Abstract
Breast cancer is one of the most frequently diagnosed cancers and the leading cause of cancer deaths among females across the world, accounting for 23 % (1.38 million) of total new cancer cases and 14 % (0.45 million) of the total cancer deaths in 2008. c-kit is expressed in mast cell growth factor, cellular migration, proliferation, melanoblasts, haematopoietic progenitors and germ cells. We have designed our study with aim to explore the c-kit gene mutations in invasive ductal carcinoma (IDC) breast. To ascertain the range of mutations in exon 11, 13 and 17 of c-kit gene in 53 cases of IDC breast, we carried out PCR-SSCP followed by DNA sequencing. The mutation frequency of c-kit gene in exon 11, 13 and 17 were 9.43 % (5/53), 1.88 % (1/53) and 3.77 % (2/53), respectively. During our mutational analysis, we have detected five missense mutations in exon 11 (Pro551Leu, Glu562Val, Leu576Phe, His580Tyr and Phe584Leu), one missense mutation in exon 13 (Ser639Pro) and two missense mutations in exon 17 (Arg796Gly and Asn822Ser). It seems that c-kit mutations might participate in breast cancer pathogenesis and may be utilized as predictive marker, since the loss of c-kit positivity is generally linked with different types of breast cancer. Further molecular studies are necessary to validate the association of c-kit gene mutation in IDC breast pathogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is one of the most frequently diagnosed cancer and the leading cause of cancer deaths among females across the world, accounting for 23 % (1.38 million) of total new cancer cases and 14 % (0.45 million) of the total cancer deaths in 2008. About half the breast cancer cases and 60 % of the deaths are estimated to occur in economically developing countries [1]. Estimated breast cancer incidence and mortality in India were 22.9 and 11.1 % (http://globocan.iarc.Fr/factsheets/cancers/breast.asp). c-kit is a type III receptor tyrosine kinase that is activated by binding to its ligand stem cell factor. In humans, the c-kit gene has been mapped on chromosome 4q11-12 and encodes a 145 KD receptor tyrosine kinase III [2]. c-kit plays an important role in the development of multiple cell types including hematopoietic cells, germ cells, melanocytes and mammary gland epithelium [3, 4]. Numerous findings have suggested the role of c-kit in the development of a series of cells including haematopoietic cells [5]. Binding of c-kit to stem cell factor initiates a type phosphorylation of tyrosine residue which is necessary factor for the activation of numerous signal transduction pathways linked with the process of apoptosis, proliferation and tumorigenesis [6]. The expression of c-kit protein has been recognized in an extensive series of malignant tumours including myeloid leukaemia [7], small cell lung cancer [8], seminomas [9] and gastrointestinal stromal tumors (GIST) [10] but often expressed in other forms of tumour including malignant melanoma [11]. Hirota et al. [10] reported that some GISTs have activating mutations which are interconnected with the activation of the c-kit gene. c-kit gene mutations in exons 8, 9, 11, 13 and 17 have been already reported in GIST, human solid tumours, human germ cell tumours and leukaemia cases [9, 12–14]. In brief, the GIST helps in activating mutations particularly agglomerated in exons 9, 11 and 13 [15, 16]. Earlier, the function of c-kit has been evaluated notably in subtypes of breast carcinoma associated with different kinds of clinical results. c-kit gene represents to be an indicator of high-grade breast carcinoma-group, frequently enclose the carcinomas with the mesenchymal and myoepithelial differentiation that involve an inadequate prognosis [17, 18]. The undifferentiated-type breast carcinomas appear to show mammary epithelial stem cell like features that may possibly be identified by the c-kit gene over-expression/mutations [18]. Various studies have shown that c-kit is highly expressed in normal breast epithelium, but its level decreases or is null in primary invasive breast cancer or breast cancer metastases [19–22]. The involvement of c-kit in ductal carcinoma in situ, a premalignant breast disease has been recently evaluated in correlation with nuclear grading [23]. As of now, adequate data is not available dealing with c-kit gene mutation in invasive ductal carcinoma (IDC) breast cases. In this study we have screened the mutations in exon 11, 13 and 17 of c-kit gene in IDC breast among the north Indian population and further explored whether the c-kit gene mutations were valuable as predictive markers in IDC Breast.
