Abstract
Background
Tropomyosin is a major allergen in crustaceans, including mud crab species, but its molecular and allergenic properties in Scylla olivacea are not well known. Thus, this study aimed to produce the recombinant tropomyosin protein from S. olivacea and subsequently investigate its IgE reactivity.
Methods and Results
The tropomyosin gene was cloned and expressed in the Escherichia coli system, followed by SDS-PAGE and immunoblotting test to identify the allergenic potential of the recombinant protein. The 855-base pair of tropomyosin gene produced was found to be 99.18% homologous to Scylla serrata. Its 284 amino acids matched the tropomyosin of crustaceans, arachnids, insects, and Klebsiella pneumoniae, ranging from 79.03 to 95.77%. The tropomyosin contained 89.44% alpha-helix folding with a tertiary structure of two-chain alpha-helical coiled-coil structures comprising a homodimer heptad chain. IPTG-induced histidine tagged-recombinant tropomyosin was purified at the size of 42 kDa and confirmed as tropomyosin using anti-tropomyosin monoclonal antibodies. The IgE binding of recombinant tropomyosin protein was reactive in 90.9% (20/22) of the sera from crab-allergic patients.
Conclusions
This study has successfully produced an allergenic recombinant tropomyosin from S. olivacea. This recombinant tropomyosin may be used as a specific allergen for the diagnosis of allergy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mud crab Scylla species is an edible swimming crab found in brackish and tidal mangrove areas. Mud crab is considered an important economic resource for most Asia–Pacific nations, including Bangladesh, Sri Lanka, India, Thailand, Vietnam, Cambodia, and Malaysia, with over 45,300 tonnes of global production in 2016 [1]. In addition to high-quality meat, delicate taste, and fast growth, mud crab aquaculture is expanding rapidly to satisfy the needs for export and local market demand for adult male crabs, berried female crabs, soft-shell crabs, and crablets [2,3,4]. However, mud crab has also been discovered as a major source of shellfish allergies due to various protein allergens. Similar to crab allergy, mud crab ingestion can cause moderate to extreme allergic reactions, including clinical disorders of the digestive system, respiratory system, and the skin, although for certain allergic patients, contact with an allergen can also cause life-threatening reactions [5].
Tropomyosin belongs to a family of phylogenetically conserved proteins in many invertebrates. It is a significant pan-allergen that has been widely reported as a major allergen in crab species [6,7,8,9,10,11,12,13,14]. Allergenicity due to the presence of tropomyosin protein in crab species has been reported, including giant mud crab Scylla serrata [6, 7], purple mud crab Scylla tranquebarica [9], green mud crab Scylla paramamosain [10], blue crab Portunus pelagicus [8, 11], red crab Charybdis feriatus [11, 12], Chinese mitten crab Eriocheir sinensis [13], and snow crab Chionoecetes opilio [14]. However, allergenic tropomyosin in orange mud crab Scylla olivacea has yet to be investigated.
Due to the prevalence of allergy events in mud crabs, the development of a reliable diagnostic protocol is important for effective detection and clinical management. Up to now, raw extracts are a routine resource for traditional first-line diagnostics, including skin prick test, oral food challenge, and serum-specific IgE test [15]. Although optimized standard procedures are commonly used in the extraction of raw extracts prior to allergenicity testing, very often such practice resulted in various drawbacks, such as unintentional removal of minor allergens, differing allergen concentrations, accidental inclusion of contaminants and undefined non-allergenic content, and poor immunogenicity of some allergens [15, 16]. With the availability of pure allergens, it is possible to characterize the allergenic properties of individual allergens [17, 18]. One way of producing pure allergen is by utilizing bacteria expression systems. Abundant recombinant tropomyosin allergen has been successfully produced for blue swimming crab (Por p 1), crucifix crab (Cha f 1), and tropomyosin homologue from Chinese mitten crab. These recombinant tropomyosin allergens exhibit desired characteristics, such as known molecular weight, high purity, quality, stability, high concentration, and retained immunoreactive [12, 19, 20]. Therefore, this research aimed to produce recombinant tropomyosin protein from mud crab S. olivacea.
IgE antibodies are the main mediators for acute allergy hypersensitivity reactions. The attachment of IgE-allergen complex to mast cells and basophils leads to the crosslinking of receptor-bound IgE-specific allergens, resulting in the activation of inflammatory mediators, such as histamines and leukotrienes [21]. At present, the most commonly used approach for detecting tropomyosin IgE-binding reactivity is the use of sera from sensitized patients to identify allergens in the allergenic protein source [6,7,8,9,10,11,12,13]. Based on the importance of allergen IgE antibody reactions that cause allergic reactions, the purpose of this study was to determine the IgE reactivity of recombinant tropomyosin from S. olivacea, the most consumed mud crab species in Malaysia [22].
Materials and methods
Crab samples
Live male S. olivacea was obtained from a local supplier in Sungai Petani, Kedah, Malaysia [22]. Species identification of S. olivacea was carried out according to Keenan et al. [23] and Fazlan et al. [24]. The morphological features investigated during identification include frontal lobe spine height and shape, cheliped carpus and propodus spines, male first gonopod shape, and carapace color of crab. Prior to extraction, the crab was frozen in a freezer for 20 min [22, 24]. Once killed, the crab meat from the abdomen was excised and preserved by completely submerged into RNAlater RNA Stabilization Reagent (Qiagen, Germany).
