Abstract
A suite of polymorphic microsatellite markers and the complete mitochondrial genome sequence was developed by next generation sequencing (NGS) for the critically endangered orange-bellied parrot, Neophema chrysogaster. A total of 14 polymorphic loci were identified and characterized using DNA extractions representing 40 individuals from Melaleuca, Tasmania, sampled in 2002. We observed moderate genetic variation across most loci (mean number of alleles per locus = 2.79; mean expected heterozygosity = 0.53) with no evidence of individual loci deviating significantly from Hardy–Weinberg equilibrium. Marker independence was confirmed with tests for linkage disequilibrium, and analyses indicated no evidence of null alleles across loci. De novo and reference-based genome assemblies performed using MIRA were used to assemble the N. chrysogaster mitochondrial genome sequence with mean coverage of 116-fold (range 89 to 142-fold). The mitochondrial genome consists of 18,034 base pairs, and a typical metazoan mitochondrial gene content consisting of 13 protein-coding genes, 2 ribosomal subunit genes, 22 transfer RNAs, and a single large non-coding region (control region). The arrangement of mitochondrial genes is also typical of Avian taxa. The annotation of the mitochondrial genome and the characterization of 14 microsatellite markers provide a valuable resource for future genetic monitoring of wild and captive N. chrysogaster populations. As found previously, NGS provides a rapid, low cost and reliable method for polymorphic nuclear genetic marker development and determining complete mitochondrial genome sequences when only a fraction of a genome is sequenced.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Next Generation Sequencing (NGS) has revolutionised the field of molecular biology through the rapid and cost-effective collection of large amounts of genomic data. While this technology has been applied widely across a variety of research disciplines [31], its utility has remained limited in the field of conservation genetics. NGS technologies provide an effective platform for the development of genetic markers that can be used to provide insight into population processes and the evolutionary history of species. This information is critical when devising optimal conservation strategies and is therefore being increasingly used to guide management decisions. More recently NGS has been used for the rapid and cost effective isolation of nuclear microsatellite markers, where it has been shown that random sequencing of a small fraction of a genome can result in a high density of potential microsatellite loci [10]. Polymorphic microsatellite loci are used widely in population genetics and are routinely used in the field of conservation genetics to identify individuals [21], determine relatedness between individuals [13], estimate gene flow and genetic structure between populations [5], determine genetic diversity estimates within populations [20] and also estimate effective [11] and census [32] population size.
Even though recent studies have demonstrated the utility of NGS for isolating microsatellite loci for species [18, 19], these studies rarely explore or utilize the bulk of NGS data. For example, a 454 sequencing analysis using only 1/8th of a 70 × 75 mm Pico Titer Plate will typically generate over 20 Mb of sequence data, yet microsatellite containing contigs often amount to less than 3 % of total sequence reads (Miller and Weeks unpubl. data). The remaining sequence data generally overlooked contains a high density of contigs of both nuclear and mitochondrial origin, many of which are potentially valuable genetic markers for systematic research. Mitochondrial DNA is particularly useful and is applied widely to explore patterns of intra- and interspecific genetic variation [5, 17]. By exploiting certain tissue types (e.g. blood or muscle), total genomic DNA extractions can contain high concentrations of mtDNA, which may then be overrepresented in NGS analyses [7]. Such an approach may allow entire mtDNA genome sequences to be generated when only a small portion of the genome is sequenced and at a fraction of the cost of traditional approaches. Here we test this method by extracting DNA from muscle tissue of the critically endangered orange-bellied parrot, Neophema chrysogaster, and undertaking a modest (1/8th of a 70 × 75 mm picoTitre Plate) 454 NGS analysis to isolate microsatellite loci and mtDNA sequences.
