently, durum wheat production in Turkey reached 4 million tons per year, with 1.257 million ha of the growing area [4]. Durum wheat is preferred mainly for pasta or macaroni products, couscous, burghul, and freekeh [5,6,7]. More emphasis has been shown in recent years on obtaining new cultivars with superior output, high grain quality, and biotic-abiotic stress resistance. Varieties generally obtain special prices in local or international marketing based on the high yield of varieties coupled with the high quality of the end product. Turkey’s total number of registered durum wheat varieties is continuously relatively increasing. Nevertheless, durum wheat has a genetic potential to accumulate heavy metals and toxic elements from soils in grains which can exceed the safety limit of the international standards [3].
The heavy metal platinum (Pt) is usually used as an electrode in car catalytic converters, so Pt pollution is the primary source of soil and roadsides dust. When Pt is forming complex ions, these complexes are fully bioavailable and highly toxic, which can accumulate in plants. Thus, we are concerned about secondary toxicity to humans from Pt plant accumulation. Considering the vast number of automobiles on the road each day and various other human activities, so Pt emissions will be significantly increased many times and contribute to an alarming increase in the accumulation of Pt [8]. Based to World Health Organization (WHO) Regional Office for Europe mean platinum concentration in grain products is 3.2 ng/g [9], and the Pt intake from the diet was at 1.44 µg/day for adults, has been reported by the Australian Federal Department of Health [10]. Therefore, it is vital to measure the Pt contents in the durum genotypes with analytical tools. However, it is still necessary to mention many disadvantages of analytical phenotypic analysis, as the process is hazardous, time- and labor-consuming, and requires expensive equipment and expertise [3].
Simple sequence repeats (SSRs) have numerous advantages, like co-dominance character, high polymorphism level, chromosome-specific, and high repeatability; it is also perfect for identifying and tracking target traits within varieties. Thus, SSRs could help durum wheat breeders as a useful molecular marker [11]. Additionally, SSR markers were plenty used for the construction of genetic linkage maps and quantitative trait loci (QTL) analysis [7, 12,13,14,15], and also for genome-wide association studies (GWAS) as a powerful tool to scrutinize the genetic architecture of complex traits. GWA studies have been widely applied in different crops: wheat, rice, maize, and Arabidopsis thaliana; it is prevalent for association studies of many traits in crops [16]. Many GWAS have used SSRs for different crops [17,18,19,20].
Recent development in molecular markers and the exploitation of QTL mapping and GWAS have also been submitted to uncover new functional allelic variants through a genome-wide scan. Elucidating phenotypic variability based on changes in DNA is an objective of numerous breeders in breeding programs generally and especially in durum wheat breeding programs. Moreover, detecting loci’s positions involving a specific trait delivers plant breeders an opportunity to apply marker-assisted selection. Durum wheat has a complex genome, so geneticists are required to simplify the complexity, Therefore, at the current time, GWAS may be a preferred option to identify favorable alleles, which can support breeders in better controlling crosses and selecting desired attributes in genotypes of durum wheat breeding programs. Association studies also provide valuable insights into the genetic architecture of quantitative traits across many unrelated genotypes [21]. Detection and identifying the association of specific genetic functional variants with phenotypic variations remains an urgent need. Therefore, association studies are an optimal option, provide a powerful tool, and have been broadly employed in plant research since it was first declared to be used in maize. Thus, association mapping has been widely utilized in many crops [22,23,24,25,26,27,28,29,30].
Turkey is a remarkable segment of the Fertile Crescent, the incipient wheat domestication and variousness center. Despite the significance of the gene pool in this essential region, there was no exploration of the accumulated platinum levels of the Turkish durum wheat gene pool. Therefore, in the present study, a diversity panel of durum wheat genotypes was used for the following objectives: (i) measure and evaluate the phenotypic Pt contents variation using ICP-MS analytical analysis for a panel of Turkish, foreign cultivars, and landraces, (ii) screening the genetic polymorphisms using SSR markers (iii) performing marker-trait association of genotypic data with Pt phenotypic trait, and alleles identification of underpinning trait variation to mine the markers linked to Pt content trait and to detect putative candidate gene loci, which can be used as marker-assisted selection (MAS) for durum wheat breeding programs.