Serum samples
A total of 22 sera from patients with crab allergy and one serum from a healthy and non-allergic subject that served as negative control were used in this study. The status of the crab allergic patients was confirmed by a positive skin prick test and/or an ImmunoCap test with a value of more than 0.35 kU/L to crab allergen. The patients' sera also showed reactivity to the 36 kDa tropomyosin band in immunoblotting of crude extract of S. olivacea, as shown in Fig. S1. Ethical approval was obtained from the Medical Research and Ethics Committee, Ministry of Health Malaysia, prior to conducting the study (approval number: NMRR-11-856-10216).
Total RNA extraction, cDNA synthesis, and PCR reamplification
Homogenization of preserved crab muscles was performed in the presence of liquid nitrogen and crushed using a mortar and pestle. Subsequent total RNA extraction was performed using the RNeasy Plus Mini Kit (Qiagen, Germany). The samples with A260/A280 (total RNA concentration) and A260/A230 (total RNA purity) ratios of 1.8–2.1 and 2.0–2.2, respectively, were used for the synthesis of cDNA using QuantiNova Reverse Transcription Kit (Qiagen, Germany). PCR reaction was conducted in a final volume of 25 μL, each comprising 12.5 μL of 2X First Base PCR Mix, 0.6 μM of forward primer (5′-ATGGACGCCATCAAGAAGAAGATG-3′), 0.6 μM of reverse primer (5′-TTAGTAGCCAGACAGTTCGCT-3′), cDNA template (50 ng), and nuclease-free water. The primer sequences were designed based on the tropomyosin sequence of S. serrata. PCR was performed in a MyCycler EP Gradient thermal cycler in the following steps: initial denaturation at 95 °C for 15 min, 35 cycles of amplification (94 °C for 1 min, 60 °C for 45 s, 72 °C for 1 min), and a final extension step of 72 °C for 7 min and maintained at 4 °C. The PCR amplified products were run on a 1.5% agarose gel electrophoresis, operating for 1 h at 75 V. Two microliters of the gel-purified PCR product (Vivantis, GF-1 AmbiClean Kit) was used as a template for the second-round PCR, prepared in the same PCR mixture and thermal cycle condition as the first-round PCR.
cDNA cloning and subcloning
Tropomyosin's cDNA was cloned into pJET1.2/blunt vector using CloneJET PCR Cloning Kit (Thermo Fisher Scientific, USA) and subsequently transformed into E. coli DH5α. Tropomyosin was amplified using 10 ng of plasmid as a template in a 25 µL of PCR reaction mix containing 12.5 µL of 2X First Base PCR Mastermix, 1.5 µL of forward subcloning primers containing BamHI sequence (5′-CGCGGATCCATGGACGCCATCAAGAAGAAGATG-3′), and 1.5 µL of reverse subcloning primers containing EcoRI sequence (5′-GCGAATTCTTAGTAGCCAGACAGTTCGCT-3′). The PCR was conducted as follows: one cycle (95 °C for 15 min) for initial denaturation; 30 cycles (94 °C for 1 min, 59 °C for 40 s, 72 °C for 1 min) for annealing and extension, and one cycle (72 °C for 7 min) for a final extension of the amplified tropomyosin. The amplified PCR products were viewed on a 1.5% agarose gel electrophoresis and purified using a QIAquick PCR purification kit (Qiagen, Germany). Three PCR colony products were sequenced by Apical Scientific, Malaysia using tropomyosin complete sequence primer of the BigDye® Terminator v3.1 Cycle Sequencing system. The identity of sequenced tropomyosin was annotated using the Basic Local Alignment Search Tool (Blastn) program (https://blast.ncbi.nlm.nih.gov/) and aligned under an alignment view of “query anchored with dots for identities”. Double restriction enzyme digestion of pRSETA insert and purified tropomyosin was carried out using FastDigest BamHI and FastDigest (Thermo Scientific, USA), which were incubated at 37 °C for 30 min before both enzymes were inactivated at 80 °C for 5 min. The digested products were purified using the GF-1 AmbiClean Kit and subsequently ligated using T4 DNA ligase (Thermo Scientific, USA) for the molar ratio over the vector of 1:1 and incubated at 22 °C for 1 h. The pRSETA-tropomyosin was transformed into One Shot® BL21(DE3) pLysS competent cells (Invitrogen, USA). The cell was heat-shocked by immediately immersed into a 42 °C water bath for 40 s and quickly placed on ice for 10 min. A volume of 950 µL of pre-warmed LB broth was added into the vial and placed in an incubator at 37 °C and gently shaken at 200 rpm for 1 h. A volume of 20 µL of bacterial suspension was spread on LB agar plates containing 35 µg/mL chloramphenicol and 100 µg/mL ampicillin and then incubated overnight at 37 °C. The sequence of tropomyosin was re-confirmed using DNA sequencing and the analyzed sequence was deposited in the NCBI database via BankIt submission (https://www.ncbi.nlm.nih.gov/WebSub/).
