Abstract
The Phytophthora genus is composed, mainly, of plant pathogens. This genus belongs to the Oomycete class, also known as “pseudo-fungi”, within the Chromista Kingdom. Phytophthora spp. is highlighted due to the significant plant diseases that they cause, which represents some of the most economically and cultural losses, such as European chestnut ink disease, which is caused by P. cinnamomi. Currently, there have been four genome assemblies placed at the National Center for Biotechnology Information (NCBI), although the progress to understand and elucidate the pathogenic process of P. cinnamomi by its genome is progressing slowly. In this review paper, we aim to report and discuss the recent findings related to P. cinnamomi and its genomic information. Our research is based on paper databases that reported probable functions to P. cinnamomi proteins using sequence alignments, bioinformatics, and biotechnology approaches. Some of these proteins studied have functions that are proposed to be involved in the asexual sporulation and zoosporogenesis leading to the host colonization and consequently associated with pathogenicity. Some remarkable genes and proteins discussed here are related to oospore development, inhibition of sporangium formation and cleavage, inhibition of flagellar assembly, blockage of cyst germination and hyphal extension, and biofilm proteins. Lastly, we report some biotechnological approaches using biological control, studies with genome sequencing of P. cinnamomi resistant plants, and gene silencing through RNA interference (iRNA).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The species of the Phytophthora genus, which belong to the Oomycetes class, are a fungus-like group of organisms that are mainly plant pathogens and are widely spread around the world (Fig. 1). Among the hosts, forest species are the most affected [1, 2]. Indeed, this genus is highlighted due to the significant diseases that they are responsible for, which represents some of the most economically and cultural losses, such as potato late blight caused by P. infestans, the black shank of tobacco by P. nicotianae, stem rot of soybean by P. sojae and, the ink disease of chestnut caused by P. cinnamomi [3,4,5].
Representation of geographical distribution of Phytophthora cinnamomi worldwide. From: European and Mediterranean Plant Protection Organization (EPPO) (2020) EPPO Global Database (available online). https://gd.eppo.int/. Access date: July 14th, 2020
The first known species of P. cinnamomi was described by Rands in 1922, the Phytophthora cinnamomi var. cinnamomi (Rands) [6]. Nevertheless, the other two varieties were described in 1993 and 2002. Kröber and Marwitz isolated and described P. cinnamomi var. parvispora in 1993 from the nursery plants Beaucarnea genus. The main differences between P. cinnamomi var. cinnamomi and P. cinnamomi var. parvispora are the smaller-sized chlamydospores and sporangia and the highest growth temperature of P. cinnamomi var. parvispora [7, 8]. However, in the recent years, a taxonomic re-evaluation has been proposed, stating that P. cinnamomi var. cinnamomi and P. cinnamomi var. parvispora are separated species (reviewed in SCANU et al. 2014). The study conducted by Scanu and collaborators used multigene phylogeny based on DNA sequences from internal transcribed spacer (ITS) and β-tubulin (nuclear gene regions) and, cox1 and cox2 (mitochondrial gene regions) combined with the study of morphological and physiological properties revealed that Phytophthora parvispora is a unique taxon [9]. A similar situation occurs with the P. cinnamomi var. robiniae (Ho), which was first isolated by Ho from Robinia pseudoacacia in China and differs from the others due to its absence of chlamydospores [10]. Nuclear and mitochondrial analysis conducted by Martin et al. support a new taxonomic classification, whereas Phytophthora robiniae would be a distinct species and, although the authors mentioned that a multilocus analysis of a larger number of isolated was in progress in 2014, to clarify this classification, up to date, there were no updates from these authors [11]. Notwithstanding that the three varieties are harmful to the ecosystems, P. cinnamomi var. cinnamomi is most studied and also the focus of our review.
Castanea sativa (Mill) or the European chestnut tree is very important for economic interests in Portugal, due to its fruits and wood exploitation. This tree is one of the leading orchards in Portugal, mainly in the mountainous regions of Trás-os-Montes. However, the infection by P. cinnamomi has been causing low productivity and economic losses for several years [12]. Moreover, this oomycete is also a concern in Brazil, since some regions have been reporting damages in some important crops, such as avocado trees (Persea americana Miller), plane trees (Platanus acerifolia), chive (Allium fistulosum L.) and lettuce (Lactuca sativa L.), with no efficient pathogen control methods [2, 13, 14].
Infection by P. cinnamomi results in wet rotting of the roots and collar of seedlings and trees in nurseries, plantations, and forests, which leads to the death of the plants [12]. The symptoms of the disease include reduced size and chlorotic leaves, thinning of the crown, and necrosis in the collar of the tree. Nevertheless, the roots are the part of the plant most infected and affected, producing a black exudate visible in the circumjacent soil. The necrosis in the main roots extends to the lateral roots and to the stem for some centimeters, which will lead to the death of the plant [12, 15, 16].
Despite the importance of pathogen control for P. cinnamomi due to its impact on economy and biodiversity, the existing approaches are limited, expensive and, mainly targeted to make the plant more tolerant of the infection [17]. The most successful option for controlling and eliminating P. cinnamomi is the treatment of plants with phosphite, but it demands repeated applications since its effectiveness declines over time [17, 18]. Nevertheless, data on long-term treatment with phosphite and its impact on the ecosystem are scarce, but it is known that in diseased habitats, phosphite treatment significantly reduced the loss of shrub cover, bare ground and sedge cover, but, it does not causes adverse/negative impacts on species assemblages and structure [19, 20]. Other options for pathogen control are the use of the fungicide fosetyl-aluminum [21], the treatment with copper salts to improve host resistance [22] and, the extract of Phlomis purpurea [23]. Furthermore, according to Dunstan et al., the most effective control is host removal, followed by fumigation and fungicide application in the soil [18]. However, the emergence of advanced techniques for plant breeding and pathogen control using molecular biology and biotechnology are promising tools for reducing the damage caused by P. cinnamomi [5].
