Abstract
Hepatocellular carcinoma is a primary malignancy of hepatocytes which accounts for 80 % of all primary liver cancers. DFNA5 has been identified as a tumor suppressor gene with an important role in several frequent forms of cancers, while little is known about its role in hepatocellular carcinoma. Through comparison of the DFNA5 protein expression in hepatocellular carcinoma cells (HepG2) with human fetal lung fibroblast cells (MRC5), we found that the DFNA5 protein expression in hepatocellular carcinoma cells was significantly lower than that in normal cells. The transfection of DFNA5 gene into HepG2 cells could increase DFNA5 protein expression, which subsequently led to inhibition of cell proliferation. Underlying mechanism study revealed that decreased proliferation was due to increased apoptosis and cell cycle arrest. In view of the important role of DFNA5 gene in carcinogenesis, these findings are expected to provide new understanding on development and treatment of human hepatocellular carcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is a primary malignancy of hepatocytes which accounts for 80 % of all primary liver cancers and ranks globally as the fourth leading cause of cancer-related death[1]. Epidemiological studies have provided overwhelming evidence that cirrhosis is the strongest and the most common risk factor for HCC, particularly cirrhosis related to hepatitis C virus and hepatitis B virus infections [2, 3]. Epigenetic changes such as DNA methylation and histone modification have recently been shown to play important roles in tumorigenesis [4]. Genomic hypomethylation increases chromosome instability [5] while localized hypermethylation decreases tumor suppressor gene expression [6], thus increasing the risk of HCC development. A number of tumor suppressor genes, including p16 INK4A [7], suppressor of cytokine signaling-1 [8], adenomatous polyposis coli [9], glutathione-S-transferase pi class gene [10] and E-cadherin [11], are silenced by DNA methylation in HCC, and this process often starts at the preneoplastic stage [12]. Moreover, genes involved in cell cycle checkpoints[13], apoptosis [14], and gene transcription [15] all have been shown to play important roles in HCC.
The human deafness, autosomal dominant 5 gene (DFNA5) was first discovered in Dutch family with autosomal dominant hearing loss [16]. Since its identification, several published papers have shown that DFNA5 is a tumor suppressor gene and is epigenetically inactivated in several types of cancer, including gastric [17], colorectal [18] and breast cancer [19]. Forced expression of DFNA5 in colorectal carcinoma cell lines can decrease cell growth and colony-forming ability [18]. In breast carcinoma, the methylation status of DFNA5 is correlated with lymph node metastasis [19]. However, whether DFNA5 gene is involved in the carcinogenesis of HCC has never been explored before. In this study, we aimed to investigate the DFNA5 expression in HCC and explore its underlying molecular mechanism.
Materials and methods
Cell lines
Human cell line HepG2 (hepatoblastoma) and human fetal lung fibroblast cell (MRC5 cell line) were obtained from the American Tissue Culture Collection (ATCC, Rockville, USA). HepG2 and MRC5 were respectively cultured in PRMI1640 medium and high glucose dulbecco’s modified eagle’s medium (DMEM), supplemented with 10 % heat-inactivated fetal bovine serum, 100 μ/ml penicillin, and 100 μg/ml streptomycin at 37 °C in a humidified incubator with 5 % CO2.
Transfection experiments
Eukaryotic cell expression plasmid pEGFP-N1-DFNA5 was purchased from Yingrui Biotech (Changsha, China). The HepG2 cells at logarithmic phase were seeded in 6-well plates at a density of 2 × 105 cells/well and grown for 24 h. Cells were transfected with pEGFP-N1-DFNA5 (with 1 μg plasmid) using Lipofectamine 2000 reagent according to the protocols provided by the manufacture(Invitrogen, Carlsbad, CA).