Phylogenetic tree, amino acid sequence, and structural composition analysis of tropomyosin
The cloned tropomyosin sequence was annotated using the Basic Local Alignment Search Tool (Blastn) program (https://blast.ncbi.nlm.nih.gov/) and aligned under an alignment view of “query anchored with dots for identities”. The tropomyosin sequence of S. serrata was acquired from the NCBI database at https://www.ncbi.nlm.nih.gov/protein/151505279. The ExPASy Amino Acid Translation (https://web.expasy.org/translate/) online tool was used to deduce the amino acid sequence of tropomyosin. Molecular Evolutionary Genetic Analysis (MEGA) X version 10.05 was used to construct the phylogenetic tree using Neighbor-Joining. Then, the prediction of the secondary protein structure was conducted using SOPMA server (NPS@:SOPMA secondary structure prediction (ibcp.fr)). Meanwhile, the prediction of the tertiary protein structure of tropomyosin was carried out using SWISS-MODEL server (http://swissmodel.expasy.org/) by applying Eriocheir sinensis tropomyosin (PDB code: 1c1gA) template as the reference gene for comparison. Remodeling of tropomyosin structure was generated using the PyMOL program (https://pymol.org/2/).
Induction, expression, and purification of recombinant tropomyosin protein
A fresh colony of BL21(DE3) PlysS-pRSETA-tropomyosin was streaked on the LB agar plate comprising 35 µg/mL chloramphenicol and 100 µg/mL ampicillin. After 18 h of incubation at 37 °C, positive bacteria clone was inoculated into fresh sterile (5 mL of LB-ampicillin-chloramphenicol broth) in a culture flask and incubated at 37 °C with shaking at 250 rpm overnight. Then, 1 mL of the overnight culture was added into 10 mL of fresh LB-ampicillin-chloramphenicol broth and incubated at 37 °C with shaking at 250 rpm until the optical density (OD) at A600 nm reached 0.6. The BL21(DE3) pLysS-pRSETA-tropomyosin protein was overexpressed with IPTG induction at a final concentration of 1 mM for 4 h at 37 °C with shaking at 250 rpm. The bacterial cells were harvested initially by centrifugation at 4000×g at room temperature for 20 min. Next, the soluble fraction was lysed using lysis buffer (25 mM Tris–HCl pH 8, 0.2 mg/mL lysozyme, 1 mM imidazole, 300 mM NaCl) at -20 °C for 1 h and sonicated (5-s pulse-on, 5-s pulse-off at the amplitude of 60°). Ten microliters of recombinant tropomyosin protein in the presence of 2X Laemmli sample buffer was denatured at 97 °C for 4 min before running on 12.0% SDS-PAGE, and the results were visualized using Image Lab 3.0 software of Gel Doc EZ Imager. The soluble fraction of expressed recombinant tropomyosin was pooled for further purification via ProBond™ affinity chromatography kit (Thermo Scientific, USA).
Immunoblotting with monoclonal anti-tropomyosin antibody
Immunoblotting with monoclonal anti-tropomyosin antibody was performed to demonstrate the presence of tropomyosin allergen in recombinant tropomyosin protein. The non-specific protein sites on the strips were blocked for 1 h with a blocking buffer (1X TBS containing 10% non-fat milk). The strip was reacted with a rat monoclonal anti-tropomyosin IgG antibody, Ab50567 (1:6,000) (Abcam, United Kingdom) in an incubator shaker at 4 °C for 16 h. After four times washed with 1X TTBS for every 5-min interval, the strip was subsequently reacted with an alkaline phosphatase goat anti-rat (IgG) secondary antibody (1:40,000) (Abcam, United Kingdom) for 30 min. After washing four times with 1X TTBS solution and one time with 1X TBS solution, the antibody binding on the strip was detected by a colorimetric alkaline phosphatase conjugate substrate kit. Before being air-dried, the strip was washed with distilled water three times every 5 min to stop color development. The immunoblot results were scanned using an imaging densitometer (BioRad, USA).
Immunoblotting with patient sera IgE
Immunoblotting with sera from crab allergic patients was performed to characterize the IgE reactivity of recombinant tropomyosin protein. Once the transferred recombinant protein was confirmed, the non-specific protein sites on the strips were blocked for 1 h with 1X TBS containing 10% non-fat milk, followed by 16 h incubation with the patient's sera (1:5) at 4 °C. The strips were washed four times with 1X TTBS for every 5-min interval and subsequently incubated for 30 min in affinity-purified biotin-labeled goat anti-human IgE (1:1,000) (KPL, USA) as a secondary antibody. The unbound secondary antibodies were washed out four times with 1X TTBS for every 5-min interval and non-specific binding of secondary antibodies was blocked using a blocking buffer for 5 min. After one time wash with 1X TTBS, the strips were incubated with streptavidin-conjugated alkaline phosphatase (1:5000) (BioRad, USA) for 30 min. After washing four times with 1X TTBS solution and one time with 1X TBS solution, the strips were incubated with alkaline phosphatase conjugate substrate that reacts with the conjugated secondary antibody for 15 min to reveal the antibody-bound recombinant tropomyosin. Immunoblotting results were scanned using an imaging densitometer (BioRad, USA). Each set of strips contained a negative control serum from a healthy and non-allergic individual and a blank strip (no serum).