In line with this, this review aims to discuss the mechanism of infection of P. cinnamomi var. cinnamomi with the latest advances concerning genomic information of this pathogen and the use of its genome to better understand its action and new promising findings.
Phytophthora cinnamomi strategies for plant infection
At the time of writing, the main questions concerning plant infection by P. cinnamomi are whether this pathogen can avoid triggering the host defence, or if it is able to suppress or overcome the host defence, or both of these actions. It is known that other species from the Phytophthora genus, such as P. infestans, P. sojae, and P. capsici have a restricted host range [24,25,26]. However, P. cinnamomi has an extensive host range and in different climate conditions, thus, another important question is which molecular factors may be responsible for this adaptation.
Phytophthora cinnamomi is a soil-borne plant pathogen that belongs to the Oomycetes class, which is a group of fungus-like microorganisms within the Kingdom Chromista [27]. This pathogen can grow in a saprophytic way, in dead organic matter, or in a parasitic way, in susceptible hosts, such as the European chestnut tree [28]. P. cinnamomi has sexual and asexual phases in its lifecycle (Fig. 2) and both of them are implicated in the process of host infection [16, 29].

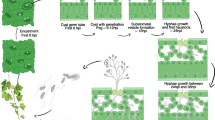
Adapted from Hardham [33]
Brief representation of the life cycle of Phytophthora cinnamomi. The infection of the host plant by P. cinnamomi begins through the connection of zoospores with the region of elongation of the plant roots. Then zoospores enter the encystment phase with the formation of biofilm from secreted proteins. Between 20–30 minutes after the zoospore encysting, the cysts germinate and give rise to hyphae. These hyphae are responsible for the production of enzymes that will degrade the cell wall of plants, such as those belonging to the Cell Wall-Degrading Enzymes (CWDEs) family. The intracellular and intercellular growth of hyphae in the root cortex towards the cortical and vascular tissues causes water stress and necrosis.
Currently, there are four genome assemblies deposited at the National Center for Biotechnology Information (NCBI) for P. cinnamomi var. cinnamomi. These four genome assemblies (https://www.ncbi.nlm.nih.gov/genome/browse#!/eukaryotes/6958/) were obtained through Whole Genome Sequencing (WGS), using Next-Generation Sequencing (NGS) [30, 31]. The samples were from three different locations in Australia and one location in New Zealand, see Table 1 for more detailed information. The Australia ecosystems have been significant affected by P. cinnamomi virulence since this pathogen decimated the Jarrah forest in Western Australia and more than 40% of the plant species present in this region are susceptible to P. cinnamomi infection [32, 33]. However, this genomic information is not complete due to the lack of gene annotation for these genome assemblies.
Nevertheless, the use of bioinformatics tools for local alignments, such as BLAST (Basic Local Alignment Search Tools) (https://blast.ncbi.nlm.nih.gov/Blast.cgi), have made it possible to propose a function for presumably some proteins of the P. cinnamomi genome, through the alignment with homolog sequences. All the following mentioned genes are summarized in Table 2.
During the asexual phase of the P. cinnamomi lifecycle sporulation and zoosporogenesis occurs, and this phase is triggered by the reduction in the availability of nutrients [33]. The asexual spores are motile and capable of forwarding movements and the resultant motile zoospores are thought to initiate the infectious process, as seen in many species of Phytophthora [3, 27]. Indeed, some proteins are proposed to be involved in asexual sporulation and zoosporogenesis and are also associated with the pathogenicity of P. cinnamomi. Gao et al. [34] reported that the silencing of the stress-associated mitogen-activated protein kinase (MAPK) gene in P. sojae, (PsMPK7), inhibited the oospore development and reduced the virulence of this pathogen; P. cinnamomi shares the same gene with P. sojae, probably with a conserved function. Similarly, Li et al. [35] showed that MAPK is up-regulated in sporulating hyphae and early infection in P. sojae (PsMPK1) and that gene silencing revealed inhibition of sporangium formation and also reduced virulence. The sporangial cleavage is an important step in the process of infection and occurs to create uninucleate zoospores from multinucleate sporangia [36]. Hua et al. [37] reported that silencing of G-protein-coupled receptor 4 with a C-terminal PIP kinase domain in P. infestans (PiGK4), a homologue gene in P. cinnamomi, inhibits the sporangial cleavage and, consequently, the infection.
The zoospores formed by sporangial cleavage are the key to the infection through their active movement due to the presence of an anterior flagellum and a posterior flagellum [28]. These flagella have the substructure of a eukaryotic flagellum (9 + 2 microtubular) and the flagellar proteins are encoded for by two genes: dynein light chain 1 (PcDL1) and radial spoke protein 6 (PcRSP6) [38, 39]. Narayan et al. [39] showed that silencing the homologue PcDLC1 gene in P. parasitica resulted in the inhibition of the flagellar assembly, therefore avoiding the infection of the potential host.