Western blotting
At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry (BD Biosciences, Heidelberg, Germany). Both of the HepG2 and MRC5 cells were washed three times and lysed in ice-cold lysis buffer for 30 min, followed by centrifuging at 12,000 rpm for 10 min at 4 °C. The sediment was discarded and the protein concentration was measured by bicinchoninic acid method. Equivalent amounts of denatured cellular lysates (20 μg) from each cell line were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis and then transferred onto a polyvinylidene membrane using electro-transfer system. The membranes were blocked with 5 % skimmed milk solution for 2 h at room temperature. Next, the membranes were incubated with primary antibodies (1:1,000 dilution), including anti-DFNA5 antibody (sigma), anti-FAS antibody, anti-FasL antibody, and anti-cleaved caspase 8 antibody (Cell Signaling, USA) at 4 °C overnight, followed by a secondary antibody (Anti-rabbit IgG, HRP-linked Antibody, 1:5,000 dilution, Cell Signaling, USA) for 1 h at room temperature. Immunoblots were developed using the enhanced chemiluminescence western blotting kit (Santa Cruz Biotechnology). β-actin protein level was measured as reference.
Tetrazolium-based colorimetric assay (MTT test)
At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted using flow cytometry (BD Biosciences, Heidelberg, Germany) and prepared into single cell suspension. The cells were seeded in 96-well plate in triplicate at a density of 2 × 103/well and grown for two days, followed by the addition of 50 μl MTT (1 mg/ml) in each well. After a 4 h culture, the supernatant was discarded, 150 μl dimethyl sulfoxide was added to each well, and the plate was shaken for 10 min to dissolve the MTT. Absorbance of each well was measured by 96-well microplate reader at 570 nm wavelength and recorded.
Measurements of cell apoptosis
At 24 or 48 h after transfection, the HepG2 cells were collected by centrifugation at 1,000 rpm for 5 min and washed twice by PBS. The precipitate was resuspended in 500 μl binding buffer and stained with Annevin V-FITC/propidium iodide (PI) (Kaiji Biotechnology, Nanjing, China) for 5–15 min in the dark at room temperature. Apoptotic cells were analyzed by flow cytometry within 1 h and apoptosis index (AI) was calculated using the following formula: AI = apoptosis cells/total cells × 100 %.
Measurement of cell cycle
At 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry and centrifuged at 1,000 rpm for 5 min, followed by washing twice with PBS. After fixation in pre-cooled 70 % ethanol at 4 °C for 2 h, the fixation buffer was discarded, the cells were resuspended in PBS and passed through 400 mesh. After that, the cells were centrifuged at 1,000 rpm for 5 min and the PBS was discarded. The cells were then stained with PI for 30 min in the dark at 4 °C. The cell cycle was analyzed by flow cytometry.
Statistical analysis
All statistical analyses were performed using t test. P-values below 0.05 were considered statistically significant. Statistics were calculated using SPPS 15.0 (SPSS Inc., Chicago, IL, USA).
Results
DFNA5 expression in HepG2 and MRC5 cell lines
Western blotting analysis using anti-DFNA5 antibody was done to evaluate the DFNA5 expression in HepG2 and MRC5 cells. We found that the DFNA5 expression level in MRC5 cells was significantly higher than that in HepG2 cells. The quantitative analysis of the expression level through scanning the image density revealed that the DFNA5 expression level in HepG2 cells was only equal to around 20 % of that in MRC5 cells (Fig. 1).
DFNA5 protein expression in HepG2 cells and MRC5 cells. HepG2 cells and MRC5 cells at logarithmic phase were collected and subjected to western blotting analysis for DFNA5 protein expression using anti-DFNA5 antibody. The DFNA5 expression in HepG2 cells was significantly lower than that in MRC5 cells. β-actin expression served as internal control. a electrophoretogram; b quantitative analysis. *compared with MRC5 cells, P < 0.05
The expression level increases after transfection of DFNA5 into HepG2 cells
Since the DFNA5 expression in HepG2 cell cline had been demonstrated to be much lower than that in normal cells, whether transfection of DFNA5 into HepG2 cells could increase the DFNA5 expression became a subject of interesting investigation. The transfection experiment was performed on HepG2 cells using lipofectamine 2000. At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry and the DFNA5 expression level was detected by western blotting. The results indicated that the DFNA5 expression level in DFNA5 transfected HepG2 cells was significantly increased at 24 h after transfection and 48 h after transfection (p < 0.05, Fig. 2).