Plant materials and methods:
The plant material in the present study originated from a wide range of ecological conditions and consists of a panel of 130 durum wheat (Triticum turgidum L.) genotypes, provided by Professor Dr. Hakan Özkan, Field Crops Department, Çukurova University, Adana, Turkey. The single-seed descent method was utilized for each genotype advanced [31]. Cultivars and landraces of the entire study were sourced from four groups: 50 local cultivars (Turkish CVs), 21 foreign cultivars (Foreign CVs), 44 landraces received from the National Genebank in Aegean Agricultural Research Institute, İzmir/Turkey (hereafter ex-situ LDs), and 15 landraces which are most prevalent among the growers and domestically grown on a momentous level, especially in southeastern Turkey (hereafter in situ LDs). The same panel was used in 2016 for a PhD study; where quite different traits were studied at that time, where agronomic, spike and some quality characteristics like thousand kernel weight, vitreousness kernel count, and test weight were analyzed. The diversity structure analysis of the same panel was recently reported for Turkish durum wheat diversity studies by Alsaleh et al. [3, 24, and 32]. Full details of these genotypes are presented in Table 1.
The genotypes were sown in the 2019/2020 growing season at the research and implementation area of the Field Crops Department of Çukurova University, Adana, Turkey (370 21” N latitude, 350 10” E longitude, and 20 m above the sea level). This area has hot summers and high humidity with Mediterranean climate. Nitrogen (180 kg/ha) and phosphorus fertilizers (60 kg/ha) were applied to the experimental plots. The experiment was established in a randomized block design with three replications. The genotypes were sown in rows at 30-cm row spacing and a length of 5 m. The trial site was maintained free from weeds and diseases by spewing herbicides and fungicides, where measurement was carried out at a normal level throughout the experiment.
DNA isolation:
A single plant was selected randomly from each genotype and used for molecular markers screening. The young plant leaves were harvested in the middle of February 2020. The leaves were immersed in liquid nitrogen and brought to the Molecular Genetic Laboratory at Science and Technology Application and Research Center (BİLTEM), Yozgat Bozok University, Yozgat-Turkey, to be stored in a deep freezer at -80 °C until the total genomic DNA was isolated according to CTAB protocol [33] with modification [34]. The extracted DNA was evaluated quantitatively and qualitatively by low agarose concentration (0.8%) gel electrophoresis. Before use, the DNA was diluted to a necessary concentration of 10 ng/µl for SSR applications.
Simple sequence repeats analysis
A group from different sources of microsatellites primers were selected to cover part somewhat of durum wheat chromosomes. Moreover, it was screened first to confirm their polymorphism level on a few genotypes. Based on the initial screening results, a total of 82 SSR primers were genotyped for the entire panel of 130 genotypes. The supplementary table briefly describes the SSR primers used and their information. PCR was used to amplify the SSRs region, as mentioned by Schuelke [35], where the M13-tailed primer method utilized a forward primer with a nucleotide extension at its 5’-end, uniform to an M13 sequencing (5-TGTAAAACGAAGGCCAGT-3), a standard length reverse of a fluorescently labeled M13 primer. The SSR fragments were scored twice for accuracy using the Gene Mapper software v3.7 (Applied Biosystems) described in the device user instructions. PCR reaction accomplished with a final volume of 12µl, contained 1X buffer, 0.125 mM dNTPs, 0.4 pmol “M13” forward primer, 0.3 pmol reverse primers, 3.0 pmol universal M13 primer labeled with one of four (6-FAM, VIC, NED or PET) fluorescent dyes, 0.12U Taq DNA polymerase, and roughly 25 ng genomic DNA. PCR amplification was performed with a primary denaturation at 94°C for 5 min; followed by 30 cycles of 94°C for 1 min, 55 to 65°C (annealing temperature depending on primers) for 1 min, 72°C for 1 min; followed by eight cycles of 94°C for 30 s, 53°C for 45 s, and 72°C for 45 s; and final extension was 72°C for 10 min. A set of four PCR products (1 µl each) labeled with various dyes was mixed with 0.25µl GeneScan-500 LIZ size standards (Applied Biosystems) and 9.86µl Hi-Di Formamide (Applied Biosystems), then denatured for 5 min at 94°C, and chilled on ice before loading. The bands fragmented on an ABI 3130xl Genetic Analyzer device (Applied Biosystems). The SSR individual bands were evaluated to represent a locus and scored as binary data, with the existence of bands marked as ‘1’ and their absence as ‘0’ because alleles scoring in such a binary type simplifies the appraisal and statistical investigation of co-dominant SSR data [36].