To achieve a successful host infection, P. cinnamomi zoospores move towards the elongation zone of plant roots and form a cluster on the plant root surface. Once there, the encysting zoospores secrete adhesins to attach to the root surface and also secrete three glycoproteins to form a mucilage-like biofilm that covers the cyst surface [33, 40]. The next step in the host infection process is cyst germination, after the zoospore encystment, and penetration and colonization of the plant host. The initial penetration possibly depends on the action of degradative enzymes responsible for degrading the components of the plant cell wall. The Cell Wall-Degrading Enzymes (CWDEs) families are known to be present in the genomes of Phytophthora spp [16, 41]. After penetration of the plant root surface, the hyphae of P. cinnamomi grow and followed by the invasion of the root cortex and blockage of xylem, resulting in water stress. P. cinnamomi may continue in a biotrophic phase and, consequently, the absence of disease symptoms, or it may turn to the necrotrophic phase; this transition is mainly influenced by the plant species and environmental conditions [42, 43].
In a study of the protein function in homologue genes, Zhang et al. [44] reported that the silencing of the PsHint1 gene from P. sojae, which encodes the histidine triad domain-containing protein, results in inhibition of cyst germination and hyphal extension. Larousse et al. [45] showed that the PPMUCL1 gene from P. parasitica encodes for mucin-like proteins, this gene is a homologue in the P. cinnamomi genome and these proteins are high-molecular-weight glycoproteins found in biofilms from cysts, and probably protect the germinated cyst against desiccation. Using techniques of gene silencing for the three genes found in P. cinnamomi genome may reduce its virulence and pathogenicity.
Effectors and elicitors for P. cinnamomi interaction with the host
Despite the importance of all the molecular factors mentioned above, the Phytophthora genus has in its genome three distinct biomolecules that are highlighted due to their mechanism of action and importance for host infection. Here we emphasize the CWDEs families, and the molecules involved in plant-pathogen interactions: effectors and elicitors. The colonization of the plant host by P. cinnamomi depends on the action of CWDEs for initial penetration. These enzymes are responsible for the degradation of components of the plant cell wall, such as cellulose, hemicellulose, and pectin [16]. Within the CWDEs, some multigene families contain one or more carbohydrate-active enzyme (CAZyme) modules, and these modules are divided in glycoside hydrolase (GH) module, auxiliary activity (AA) module, carbohydrate esterase (CE) module, polysaccharide lyase (PL) model and non-catalytic carbohydrate-binding module (CBM) [16, 41, 46,47,48].
An effector molecule facilitates the establishment of the infection and, consequently, the establishment of the disease. An elicitor molecule, or avirulence factor, is an effector recognized by the plant and elicits a defence response from the host. Effectors are mainly proteins, such as CWDEs, however, elicitors exist in a great diversity of molecules, such as proteins, lipids, carbohydrates, and may be a molecule resulting from the digestion of the host cell wall, referred to as pathogen-associated molecular patterns (PAMPs) or microbe-associated molecular patterns (MAMPs) [16, 49,50,51,52].
Phytophthora cinnamomi effectors are classified as either attack strategies or defence strategies. Among the main identified effector proteins for attack strategies in the genome of P. cinnamomi are elicitins, a family of low molecular weight proteins that trigger the defence mechanisms of some plant hosts [16]. Although their biological role in the infectious process remains not fully clarified, it is known that they act as carriers of sterols. Cinnamomins are the most studied elicitin from P. cinnamomi, and the silencing of both the α- and β-cinnamomin gene reduces the pathogen’s ability to penetrate and colonize the roots of C. sativa [16, 53].
Small cysteine-rich (SCR) effector proteins are also produced by P. cinnamomi, although their function has not been elucidated, although the silencing of the encoding SCR gene (Scr96) in P. cactorum reduced pathogen virulence and made it more sensitive to oxidative stress [54]. GP42 transglutaminase is present in the P. cinnamomi genome and was identified by Martins et al. [55], its main function is to catalyze the acyl transfer reaction that increases the peptide bonds resistance against proteolytic degradation.
The group of effectors named CRN crinklers is present in most of the identified species of the Phytophthora genus. They are responsible for crinkling the leaves and result in host cells becoming necrotic and suppresses the plant’s defence [56, 57], but more studies are needed to clarify the functions of CRN proteins, because they may be the key to understanding the ability of P. cinnamomi to infect a wide range of host plant species [16]. Another group of effectors that may be involved in plant cell necrosis, is the Nep1-like proteins (NLPs), which may contribute to pathogen virulence by inducing a plant defence response and/or by acting as toxins [58, 59].
The best-studied and clarified effectors proteins within the Phytophthora genus is the RxLR effector family. These molecules are characterized by a signal peptide followed by an N-terminal conserved Arginine – any amino-acid residue – Leucine – Arginine (RxLR) motif [60]. The most common mechanism of action of RxLR effectors is the modulation of host defense by suppression of host immunity through distinct ways [61]. One example of RxLR effector is Avr3a of P. infestans, that confers avirulence to strains of this pathogen, once this protein binds and stabilizes the CMPG1 (E3 ubiquitin ligase protein CMG1) to suppress BAK1/SERK3-regulated immunity during the biotrophic phase of P. infestans infection [62, 63]. Recently, the effector Avr3a gene was identified and characterized in silico in genomic sequences of P. cinnamomi deposited at NCBI by Branco and Choupina [64]. Similarly, Dai et al. identified an RxLR effector, avirulence homolog protein 87 (Avh87), in the genome of P. cinnamomi using bioinformatics tools. In this study, they also characterized the suppressing activity of pro-apoptotic protein BAX and elicitin protein INF1-mediated cell death using Nicotiana benthamiana as a model [65].