DFNA5 protein expression in DFNA5 transfected HepG2 cells. At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry. HepG2 cells were collected and subjected to western blotting analysis for DFNA5 protein expression using anti-DFNA5 antibody. The DFNA5 expression level in HepG2 cells significantly increased after transfection. a electrophoretogram; b quantitative analysis. *compared with 0 h, P < 0.05
Effect of transfection of DFNA5 into HepG2 cells on cell proliferation and apoptosis
To investigate how the transfection of DFNA5 into HepG2 cells affects the cell proliferation, we performed the transfection experiment as mentioned above and compared the viability of DFNA5 transfected HepG2 with the viability of non-transfected HepG2 cells through MTT assay at 24 or 48 h after incubation. As expected, the viability of non-transfected HepG2 cells didn’t show much difference between 24 and 48 h (90–88 %), while DFNA5 transfected HepG2 cells showed a decreasing viability after transfection in a time-dependent manner. The viability of transfected cells decreased from 86 to 60 %, which was significantly lower than that of non-transfected cells (p < 0.05, Fig. 3).
Cell viability decreased 48 h after transfection of DFNA5 gene into HepG2 cells. At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry. MTT assay was performed to analyze cell viability. The viability of non-transfected HepG2 cells didn’t show much difference between 24 h and 48 h (90 %-88 %), while the viability of DFNA5 transfected cells decreased from 86 % to 60 %, which was significantly lower than that of non-transfected cells (*p < 0.05)
Simultaneously, apoptosis experiments was also conducted on HepG2 cells at 48 h after transfection, and the results showed that the apoptotic rate in the DFNA5 transfected HepG2 cells was around 30 %, which was significantly higher than that in non-transfected cells with 5 % apoptotic rate (Fig. 4).
Effect of transfection of DFNA5 gene into HepG2 cells on cell apoptosis. At 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry and stained with Annevin V-FITC/propidium iodide to detect apoptosis. The cell apoptotic rate in the DFNA5 transfected HepG2 cells (b) was strikingly higher that in non-transfected cells (a)
Changes in apoptosis-related proteins after transfection of DFNA5 into HepG2 cells
We have demonstrated that the transfection of DFNA5 into HepG2 cells could significantly increase apoptosis rate, while we are not clear how the apoptosis related proteins mediate the apoptosis process. Here, the expressions of Fas, Fas ligand (FasL), and capase 8, were analyzed by western blotting in the DFNA5 transfected HepG2 cells. We found that the expressions of Fas and caspase 8 were significantly increased after transfection in a time-dependent manner, while the expression of FasL was reduced after transfection in a time-dependent manner (Fig. 5).
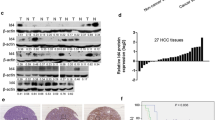
Effect of transfection of DFNA5 gene into HepG2 cells on Fas, Fas L and caspase 8 protein expression. At 24 or 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry and subjected to western blotting analysis for DFNA5 protein expression using anti-Fas antibody, anti-FasL antibody, and anti-cleaved caspase 8 antibody. The expression of Fas and caspase 8 significantly increased after transfection in a time-dependent manner, while the expression of FasL decreased after transfection in a time-dependent manner. a electrophoretogram; b quantitative analysis. *compared with 0 h, P < 0.05
Effect of transfection of DFNA5 into HepG2 cells on cell cycle
To determine the nature of decreased cell proliferation in DFNA5 transfected HepG2 cells, we performed cell cycle analysis through flow cytometry. As shown in Fig. 6, transfection of DFNA5 into HepG2 cells enhanced G2/M arrest with 31.3 % of population arrested in the G2/M, compared to 6.2 % of population arrested in the G2/M in the non-transfected HepG2 cells. So the increased G2/M arrest in transfected HepG2 cells might account for the decreased cell proliferation.