Phenotyping
Three spikes, “one from each replication” for each genotype, were selected randomly and harvested manually from the experimental plots in Adana location at the beginning of June 2020. Manually the collected spikes were also threshed, and the harvested grains were kept in paper bags in a dry place. However, to reduce the cost of analytical analysis, seeds from three replications of each genotype were mixed, milled, and dried in an oven. The digestion step was used by dissolving 0.5 g of mixed flour into an acidic solution, based on the “HPR-FO-52” procedure for wheat flour by SK-10 high-pressure rotor microwave digestion system (ETHOS EASY Milestone, Italy). Once the digestion process ended, the samples were cooled to room temperature; each was diluted with 10% v/v nitric acid up to 20 ml. To estimate Pt content, Inductively Coupled Plasma Mass Spectrometry (ICP-MS) (Thermo Scientific ICAPQC, USA) was used, with the following settings: 1550 W for radiofrequency power, 0.96 L/min for nebulizer gas, 0.88 L/min for a plasma gas, 3.01 bar for nebulizer pressure, dwell time 0.01 ms, and spray chamber temperature 3.7oC. Between injections, the sampler probe was rinsed for a half minute with ultrapure water, followed by washing for 50 s with 2% HNO3 and rinsing with ultrapure water for 50 s. For results accuracy, each measurement was repeated three times for the whole samples and the standards. The digestion step and the Pt measurements (ICP-MS) activities were done at the laboratories of “BİLTEM” Yozgat Bozok University, Yozgat, Turkey [3].
Statistical analysis of phenotypic and molecular data
Based on sources of genotypes, the studied genotypes were classified into four groups for analysis of variance (ANOVA). The first group involved released Turkish cultivars; the second was foreign cultivars, while ex-situ and in-situ landraces were the third and fourth. The variance analysis for the investigated Pt toxic element and the phenotypic frequency distribution was undertaken using Excel software. The proportion of the phenotypic variation explained by Pt content by each marker was estimated by the relevant R2 in TASSEL 5 [37]. The significant association’s levels were detected firstly based on the Bonferroni threshold for multiple testing and adjusted corrective threshold. For instance, the 5% Bonferroni threshold for multiple comparisons was considered [38].
Results
Phenotypic variations
The genotypes generally showed low and nontoxic Pt levels by ICP-MC analysis; Pt content variation varied from ≤ 0.001 to 0.725 µg/kg, with a mean of 0.020 µg/kg; one of the foreign CVs showed the highest value but was far from the risky limit (˃3.2 ng/g) (Table 2).
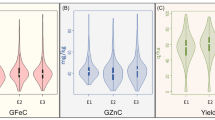
The frequency distribution of grain Pt concentrations for the whole panel is illustrated in Fig. 1a; as the genotypes were categorized into four groups, the average Pt contents for the Turkish and foreign CVs groups were the highest at 0.44 and 0.43 µg/kg, respectively, while it was lower for ex-situ and in-situ LDs at 0.03 and 0.02 µg/kg, respectively (Fig. 1).
(a): illustrates the frequency distribution of grain Pt concentrations for the whole panel; 1-(b): illustrates the frequency distribution of grain Pt concentrations among groups and for each group separately; 1-(c): illustrates the number of genotypes frequency distribution for each group separately
Figure 1-(a): illustrates the frequency distribution of grain Pt concentrations for the whole panel; 1-(b): illustrates the frequency distribution of grain Pt concentrations among groups and for each group separately; 1-(c): illustrates the number of genotypes frequency distribution for each group separately.