The main effectors involved in pathogen defence strategies are molecules that present a diversity of mechanisms, such as suppression, deactivation, and tolerance related to the plant defence. One of the pathogen defences is protection against reactive oxygen species (ROS) that may occur as part of the plant defence response. In line with this, the P. cinnamomi genome contains three superoxide dismutases (SODs) and three catalase genes, and also contains genes for the signal transduction pathways [34, 66,67,68].
Another important effector for defence against pathogens are inhibitors of plant endoglucanases (GIPs), which are enzymes released by the plant that degrade β-1,3-glucans, key components of Phytophthora spp cell wall [1]. A crucial P. cinnamomi defence is protease inhibitors since plant proteases are important in plant immunity because they are involved in pathogen detection, activation of defence responses, and degradation of pathogen proteins. P. cinnamomi, like other species of the Phytophthora genus, produces three families of proteases inhibitor: glucanase inhibitor proteins (GIPs), Kazal-like protease inhibitors (EPI1-4), and cystatin-like protease inhibitors (EPIC1-4) [1, 69, 70]. For a better understanding of the effectors addressed in this review, we recommend the reading of Hardham and Blackman.
How biotechnology can be useful to understand P. cinnamomi and find a solution to environmental problems
The technological advances of the last years promoted the use of new molecular biology and biotechnology techniques to elucidate the molecular factors involved in the pathogenicity of P. cinnamomi. Meyer et al. used the dual RNA-sequencing, a technique that allows the simultaneous detection of pathogen and host transcripts during infection, to better understand the interaction between P. cinnamomi and Eucalyptus nitens factors in the infectious process. The main results revealed that the E. nitens PR-9 gene may be a common target for the CRN effector of the pathogen since a high expression of crinkler effector of P. cinnamomi and a down-regulation of a PR-9 gene in E. nitens were found [71]. In another study, the contribution of the β-cinnamomin in the P. cinnamomi virulence was established by using immunodepletion tests with Lupinus angustifolius. Moreover, the same study revealed that β-cinnamomin is secreted at different life stages of P. cinnamomi, through the use of a β-cinnamomin immune-labeling [72]. Furthermore, recent studies using bioinformatics approaches, heterologous protein expression system and molecular biology techniques have characterized necrosis-inducing Phytophthora protein 1 (NPP1) elicitin and an endo-1,3-β-d-glucosidase of P. cinnamomi [73, 74].
Identification and diagnostic
P. cinnamomi is a soil-borne pathogen that can produce oospores in host roots and soil, which makes this oomycete a persistent and difficult pathogen to manage [28]. The current control measures are soil sanitation and crop rotation, however, P. cinnamomi also is resistant to the most used fungicides and oomyceticides [28, 42]. In line with this, early detection is the most important step in the management of P. cinnamomi diseases, and approaches that use DNA identification are the most reliable [16]. Several variations of polymerase chain reaction (PCR) are available for P. cinnamomi identification, the most recent proposed is a loop-mediated isothermal amplification (LAMP) of DNA using specific primers designed for a new target gene (Pcinn100006) developed by Dai et al. [75]. The novel assay presented higher accuracy and a shorter period, which can be considered a promising diagnostic tool when compared to the conventional PCR-based and culture-dependent assay.
Biological control
The utilization of techniques in biological-control has grown in recent years in numerous biotechnology applications. The use of other organisms to control the presence and the infection by P. cinnamomi has been tested and the results are promising, although more studies are needed. Bosso et al. showed that Byssochlamys nivea and Scopulariopsis brumptii in laboratory studies were able to inhibit the growth of P. cinnamomi and P. cambivora and reduce the mortality of chestnut plants [76]. Supporting the use of biological-control for P. cinnamomi, Méndez-Bravo et al. reported that two rhizobacteria, closely related to Bacillus acidiceler, were able to inhibit P. cinnamomi growth in vitro by 76%, suggesting that these bacteria could be used for biological control of oomycetes [77]. Lastly, Trzewik et al. reported a practical possibility of biological protection against P. cinnamomi or Piriformospora indica, an endomycorrhizal-like fungus, in two cultivars of rhododendron plants (‘Nova Zembla’ and ‘Alfred’) [78].
Resistant plants
It is known that a wide range of plants have resistance to P. cinnamomi infection, among them are Castanea crenata and Castanea dentata (see Appendix 1 [79]). In Portugal, a chestnut breeding program was initiated in 2006 to introduce the resistant genes from C. crenata into C. sativa through crossing both species [80]. However, the identification of the genes involved in the resistance of C. crenata are still not fully clarified. Regardless, studies have reported that C. crenata and C. sativa have the same gene expression of eight genes studied, but C. sativa has a lower and delayed expression of these genes when infected by P. cinnamomi when compared to C. crenata, which may be related to the sensitivity of this species to this pathogen [81]. In this context, the findings reported could be useful for the development of new strategies to control the diseases caused by P. cinnamomi, as an early selection of resistant genotypes, or even engineering C. sativa to obtain better gene promoters.
Gene silencing
Ultimately, the use of molecular biology techniques of gene silencing could be a useful tool to reduce the virulence of P. cinnamomi and thus decrease the effects of the disease. One of the most promising tools for gene silencing is RNA-mediated gene silencing through RNA interference (RNAi). In RNAi, the gene silencing occurs by the inhibition of RNAm translate [82]. Chahed proposed gene silencing using RNAi of a glucanase inhibitor protein (gip gene) from P. cinnamomi, which probably made the pathogen more susceptible to the plant host defence. The results were promising since the chestnuts infected by the transformed P. cinnamomi had a smaller percentage of wilting leaves and root necrosis [83]. Pereira also proposed a transformed P. cinnamomi to decrease the virulence and pathogenicity of the oomycete. In this study, the genetic construction for gene silencing using RNAi was developed for the Avr3a effector from P. cinnamomi [84].