Materials and methods
Breast specimen
Our study group included 53 cases of IDC diagnosed by fine needle aspiration biopsy yielded positive cytology using quantitative, qualitative and histological grade according to Elston and Ellis [24] and Haagensen et al. [25]. Specimens were collected from the department of Pathology, Era’s Lucknow Medical College and other hospital of Lucknow, Uttar Pradesh, India.
DNA isolation
The genomic DNA was extracted by DNA extraction kit (Fermentas, Germany). The quality of DNA was checked on 1 % agarose gel and stored at −20 °C.
Polymerase chain reaction and single-strand conformational polymorphism
Polymerase chain reaction (PCR) was performed with 30 μl PCR reaction mixture containing 100 ng of template DNA, 10 pmol of each primer, 1X PCR master mix reaction and 0.5 units of Taq polymerase enzyme (Fermentas, Germany) in an MJ Mini Thermocycler (Bio-Rad, UK). The cycling conditions was initial denaturation step at 96 °C for 5 min, denaturation at 96 °C for 30 s, followed by annealing at 56 °C for 30 s, and extension at 72 °C for 30 s, repeated for 35 cycles followed by a final extension step at 72 °C for 9 min using the primers for exon 11 forward 5- ATTATTAAAAGGTGATCTATTTTTC -3 and reverse 5- ACTGTTATGTGTACCCAAAAAG -3, exon 13 forward 5- ATCAGTTTGCCAGTTGTGCT -3 and reverse 5- TTTATAATCTAGCATTGCC -3 and for exon 17 forward 5- TTCACTCTTTACAAGTTAAAATG -3 and reverse 5- GGACTGTCAAGCAGAGAATG -3. Single-strand conformational polymorphism analysis was performed according to Orita et al. [26] with some modifications. 20 μl of amplified PCR product with equal volume of denaturing dye were denatured at 96 °C for 6 min and immediately snap-cooled. These were loaded on 10 % polyacrylamide gel which was run in pre-cooled 1X TBE (Tris–Borate EDTA) buffer for 15 h at 130 Volt. The gel tank was placed in a cold room at 4 °C. After electrophoresis silver staining was done. Electrophoresis mobility shift in single strand DNA of the cases and their wild type homologous were comparatively evaluated by the gel patterns [26].
Analysis of sequencing
Those samples which showed shift in mobility compared to wild types were further sequenced using an automated sequencer, ABI 3730XL DNA Analyser (Applied Biosystems, Foster city, California, USA). Mutations were reconfirmed by sequencing amplicons in both the directions and in the independent second samples, too. Sequences were analysed using BLAST (National Center for Biotechnology Information) and BioEdit software.
Results
53 cases diagnosed as IDC categorised as 28 cases of grade I, 15 cases of grade II and 10 samples of grade III. The mean age of cases were 55 years with SD ± 6.75, ages ranging between 40 and 70 years. IDC breast tumour sizes varied from 0.5 to 2.2 cm.The estrogen receptor (ER) was positive for 29 samples and negative for 24 samples. The human epidermal growth factor receptor 2 (HER2) was 2+ (borderline) for 13 samples, 1+ (negative) for 19 samples and 0 (negative) for 11 samples. We could not performed Her2 analysis in 10 samples. Eight missense mutations were detected in six IDC breasts cancer cases in exons 11, 13 and 17 (Figs. 1 and 2). Five missense mutations were detected in exon 11 (Pro551Leu, Glu562Val, Leu576Phe, His580Tyr and Phe584Leu), one missense mutation in exon 13 (Ser639Pro) and two missense mutations in exon 17 (Arg796Gly and Asn822Ser). Details of the clinical data and missense mutations detected in exons 11, 13 and 17 are shown in Table 1.