Genetic variations and marker-trait associations
In the present study, 82 SSR primers genotyped across the 130 genotypes showed 780 polymorphic markers. The frequency for allele “1” ranged from 0.023 to 0.992, while for allele “0” ranged from 0.008 to 0.969. Many markers have a low allele frequency of 0.05, which is considered non-useful. So the 780 markers were filtered based on minor allele frequency (MAF) value, where markers with MAF < 0.05 were ignored, and those with MAF ≥ 0.05 were kept; therefore, among 780 SSR markers, only 337 markers remained and were utilized for current GWAS. However, to detect a highly significant association, and to reduce false spurious or positive associations, the population structure (Q) and kinship (K) were figured first, then used as covariates in a mixed linear model (MLM + Q + K). By conducting GWAS in the present study, five marker-trait associations (MTAs) were detected and significantly associated with Pt contents using a Bonferroni correction at p < 0.05 [38] (Table 3; Fig. 2). The Manhattan plot illustrated in Fig. 2 shows significant SSRs markers associated with Pt content (with 5% Bonferroni correction threshold; p < 0.01; and MAF ≥ 5%).
Discussion
Recently, improving low toxicity varieties has become an important goal in crop breeding naturally. Breeding wheat genotypes with enhanced quality, especially for common toxic elements, along with a set of desirable agronomic and desired traits, also became one of the main priorities of durum wheat breeding programs, as durum wheat is an important economic crop. Generally, durum wheat is not only a significant crop for food security but furthermore has higher prices than bread wheat [39]. The extensive genetic diversity in Turkish durum wheat landraces could generally be a potential gene pool for durum wheat improvements. Identifying and using low Pt contents accessions from the current panel could overcome the severe effect of accumulated toxicity [32]. Nevertheless, despite the importance of this issue, no previous studies on Pt accumulation have been reported in Turkish durum wheat germplasm. Therefore, in the present study, Pt contents were assessed in a varied panel of genotypes, including historical cultivars from Turkey and different countries, landraces originating from a broad area of ecological conditions, whether ex-situ LDs from the Izmir gene bank, or locally adapted “in-situ LDs” gathered from diverse sources.
Although the phenotypic analysis showed variations for the Pt trait, most of the genotypes (51.5%) exhibited values ≤ 0.001 µg/kg. The highest Pt contents in the studied germplasms were within the safe range; it was less than the maximum level (3.2 ng/g) of grain Pt contents determined by international standards. Regarding the four groups formed in the studied panel, significant variations in the mean Pt content were found among the four different groups. The highest average grain accumulated Pt content (0.044 µg/kg) was found in Turkish cultivars, while the average Pt contents among studied groups were in the following order: in-situ LDs < ex-situ LDs < foreign cultivars < Turkish cultivars. While the in-situ group has an average Pt content of (0.02 µg/kg), and (0.03 µg/kg) for ex-situ landraces, both groups showed significantly low nontoxic Pt contents. Therefore, landraces could be valuable and vital candidate sources for developing durum wheat cultivars having low Pt contents (Table 2; Fig. 1).
Conventional plant breeding methods have significantly contributed to crop improvement. However, they have not been hurriedly progressing as required for current needs in picking out complex traits, whether morphological or agronomic characteristics. Therefore, to overcome the difficulties of conventional breeding and for disadvantages associated with chemical analytical phenotype profiling of Pt content trait. Unearthing the loci and alleles related to accumulating Pt is suggested to have more practical importance for the genetic improvement of the low toxic crops through molecular breeding using GWAS. The exploitation of recent development in molecular markers and their essential advantages for GWAS has been conducted to find novel functional allelic variants through a genome-wide scan. It is incredibly significant to find associations related to accumulated Pt to develop low toxic cultivars. So, a genome-wide association study is recommended, as GWAS has been used plenty for different crops recently. Therefore, to explore the genetic factors associated with the accumulated Pt, the phenotypic values of accumulated Pt contents with the genotypic data of SSR markers were used to conduct GWAS analysis.