Conclusions
Further research of P. cinnamomi will necessitate the utilization of the new techniques of molecular biology, bioinformatics, and biotechnology. The identification of key genes related to the pathogenicity, the action of the proteins encoded by these genes, and the complete mechanism of infection by P. cinnamomi is necessary for further advances. The development of techniques for regarding genome sequencing, exome sequencing, and gene silencing are essential tools to understand P. cinnamomi and also to understand the mechanisms of resistance of the resistant plants.
References
Martins IM, Martins F, Belo H et al (2014) Cloning, characterization and in vitro and in planta expression of a glucanase inhibitor protein (GIP) of Phytophthora cinnamomi. Mol Biol Rep 41:2453–2462. https://doi.org/10.1007/s11033-014-3101-1
Santos F, Santos ÁF dos (2019) Morte de árvores de plátano causada por Phytophthora cinnamomi. Summa Phytopathol 45:179–185. https://doi.org/10.1590/0100-5405/187118
Kamoun S, Furzer O, Jones JDG et al (2015) The Top 10 oomycete pathogens in molecular plant pathology. Mol Plant Pathol 16:413–434. https://doi.org/10.1111/mpp.12190
Rani DGS (2008) Advances in Soil Borne Plant Diseases. New India Publishing, New Delhi
Choupina AB, Estevinho L, Martins IM (2014) Scientifically advanced solutions for chestnut ink disease. Appl Microbiol Biotechnol 98:3905–3909
Rands RD (1922) Streepkanker van kaneel, veroorzaakt door Phytophthora cinnamomi n. sp. Ruygrok, Batavia
Erwin DC, Ribeiro OK (1996) Phytophthora Diseases Worldwide. Plant Pathol 47:224–225. https://doi.org/10.1046/j.1365-3059.1998.0179a.x
Kröber H, Marwitz R (1993) Phytophthora tentaculata sp. nov. und Phytophthora cinnamomi var. parvispora var. nov., zwei neue Pilze von Zierpflanzen in Deutschland / Phytophthora tentaculata sp. nov. and Phytophthora cinnamoml var. parvispora var. nov., two new fungi from ornamental plants in Germany. Zeitschrift für Pflanzenkrankheiten und Pflanzenschutz / J. Plant Dis. Prot. 100:250–258
Scanu B, Hunter GC, Linaldeddu BT et al (2014) A taxonomic re-evaluation reveals that Phytophthora cinnamomi and P. cinnamomi var. parvispora are separate species. For Pathol 44:1–20. https://doi.org/10.1111/efp.12064
Ho HH (2002) Phytophthora cinnamomi var. robiniae var. nova on black locust in jiangsu province of China. Mycotaxon 82:391–396
Martin FN, Blair JE, Coffey MD (2014) A combined mitochondrial and nuclear multilocus phylogeny of the genus Phytophthora. Fungal Genet Biol 66:19–32. https://doi.org/10.1016/j.fgb.2014.02.006
Vannini A, Vettraino AM, Lellis SC de (2001) Ink disease in chestnuts: impact on the European chestnut. For Snow Landsc Res 76:345–350
Santos MVO dos, Araújo DCS de, Magalhães DMA et al (2014) Phytophthora cinnamomi na rizosfera de cultivos agrícolas no Sul da Bahia. Summa Phytopathol 40:92–92. https://doi.org/10.1590/s0100-54052014000100017
Silva SR da, Canturias-Avilés T, Neto HB et al (2016) Management of root rot in avocado trees. Rev Bras Frutic 38:e–e175
Fernandes CT (1953) A luta contra a “doença da tinta” dos castanheiros no Norte de Portugal. Direção Geral dos Serviços Florestais e Aquícolas 20:153–158
Hardham AR, Blackman LM (2018) Phytophthora cinnamomi. Mol Plant Pathol 19:260–285. https://doi.org/10.1111/mpp.12568
Sena K, Crocker E, Vincelli P, Barton C (2018) Phytophthora cinnamomi as a driver of forest change: Implications for conservation and management. For Ecol Manage 409:799–807. https://doi.org/10.1016/j.foreco.2017.12.022
Dunstan WA, Rudman T, Shearer BL et al (2010) Containment and spot eradication of a highly destructive, invasive plant pathogen (Phytophthora cinnamomi) in natural ecosystems. Biol Invasions 12:913–925. https://doi.org/10.1007/s10530-009-9512-6
Barrett S, Rathbone D (2018) Long-term phosphite application maintains species assemblages, richness and structure of plant communities invaded by Phytophthora cinnamomi. Austral Ecol 43:360–374. https://doi.org/10.1111/aec.12574
Romero MA, González M, Serrano MS, Sánchez ME (2019) Trunk injection of fosetyl-aluminium controls the root disease caused by Phytophthora cinnamomi on Quercus ilex woodlands. Ann Appl Biol 174:313–318. https://doi.org/10.1111/aab.12503
González M, Caetano P, Sánchez ME (2017) Testing systemic fungicides for control of Phytophthora oak root disease. For Pathol 47:e12343. https://doi.org/10.1111/efp.12343
Keast D, Tonkin C, Sanfelieu L (1985) Effects of copper salts on growth and survival of Phytophthora cinnamomi in vitro and on the antifungal activity of actinomycete populations from the roots of Eucalyptus marginata and Banksia grandis. Aust J Bot 33:115–129. https://doi.org/10.1071/BT9850115