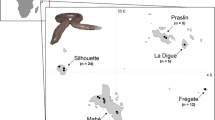
Neophema chrysogaster is endemic to south-eastern Australia, with a former range on the mainland extending from Adelaide (South Australia) through coastal Victoria and as far north as Sydney (New South Wales), and Tasmania extending along the west and south coasts, east to Bruny Island [6]. Since the 1920s, the species has suffered a steady decline in the wild, with major threats including the degradation and loss of habitat, and introduced predators and competitors [6], and the wild population is currently estimated to be less than 50 individuals (R. Pritchard, pers. comm.). A captive breeding program was initiated in the late 1980s, but to date this program has largely been unsuccessful in returning birds back to the wild. The orange-bellied parrot is protected by both State and Commonwealth legislation throughout its range, including a listing as ‘Endangered’ under the Environment Protection and Biodiversity Conservation (EPBC) Act 1999. The International Union for Conservation of Nature and Natural Resources (IUCN) lists the orange-bellied parrot as ‘Critically Endangered’.
Microsatellite loci were developed primarily to help inform and monitor conservation efforts both in the captive breeding colony and the remaining wild populations. Similarly, we aimed to target mitochondrial DNA in our NGS approach so that future studies could use this resource to gain a better understanding of the evolutionary history of Neophema chrysogaster.
Materials and methods
Next-generation sequencing
The 454 next generation sequencing platform was used to identify microsatellite and mitochondrial markers for N. chrysogaster. Approximately 10 μg of genomic DNA was extracted from muscle tissue from a single specimen using a QIAGEN DNA Easy kit (Qiagen). DNA was subsequently processed by the Australian Genome Research Facility (AGRF) where it was nebulized, ligated with 454 sequencing primers and tagged with a unique oligo sequence allowing sequences to be separated from pooled species DNA sequences using post-run bioinformatic tools. The DNA sample was analyzed using high throughput DNA sequencing on 1/8th of a 70 × 75 mm Pico Titer Plate using the Roche GS FLX (454) system [15].
Microsatellite isolation and characterisation
Unique sequence contigs possessing microsatellite motifs were identified using the software QDD version 2 [16]. Primer 3 [29] was used to design optimal primer sets for each unique contig where possible. A selection of 55 contigs including di-, tri-, and tetra-nucleotide repeats, were used for subsequent analysis. Loci were screened for polymorphism using template DNA from eight individuals, representing three temporally spaced samples (1992, 2002, and 2005) from the wild population at Melaleuca, Tasmania. Loci were pooled into groups of four, labeled with unique fluorophores (FAM, NED, VIC, PET) and co-amplified by multiplex PCR using a Qiagen multiplex kit (Qiagen) and an Eppendorf Mastercycler S gradient PCR machine following the protocol described by Blacket et al. [3]. Genotyping was subsequently performed using an Applied Biosystems 3,730 capillary analyzer (http://www.agrf.org.au) and product lengths were scored manually and assessed for polymorphisms using GeneMapper version 4.0 (Applied Biosystems).
Polymorphic loci were selected, pooled into two groups for multiplexing based on observed locus specific allele size ranges, and further characterized using DNA from 40 individuals sampled in 2002 from the Melaleuca wild population. Microsatellite profiles were again examined using GeneMapper version 4.0 and alleles scored manually. The Excel Microsatellite Toolkit [24] was then used to estimate expected (H E) and observed (H O) heterozygosities and number of alleles (NA), while examination of conformation to Hardy–Weinberg equilibrium (HWE), the inbreeding coefficient (F IS) and linkage disequilibrium estimates between all pairs of loci was conducted using GENEPOP version 4 [26]. Where necessary, significance values were adjusted for multiple comparisons using Bonferroni corrections [28]. Finally, all loci were assessed using MICRO-CHECKER to check for null alleles and scoring errors [34]. The frequency of null alleles per locus was obtained using the ‘Brookfield 1’ formula, as evidence of null homozygotes across loci was not observed [2].