Results
Annotation of recombinant tropomyosin gene
PCR reamplification of the cloned sequence had been identified as tropomyosin with a product size of 855 base pairs (Fig. S2). The cloned sequence of S. olivacea clones was 99.18% homologous to the tropomyosin sequence of S. serrata, indicating that the tropomyosin DNA sequence is strongly conserved between mud crab species. The S. olivacea tropomyosin sequence was deposited in the NCBI database with the appointed accession number of MN218638. Moreover, the presence of mismatched nucleotides between the cDNAs of S. serrata tropomyosin and the S. olivacea tropomyosin at seven loci of 35 (A to T), 339 (G to A), 342 (T to C), 542 (A to G), 560 (A to G), 801 (A to G), and 852 (T to C) indicated the existence of single-nucleotide polymorphism within the tropomyosin coding region (Fig. 1). The terminal sequences of the proteins in vivo might differ from the sequences provided due to the predetermined sequence of the primers used for amplification.
Annotation of tropomyosin protein, cluster analysis, and determinant of protein structure
The cloned sequence of S. olivacea was translated to a polypeptide of 284 amino acids with a predicted molecular weight of 32.776 kDa, producing a highly best match of 41 sequences ranging from 88.02 to 95.77% for crustaceans, 79.23 to 81.69% for arachnids, 81.69 to 83.36% for insects, and 88.03% for K. pneumoniae. The evolution relationship showed that the S. olivacea tropomyosin shared the same cluster with invertebrate phyla known as an arthropod, which consists of the species from subphylum Crustacea, Arachnida, and Insecta (Fig. 2). Interestingly, S. olivacea tropomyosin was discovered to share the same cluster with bacteria K. pneumoniae, which had the longest phylogenetic tree branch, indicating a new evolutionary relationship between both unrelated species. The predicted structural composition of tropomyosin was predominantly made up of 89.44% alpha-helix folding and a tertiary structure of a two-chain alpha-helical coiled-coil structure made up of a homodimer heptad chain (Fig. 3).
Phylogenetic tree of tropomyosin amino acid sequences between S. olivacea and other species. Description of tropomyosin of each species was presented as follows: Common name-Scientific Name GenBank accession No. (Subphylum) percentage of sequence similarity against tropomyosin sequence of S. olivacea
3-D structure of tropomyosin of S. olivacea generated using the PyMOL program (https://pymol.org/2/). The structure was predicted by using SWISS-MODEL server (http://swissmodel.expasy.org/) based on Eriocheir sinensis tropomyosin (PDB code: 1c1gA) template as the reference gene
Detection of recombinant tropomyosin protein
SDS-PAGE analysis of the protein fraction of recombinant tropomyosin from S. olivacea revealed that IPTG-induced culture displayed protein overexpression in a soluble fraction at 42 kDa (Figs. S3 and 4a). To confirm the presence of tropomyosin in recombinant samples, immunoblotting analysis was performed using a commercial monoclonal anti-tropomyosin antibody (MAC-141), producing a protein band at 42 kDa (Fig. 4b) and indicating that the sample contained tropomyosin epitopes.
Allergenicity of recombinant tropomyosin protein
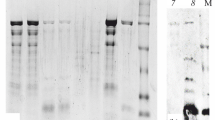
Immunoblotting results of recombinant tropomyosin protein exhibited positive reactivity with IgE in sera from 22 crab-allergic patients (Fig. 5). The results showed that almost all patients' sera IgE reacted with recombinant tropomyosin protein, producing a single protein band at 42 kDa. The absence of a protein band on the negative control sample showed that the positive reaction was not due to contamination. The presence of a positive allergen-specific IgE antibody reaction confirmed that recombinant tropomyosin protein from S. olivacea was an allergenic protein. The immunoblot results also showed that two (patient 6 and 10) out of 22 patients' sera did not react to recombinant tropomyosin protein. As other patients' sera reacted with the recombinant protein, the IgE reactivity of the protein did not cause the absence of antigen–antibody response, but this observation was possibly due to the low concentration of specific IgE antibodies to tropomyosin in the sera. This finding is in accordance with the results of immunoblotting of crude extract (Fig. S1) that demonstrated a faint band, indicating a weak IgE reactivity of sera from patient 6 and 10. Overall, the results revealed that the IgE in sera binding with recombinant tropomyosin protein from S. olivacea was present in 90.9% (20/22) of the sera analyzed, suggesting that tropomyosin is a major allergen.
Discussion
Tropomyosin is a well-known allergen in many crustacean species. Other studies have reported that the 855-base pair sequence of tropomyosin has been detected in crucifix crab Charybdis feriatus [12], Chinese mitten crab Eriocheir sinensis [20], and blue swimming crab Portunus trituberculatus [25] using tropomyosin-specific primers. The interspecies tropomyosin sequence homology between cockroach species and S. olivacea found in this study is consistent with findings from other researchers. For instance, according to Jeong et al. [26], the dusky brown cockroach tropomyosin was found to be 98.5% identical to the American and German cockroach tropomyosin. The findings suggested the highly conserved nature of tropomyosin gene, hence offering the advantages of being used as a molecular marker for ancestry study, especially among species within arthropod phylum [27], including mud crab species [28]. Single-nucleotide polymorphism was observed between the tropomyosin gene sequences of S. olivacea and S. serrata. Previously, polymorphism at the tropomyosin gene region was also identified in other invertebrates, such as American house dust mite, Dermatophagoides farinae [29], and giant freshwater shrimp Macrobrachium rosenbergii [30]. Nucleotide polymorphism that occurred between giant freshwater shrimp and other shrimp tropomyosin, such as white leg shrimp Litopenaeus vannamei, giant tiger prawn Penaeus monodon, and brown shrimp Penaeus aztecus, has the potential to affect the biological efficacy in the diagnosis and immunotherapy of allergen [31, 32].