Neves D, Caetano P, Oliveira J et al (2014) Anti-Phytophthora cinnamomi activity of Phlomis purpurea plant and root extracts. Eur J Plant Pathol 138:835–846. https://doi.org/10.1007/s10658-013-0357-6
Cooke D, Andersson B (2013) Phytophthora infestans and Potato Late Blight in Europe. In: Lamour K (ed) Phytophthora: a global perspective, 1st ed. CABI, pp 59–68
Dorrance A (2013) Phytophthora sojae on Soybean. In: Lamour K (ed) Phytophthora: a global perspective, 1st ed. CABI, pp 79–86
Sanogo S, Bosland P (2013) Biology and Management of Phytophthora capsici in the Southwestern USA. In: Lamour K (ed) Phytophthora: a global perspective, 1st ed. CABI, pp 87–95
Beakes GW, Glockling SL, Sekimoto S (2012) The evolutionary phylogeny of the oomycete “fungi”. Protoplasma 249:3–19. https://doi.org/10.1007/s00709-011-0269-2
O’Gara E, Howard K, Wilson B, Hardy Ges (2005) Management of Phytophthora cinnamomi for Biodiversity Conservation in Australia: Part 1 ñ A Review of Current Management. A report funded by the Commonwealth Government Department of the Environment and Heritage by the Centre for Phytophthora Science and. Western Australia
Zentmyer GA (1980) Phytophthora cinnamomi and the diseases it causes. American Phytopathological Society(No. 10)
Longmuir AL, Beech PL, Richardson MF (2018) Draft genomes of two Australian strains of the plant pathogen, Phytophthora cinnamomi. F1000Research 6:1–12. https://doi.org/10.12688/f1000research.12867.2
Studholme DJ, McDougal RL, Sambles C et al (2016) Genome sequences of six Phytophthora species associated with forests in New Zealand. Genomics Data 7:54–56. https://doi.org/10.1016/j.gdata.2015.11.015
Shearer BL, Crane CE, Barrett S, Cochrane A (2007) Phytophthora cinnamomi invasion, a major threatening process to conservation of flora diversity in the South-west Botanical Province of Western Australia. Aust J Bot 55:225. https://doi.org/10.1071/BT06019
Hardham AR (2005) Phytophthora cinnamomi. Mol Plant Pathol 6:589–604. https://doi.org/10.1111/j.1364-3703.2005.00308.x
Gao J, Cao M, Ye W et al (2015) PsMPK7, a stress-associated mitogen-activated protein kinase (MAPK) in Phytophthora sojae, is required for stress tolerance, reactive oxygenated species detoxification, cyst germination, sexual reproduction and infection of soybean. Mol Plant Pathol 16:61–70. https://doi.org/10.1111/mpp.12163
Li A, Zhang M, Wang Y et al (2014) PsMPK1, an SLT2-type mitogen-activated protein kinase, is required for hyphal growth, zoosporogenesis, cell wall integrity, and pathogenicity in Phytophthora sojae. Fungal Genet Biol 65:14–24. https://doi.org/10.1016/j.fgb.2014.01.003
Tani S, Judelson H (2006) Activation of zoosporogenesis-specific genes in Phytophthora infestans involves a 7-nucleotide promoter motif and cold-induced membrane rigidity. Eukaryot Cell 5:745–752. https://doi.org/10.1128/EC.5.4.745-752.2006
Hua C, Meijer HJGG, De Keijzer J et al (2013) GK4, a G-protein-coupled receptor with a phosphatidylinositol phosphate kinase domain in Phytophthora infestans, is involved in sporangia development and virulence. Mol Microbiol 88:352–370. https://doi.org/10.1111/mmi.12190
Hardham AR (1987) Microtubules and the flagellar apparatus in zoospores and cysts of the fungus Phytophthora cinnamomi. Protoplasma 137:109–124. https://doi.org/10.1007/BF01281146
Narayan RD, Blackman LM, Shan W, Hardham AR (2010) Phytophthora nicotianae transformants lacking dynein light chain 1 produce non-flagellate zoospores. Fungal Genet Biol 47:663–671. https://doi.org/10.1016/j.fgb.2010.04.008
Robold AV, Hardham AR (2005) During attachment Phytophthora spores secrete proteins containing thrombospondin type 1 repeats. Curr Genet 47:307–315. https://doi.org/10.1007/s00294-004-0559-8
Blackman LM, Cullerne DP, Hardham AR (2014) Bioinformatic characterisation of genes encoding cell wall degrading enzymes in the Phytophthora parasitica genome. BMC Genom 15:. https://doi.org/10.1186/1471-2164-15-785
Crone M, McComb JA, O’Brien PA, Hardy GESJ (2013) Survival of Phytophthora cinnamomi as oospores, stromata, and thick-walled chlamydospores in roots of symptomatic and asymptomatic annual and herbaceous perennial plant species. Fungal Biol 117:112–123. https://doi.org/10.1016/j.funbio.2012.12.004
Crone M, McComb JA, O’Brien PA, Hardy GESJ (2013) Annual and herbaceous perennial native Australian plant species are symptomless hosts of Phytophthora cinnamomi in the Eucalyptus marginata (jarrah) forest of Western Australia. Plant Pathol 62:1057–1062. https://doi.org/10.1111/ppa.12016
Zhang X, Zhai C, Hua C et al (2016) PsHint1, associated with the G-protein α subunit PsGPA1, is required for the chemotaxis and pathogenicity of Phytophthora sojae. Mol Plant Pathol 17:272–285. https://doi.org/10.1111/mpp.12279