Effect of transfection of DFNA5 gene into HepG2 cells on cell cycle. At 48 h after transfection, the fluorescence-labeled HepG2 cells were sorted by flow cytometry and subjected to cell cycle analysis. Transfection of DFNA5 into HepG2 cells enhanced G2/M arrest with 31.3 % of population arrested in the G2/M (b), compared to 6.2 % of population arrested in the G2/M in the non-transfected HepG2 cells (a)
Discussion
Aberrant methylation of DFNA5 has been recently detected in gastric, colorectal and breast cancer [17–20], which leads to the down-regulation of DFNA5 compared with matched normal tissues or cells. Our study was the first to investigate the role of DFNA5 in HCC. As expected, the DFNA5 expression level was also significantly lower in HCC cell than that in normal human MRC5 cell. In order to explore the underlying molecular mechanism of DFNA5 in HCC cell, we transfected DFNA5 into HepG2 cell and found that the DFNA5 expression level was significantly increased after transfection. Subsequent studies indicated that DFNA5 overexpression could inhibit HCC cell proliferation through induction of cell apoptosis and G2/M arrest. These results are in accordance with a recent study described by de Beeck et al. [21]. They report that DFNA5 is composed of two domains: the first region induces apoptosis when transfected in HEK293T cells and the second region masks and probably regulates this apoptosis inducing capability [21].
Apoptosis is the most common forms of eukaryotic cell death, acting as a physiological suicide mechanism to preserve homeostasis, and occurs naturally during tissue turnover [22]. Any decrease in apoptotic rate may disturb this homeostasis and result in cancer formation and progression [23]. Fas is a member of a subgroup of the tumor necrosis factor receptor superfamily that contain an intracellular death domain and can initiate apoptosis signaling [24]. It has been reported that HCC tissues express Fas less frequently and more weakly than noncancerous tissues [25]. FasL is normally expressed by T lymphocytes when they are activated [26]. FasL has been reported to be high expressed in many human tumor cells and promotes apoptosis of T Lymphocytes [27], ultimately leading to immune evasion of tumor cells [28]. Low expression of Fas and high expression of FasL cause tumor metastasis [29]. Caspase-8 is an aspartate-specific cysteine protease and present in healthy cells as an inactive proenzyme. Through complex proximity-driven conformational changes, procaspase-8 molecules within the death-inducing signaling complex (DISC) gain full enzymatic activity [30]. This active caspase-8 can leave the DISC to target its specific substrates within the cytosol, such as effector caspases (caspase-3), which is critical for Fas-mediated apoptosis [31]. A characteristic feature of human cancer is their inability to undergo apoptosis in response to apoptotic stimuli, and the inactivation of caspase-8 by genetic or epigenetic mechanisms is one mechanism to evade apoptosis [32, 33]. In this study, we transfected DFNA5 into HepG2 cells and increased the Fas and caspase-8 protein expression, while decreased FasL protein expression, which finally contribute to the induction of apoptosis and prevention of tumor counterattack.
It is known that cell growth is controlled by cell cycle progression, a highly regulated process. The standard cell cycle is divided into four non-overlapping phases, namely G1, S, G2 and M phases in sequence. Each phase has checkpoints that cause cell cycle arrest and activation of repair mechanisms. Unlike normal cells that rely on the G1 checkpoint to protect against DNA damage, cancer cells are more dependent on G2 checkpoint for DNA damage repair [34]. Our study demonstrated that transfection of DFNA5 into HepG2 cells induced G2/M arrest, which leads to inhibition of cell proliferation [35]. While the underlying mechanism on how the transfection of DFNA5 into HepG2 cells to induce G2/M arrest requires further investigations.