The new approach to association studies uses natural germplasm and historical recombination “as genotypes panel used with the present study” with several advantages such as increased resolution, more overall allele coverage, time-saving, and gene tagging is economically [40]. Besides, no efforts have been reported previously for selecting durum wheat genotypes based on molecular markers for reduced Pt contents. Thus, GWAS was conducted in the present study, which sought to detect genetic factors that control accumulated Pt. Detecting alleles at particular loci associated with Pt content reduces the research time of phenotype detection of durum wheat breeders to develop new varieties with low Pt content. Generally, SNPs markers were widely used for most GWA studies because microsatellites resources were previously limited and considerably less widely exploited. However, this obstacle has been overcome as now thousands approximately are highly polymorphic microsatellites available. Many studies have acknowledged using microsatellites for GWA studies, where a single SSR catches a more genomic region than a single SNP. SSR also supplies additional benefits, such as higher information content, a minor inter-population variability, and significant intrinsic functional relevance. Therefore, SSRs primers were used in the present study for their numerous advantages [41].
Based on the proportion for genotype scored either “1” or “0”, rather than “- for missing data, the initial score showed 780 SSR markers. However, for GWAS, scored markers have been filtered based on two criteria for quality control: 1- selecting markers with zero missing scores, 2- disregarding all makers with MAF < 0.05%. Finally, 337 markers only remain and distributed over A and B of the durum wheat genome; therefore, GWAS in the current study utilized 337 markers. However, to enhance the strength and robustness of association analysis, many previous studies used the mixed linear model (MLM). Additionally, to reduce false or spurious associations, population structure (Q) and kinship (K) were calculated, and both were used as covariates in a mixed linear model (MLM) for the associations. So, (MLM + Q + K) model was used to reduce the false positives and detect the high significance of associations. Significance levels were considered and established using a Bonferroni modification at p < 0.05 based on the number of independent tests determined. Any result below the corrected < 0.05 p-value has been considered significant, so based on this regard GWAS analysis revealed that five MTAs “gwm335-bp270”, “wmc612-bp306”, “gwm156-bp308”, “wmc532-bp204”, and “wmc612-bp294” were significantly associated with accumulated low grain Pt content and explained phenotypic variation, with values ranging from 14.4–23.1%. Chromosome 5B harbors the significant MTA (gwm335-bp270) with the highest explanation value of total phenotypic variance (23.1%).
The detected MTAs were located apart, so they were considered different QTL. Chromosome 3B was found to harbor three QTL, as three MTAs were detected (wmc612-bp306, gwm156-bp308, and wmc612-bp294) and explained 22.5, 18.9, and 14.4% of total phenotypic variance, respectively. Generally, chromosomes 3 A, 3B, and 5B showed significant results and were found to harbor the significant QTL, which were spread 1, 3, and 1 QTL, respectively. The detected QTL were distributed in genomes (A) and (B). Nevertheless, genome (B) significantly has a higher harbor to QTL and acquired four QTL (80%) which were located in the (B) genome when compared to the genome (A), which had only one (20%) of detected QTL. Therefore, these chromosomal regions are considered hotspots for Pt accumulation in durum wheat. So, the present study indicates that genome (B) regions are likely related to the inheritance control of Pt content. The five Pt-associated markers detected in the present study were not previously reported for any of the durum wheat gene pools, so this marker’s associations will be considered novel QTL (Table 3; Fig. 2). These chromosomes and the respective QTL may be exploited by marker-assisted selection to improve durum wheat cultivars with low Pt contents. In the present GWAS, it is necessary to mention that several QTL were detected at α = 0.01 and explained nearly up to 19% of the total phenotypic variance but were considered a false positive and thus not considered or included in the current study.