Larousse M, Govetto B, Séassau A et al (2014) Characterization of PPMUCL1/2/3, three members of a new oomycete-specific mucin-like protein family residing in Phytophthora parasitica biofilm. Protist 165:275–292. https://doi.org/10.1016/j.protis.2014.03.003
Götesson A, Marshall JS, Jones DA, Hardham AR (2002) Characterization and evolutionary analysis of a large polygalacturonase gene family in the oomycete plant pathogen Phytophthora cinnamomi. Mol Plant-Microbe Interact 15:907–921. https://doi.org/10.1094/MPMI.2002.15.9.907
Larroque M, Barriot R, Bottin A et al (2012) The unique architecture and function of cellulose-interacting proteins in oomycetes revealed by genomic and structural analyses. BMC Genom 13:. https://doi.org/10.1186/1471-2164-13-605
Ospina-Giraldo MD, Griffith JG, Laird EW, Mingora C (2010) The CAZyome of Phytophthora spp.: A comprehensive analysis of the gene complement coding for carbohydrate-active enzymes in species of the genus Phytophthora. BMC Genom 11:525. https://doi.org/10.1186/1471-2164-11-525
Dou D, Kale SD, Wang X et al (2008) RXLR-mediated entry of Phytophthora sojae effector Avr1b into soybean cells does not require pathogen-encoded machinery. Plant Cell 20:1930–1947. https://doi.org/10.1105/tpc.107.056093
Dou D, Kale SD, Wang X et al (2008) Conserved C-terminal motifs required for avirulence and suppression of cell death by Phytophthora sojae effector Avr1b. Plant Cell 20:1118–1133. https://doi.org/10.1105/tpc.107.057067
Du Y, Mpina MH, Birch PRJ et al (2015) Phytophthora infestans RXLR effector AVR1 interacts with exocyst component Sec5 to manipulate plant immunity. Plant Physiol 169:1975–1990. https://doi.org/10.1104/pp.15.01169
Chang YH, Yan HZ, Liou RF (2015) A novel elicitor protein from Phytophthora parasitica induces plant basal immunity and systemic acquired resistance. Mol Plant Pathol 16:123–136. https://doi.org/10.1111/mpp.12166
Horta M do RG (2006) Estudo do papel biológico das elicitinas de Phytophthora cinnamomi e do seu envolvimento no processo infeccioso. Universidade do Algarve
Chen XR, Li YP, Li QY et al (2016) SCR96, a small cysteine-rich secretory protein of Phytophthora cactorum, can trigger cell death in the Solanaceae and is important for pathogenicity and oxidative stress tolerance. Mol Plant Pathol 17:577–587. https://doi.org/10.1111/mpp.12303
Martins IM, Matos M, Costa R et al (2014) Transglutaminases: Recent achievements and new sources. Appl Microbiol Biotechnol 98:6957–6964
Torto TA, Li S, Styer A et al (2003) EST Mining and Functional Expression Assays Identify Extracellular Effector Proteins From the Plant Pathogen Phytophthora. Genome Res 13:1675–1685. https://doi.org/10.1101/gr.910003
Stam R, Jupe J, Howden AJM et al (2013) Identification and Characterisation CRN Effectors in Phytophthora capsici Shows Modularity and Functional Diversity. PLoS One 8:e59517. https://doi.org/10.1371/journal.pone.0059517
Oome S, Van Den Ackerveken G (2014) Comparative and functional analysis of the widely occurring family of Nep1-like proteins. Mol Plant-Microbe Interact 27:1081–1094. https://doi.org/10.1094/MPMI-04-14-0118-R
Santhanam P, Van Esse HP, Albert I et al (2013) Evidence for functional diversification within a fungal Nep1-like protein family. Mol Plant-Microbe Interact 26:278–286. https://doi.org/10.1094/MPMI-09-12-0222-R
Morgan W, Kamoun S (2007) RXLR effectors of plant pathogenic oomycetes. Curr Opin Microbiol 10:332–338
Anderson RG, Deb D, Fedkenheuer K, McDowell JM (2015) Recent progress in RXLR effector research. Mol. Plant-Microbe Interact 28:1063–1072
Chaparro-Garcia A, Schwizer S, Sklenar J et al (2015) Phytophthora infestans RXLR-WY Effector AVR3a Associates with Dynamin-Related Protein 2 Required for Endocytosis of the Plant Pattern Recognition Receptor FLS2. PLoS One 10:e0137071. https://doi.org/10.1371/journal.pone.0137071
Bos JIB, Armstrong MR, Gilroy EM et al (2010) Phytophthora infestans effector AVR3a is essential for virulence and manipulates plant immunity by stabilizing host E3 ligase CMPG1. Proc Natl Acad Sci U S A 107:9909–9914. https://doi.org/10.1073/pnas.0914408107
Branco I, Choupina AB (2020) In Silico Characterization of the Phytopathogenic Efector, avr3a, from Phytophthora cinnamomi. J Basic Appl Sci 16:20–30
Dai T, Wang A, Yang X et al (2020) PHYCI_587572: An RxLR Effector Gene and New Biomarker in A Recombinase Polymerase Amplification Assay for Rapid Detection of Phytophthora cinnamomi. Forests 11:306. https://doi.org/10.3390/f11030306
Blackman LM, Mitchell HJ, Hardham AR (2005) Characterisation of manganese superoxide dismutase from Phytophthora nicotianae. Mycol Res 109:1171–1183. https://doi.org/10.1017/S0953756205003771