Due to high amino acid sequence conservation and similar nature of tropomyosin sequences between S. olivacea and other species from crustaceans, arthropods, bivalves, and bacteria, we believed that tropomyosin could cause cross-reactivity allergic reaction. Cross-reactivity is one of the crucial characters of major shellfish allergens. Researchers observed that allergens with at least 70% of amino acid sequence identity are likely to be cross-reactive [32]. Cross-reactivity among crustaceans has been frequently reported, with most of the findings supported that the allergenic cross-reactivity occurred among the crustacean species, including between crab and shrimp, and crab and prawn [11, 19, 33, 34]. For instance, cross-reactivity of crab tropomyosin was reported among species of bivalves, including blue mussel Mytilus edulis, Sydney rock oyster Saccostrea glomerata, southern calamari Sepioteuthis australis, and saucer scallop Amusium balloti, as examined through IgE inhibition immunoblotting [35]. In addition, cross-reactivity occurrence was also reported between crustaceans and other arthropods, including insects and arachnids. It was found that cross-reactivity between mites and shrimp could react to tropomyosin from the house dust mite tropomyosin in a shrimp-sensitive patient [36]. Allergy sensitization is believed to have occurred first due to inhalation of mite allergens and then ultimately due to the ingestion of crustaceans [37].
Interestingly, in this study, tropomyosin gene sequences of S. olivacea showed very high homology with a human intestinal bacterium called K. pneumoniae. K. pneumoniae is a Gram-negative bacterium with very high homology with S. olivacea tropomyosin, which has not yet been experimentally discovered by other studies. Up to now, only one study reported cross-reactivity relatedness between the shellfish and tropomyosin bacteria in polluted water [38]. K. pneumoniae and Pseudomonas spp. were detected in the water from the gutter, which could be responsible for respiratory symptoms among trout processing workers [38]. This finding revealed the risk of bacterial infection due to the unprotected handling of different seafood products in various stages of manufacturing process [39]. This finding also demonstrated the likelihood of pneumonia as a cause of allergy upon exposure; thus, the existence of the highest possible cross-reactivity is due to the high amino acid sequence of tropomyosin. However, future studies should be undertaken to validate the allergenicity properties of K. pneumoniae.
In the present findings, S. olivacea tropomyosin three-dimensional structure predicted continuous coiled-coil dimers made up of two parallel helical molecules that wound around each other. The investigated coiled-coil interaction arrangement of two alpha-helices composed of a repeated heptad sequence of each seven amino acid residues, where all positions are solvent-exposed, except for the inner position of hydrophobic amino acid residues [40]. This revealed that protein structure is a cause of the allergenicity of tropomyosin, particularly due to the existence of repetitive patterns [41, 42]. In addition, the IgE antibodies-epitope sites on unexposed hydrophobic amino acid residues have the capability to mask enzymatic digestion and thus maintain the allergenicity of tropomyosin [40]. It should be noted that this study only used a program to predict the secondary structure of this recombinant protein. Thus, a further study to evaluate the secondary structure composition via circular dichroism is needed to confirm the correct folded structure of the recombinant tropomyosin.
Recombinant-produced tropomyosin protein exhibited higher molecular mass than predicted. Similar results have also been demonstrated for recombinant tropomyosin from Korean storage mite Tyrophagus putrescentiae [43], non-biting midge Chironomus kiiensis [26], and German cockroach Blattella germanica [44]. Previous studies also revealed that the molecular mass of the tropomyosin protein molecule in crab species ranged from 34 to 39 kDa [6,7,8,9,10,11,12,13,14, 19, 20]. The recombinant tropomyosin from S. olivacea contains 38 more amino acids at the N-terminus, plus a polyhistidine-tag molecule is expected to increase its molecular mass. Some also proposed that the larger-than-expected size of tropomyosin may be attributed to the peculiar elongated structure of this molecule. Tropomyosin is an elongated two-stranded protein with a structure of full-length dimeric-α-helical coiled-coil that has these unique migratory properties [45].
This study has successfully recognized the recombinant tropomyosin of S. olivacea using MAC-141, a commercial rat monoclonal anti-tropomyosin antibody capable of detecting tropomyosin in various invertebrates [46]. Other studies have also shown that MAC-141 recognized recombinant tropomyosin proteins from black tiger prawn P. monodon [46], banana prawn Fenneropenaeus merguiensis [47], and king prawn Melicertus latisulcatus [48]. It is predicted that the sequence part of QAMKLEKDNAM in MAC-141 is the monoclonal antibody binding epitopes capable of recognizing tropomyosin allergens from crustaceans, molluscs, and nematode worms [46, 49].
Tropomyosin protein isoforms of different allergenicity have been found in various tissues and development stages, like the oriental migratory locust Locusta migratoria [50] and brown shrimp Penaeus aztecus [51]. Therefore, the differences in IgE-binding frequency in the current sample may be due to different amino acids serving different IgE-epitope regions. Also, cross-reactivity with other invertebrates in polysensitized populations and genetic variants causing particular sensitizations may lead to different allergenicity [52]. Meanwhile, the results for S. olivacea have been novel as this is the first research to prove that tropomyosin is a major allergen in S. olivacea. This finding is consistent with previous studies using a crude extract that identified reactive tropomyosin to be higher than 70% of the crab-sensitive subject as a major allergen in S. serrata, S. paramamosain, and S. tranquebarica [6, 7, 9, 10]. Two critical factors attributed to tropomyosin as a major allergen are its natural primary function and structural stability. Tropomyosin is a highly conserved actin-binding protein that resides in both vertebrates and invertebrates' muscle and non-muscle cells, playing a central function in muscle contraction. Due to its primary role in muscle function, it is far more abundant than other known shellfish allergens. However, further IgE reactivity tests, such as inhibition assay for IgE binding, should be performed using the recombinant tropomyosin to better define the allergenic characteristics of the recombinant tropomyosin.
