Abstract
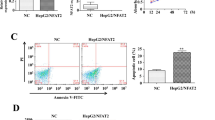
Purpose: This study aims to explore the differential expression of SFN gene and its regulatory role in different hepatocarcinoma cells, and the impact on hepatocarcinoma. Materials and Methods: High and low SFN expression cells were screened by qRT-PCR and western blotting methods. SFN over expression and interference vectors were constructed. Cell viability was detected by CCK8 kit, cell cycle and apoptosis were detected by flow cytometry. Cell invasion and migration were detected. CCNB1 and CDK1 expression levels were detected by qRT-PCR and Western blotting methods. Results: The high SFN expression BEL7402 cells and the low SFN expression Hep3B cells were screened from Hep3B, HepG2, and BEL7402 cells. The activity of Hep3B cells overexpression vector SFN-pcDNA3.1(+) decreased and apoptosis increased, the ratio of G0/G1 decreased and the ratio of S phase increased. The activity of BEL7402 cells transfected with SFN siRNA decreased and apoptosis increased, the ratio of G0/G1 decreased and the ratio of G2/M increased. Interference and overexpression vectors have little effect on the invasion and migration of the two cells. The expression of CDK1 in Hep3B cells decreased significantly, the expression of CDK1 and CCNB1 in BEL7402 cells increased significantly. Conclusions: The differentially expressed SFN gene can regulate the growth of the two hepatocarcinoma cells, high expression of SFN gene can inhibit their growth. The mechanism may be achieved by regulating CCNB1 and CDK1 expression.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
European Association for the Study of the Liver (2018) EASL Clinical Practice Guidelines: Management of hepatocellular carcinoma. J. Hepatol. 69: 182–236.
Bray, F., J. Ferlay, I. Soerjomataram, R. L. Siegel, L. A. Torre, and A. Jemal (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. Cancer. J. Clin. 68: 394–424.
Omata, M., A. L. Cheng, N. Kokudo, M. Kudo, J. M. Lee, J. Jia, R. Tateishi, K. H. Han, Y. K. Chawla, S. Shiina, W. Jafri, D. A. Payawal, T. Ohki, S. Ogasawara, P. J. Chen, C. R. A. Lesmana, L. A. Lesmana, R. A. Gani, S. Obi, A. K. Dokmeci, and S. K. Sarin (2017) Asia-Pacific clinical practice guidelines on the management of hepatocellular carcinoma: a 2017 update. Hepatol. Int. 11: 317–370.
Yuan-Dong, S., Z. Hao, X. Hui-Rong, L. Jing-Zhou, W. Hui-Yong, H. Jian-Jun, and J. M. Yu (2019) Combination therapy: Meta-analysis of the effects of TACE and cryoablation on hepatocellular carcinoma. Medicine (Baltimore). 98: e18030.
Sanmamed, M. F. and L. Chen (2019) A paradigm shift in cancer immunotherapy: From enhancement to normalization. Cell. 176: 677.
Leffers, H., P. Madsen, H. H. Rasmussen, B. Honoré, A. H. Andersen, E. Walbum, J. Vandekerckhove, and J. E. Celis (1993) Molecular cloning and expression of the transformation sensitive epithelial marker stratifin. A member of a protein family that has been involved in the protein kinase C signalling pathway. J. Mol. Biol. 231: 982–998.
Nakamura, Y., K. Oshima, Y. Naoi, T. Nakayama, S. J. Kim, K. Shimazu, A. Shimomura, N. Maruyama, Y. Tamaki, and S. Noguchi (2012) 14-3-3sigma expression is associated with poor pathological complete response to neoadjuvant chemotherapy in human breast cancers. Breast Cancer Res. Treat. 134: 229–236.
Zhou, R., Z. Shao, J. Liu, W. Zhan, Q. Gao, Z. Pan, L. Wu, L. Xu, Y. Ding, and L. Zhao (2018) COPS5 and LASP1 synergistically interact to downregulate 14-3-3sigma expression and promote colorectal cancer progression via activating PI3K/AKT pathway. Int. J. Cancer. 142: 1853–1864.
Umbricht, C. B., E. Evron, E. Gabrielson, A. Ferguson, J. Marks, and S. Sukumar (2001) Hypermethylation of 14-3-3 sigma (stratifin) is an early event in breast cancer. Oncogene. 20: 3348–3353.
Itoguchi, N., T. Nakagawa, Y. Murata, D. Li, A. Shiba-Ishii, Y. Minami, and M. Noguchi (2015) Immunocytochemical staining for stratifin and OCIAD2 in bronchial washing specimens increases sensitivity for diagnosis of lung cancer. Cytopathology. 26: 354–361.
Zeng, H., Y. Hui, W. Qin, P. Chen, L. Huang, W. Zhong, L. Lin, H. Lv, and X. Qin (2020) High-throughput sequencing-based analysis of gene expression of hepatitis B virus infection-associated human hepatocellular carcinoma. Oncol. Lett. 20: 18.