Blackman LM, Hardham AR (2008) Regulation of catalase activity and gene expression during Phytophthora nicotianae development and infection of tobacco. Mol Plant Pathol 9:495–510. https://doi.org/10.1111/j.1364-3703.2008.00478.x
Li A, Weng Y, Tao K et al (2010) PsSAKI, a stress-activated MAP kinase of phytophthora sojae, is required for zoospore viability and infection of soybean. Mol Plant-Microbe Interact 23:1022–1031. https://doi.org/10.1094/MPMI-23-8-1022
Chinnapun D, Tian M, Day B, Churngchow N (2009) Inhibition of a Hevea brasiliensis protease by a Kazal-like serine protease inhibitor from Phytophthora palmivora. Physiol Mol Plant Pathol 74:27–33. https://doi.org/10.1016/j.pmpp.2009.08.005
Kaschani F, van der Hoorn RAL (2011) A model of the C14-EPIC complex indicates hotspots for a protease-inhibitor arms race in the oomycete-potato interaction. Plant Signal Behav 6:109. https://doi.org/10.4161/psb.6.1.14190
Meyer FE, Shuey LS, Naidoo S et al (2016) Dual RNA-sequencing of Eucalyptus nitens during Phytophthora cinnamomi challenge reveals pathogen and host factors influencing compatibility. Front Plant Sci 7:191. https://doi.org/10.3389/fpls.2016.00191
Islam MT, Hussain HI, Russo R et al (2019) Functional analysis of elicitins and identification of cell wall proteins in Phytophthora cinnamomi. Physiol Mol Plant Pathol 107:21–32. https://doi.org/10.1016/j.pmpp.2019.04.003
Costa R, Domínguez A, Choupina A (2020) Cloning and expression analysis of an endo-1,3-β-d-glucosidase from Phytophthora cinnamomi. Mol Biol Rep 47:935–942. https://doi.org/10.1007/s11033-019-05185-9
Martins IM, Meirinho S, Costa R et al (2019) Cloning, characterization, in vitro and in planta expression of a necrosis-inducing Phytophthora protein 1 gene npp1 from Phytophthora cinnamomi. Mol Biol Rep 46:6453–6462. https://doi.org/10.1007/s11033-019-05091-0
Dai T, Yang X, Hu T et al (2019) A Novel LAMP Assay for the Detection of Phytophthora cinnamomi Utilizing a New Target Gene Identified from Genome Sequences. Plant Dis 103:3101–3107. https://doi.org/10.1094/PDIS-04-19-0781-RE
Bosso L, Scelza R, Varlese R et al (2016) Assessing the effectiveness of Byssochlamys nivea and Scopulariopsis brumptii in pentachlorophenol removal and biological control of two Phytophthora species. Fungal Biol 120:645–653. https://doi.org/10.1016/j.funbio.2016.01.004
Méndez-Bravo A, Cortazar-Murillo EM, Guevara-Avendaño E et al (2018) Plant growth-promoting rhizobacteria associated with avocado display antagonistic activity against Phytophthora cinnamomi through volatile emissions. PLoS One 13:1–18. https://doi.org/10.1371/journal.pone.0194665
Trzewik A, Maciorowski R, Klocke E, Orlikowska T (2020) The influence of Piriformospora indica on the resistance of two rhododendron cultivars to Phytophthora cinnamomi and P. plurivora. Biol Control 140:104121. https://doi.org/10.1016/j.biocontrol.2019.104121
Groves E, Hardy G, McComb J (2020) Compile list of Western Australian Natives susceptible and resistant to Phytophthora cinnamomi. Murdoch University
Santos C, Nelson CD, Zhebentyayeva T et al (2017) First interspecific genetic linkage map for Castanea sativa x Castanea crenata revealed QTLs for resistance to Phytophthora cinnamomi. PLoS One 12:e0184381. https://doi.org/10.1371/journal.pone.0184381
Santos C, Duarte S, Tedesco S et al (2017) Expression Profiling of Castanea Genes during Resistant and Susceptible Interactions with the Oomycete Pathogen Phytophthora cinnamomi Reveal Possible Mechanisms of Immunity. Front Plant Sci 8:515. https://doi.org/10.3389/fpls.2017.00515
Lenz G (2005) The RNA interference revolution. Brazilian J Med Biol Res 38:1749–1757. https://doi.org/10.1590/S0100-879X2005001200003
Chahed A (2016) Silencing the gip gene of Phytophthora cinnamomi by iRNA and studying the subcellular localization of GIP and NPP1. Instituto Politécnico de Bragança
Pereira NAF da N (2017) Caracterização de fatores moleculares implicados na patogenicidade de Phytophthora cinnamomi. Instituto Politécnico de Bragança
Acknowledgements
The authors are grateful to the Foundation for Science and Technology (FCT, Portugal) and FEDER under Programme PT2020 for financial support to CIMO (UID/AGR/00690/2019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there is no conflict of interest in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
de Andrade Lourenço, D., Branco, I. & Choupina, A. Phytopathogenic oomycetes: a review focusing on Phytophthora cinnamomi and biotechnological approaches. Mol Biol Rep 47, 9179–9188 (2020). https://doi.org/10.1007/s11033-020-05911-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05911-8