Amino acid sequences of the exons 11, 13 and 17 of c-kit gene. The wild-type sequence is shown above in the cases. Missense mutations are shown in shade filled square and detected in cases no. 03, 11, 24, 29, 37 and 45. The sequence at codons: a codon 550–591 (exon 11) b codon 627–664 (exon 13) c codon 788–828 (exon 17)
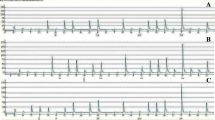
Nucleotides sequence chromatogram of c-kit gene missense mutations: a exon 11 nucleotides C → T, A → T, C → T, C → T, T → C and resulting in the amino-acid substitution Pro551Leu, Glu562Val, Leu576Phe, His580Tyr and Phe584Leu b exon 13 nucleotides T → C and resulting in the amino-acid substitution Ser639Pro c exon 17 nucleotides A → G, A → G and resulting in the amino-acid substitution Arg796Gly and Asn822Ser
Discussion
Breast cancer is a heterogeneous disease, encompassing a number of distinct biological entities. c-kit gene is involved in the development of multiple cell types together with haematopoietic cells, germ cells, mast cells and melanocytes. Among the north Indian population this is the first report stating mutations in the exons 11, 13 and 17 of c-kit gene in IDC breast. Ulivi et al. [27] reported a progressive decrease in c-kit gene mutations from normal mammary epithelium to carcinoma in situ and an almost complete loss of proto-oncogene mutation in case of invasive breast carcinoma. Supplementarily, a substantial mutation of c-kit gene and its ligand stem cell factor has been detailed in normal breast epithelium, depicting that an autocrine stimulation of c-kit gene by the stem cell factor is significant in the maintenance of differentiation of mammary epithelium, while a successive decline in both factors play along malignant transformation [28]. Remarkable to quote that 1–14 % of breast cancers has been reported which involve c-kit gene mutations in malignant breast epithelium [4, 19–22, 29]. c-kit gene mutation confirms a considerable relationship with high tumour grade and is identified in 82 % of progressive metastatic breast cancers [30]. Expression of c-kit gene in adenoid cystic carcinoma of the breast associated with an encouraging analysis has been reported in prior studies [31].
In our study c-kit gene missense mutations frequency was 11.32 % (6/53) IDC breasts. We have detected eight missense mutations in exons 11, 13 and 17 in 53 IDC breasts in our study Fig. 1. In particular five missense mutation detected in exon 11 (Pro551Leu, Glu562Val, Leu576Phe, His580Tyr and Phe584Leu), one missense mutation in exon 13 (Ser639Pro) and two missense mutation in exon 17 (Arg796Gly and Asn822Ser). c-kit gene mutation frequency in exon 11, 13 and 17 were 9.43 % (5/53), 1.88 % (01/53), and 3.77 % (2/53), respectively. Mutation at codon 584, 639 was detected in IDC grade I, Mutation at codon 562, 576 and 796 was detected in IDC grade II, While Mutation at codon 551, 580 and 822 was found in IDC grade III. The eight mutations detected in our study were found in IDC breast cases with positive ER status and negative HER2 status. The c-kit gene missense mutations at codon 551, 562, 576, 580, 584, 639, 796 and 822 were detected in our study has been reported, previously [32, 33] as shown in Fig. 2 and Table 2 but the substituted amino acid were different form the reported one.
It seems that c-kit mutations might participate in breast cancer pathogenesis and may be utilized as predictive marker, since the loss of c-kit positivity is generally linked with different types of breast cancer. Further molecular studies are necessary to validate the association of c-kit gene mutation in IDC breast pathogenesis.