Shepherd, R., S. A. Forbes, D. Beare, S. Bamford, C. G. Cole, S. Ward, N. Bindal, P. Gunasekaran, M. Jia, C. Y. Kok, K. Leung, A. Menzies, A. P. Butler, J. W. Teague, P. J. Campbell, M. R. Stratton, and P. A. Futreal (2011) Data mining using the Catalogue of Somatic Mutations in Cancer BioMart. Database (Oxford). 2011: bar018.
He, Z. X., P. Xiang, J. P. Gong, N. S. Cheng, and W. Zhang (2016) Radiofrequency ablation versus resection for Barcelona clinic liver cancer very early/early stage hepatocellular carcinoma: a systematic review. Ther. Clin. Risk Manag. 12: 295–303.
Deisboeck, T. S. and Z. Wang (2008) A new concept for cancer therapy: out-competing the aggressor. Cancer Cell Int. 8: 19.
Xie, B., S. Wang, N. Jiang, and J. J. Li (2019) Cyclin B1/CDK1-regulated mitochondrial bioenergetics in cell cycle progression and tumor resistance. Cancer Lett. 443: 56–66.
He, J., S. Yu, C. Guo, L. Tan, X. Song, M. Wang, J. Wu, Y. Long, D. Gong, R. Zhang, Z. Cao, Y. Li, and C. Peng (2019) Polyphyllin I induces autophagy and cell cycle arrest via inhibiting PDK1/Akt/mTOR signal and downregulating cyclin B1 in human gastric carcinoma HGC-27 cells. Biomed. Pharmacother. 117: 109189.
Choi, H. J. and B. T. Zhu (2019) Upregulated cyclin B1/CDK1 mediates apoptosis following 2-methoxyestradiol-induced mitotic catastrophe: Role of Bcl-XL phosphorylation. Steroids. 150: 108381.
Qian, J. Y., J. Gao, X. Sun, M. D. Cao, L. Shi, T. S. Xia, W. B Zhou, S. Wang, Q. Ding, and J. F. Wei (2019) KIAA1429 acts as an oncogenic factor in breast cancer by regulating CDK1 in an N6-methyladenosine-independent manner. Oncogene. 38: 6123–6141.
Zhang, R., H. Shi, F. Ren, M. Zhang, P. Ji, W. Wang, and C. Liu (2017) The aberrant upstream pathway regulations of CDK1 protein were implicated in the proliferation and apoptosis of ovarian cancer cells. J. Ovarian. Res. 10: 60.
Yang, H., Y. Y. Wen, R. Zhao, Y. L. Lin, K. Fournier, H. Y. Yang, Y. Qiu, J. Diaz, C. Laronga, and M. H. Lee (2006) DNA damage-induced protein 14-3-3 sigma inhibits protein kinase B/Akt activation and suppresses Akt-activated cancer. Cancer Res. 66: 3096–3105.
Yang, H., R. Zhao, and M. H. Lee (2006) 14-3-3sigma, a p53 regulator, suppresses tumor growth of nasopharyngeal carcinoma. Mol. Cancer Ther. 5: 253–260.
Lodygin, D. and H. Hermeking (2005) The role of epigenetic inactivation of 14-3-3sigma in human cancer. Cell Res. 15: 237–246.
Winter, M., D. Lodygin, B. Verdoodt, and H. Hermeking (2016) Deletion of 14-3-3sigma sensitizes mice to DMBA/TPA-induced papillomatosis. Oncotarget. 7: 46862–46870.
Li, Y. L., L. Liu, Y. Xiao, T. Zeng, and C. Zeng (2015) 14-3-3σ is an independent prognostic biomarker for gastric cancer and is associated with apoptosis and proliferation in gastric cancer. Oncol. Lett. 9: 290–294.
Acknowledgement
This study was supported by research project of Guangxi Health Committee (Z20200814).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Ying Hui and Xue Qin; Execution of experiments: Hao Zeng, Ying Hui and Wenzhou Qin; Data analysis: Peisheng Chen and Ying Hui; Discussion of results: Hao Zeng, Lifang Huang, Wenfu Zhong and Xue Qin; Wrote the manuscript: Liwen Lin, Hui Lv and Hao Zeng, Critical reading of manuscript: Ying Hui, Yi Feng, and Xue Qin.
Corresponding authors
Additional information
Disclosure Statement
The authors declare no conflicts of interest.
Neither ethical approval nor informed consent was required for this study.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hui, Y., Zeng, H., Feng, Y. et al. Regulatory Role of SFN Gene in Hepatocellular Carcinoma and Its Mechanism. Biotechnol Bioproc E 26, 375–383 (2021). https://doi.org/10.1007/s12257-020-0292-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-020-0292-2