Conclusions
Recombinant allergen production has gained attention in past years due to the potential as a specific allergen source for diagnosis of specific allergy causal, and also as an initiative allergy treatment in the form of vaccines. In this study, the recombinant tropomyosin from mud crab S. olivacea has been successfully expressed in E. coli system at 42 kDa, and its IgE reactivity has been proved through the reaction of 90.9% of patients allergic to crab. For diagnosis purposes, the produced recombinant tropomyosin is ready to be tested on the patient with allergy to mud crab S. olivacea. Future vaccine work also involves the production of wild-type recombinant tropomyosin of S. olivacea with diminished IgE-epitope reactivity.
Data availability
Data deposition (mud crab tropomyosin identification number) is made in the GenBank repository found at https://www.ncbi.nlm.nih.gov/. Other data is available from the authors for non-commercial purposes on reasonable request.
References
Azra MN, Ikhwanuddin M (2016) A review of maturation diets for mud crab genus Scylla brood stock: present research, problems and future perspective. Saudi J Biol Sci 23(2):257–267. https://doi.org/10.1016/j.sjbs.2015.03.011
Devi PL, Joseph A (2015) Taxonomy of mud crabs of the genus Scylla (De Haan) from Cochin Backwaters, Kerala India. Indian J Fish 62(3):45–51
Waiho K, Mustaqim M, Fazhan H et al (2015) Mating behavior of the orange mud crab, Scylla olivacea: the effect of sex ratio and stocking density on mating success. Aquac Rep 2:50–57. https://doi.org/10.1016/j.aqrep.2015.08.004
Ikhwanuddin M, Muhd-Farouk H, Memon AJ et al (2014) Sperm viability assessment over elapsing time maintained at 2 degrees C of orange mud crab, Scylla olivacea (Herbst, 1796). Pak J Biol Sci 17(9):1069–1073. https://doi.org/10.3923/pjbs.2014.1069.1073
Lopata AL, Kamath S (2012) Allergy diagnosis: gaps and needs. Curr Allergy Clin Immunol 25(2):60–67
Misnan R, Abdul Rahman NI, Yadzir ZHM et al (2015) Characterization of major allergens of local mud crab (Scylla serrata). Sci Res J 12(1):1. https://doi.org/10.24191/srj.v12i1.5434
Nurul Izzah A, Rosmilah M, Zailatul Hani MY (2015) Identification of major and minor allergens of mud crab (Scylla Serrata). Med Health 10(2):90–97
Rosmilah M, Shahnaz M, Zailatul HMY et al (2012) Identification of tropomyosin and arginine kinase as major allergens of Portunus pelagicus (blue swimming crab). Trop Biomed 29(3):467–478
Jasim HA, Misnan R, Yadzir ZHM et al (2021) Identification of common and novel major crab allergens in Scylla tranquebarica and the allergen stability in untreated and vinegar-treated crab. Iranian J Allergy Asthma Immunol 20(1):76–87. https://doi.org/10.18502/ijaai.v20i1.5414
Nasrat M, Misnan R, Yadzir ZHM et al (2018) Comparison of protein profiles and allergenicity of different body parts and genders of Scylla paramamosain. Int J Sci Environ Technol 7(5):1759–1765
Misnan R, Murad S, Yadzir ZHM et al (2012) Identification of the major allergens of Charybdis feriatus (red crab) and its cross-reactivity with Portunus pelagicus (blue crab). Asian Pac J Allergy Immunol 30(4):285–293
Leung PSC, Chen YC, Gershwin ME (1998) Identification and molecular characterization of Charybdis feriatus tropomyosin, the major crab allergen. J Allergy Clin Immunol 102(5):847–852. https://doi.org/10.1016/s0091-6749(98)70027-2
Liu GM, Cao MJ, Huang YY et al (2010) Comparative study of in vitro digestibility of major allergen tropomyosin and other food proteins of Chinese mitten crab (Eriocheir sinensis). J Sci Food Agric 90(10):1614–1620. https://doi.org/10.1002/jsfa.3988
Rahman AMA, Lopata AL, O’Hehir RE et al (2010) Characterization and de novo sequencing of snow crab tropomyosin enzymatic peptides by both electrospray ionization and matrix-assisted laser desorption ionization QqToF tandem mass spectrometry. J Mass Spectrom 45(4):372–381. https://doi.org/10.1002/jms.1721
Ansotegui IJ, Meliolo G, Canonica GW et al (2020) IgE allergy diagnostics and other relevant tests in allergy, a World Allergy Organization position paper. World Allergy Org J 13(2):100080. https://doi.org/10.1016/j.waojou.2019.100080
Chapman MD, Smith AM, Vailes LD et al (2000) Recombinant allergens for diagnosis and therapy of allergic disease. J Allergy Clin Immunol 106(3):409–418. https://doi.org/10.1067/mai.2000.109832
Larsen JM, Bang-Berthelsen CH, Qvortrup K et al (2020) Production of allergen-specific immunotherapeutic agents for the treatment of food allergy. Crit Rev Biotechnol 40(6):881–894. https://doi.org/10.1080/07388551.2020.1772194