References
Jemal A, Bray F, Melissa M, Ferlay J, Ward E, Forma D (2011) Global cancer statistics. CA Cancer J Clin 61:69–90
Yarden Y, Kuang WJ, Yang-Feng T, Coussens L, Munemitsu S, Dull TJ, Chen E, Schlessinger J, Francke U, Ullrich A (1987) Human proto-oncogene c-kit: a new cell surface receptor tyrosine kinase for an unidentified ligand. EMBO J 6:3341–3351
Gibson PC, Cooper K (2002) CD117 (KIT): a diverse protein with selective applications in surgical pathology. Adv Anat Pathol 9:65–69
Matsuda R, Takahashi T, Nakamura S et al (1993) Expression of the c-kit protein in human solid tumors and in corresponding fetal and adult normal tissues. Am J Pathol 142:339–346
Reilly JT (2002) Class III receptor tyrosine kinases: role in leukaemogenesis. Br J Haematol 116:744–757
Heinrich MC, Rubin BP, Longley BJ, Fletcher JA (2002) Biology and genetic aspects of gastrointestinal stromal tumors: KIT activation and cytogenetic alterations. Hum Pathol 33:484–495
Wang C, Curtis J, Geissler E, McCulloch EA, Minden MD (1989) The expression of the proto-oncogene c-kit in the blast cells of acute myeloblastic leukemia. Leukemia 3:699–702
Plummer H III, Catlett J, Leftwich J, Armstrong B, Carlson P, Huff T, Krystal G (1993) C-myc expression correlates with suppression of c-kit proto-oncogene expression in small cell lung cancer cell lines. Cancer Res 53:4337–4342
Tian Q, Frierson HF Jr, Krystal GW, Moskaluk CA (1999) Activating c-kit gene mutations in human germ cell tumors. Am J Pathol 154:1643–1647
Hirota S, Isozaki K, Moriyama Y (1998) Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 279:577–580
Lassam N, Bickford S (1992) Loss of c-kit expression in cultured melanoma cells. Oncogene 7:51–56
Hou YY, Tan YS, Sun MH, Wei YK, Xu JF, Lu SH, A-Ke-Su SJ, Zhou YN, Gao F, Zheng AH, Zhang TM, Hou WZ, Wang J, Du X, Zhu XZ (2004) c-kit gene mutation in human gastrointestinal stromal tumors. World J Gastroenterol 10:1310–1314
Sihto H, Sarlomo-Rikala M, Tynninen O, Tanner M, Andersson LC, Franssila K, Nupponen NN, Joensuu H (2005) KIT and platelet-derived growth factor receptor alpha tyrosine kinase gene mutations and KIT amplifications in human solid tumors. J Clin Oncol 23:49–57
Gari M, Goodeve A, Wilson G, Winship P, Langabeer S, Linch D, Vandenberghe E, Peake I, Reilly J (1999) c-kit proto-oncogene exon 8 in-frame deletion plus insertion mutations in acute myeloid leukaemia. Br J Haematol 105:894–900
Miettinen M, Lasota J (2001) Gastrointestinal stromal tumors—definition, clinical, histological, immunohistochemical, and molecular genetic features and differential diagnosis. Virchows Arch 438:1–12
Lux ML, Rubin BP, Biase TL, Chen CJ, Maclure T, Demetri G, Xiao S, Singer S, Fletcher CD, Fletcher JA (2000) KIT extracellular and kinase domain mutations in gastrointestinal stromal tumors. Am J Pathol 156:791–795
Tsuda H, Morita D, Kimura M, Shinto E, Ohtsuka Y, Matsubara O, Inazawa J, Tamaki K, Mochizuki H, Tamai S, Hiraide H (2005) Correlation of KIT and EGFR overexpression with invasive ductal breast carcinoma of the solid-tubular subtype, nuclear grade 3 and mesenchymal of myoepithelial differentiation. Cancer Sci 96:48–53
Tsuda H, Tani Y, Weisenberger J, Kitada S, Hasegawa T, Murata T, Tamai S, Hirohashi S, Matsubara O, Natori T (2005) Frequent KIT and epidermal growth factor receptor over expressions in undifferentiated-type breast carcinomas with ‘stem-cell-like’ features. Cancer Sci 96:333–339