Although the phenotypic data in this work were from a single environment only, the GWAS results identified several significant QTL of accumulated Pt that provide potential candidates for durum wheat improvement programs and may prove helpful in marker-assisted breeding. However, this work in which a unique panel of durum wheat was described and characterized concerning Pt accumulation was a fundamental and necessary study. There have also been no QTL reported previously related to Pt contents in durum wheat. Therefore, it is the first and vital GWA study for platinum toxicity in durum wheat. The SSR markers associated with Pt contents within this study could be a starting point, helpful gesture, and valuable resource for durum wheat breeders for marker-assisted selection, resulting in new durum wheat genotypes with low Pt contents. The detected markers in the current study could also be integrated into genomic selection strategy. This work may be considered a jump-starting process to help durum wheat breeding programs with introgression or selection processes. Nevertheless, detecting stable loci remains an urgent necessity. More functional genomic research is still pivotal and remains a critical need to validate the effect of the detected nominee QTL on Pt contents in several different environments.
References
Nachit MM (1998) Durum breeding research to improve dry-land productivity in the Mediterranean region. In: Rao SC, Ryan J et al (eds) SEWANA durum research network. ICARDA editions, pp 1–15
Baloch FS, Alsaleh A, Shahid MQ, Çiftçi V, Sáenz de Miera LE, Aasim M, Nadeem MA, Aktaş H, Özkan H, Hatipoğlu R (2017) A whole genome DArTseq and SNP analysis for genetic diversity assessment in durum wheat from Central Fertile Crescent. PLoS ONE 12(1):e0167821. https://doi.org/10.1371/journal.pone.0167821
Alsaleh A, Baloch FS, Sesiz U, Nadeem MA, Hatipoğlu R, Erbakan M, Özkan H (2022) Marker-assisted selection and validation of DNA markers associated with cadmium content in durum wheat germplasm. Crop & Pasture Science. https://doi.org/10.1071/CP21484
TUIK (2020) Turkish Statistical Institute. Available at http://www.turkstat.gov.tr/
Nachit M, Elouafi I, Pagnotta MA, El-Saleh A, Iacono E, Labhilili M, Asbati A, Azrak M, Hazzam H, Benscher D, Khairallah M, Ribaut JM, Tanzarella OA, Porceddu E, Sorrells ME (2001) Molecular linkage map for an intraspecific recombinant inbred population of durum wheat (Triticum turgidum L. var. durum). Theor Appl Genet 102:177–186
Alsaleh A, Baloch FS, Derya M, Azrak M, Kilian B, Nachit MM, Özkan H (2015) Genetic Linkage map of Anatolian Durum Wheat Derived from a Cross of Kunduru-1149 x Cham1. 33:209–220. Plant Molecular Biology Reporter10.1007/s11105-014-0749-6. 2
Farouk I, Alsaleh A, Motowaj J, Gaboun F, Belkadi B, Maltouf AF, Kehel Z, Elouafi I, Nsarellah N, Nachit MM (2021) Detection of grain yield QTLs in the durum population Lahn/Cham1 tested in contrasting environments. Turkish J Biology 45(1):65–78. doi:https://doi.org/10.3906/biy-2008-41
Oke S, Kikkert J, Vasiluk L, Hale B (2013) A study of platinum accumulation in radish (Raphanus sativus) and durum wheat (Triticum durum) plants. Studies by Undergraduate Researchers at Guelph (SURG). Journal 6(2):66–70
WHO (2000) Platinum, Chap. 6.11, Air Quality Guidelines - Second Edition, WHO Regional Office for Europe, Copenhagen, Denmark, 2000, p:1-133
Vaughan GT, Florence TM (1992) Platinum in the human diet, blood, hair and excreta. Sci Total Environ 111:47–58
Nadeem MA, Nawaz AA, Shahid QM, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F, Alsaleh A, Labhane N, Özkan H, Chung G, Baloch FS (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32(2):261–285. DOI: https://doi.org/10.1080/13102818.2017.1400401