Mitochondrial assembly and annotation
Sequence reads in SFF format were edited by trimming 454 primer tags using the Roche software. Sequence assembly of genomic sequence contigs was achieved by ‘de novo assembly’ using the assembly software MIRA and default 454 parameters [4]. Annotation of the mitochondrial genome was determined using the DOGMA online software using the default parameters including a 5× parallel BLAST search option (http://dogma.ccbb.utexas.edu). DOGMA estimated gene positions, codon usages, transcriptional orientations, and where relevant, secondary structures. All alignments were confirmed by visual inspection with reference genome sequences. Genome annotation was exported to SEQUIN and submitted to Genbank (accession number JX133087). The software OGDRAW [14] was used to provide a visual depiction of the N. chrysogaster mitochondrial DNA gene content and orientation.
Results and discussion
Next-generation sequencing and de novo genome assembly
A total of 73,522 sequence reads covering 24.7 Mb of the N. chrysogaster genome was obtained by NGS. Previous studies indicate that these figures are not excessive as these are commonly achieved by NGS analyses using only 1/16th of a 70 × 75 mm picoTitre Plate [18, 19]. Nonetheless this data represents ~2 % of the ~1.5 Gb parrot genome [1]. MIRA assemblies indicate that approximately 6.5 % of the total sequence reads are of mitochondrial origin (4,765 reads). De novo assembly of mtDNA sequence contigs revealed complete genome coverage with a mean coverage of 116-fold (range 89–142).
Microsatellite isolation and characterization
A total of 1,130 unique sequence contigs possessing microsatellite motifs were identified by QDD analysis, of which 883 contigs were found to possess optimal priming sites. Initially, 55 contigs were screened for polymorphism, with 39 containing di-nucleotide repeats, 12 containing tri-nucleotide repeats, and 4 containing tetra-nucleotide repeat motifs. The screening analysis found 14 loci to be polymorphic, 26 were monomorphic and 15 failed to amplify.
The majority of the 14 polymorphic loci were characterized by low to moderate genetic variation, with an average of 2.79 alleles per locus (range = 2–8 alleles) and heterozygosity estimates ranging between 0.06 and 0.74 (mean = 0.53). Linkage disequilibrium analyses confirmed maker independence (indicating no evidence of significant linkage between loci), while MICRO-CHECKER analyses revealed no evidence of null alleles or scoring issues across loci. All loci were found to conform with Hardy–Weinberg expectations and estimates of F IS indicate no significant evidence of heterozygote excess or deficit. HWE and F IS estimates for marker OBP55 are high, however, following Bonferroni corrections these were not significant. Table 1.
Mitochondrial genome of N. chrysogaster
Genome composition
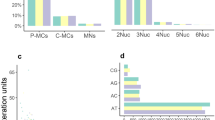
The mitochondrial genome of N. chrysogaster is a circular molecule 18,034 bp in length, and characterized by a typical metazoan gene composition; 13 protein-coding genes, 2 ribosomal subunit genes (rRNA), and 22 transfer RNA genes (trn) (Fig. 1; Table 2). The gene arrangement, including respective transcriptional polarities of genes, is typical of avian species and identical to those taxa described in Table 3. Five gene pairs were found overlapping by up to 6 bp (Table 2), a characteristic that has been reported for other animal mtDNAs including birds [35]. The majority-strand (α) encodes 28 genes, while the minority-strand (β) encodes 9 genes (Table 2). The nucleotide composition of the α-strand is 5,498 adenine (30.5 %), 6,014 cytosine (33.3 %), 2,546 guanine (14.1 %), and 3,985 thymine (22.1 %). While A–T biases of higher magnitude are commonly observed in other taxonomic groups such as arthropods and nematodes, more modest biases are common in birds, mammals and fish [30]. Bias to cytosine on the α-strand is a common feature of metazoan mtDNAs and is a feature that appears associated with the duration of single-stranded state of ‘heavy-stranded’ genes during mtDNA replication [27, 30].