References
Abrams P, Marsh JW (2010) Current approach to hepatocellular carcinoma. Surg Clin North Am 90(4):803–816. doi:S0039-6109(10)00045-9 10.1016/j.suc.2010.04.010
Tan YJ (2011) Hepatitis B virus infection and the risk of hepatocellular carcinoma. World J Gastroenterol 17(44):4853–4857. doi:10.3748/wjg.v17.i44.4853
Cabibbo G, Craxi A (2010) Epidemiology, risk factors and surveillance of hepatocellular carcinoma. Eur Rev Med Pharmacol Sci 14(4):352–355
Toyota M (2005) Issa J Epigenetic changes in solid and hematopoietic tumors. Seminars in oncology, In, p 521
Zhang YJ, Wu HC, Yazici H, Yu MW, Lee PH, Santella RM (2012) Global hypomethylation in hepatocellular carcinoma and its relationship to aflatoxin B1 exposure. World J hepatol 4(5):169
Nomoto S, Kinoshita T, Kato K, Otani S, Kasuya H, Takeda S, Kanazumi N, Sugimoto H, Nakao A (2007) Hypermethylation of multiple genes as clonal markers in multicentric hepatocellular carcinoma. Bri J cancer 97(9):1260–1265
Saelee P, Petmitr S, Wongkham S, Chariyalertsak S, Homcha-cm P (2010) Detection of DNA methylation of p16INK4a in hepatocellular carcinoma by methylation specific-polymerase chain reaction. Thai Cancer J 29(2):72–79
Okochi O, Hibi K, Sakai M, Inoue S, Takeda S, Kaneko T, Nakao A (2003) Methylation-mediated silencing of SOCS-1 gene in hepatocellular carcinoma derived from cirrhosis. Clin Cancer Res 9(14):5295–5298
Jain S, Chang TT, Hamilton JP, Lin SY, Lin YJ, Evans AA, Selaru FM, Lin PW, Chen SH, Block TM (2011) Methylation of the CpG Sites only on the sense strand of the APC gene is specific for hepatocellular carcinoma. PLoS ONE 6(11):e26799
Jain S, Chen S, Chang KC, Lin YJ, Hu CT, Boldbaatar B, Hamilton JP, Lin SY, Chang TT, Chen SH (2012) Impact of the location of CpG methylation within the GSTP1 gene on its specificity as a DNA marker for hepatocellular carcinoma. PLoS ONE 7(4):e35789
Lim SO, Gu JM, Kim MS, Kim HS, Park YN, Park CK, Cho JW, Park YM, Jung G (2008) Epigenetic changes induced by reactive oxygen species in hepatocellular carcinoma: methylation of the E-cadherin promoter. Gastroenterology 135(6):2128–2140
Lee S, Lee HJ, Kim JH, Lee HS, Jang JJ, Kang GH (2003) Aberrant CpG island hypermethylation along multistep hepatocarcinogenesis. Am J Pathol 163(4):1371
Wang X, Meng X, Sun X, Liu M, Gao S, Zhao J, Pei F, Yu H (2009) Wnt/β-catenin signaling pathway may regulate cell cycle and expression of cyclin A and cyclin E protein in hepatocellular carcinoma cells. Cell Cycle 8(10):1567–1570
Augello C, Caruso L, Maggioni M, Donadon M, Montorsi M, Santambrogio R, Torzilli G, Vaira V, Pellegrini C, Roncalli M (2009) Inhibitors of apoptosis proteins (IAPs) expression and their prognostic significance in hepatocellular carcinoma. BMC Cancer 9(1):125
Tanaka S, Shiraha H, Nakanishi Y, Nishina SI, Matsubara M, Horiguchi S, Takaoka N, Iwamuro M, Kataoka J, Kuwaki K (2012) Runt-related transcription factor 3 reverses epithelial–mesenchymal transition in hepatocellular carcinoma. Int J Cancer 131:2537–2546
Van Laer L, Huizing EH, Verstreken M, Van Zuijlen D, Wauters JG, Bossuyt PJ, Van de Heyning P, McGuirt WT, Smith RJH, Willems PJ (1998) Nonsyndromic hearing impairment is associated with a mutation in DFNA5. Nat Genet 20(2):194–197
Akino K, Toyota M, Suzuki H, Imai T, Maruyama R, Kusano M, Nishikawa N, Watanabe Y, Sasaki Y, Abe T (2006) Identification of DFNA5 as a target of epigenetic inactivation in gastric cancer. Cancer Sci 98(1):88–95
Kim M, Chang X, Yamashita K, Nagpal J, Baek J, Wu G, Trink B, Ratovitski E, Mori M, Sidransky D (2008) Aberrant promoter methylation and tumor suppressive activity of the DFNA5 gene in colorectal carcinoma. Oncogene 27(25):3624–3634