Zhang W, Chao S, Manthey F, Chicaiza O, Brevis JC, Echenique V, Dubcovsky J (2008) QTL analysis of pasta quality using a composite microsatellite and SNP map of durum wheat. Theor Appl Genet 117:1361–1377
Maccaferri M, Sanguineti MC, Corneti S, Araus O, Ben JA, Salem M, Deambrogio JBE, Del Moral G, Demontis A, El-Ahmed A, Maalouf F, Machlab H, Martos V, Moragues M, Motawaj J, Nachit M, Nserallah N, Ouabbou H, Royo C, Slama A, Tuberosa R (2008) Quantitative trait loci for grain yield and adaptation of durum wheat Triticum durum Desf. across a wide range of water availability. Genetics 178:289511
Baloch FS, Alsaleh A, Andeden E, Derya M, Hatipoğlu R, Nachit M, Özkan H (2016) High levels of segregation distortion in the molecular linkage map of bread wheat representing WANA (West Asia and North Africa) region. Turkish Journal of Agriculture and Forestry. (2016) 40: 352–364 DOI: https://doi.org/10.3906/tar-1508-27
Alsaleh A, Baloch FS, Azrak M, Hamwieh A, Cömertpay G, Hatipoğlu R, Nachit M, Özkan H (2019) Identification of chromosomal regions in the genetic control of quality traits in durum wheat (Triticum turgidum L.) from the Fertile Crescent. Turkish J Agric Forestry 43(3):334–350. doi:https://doi.org/10.3906/tar-1807-83
Yan X, Zhao L, Ren Y, Dong Z, Cui D, Chen F (2019) Genome-wide association study revealed that the TaGW8 gene was associated with kernel size in Chinese bread wheat. Sci Rep 9, 2702 (2019). https://doi.org/10.1038/s41598-019-38570-2.
Stich B, Melchinger AE, Frisch M, Maurer HP, Heckenberger M, Reif JC (2005) Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor Appl Genet. 2005; 111: 723–730. https://doi.org/10.1007/s00122-005-2057-x PMID: 15997389
Nie X, Huang C, You C et al (2016) Genome-wide SSR-based association mapping for fiber quality in nation-wide upland cotton inbreed cultivars in China. BMC Genomics 17, 352 (2016). https://doi.org/10.1186/s12864-016-2662-x.
Ukoskit K, Posudsavang G, Pongsiripat N, Chatwachirawong P, Klomsa-Ard P, Poomipant P (2018) Detection and validation of EST-SSR markers associated with sugar-related traits in sugarcane using linkage and association mapping. Genomics. 2019; 111: 1–9. https://doi.org/10.1016/j.ygeno.2018. 03.019 PMID: 29608956
Barreto FZ, Rosa J, Balsalobre TWA, Pastina MM, Silva RR, Hoffmann HP et al (2019) A genome-wide association study identified loci for yield component traits in sugarcane (Saccharum spp.). PLoS ONE 14(7):e0219843. https://doi.org/10.1371/journal.pone.0219843
Cane MA, Maccaferri M, Nazemi G, Salvi S, Francia R, Colalongo C, Tuberosa R (2014) Association mapping for root architectural traits in durum wheat seedlings as related to agronomic performance. Mol Breeding 34(4):1629–1645
Maccaferri M, Sanguineti MC, Demontis A, El-Ahmed A, Moral LGD, Maalouf F, Nachit M, Nserallah N, Ouabbou H, Rhouma S et al (2011) Association mapping in durum wheat grown across a broad range of water regimes. J. Exp. Bot. 2011, 62, 409–438
Zhang P, Liu X, Tong H, Lu Y, Li J (2014) Association Mapping for Important Agronomic Traits in Core Collection of Rice (Oryza sativa L.) with SSR Markers. PLoS ONE 9(10):e111508. 10.1371/ journal.pone.0111508
Alsaleh A (2016) Genome wide association mapping (GWAS) for important agronomic and quality traits in durum wheat (Triticum durum L.). PhD dissertation, Department of Biotechnology, Institute of Natural and Applied Science, Çukurova University, Adana, Turkey. 2016. http://libratez.cu.edu.tr/tezler/10993.pdf
Pawar S, Pandit E, Mohanty IC, Saha D, Pradhan SK (2021) Population genetic structure and association mapping for iron toxicity tolerance in rice. PLoS ONE 16(3):e0246232. https://doi.org/10.1371/journal.pone.0246232