Gene map of the Neophema chrysogaster mitochondrial genome. COI-III indicates cytochrome c oxidase subunits 1–3; cyt b, cytochrome b; ATP6–8, ATPase subunits 6 and 8; ND1–6/4L, NADH dehydrogenase subunits 1–6/4L. Transfer RNA genes are designated by single-letter amino acid codes (Table 2.) Protein-coding and RNA genes on the outside circle are transcribed in a clockwise direction (α-strand), those encoded on the β-strand are shown on the inside circle of the molecule
A total of 2,662 noncoding nucleotides are evident in the genome, with 158 bp at 24 intergenic regions and a large 2,504 bp noncoding region (Table 2). The large noncoding region found represents the putative control region on the basis of its relative position between the trnQ and trnF which is typical of birds, and sequence characteristics (A + T-rich, noncoding). The N. chrysogaster putative control region is notably larger than those reported for species given in Table 3, however, control region length variations are common among avian species and other metazoan groups [35]. Gene lengths and A + T base compositions of the N. chrysogaster α-strand, protein-coding, rRNA, and trn genes, as well as the putative control region, are displayed in Table 3.
Protein-coding genes
All protein-coding genes except for ND6 are encoded by the α-strand (Table 2), with overlapping nucleotides observed at the ATP6 and 8, and NAD4 and NAD4L gene boundaries (Table 2). Overlaps at these particular gene boundaries are a common feature of metazoan mitochondrial genomes [35], and have been validated by surveys of bicistronic transcripts and protein characteristics [9, 23]. Translation initiation and termination codons of the N. chrysogaster 13 protein-coding genes are summarized in Table 2. The standard methionine (ATN) initiation codon was inferred for 12 of the 13 genes while the ND5 gene appears to use a valine (GTG) codon, a nonstandard codon used in other metazoans including birds [8, 35]. Open reading frames are terminated with the typical TAA and TAG codons for all genes except for COIII and NAD4. We suggest that these genes are characterized by truncated termination codons (T) with the production of the TAA termini being created by post-transcriptional polyadenylation [23, 33]. This is a common feature reported for other metazoan mt genomes [12, 17, 22].
Ribosomal and transfer RNA genes
The rRNA gene boundaries were estimated by BLAST sequence alignments implemented in DOGMA, with a high degree of conservatism at the beginning and end of the respective genes across avian taxa. Both ribosomal subunit genes are encoded by the α-strand with the rrnS (12S) gene separating trnF and trnV, and the rrnL (16S) gene separating the trnV, trnL(uac). The genomic position and transcriptional polarity of the rRNA genes is typical of avian species (Table 3).
A total of 22 trn genes corresponding with the standard set of metazoan genes were identified on the basis of their respective anticodons and secondary structures (Table 2). Gene lengths and anticodon sequences are largely congruent with those described for other avian species described in Table 3. All genes can be folded into the canonical cloverleaf structure except for trnS(gcu) and trnK which lack the DHU arm, instead replaced with unpaired loops 8 and 13 bases in length, respectively. Replacements loops are commonly observed in metazoan trnS genes [35].
Conclusion
The NGS approach using the 454 sequencing platform was successful in isolating 1,130 microsatellite containing contigs for N. chrysogaster from a total of 73,522 sequence contigs that covered approximately 24.7 Mb of the genome. While birds are thought to have inherently low numbers of microsatellite loci [25], we successfully developed 14 polymorphic microsatellite markers that will be a valuable resource for devising effective conservation strategies for the species. These markers can be used to determine changes in genetic variation, relatedness, inbreeding, gene flow, genetic structure, effective population size and past population processes in both the wild and captive populations. They should also prove integral in guiding captive breeding programs, determining success of reintroductions, and assigning parentage in the wild. We genotyped all 14 loci from blood samples collected from 40 wild birds at Melaleuca in 2002, showing moderate to low levels of genetic variation as measured by estimates of heterozygosity and allelic richness. In 2004 the estimated population size of N. chrysogaster in the wild was thought to be less than 150 birds [6]. However, substantial declines have occurred since then and it is now estimated that less than 50 N. chrysogaster currently persist in the wild (R. Pritchard, pers. comm.). This highlights the importance of ongoing genetic monitoring of both wild and captive populations to inform on-going conservation efforts for this important and iconic species.