Bird JA, Lack G, Perry TT (2015) Clinical management of food allergy. J Allergy Clin Immunol Pract 3(1):1–11. https://doi.org/10.1016/j.jaip.2014.06.008
Abramovitch JB, Kamath S, Varese N et al (2013) IgE reactivity of blue swimmer crab (Portunus pelagicus) tropomyosin, Por p 1, and other allergens; cross-reactivity with black tiger prawn and effects of heating. PLoS ONE 8(6):1–13. https://doi.org/10.1371/journal.pone.0067487
Liang Y-L, Cao M-J, Su W-J et al (2008) Identification and characterisation of the major allergen of Chinese mitten crab (Eriocheir sinensis). Food Chem 111(4):998–1003. https://doi.org/10.1016/j.foodchem.2008.05.023
Alli SJ, Tsai M (2016) IgE and mast cells in allergic disease. Natl Inst Health 18(5):693–704. https://doi.org/10.1038/nm.2755
Azemi NFH, Misnan M, Keong BP et al (2020) Reference gene and tropomyosin expression in mud crab Scylla olivacea, Scylla paramamosain and Scylla tranquebarica. Mol Biol Rep 47:9765–9777. https://doi.org/10.1007/s11033-020-05966-7
Keenan CP, Davie PJJ, Mann DL (1998) A revision of the genus Scylla De Haan, 1833 (Crustacea: Decapoda: Brachyura: Portunidae). Raffles Bull Zool 46(1):217–245
Fazhan H, Waiho K, Quinitio E et al (2020) Morphological descriptions and morphometric discriminant function analysis reveal an additional four groups of Scylla spp. Peer J 8:e8066. https://doi.org/10.7717/peerj.8066
Zailatul HMY, Leecyous B, Amelia Suhana Z (2020) Isolation and cloning of tropomyosin and arginine kinase from tiger prawn Penaeus monodon and blue swimming crab Portunus trituberculatus. J Sci Math Lett 8(2):2019–2021. https://doi.org/10.37134/jsml.vol8.2.5.2020
Jeong KY, Yum HY, Lee IY et al (2004) Molecular cloning and characterization of tropomyosin, a major allergen of Chironomus kiiensis, a Dominant Species of Nonbiting Midges in Korea. Clin Diagn Lab Immunol 11(2):320–324. https://doi.org/10.1128/cdli.11.2.320-324.2004
Gonza´lez-Ferna´ndez J, Rodero M, Daschner A (2014) New insights into the allergenicity of tropomyosin: a bioinformatics approach. Mol Biol Rep 2014(41):6509–6517. https://doi.org/10.1007/s11033-014-3534-6
Wang X, Li L, Xu F et al (2011) Tropomyosin is a nice marker gene for phylogenetic analysis of molluscs. Mol Biol Rep 38(7):4589–4593. https://doi.org/10.1007/s11033-010-0591-3
Cui Y, Zhou Y, Wang Y et al (2013) The Group 10 allergen of Dermatophagoides farinae (Acari: Pyroglyphidae): cDNA cloning, sequence analysis, and expression in Escherichia coli BL21. J Med Entomol 50(1):205–208. https://doi.org/10.1603/me12019
Kumjim S, Jirapongsananuruk O, Piboonpocanun S (2016) Cloning and characterization of recombinant tropomyosin of giant freshwater shrimp M. rosenbergii to determine major allergens causing allergic reactions among shrimp-allergic children. Asian Pacific J Allergy Immunol 34(3):229–235. https://doi.org/10.12932/AP0698
Piboonpocanun S, Malainual N, Jirapongsananuruk O et al (2006) Genetic polymorphisms of major house dust mite allergens. Clin Exp Allergy 36(4):510–516. https://doi.org/10.1111/j.1365-2222.2006.02464.x
Aalberse RC (2000) Structural biology of allergens. J Allergy Clin Immunol 106(2):228–238. https://doi.org/10.1007/s11033-012-2108-8
Leung PSC, Chow WK, Duffey S (1996) IgE reactivity against a cross-reactivity allergen in crustacea and mollusca: evidence for tropomyosin as the common allergen. J Allergy Clin Immunol 98(5):954–961. https://doi.org/10.1016/s0091-6749(96)80012-1
Zhang Y, Matsuo H, Morita E (2006) Cross-reactivity among shrimp, crab and scallops in a patient with a seafood allergy. J Dermatol 33(3):174–177. https://doi.org/10.1111/j.1346-8138.2006.00040.x
Rolland JM, Varesa NP, Abramovitch JB et al (2018) Effect of heat processing on IgE reactivity and cross-reactivity of tropomyosin and other allergens of Asia-Pacific mollusc species: identification of novel Sydney rock oyster tropomyosin Sac g 1. Mol Nutr Food Res 62(14):e1800148. https://doi.org/10.1002/mnfr.201800148
Ayuso R, Reese G, Leong-Kee S et al (2002) Molecular basis of arthropod cross-reactivity: IgE-binding cross-reactive epitopes of shrimp, house dust mite and cockroach tropomyosins. Int Arch Allergy Immunol 129(1):38–48. https://doi.org/10.1159/000065172
Shafique RH, Inam M, Ismail M et al (2012) Group 10 allergens (tropomyosins) from house-dust mites may cause covariation of sensitization to allergens from other invertebrates. Allergy Rhinol 3(2):74–90. https://doi.org/10.2500/ar.2012.3.0036