Zhu G, Hou S, Song X, Wang X, Wanf W, Chen Q, Guo W (2021) Genome-wide association analysis reveals quantitative trait loci and candidate genes involved in yield components under multiple field environments in cotton (Gossypium hirsutum). BMC Plant Biol 21:250. https://doi.org/10.1186/s12870-021-03009-2
Ali F, Nadeem MA, Habyarimana E, Altaf MT, Barut M, Kurt C, Chaudhary HJ, Khalil IH, Yildiz M, Cömertpay G, Shahid MQ, Baloch FS (2021) Identification of genetic basis associated with agronomic traits in a global safflower panel using genome-wide association study. Turk J Agric For 45:834–849. doi:https://doi.org/10.3906/tar-2105-55
Ahmed SM, Alsamman AM, Jighly A, Mubarak MH, Al-Shamaa K, Istanbuli T et al (2021) Genome-wide association analysis of chickpea germplasms differing for salinity tolerance based on DArTseq markers. PLoS ONE 16(12):e0260709. https://doi.org/10.1371/journal.pone.0260709
Baloch FS, Nadeem MA (2022) Unlocking the genomic regions associated with seed protein contents in Turkish common bean germplasm through genome-wide association study. Turkish J Agric Forestry 46(1):113–128. doi:https://doi.org/10.3906/tar-2104-63
Baloch FS, Nadeem MA, Sönmez F, Habyarimana E, Mustafa Z, Karaköy T, Comertpay G, Alsaleh A, Çiftçi V, Sun S, Chung G (2022) Magnesium- A Forgotten element: phenotypic and genome wide association study in Turkish common bean germplasm. Front. Genet. - Plant Genomics, Accepted: 21 Feb 2022. https://doi.org/10.3389/fgene.2022.848663
Goulden CH (1939) Methods of statistical analysis. John Wiley &Sons, Inc
Alsaleh A, Bektas H, Baloch FS, Nadeem MA, Özkan H (2022) Turkish durum wheat conserved in situ unveil a new hotspot of unexplored genetic diversity. Crop Science, Accept (25-Jan-2022). https://doi.org/10.1002/csc2.20723
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bull 19:11–15
Baloch FS, Alsaleh A, de Miera LES, Hatipoğlu R, Çiftçi V, Karaköy T, Yıldız M, Özkan H (2015) DNA based iPBS-retrotransposon markers for investigating the population structure of pea (Pisum sativum) germplasm from Turkey. Biochem Syst Ecol 61:244–252. https://doi.org/10.1016/j.bse.2015.06.017
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18(2):233–234. https://doi.org/10.1038/72708
Kaya HB, Cetin O, Kaya HS, Sahin M, Sefer F, Tanyolac B (2016) Association Mapping in Turkish Olive Cultivars Revealed Significant Markers Related to Some Important Agronomic Traits Biochem Genet. https://doi.org/10.1007/s10528-016-9738-9
Bradbury P, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diversesamples. Bioinformatics 23:2633–2635
Kaler AS, Purcell LC (2019) Estimation of a significance threshold for genome-wide association studies. BMC Genomics 20, 618 (2019). https://doi.org/10.1186/s12864-019-5992-7
Sall AT, Chiari T, Legesse W, Ahmed S, Ortiz R, van Ginkel M, Bassi FM (2019) Durum wheat (Triticum durum Desf.) origin, cultivation, and potential expansion in sub Saharan Africa. Agronomy 9(263). https://doi.org/10.20944/preprints04.0149.v1
Flint-Garcia SA, Thornsberry JM, Buckler ES (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Bailey-Wilson JE, Almasy L, de Andrade M et al (2005) Genetic Analysis Workshop 14: microsatellite and single-nucleotide polymorphism marker loci for genome-wide scans. BMC Genet 6:S1. https://doi.org/10.1186/1471-2156-6-S1-S1
Funding
Author thank Yozgat Bozok University, Scientific Research Projects Unit (Project No. 6602c-BİLTEM/19–323), Yozgat, Turkey, for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by the author.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Alsaleh, A. SSR-based genome-wide association study in turkish durum wheat germplasms revealed novel QTL of accumulated platinum. Mol Biol Rep 49, 11289–11300 (2022). https://doi.org/10.1007/s11033-022-07720-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07720-7