Interestingly, despite sequencing only a fraction of the nuclear genome (approx 2 %), we were able to obtain an average coverage of 116-fold of the N. chrysogaster complete mitochondrial genome sequence. Extracting DNA from muscle tissue, which is inherently rich in mitochondria [7], likely resulted in an overrepresentation of sequence contigs of mitochondrial origin in the NGS analysis. We have demonstrated that by targeting specific tissues, the NGS analysis is a rapid and cost effective method for not only developing nuclear microsatellite markers, but also sequencing entire mitochondrial genomes. Combined these genetic markers are an extremely valuable resource for investigating the population genetics and evolutionary histories of endangered species, that in turn provides a framework for establishing effective conservation strategies.
References
Andrews CB, Gregory TR (2009) Genome size is inversely correlated with relative brain size in parrots and cockatoos. Genome 52:261–267
Brookfield JFY (1996) A simple new method for estimating null allele frequency from heterozygote deficiency. Mol Ecol 5:453–455
Blacket MJ, Robin C, Good RT, Lee SF, Miller AD (2012) Universal primers for fluorescent labelling of 1 PCR fragments: an efficient and cost effective approach to genotyping by fluorescence. Mol Ecol. doi:10.1111/j.1755-0998.2011.03104.x
Chevreux B, Wetter T, Suhai S (1999) Genome sequence assembly using trace signals and additional sequence information computer science and biology: Proceedings of the german conference on bioinformatics (GCB) 99, pp. 45–56
Coleman RA, Pettigrove V, Raadik TA, Hoffmann AA, Miller AD, Carew ME (2010) Microsatellite markers and mtDNA data indicate two distinct groups in dwarf galaxias, Galaxiella pusilla (Mack) (Pisces:Galaxiidae), a threatened freshwater fish from south eastern Australia. Conserv Genet 11:1911–1928
Commonwealth of Australia (2005) Orang-bellied parrot recovery plan. Commonwealth of Australia, Canberra
Dalziel AC, Morre SE, Moyes CD (2005) Mitochondrial enzyme content in the muscles of high-performance fish: evolution and variation among fiber types. Am J Physiol Regul Integr Comp Physiol 228:R163–R172
Desjardins P, Morais R (1991) Nucleotide sequence and evolution of coding and noncoding regions of a quail mitochondrial genome. J Mol Evol 32(2):153–161
Fearnley IM, Walker JE (1986) Two overlapping genes in bovine mitochondrial DNA encode membrane components of ATP synthase. EMBO 5:2003–2008
Gardner MG, Fitch AJ, Bertozzi T, Lowe AJ (2011) Rise of the machines: recommendations for ecologists when using next generation sequencing for microsatellite development. Mol Ecol Resour 11(6):1093–1101
Gomez-Uchida D, Banks MA (2006) Estimation of effective population size for the long-lived darkblotched rockfish Sebastes crameri. J Hered 97(6):603–606
Ki JS, Hwang DS, Park TJ, Han SH, Lee JS (2010) A comparative analysis of the complete mitochondrial genome of the Eurasian otter Lutra lutra (Carnivora: Mustelidae). Mol Biol Rep 37(4):1943–1955
Larson S, Christiansen J, Griffing D, Ashe J, Lowry D, Andrews K (2011) Relatedness and polyandry of sixgill sharks, Hexanchus griseus, in an urban estuary. Conserv Genet 12(3):679–690
Lohse M, Drechsel O, Bock R (2007) Organellar genome DRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr Genet 52:267–274
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Meglécz E, Costedoat C, Dubut V, Gilles A, Malausa T, Pech N, Martin JF (2010) QDD: a user-friendly program to select microsatellite markers and design primers from large sequencing projects. Bioinformatics 26(3):403–404