Sherson D, Hansen I, Sigsgaard T (1989) Occupationally related respiratory symptoms in trout-processing workers. Allergy 44(5):336–341. https://doi.org/10.1111/j.1398-9995.1989.tb00455.x
Jeebhay MF, Robins TG, Seixas N et al (2005) Environmental exposure characterization of fish processing workers. Ann Occup Hyg 49(5):423–437. https://doi.org/10.1093/annhyg/meh113
James JK, Pike DH, Khan IJ et al (2018) Structural and dynamic properties of allergen and non-allergen forms of tropomyosin. Structure 26(7):997–1006. https://doi.org/10.1016/j.str.2018.05.002
Pekar J, Ret D, Untersmayr E (2018) Stability of allergens. Mol Immunol 100:14–20. https://doi.org/10.1016/j.molimm.2018.03.017
Verhoeckx KCM, Vissers YM, Baumert JL et al (2015) Food processing and allergenicity. Food Chem Toxicol 80:223–240. https://doi.org/10.1016/j.fct.2015.03.005
Jeong KY, Lee H, Lee SK et al (2007) Molecular cloning and the allergenic characterization of tropomyosin from Tyrophagus putrescentiae. Protein Pept Lett 14(5):431–436. https://doi.org/10.2174/092986607780782777
Jeong KY (2004) Characterization of allergenic properties of German cockroach tropomyosin using recombinant proteins. Anim Cells Syst 8:73–73
Asturias JA, Gómez-Bayón N, Arilla MC et al (1999) Molecular characterization of American cockroach tropomyosin (Periplaneta americana allergen 7), a Cross-reactive Allergen. J Immunol 162(7):4342–4348
Kamath SD, Rahman AMA, Komoda T et al (2013) Impact of heat processing on the detection of the major shellfish allergen tropomyosin in crustaceans and molluscs using specific monoclonal antibodies. Food Chem 141(4):4031–4039. https://doi.org/10.1016/j.foodchem.2013.06.105
Faisal M, Vasiljevic T, Donkor ON (2019) Effects of selected processing treatments on antigenicity of banana prawn (Fenneropenaeus merguiensis) tropomyosin. Int J Food Sci Technol 54(1):183–193. https://doi.org/10.1111/ijfs.13922
Koeberl M, Kamath SD, Saptarshi SR et al (2014) Auto-induction for high yield expression of recombinant novel isoallergen tropomyosin from king prawn (Melicertus latisulcatus) for improved diagnostics and immunotherapeutics. J Immunol Methods 415:6–16. https://doi.org/10.1016/j.jim.2014.10.008
Asnoussi A, Aibinu IE, Gasser RB et al (2017) Molecular and immunological characterisation of tropomyosin from Anisakis pegreffii. Parasitol Res 116(12):3291–3301. https://doi.org/10.1007/s00436-017-5642-4
Krieger J, Raming K, Knipper M et al (1990) Cloning, sequencing and expression of locust tropomyosin. Insect Biochem 20(2):173–184. https://doi.org/10.1016/0020-1790(90)90010-R
Ayuso R, Lehrer SB, Reese G (2002) Identification of continuous, allergenic regions of the major shrimp allergen Pen a 1 (Tropomyosin). Int Arch Allergy Immunol 127(1):27–37. https://doi.org/10.1159/000048166
Jeong KY, Lee J, Lee IY et al (2003) Allergenicity of recombinant Bla g 7, German cockroach tropomyosin. Allergy Eur J Allergy Clin Immunol 58(10):1059–1063. https://doi.org/10.1034/j.1398-9995.2003.00167.x
Acknowledgements
The authors wish to thank the Ministry of Higher Education Malaysia (MOE) for the financial funding of this research through the Fundamental Research Grant Scheme (FRGS 2016-0085-102-02).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Consent for publication
All authors have read, commented on, and approved the manuscript.
Ethical approval
Compliance with ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11033_2021_6661_MOESM1_ESM.tif
IgE immunoblotting of crude extract of Scylla olivacea. Lane M, Vivantis Chromatein Prestained Protein Ladder (Vivantis Technologies, USA); lane N, immunoblot using serum from a non-allergic individual; and lane B, blank; lanes 1-22, immunoblots showing binding of IgE from different serum samples.Supplementary file1 (TIF 1669 kb)
11033_2021_6661_MOESM2_ESM.tif
Agarose gel electrophoresis of PCR re-amplification of tropomyosin gene of S. olivacea (o). Marker using 100 bp DNA ladder (Thermo Scientific, USA). Supplementary file2 (TIF 748 kb)
11033_2021_6661_MOESM3_ESM.tif
Overexpression of recombinant tropomyosin protein from S. olivacea. S; soluble, IS; insoluble. Supplementary file3 (TIF 5958 kb)
Rights and permissions
About this article
Cite this article
Azemi, N.F.H., Misnan, R., Keong, B.P. et al. Molecular and allergenic characterization of recombinant tropomyosin from mud crab Scylla olivacea. Mol Biol Rep 48, 6709–6718 (2021). https://doi.org/10.1007/s11033-021-06661-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06661-x