Kim MS, Lebron C, Nagpal JK, Chae YK, Chang X, Huang Y, Chuang T, Yamashita K, Trink B, Ratovitski EA (2008) Methylation of the DFNA5 increases risk of lymph node metastasis in human breast cancer. Biochem Biophy Res Commu 370(1):38–43
Fujikane T, Nishikawa N, Toyota M, Suzuki H, Nojima M, Maruyama R, Ashida M, Ohe-Toyota M, Kai M, Nishidate T (2010) Genomic screening for genes upregulated by demethylation revealed novel targets of epigenetic silencing in breast cancer. Breast Cancer Res Tr 122(3):699–710
De Beeck KO, Van Camp G, Thys S, Cools N, Callebaut I, Vrijens K, Van Nassauw L, Van Tendeloo VFI, Timmermans JP, Van Laer L (2011) The DFNA5 gene, responsible for hearing loss and involved in cancer, encodes a novel apoptosis-inducing protein. Eur J Human Genet 19(9):965–973
Searle J, Harmon B, Bishop C, Kerr J (2008) The significance of cell death by apoptosis in hepatobiliary disease. J Gastroenterol Hepatol 2(1):77–96
Brown JM, Attardi LD (2005) The role of apoptosis in cancer development and treatment response. Nat Rev Cancer 5(3):231–237
Kaufmann T, Strasser A, Jost P (2011) Fas death receptor signalling: roles of Bid and XIAP. Cell Death Differ 19(1):42–50
Higaki K, Yano H, Kojiro M (1996) Fas antigen expression and its relationship with apoptosis in human hepatocellular carcinoma and noncancerous tissues. Am J Pathol 149(2):429
Kägi D, Ledermann B, Bürki K, Seiler P, Odermatt B, Olsen KJ, Podack ER, Zinkernagel RM, Hengartner H (1994) Cytotoxicity mediated by T cells and natural killer cells is greatly impaired in perforin-deficient mice. Nature 369:31–37
Gastman BR, Atarashi Y, Reichert TE, Saito T, Balkir L, Rabinowich H, Whiteside TL (1999) Fas ligand is expressed on human squamous cell carcinomas of the head and neck, and it promotes apoptosis of T lymphocytes. Cancer Res 59(20):5356–5364
Ryan AE, Shanahan F, O’Connell J, Houston AM (2006) Fas ligand promotes tumor immune evasion of colon cancer in vivo. Cell Cycle 5(3):246–249
Kykalos S, Mathaiou S, Karayiannakis AJ, Patsouras D, Lambropoulou M, Simopoulos C (2012) Tissue expression of the proteins fas and fas ligand in colorectal cancer and liver metastases. J Gastrointest Cancer 43(2):224–228
Boatright KM, Renatus M, Scott FL, Sperandio S, Shin H, Pedersen IM, Ricci JE, Edris WA, Sutherlin DP, Green DR (2003) A unified model for apical caspase activation. Mol Cell 11(2):529–541
Stennicke HR, Jürgensmeier JM, Shin H, Deveraux Q, Wolf BB, Yang X, Zhou Q, Ellerby HM, Ellerby LM, Bredesen D (1998) Pro-caspase-3 is a major physiologic target of caspase-8. J Biological Chem 273(42):27084–27090
Fulda S (2009) Caspase-8 in cancer biology and therapy. Cancer Lett 281(2):128–133
Lavrik I, Krammer P (2011) Regulation of CD95/Fas signaling at the DISC. Cell Death Differ 19(1):36–41
Bucher N, Britten C (2008) G2 checkpoint abrogation and checkpoint kinase-1 targeting in the treatment of cancer. Bri J cancer 98(3):523–528
Zhang L, Ruan J, Yan L, Li W, Wu Y, Tao L, Zhang F, Zheng S, Wang A, Lu Y (2012) Xanthatin induces cell cycle arrest at G2/M checkpoint and apoptosis via disrupting NF-κB pathway in A549 non-small-cell lung cancer cells. Molecules 17(4):3736–3750
Acknowledgments
This project is supported by Grants from the National Natural Science Foundation of China (No. 30872510, 81200320, 81272534, 81260349); the Natural Science Foundation of Hubei Province (No. 2008CDB127), and the Natural Science Foundation of Shanghai (No. 064119620 10411968400).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Cong-Jun Wang and Dong-Wei Shen are co-first authors.
Rights and permissions
About this article
Cite this article
Wang, CJ., Tang, L., Shen, DW. et al. The expression and regulation of DFNA5 in human hepatocellular carcinoma DFNA5 in hepatocellular carcinoma. Mol Biol Rep 40, 6525–6531 (2013). https://doi.org/10.1007/s11033-013-2581-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-013-2581-8