Natali PG, Nicotra MR, Sures I, Mottolese M, Botti C, Ullrich A (1992) Breast cancer is associated with loss of the c-kit oncogene product. Int J Cancer 52:713–717
Ko CD, Kim JS, Ko BG, Son BH, Kang HJ, Yoon HS, Cho EY, Gong G, Ahn SH (2003) The meaning of the c-kit protooncogene product in malignant transformation in human mammary epithelium. Clin Exp Metastasis 20:593–597
Tsuura Y, Suzuki T, Honma K, Sano M (2002) Expression of c-kit protein in proliferative lesions of human breast: sexual difference and close association with phosphotyrosine status. J Cancer Res Clin Oncol 128:239–246
Chui X, Egami H, Yamashita J, Kurizaki T, Ohmachi H, Yamamoto S, Oqawa M (1996) Immunohistochemical expression of the c-kit proto-oncogene product in human malignant and non-malignant breast tissues. Br J Cancer 73:1233–1236
Diallo R, Rody A, Jackish C, Ting E, Schaefer KL, Kissler S, Karn T, Geddert H, Engels K, Kaufmann M, Gabbert HE, Shroyer KR, Poremba C (2006) c-kit expression in ductal carcinoma in situ of the breast: co-expression with HER-2/neu. Hum Pathol 37:205–211
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 1991:403–410
Haagensen CD, Lane N, Lattes R, Rodian C (1978) Lobular neoplasia (so-called lobular carcinoma in situ) of the breast. Cancer 42:737–769
Orita M, Iwahana H, Kanazawa H, Hayashi K, Sekiya T (1989) Detection of polymorphisms of human DNA by gel electrophoresis as single-strand conformation polymorphism. Proc Natl Acad Sci USA 86:2766–2770
Ulivi P, Zoli W, Medri L, Amadori D, Saragoni L, Barbanti F, Calistri D, Silvestrini R (2004) c-kit and SCF expression in normal and tumor breast tissue. Breast Cancer Res Treat 83:33–42
Kondi-Pafiti A, Arkadopoulos N, Gennatas C, Michalaki V, Frangou-Plegmenou M, Chatzipantelis P (2010) Expression of c-kit in common benign and malignant breast lesions. Tumori 96:978–984
Simon R, Panussis S, Maurer R, Spichtin H, Glatz K, Tapia C, Mirlacher M, Rufle A, Torhorst J, Sauter G (2004) KIT (CD117)- positive breast cancers are infrequent and lack KIT gene mutations. Clin Cancer Res 10:1075–1079
Palmu S, Soderstrom KO, Quazi K, Isola J, Salminen E (2002) Expression of c-kit and HER-2 tyrosine kinase receptors in poorprognosis breast cancer. Anticancer Res 22:411–414
Azoulay S, Laé M, Fréneaux P, Merle S, Al Ghuzlan A, Chnecker C, Rosty C, Klijanienko J, Sigal-Zafrani B, Salmon R, Fourquet A, Sastre-Garau X, Vincent-Salomon A (2005) KIT is highly expressed in adenoid cystic carcinoma of the breast, a basal-like carcinoma associated with a favourable outcome. Mod Pathol 18:1623–1631
Tetsu O, Phuchareon J, Chou A, Cox DP, Eisele DW, Jordan RCK (2010) Mutations in the c-kit gene disrupt mitogen-activated protein kinase signaling during tumor development in adenoid cystic carcinoma of the salivary glands. Neoplasia 12:708–717
Lizette V, Hongyan L, Al-Quran SZ, Dominique PC, Hui-Jia D, Chen L (2009) Identification of c-kit gene mutations in primary adenoid cystic carcinoma of the salivary gland. Mod Pathol 22:1296–1302
Acknowledgments
This study was supported by intramural project grant from Era’s Lucknow Medical College and Hospital, Lucknow, Uttar Pradesh, India.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hussain, S.R., Babu, S.G., Raza, S.T. et al. Screening of the c-kit gene missense mutation in invasive ductal carcinoma of breast among north Indian population. Mol Biol Rep 39, 9139–9144 (2012). https://doi.org/10.1007/s11033-012-1786-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1786-6