Miller AD, Austin CM (2006) The complete mitochondrial genome of the mantid shrimp Harpiosquilla harpax, and a phylogenetic investigation of the Decapoda using mitochondrial sequences. Mol Phylogenet Evol 38:565–574
Miller AD, Versace VL, Matthews T, Bowie KC (2012) The development of 20 microsatellite loci for the Australian marine mollusk, Donax deltoides, through next generation DNA sequencing. Conserv Genet Resour 4(2):257–260
Miller AD, van Rooyen A, Ayres RM, Raadik TA, Fairbrother P, Weeks AR (2012) The development of 24 polymorphic microsatellite loci for the endangered barred galaxias, Galaxias fuscus, through next generation DNA sequencing. Conserv Genet Resour. doi:10.1007/s12686-012-9605-x
Mitrovski P, Heinze DA, Broome L, Hoffmann AA, Weeks AR (2007) High levels of variation despite genetic fragmentation in populations of the endangered mountain pygmy-possum, Burramys parvus, in alpine Australia. Mol Ecol 16:75–87
Mondol S, Navya R, Athreya V, Sunagarl K, Selvaraj VM, Ramakrishnan U (2009) A panel of microsatellites to individually identify leopards and its application to leopard monitoring in human dominated landscapes. BMC Genet 10:79
Oh DJ, Kim JY, Lee JA, Yoon WJ, Park SY, Jung YH (2007) Complete mitochondrial genome of the rabbitfish Siganus fuscescens (Perciformes, Siganidae). DNA Seq 18(4):295–301
Ojala D, Montoya J, Attardi G (1981) tRNA punctuation model of RNA processing in human mitochondria. Nature 290:470–474
Park SDE (2001) Trypanotolerance in west african cattle and the population genetic effects of selection. PhD Thesis, University of Dublin
Primmer CG, Raudsepp T, Chowdhary BP, Moller AP, Ellegren H (1997) Low frequency of microsatellites in the avian genome. Genome Res 7:471–482
Raymond M, Rousset F (1995) An exact test for population differentiation. Evolution 49:1280–1283
Reyes A, Gissi C, Pesole G, Saccone C (1998) Asymmetrical directional mutation pressure in the mitochondrial genome of mammals. Mol Biol Evol 15:957–966
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43(1):223–225
Rozen S, Skaletsky HJ (2000) Primer 3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, NJ, pp 365–386
Saccone C, De Giorgi C, Gissi C, Pesole G, Reyes A (1999) Evolutionary genomics in metazoa: the mitochondrial DNA as a model system. Gene 238:195–209
Schuster SC (2008) Next-generation sequencing transforms today’s biology. Nat Methods 5:16–18
Skaug HJ (2001) Allele-sharing methods for estimation of population size. Biometrics 57:750–756
Temperley RJ, Wydro M, Lightowlers RN, Chrzanowska- Lightowlers ZM (2010) Human mitochondrial mRNAs–like members of all families, similar but different. Biochim Biophys Acta 1797(6–7):1081–1085
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538
Wolstenholme DR (1992) Animal mitochondrial DNA: structure and evolution. Int Rev Cytol 141:173–216
Acknowledgments
We thank Healesville Sanctuary for providing the N. chrysogaster specimen used to extract DNA for the NGS approach. Neil Murray is thanked for providing N. chrysogaster samples used for the characterization of microsatellite loci.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miller, A.D., Good, R.T., Coleman, R.A. et al. Microsatellite loci and the complete mitochondrial DNA sequence characterized through next generation sequencing and de novo genome assembly for the critically endangered orange-bellied parrot, Neophema chrysogaster . Mol Biol Rep 40, 35–42 (2013). https://doi.org/10.1007/s11033-012-1950-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1950-z